Abstract
Both HDAC1 and HDAC2 are class I deacetylases acting as erasers of lysine-acetyl marks on histones and non-histone proteins. Several histone deacetylase inhibitors, either endogenous to the cell, such as the ketogenic β-hydroxybutyrate metabolite, or exogenous, such as butyrate, a microbial-derived metabolite, regulate HDAC activity. Different combinations of intestinal epithelial cell (IEC)-specific Hdac1 and/or Hdac2 deletion differentially alter mucosal homeostasis in mice. Thus, HDAC1 and HDAC2 could act as sensors and transmitters of environmental signals to the mucosa. In this study, enteroid culture models deleted for Hdac1 or Hdac2 were established to determine IEC-specific function as assessed by global transcriptomic and proteomic approaches. Results show that Hdac1 or Hdac2 deficiency altered differentiation of Paneth and goblet secretory cells, which sustain physical and chemical protection barriers, and increased intermediate secretory cell precursor numbers. Furthermore, IEC Hdac1- and Hdac2-dependent common and specific biological processes were identified, including oxidation-reduction, inflammatory responses, and lipid-related metabolic processes, as well as canonical pathways and upstream regulators related to environment-dependent signaling through steroid receptor pathways, among others. These findings uncover unrecognized regulatory similarities and differences between Hdac1 and Hdac2 in IEC, and demonstrate how HDAC1 and HDAC2 may complement each other to regulate the intrinsic IEC phenotype.
Similar content being viewed by others
Introduction
The small intestinal epithelium is composed of a single row of epithelial cells divided in proliferative crypt and differentiated villus compartments1. Crypt-located reserve intestinal stem cells sustain epithelial renewal by dividing in columnar stem cells generating transit-amplifying cells. These cells further segregate in absorptive enterocytes and secretory progenitor cells, precursors of Paneth, goblet and enteroendocrine cells2. Each differentiated cell lineage contributes to small intestinal functions, notably by establishing physical and chemical barriers between the host and the luminal diet and microbial content, and by providing a sensing and transmitting interface between the lumen and the mucosal immune system3. Indeed, in addition to absorptive and digestive functions, enterocytes, the most abundant intestinal epithelial cells (IEC), achieve selective barrier permeability through tight junction interactions between intestinal epithelial cells4,5,6. Enterocytes also participate in the chemical barrier by expressing transmembrane mucins as well as cytokines and antimicrobial proteins, in response to the microbial environment7. Goblet cells produce the mucus layer preventing bacterial adhesion to the epithelium as well as various antimicrobial proteins, and deliver luminal antigens to dendritic cells6,8. Crypt-located Paneth cells support the stem cell niche and produce different constitutive or inducible antimicrobial proteins to insure epithelial protection9,10. Many signaling pathways, including the Wnt and Notch pathways, regulate intestinal stem cell maintenance, renewal and differentiation11,12. Intestinal homeostasis is secured by interdependent communication signals between the intestinal mucosal system along with IEC, the luminal environment with diet-derived and microbial products, as well as the microbiota. However, alterations in the intestinal environment or the immune system, in conjunction with genetic susceptibilities, may lead to intestinal defects, including inappropriate inflammatory responses3.
Environmental changes are transmitted to the cell through epigenetic modifications of histones13. Acetylation is one epigenetic signal implicated as environmental sensor. Acetyltransferase readers add an acetyl group to histones on lysines, leading to alterations in DNA-histone interactions or to the production of acetyl marks recognized by bromodomain-containing regulators14. Histone acetylation efficiency is regulated in part by variations in mitochondrial and nucleo-cytosolic acetyl-CoA levels as a result of the cellular metabolic state15,16. Lysine acetylation is also controlled by histone deacetylase (HDAC) erasers that remove acetyl groups from histones and non-histone proteins. Endogenous HDAC activity is inhibited by metabolites including β-hydroxybutyrate17,18, L-carnitine19 and sphingosine-1-phosphate20, as well as diet- and bacterially-derived metabolites, such as butyrate21,22,23,24. Among HDACs, HDAC1 and HDAC2 are zinc-dependent class I deacetylases associated with Sin3A, CoREST and NuRD protein complexes regulating transcription, DNA replication and DNA repair, among others25,26,27. Hdac1 deletion in mice leads to embryonic lethality28 while Hdac2 deficiency results in perinatal lethality stemming from heart defects29. In contrast to no apparent defects or subtle phenotypic alterations in many Hdac1 or Hdac2 tissue-specific deletion models, dual Hdac1 and Hdac2 deficiency triggers extensive differentiation and proliferation alterations in most tissues25. Of note, gene-dosage experiments in mice have indicated different sensitivities to Hdac1 or Hdac2 expression levels. For example, murine brain is altered in mice with one allele of Hdac1 without Hdac2 in neural cells, as opposed to neural cells with one allele of Hdac2 without Hdac130. Similarly, differentiation is altered in mice with one allele of Hdac2 without Hdac1 in epidermal cells, as opposed to epidermal cells with one allele of Hdac1 without Hdac231.
In the intestine, IEC-specific Hdac1 and Hdac2 villin-Cre-induced deletion results in increased proliferation, goblet and Paneth cell loss, polarity disruption, activation of Notch, Stat3 and mTOR pathways, as well as increased susceptibility to DSS-induced colitis32,33. While Hdac2 IEC-specific deletion does not alter intestinal homeostasis, Hdac2 deficiency protects against DSS-induced colitis33. In addition, short-term deletion of both Hdac1 and Hdac2 in IEC with the Ah-Cre model leads to proliferation arrest34,35, accompanied by DNA damage responses35. Finally, in contrast to mice with one Hdac1 allele without Hdac2, villin-Cre mice with one allele of Hdac2 without Hdac1 display homeostatic defects similar to double villin-Cre Hdac1 and Hdac2 knockout phenotypic alterations35.
These results suggest that HDAC1 and HDAC2 display redundant and specific functions in intestinal epithelial cells that could alter IEC interactions with the environment, leading to modifications in intestinal homeostasis. In this study, to determine the intrinsic IEC-specific role of HDAC1 and HDAC2, we have established enteroid culture models deleted for Hdac1 or Hdac2 and assessed the IEC phenotype by transcriptomic and proteomic approaches. We show that Hdac1 or Hdac2 deficiency alters differentiation of secretory cells that sustain physical as well as chemical protection barriers, and increases the number of intermediate secretory cell precursors. We identify IEC Hdac1- and Hdac2-dependent common and specific biological processes, including oxidation-reduction and lipid-related metabolic processes, as well as canonical pathways and upstream regulators related to environment-dependent signaling through steroid receptor pathways, among others. These findings uncover unrecognized regulatory similarities and differences between Hdac1 and Hdac2 in IEC, and demonstrate how HDAC1 and HDAC2 may complement each other to regulate the intrinsic IEC phenotype.
Results
To determine the intrinsic effect of HDAC1 and HDAC2 on IEC, we generated enteroid cultures from isolated jejunal crypts of villin-Cre control, Hdac1 and Hdac2 knockout mice32. RT-PCR (Fig. 1A) and Western blot analysis (Fig. 1B), as well as immunofluorescence (Supplementary Fig. S1) confirmed Hdac1 or Hdac2 deletion at both the RNA and protein levels in the respective enteroid cultures. As opposed to other cell models with Hdac1 or Hdac2 deletions25, enteroid-specific Hdac1 or Hdac2 deletion did not lead to significantly increased protein levels of the remaining HDAC (Fig. 1B). As assessed by a colorimetric assay, we observed significantly reduced nuclear deacetylase activity in Hdac1-depleted enteroids (12% decrease, p = 0.0424), but no significant differences in HDAC activity in Hdac2-depleted enteroids (p = 0.0896), as opposed to a 80.2% HDAC activity reduction upon TSA treatment (Fig. 1C).
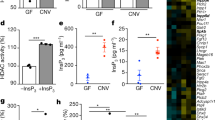
Hdac1 or Hdac2 depletion reduces enteroid histone deacetylase activity. (A) Total RNA was isolated from 5-day cultured control, Hdac1- and Hdac2-deficient enteroids. Hdac1 and Hdac2 mRNA levels were determined by semi-quantitative RT-PCR, with Gapdh as a loading control. (B) Total protein extracts from control, Hdac1- and Hdac2-deficient enteroids were separated on SDS-PAGE gels for Western blot analysis, and selected proteins were revealed with specific antibodies against HDAC1 and HDAC2, and against β-ACTIN as a loading control. Cropped images for HDAC1 and β-ACTIN are from immunoblotting experiments on the same membrane. Cropped images for HDAC2 and β-ACTIN are from immunoblotting experiments on another membrane. Full-length blots are presented in Supplementary Fig. S8 (n = 3). (C) 7.5 µg of nuclear proteins from control, Hdac1- and Hdac2-deficient enteroids were used to measure deacetylase activity with a colorimetric HDAC assay kit. Treatment with Trichostatin A, a pan-inhibitor of HDAC activity, was used as a control (n = 3; 2 or 3 wells for each). Results represent the mean ± SD (*p ≤ 0.05; ****p ≤ 0.0001).
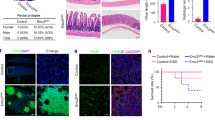
While there were no statistical differences in enteroid size, as assessed by surface area measurement (Fig. 2A; p > 0.1, one-way ANOVA), loss of Hdac1 or Hdac2 resulted in differences in enteroid structure, as determined by enhanced crypt budding per enteroid (Fig. 2B,C). Proliferation was not significantly altered, as assessed by BrdU pulse labeling (Supplementary Fig. S2). As HDAC1 and HDAC2 regulate DNA repair induced by double-strand breaks (DSB)36, we verified the expression of the DSB repair marker, γ-H2AX. A significant increase in the number of γ-H2AX stained nuclei in both Hdac1- or Hdac2-deficient enteroids was observed by immunofluorescence with a phospho-γ-H2AX antibody (Fig. 2D,E; Supplementary Fig. S3).
Hdac1 or Hdac2 depletion differently alters enteroid homeostasis. (A) Surface measurement of 5-day cultured control, Hdac1- and Hdac2-deficient enteroids (n = 3; 3–6 wells, and 50–70 enteroids for each). (B) Structural assessment of 5-day cultured control, Hdac1- and Hdac2-deficient enteroids by bud number scoring (n = 3; 3–6 wells for each): no bud (sphere), one bud, two buds, three or more buds. Results represent the mean ± SEM (*p ≤ 0.05; ****p ≤ 0.001 for more than 3 buds). (C) Representative micrographs of 5-day cultured control, Hdac1- and Hdac2-deficient enteroids. The scale bar indicates the relative size (bar = 200 μm). (D) Representative images of control, Hdac1- and Hdac2-deficient enteroids labeled with an antibody against phosphorylated γ-H2AX. Nuclei are stained with DAPI. γ-H2AX labeled cells are indicated by arrows. Magnification: 10X . For inserts, Magnification: 20X. (E) Number of γ-H2AX labeled nuclei in Hdac1- or Hdac2-depleted enteroids relative to control enteroids (n = 2). 15 independent fields (total surface: 7.8 mm2) (n = 2). Results represent the mean ± SD (*p ≤ 0.05; ***p ≤ 0.005).
To determine the intrinsic effect of Hdac1 or Hdac2 deletion on enteroid differentiation, H&E staining demonstrated an increase in the numbers of goblet cells associated with granules, in both enteroid mutants (Fig. 3A, see arrows and insert). Simultaneous staining with Alcian Blue to label goblet cells, and with Best’s Carmine to label Paneth cells, showed a significant increase in the number of goblet cells in both mutant enteroids, and in the number of Paneth cells in Hdac1-deleted enteroids (Fig. 3B,C). A significant augmentation in the number of intermediate cells displaying both goblet and Paneth cell labels was also observed (Fig. 3C, arrows and insert; Supplementary Fig. S4). Intermediate cells are considered precursors of both Paneth and goblet cell lineages in the small intestine37,38. The expression of differentiation markers was measured by qPCR. Different patterns of gene expression were observed in mutant organoids. While mRNA expression of stem cell marker Lgr5 was decreased in Hdac1-deficient enteroids (Fig. 4A), the expression of the enteroendocrine cell marker ChgA, as well as goblet cell markers Zg16 and Retnlb was increased in both Hdac1- or Hdac2-depleted enteroids (Fig. 4B,E). Augmented expression of enterocyte gene Sis, Paneth cell markers Lyz2 and Ang4, and goblet cell markers Muc2 and Muc3 was observed in Hdac2-deficient enteroids, in contrast to Hdac1-deleted enteroids. These data suggest that epithelial differentiation is altered after Hdac1 or Hdac2 deletion in enteroids, and that HDAC1 and HDAC2 are not equivalent in their regulatory function.
Hdac1 or Hdac2 depletion modifies enteroid differentiation. Control, Hdac1- and Hdac2-deficient enteroid sections were stained with Hematoxylin and Eosin (H&E) (A) or with a combination of Alcian blue and Best’s Carmine, respectively for goblet cells and Paneth cells (B). Magnification: 40X. For inserts, Magnification: 60X. Arrow: intermediate cell. (C) Number of goblet cells, Paneth cells and intermediate cells in Hdac1- or Hdac2-depleted enteroids relative to control enteroids (n = 3, 25–35 enteroids per experiment). Results represent the mean ± SEM (*p ≤ 0.05; **p ≤ 0.01).
Hdac1 or Hdac2 depletion alters enteroid differentiated gene expression levels. Total RNA was isolated from 5-day cultured control, Hdac1- and Hdac2-deficient enteroids. Expression levels of stem cell marker Lgr5 (A), enteroendocrine cell marker ChgA (B), enterocyte marker Si (C), Paneth cell markers Lys2 and Ang4 (D), and goblet cell markers Zg16, Retnlb, Muc2, Muc3, Tff3 (E) were determined by qPCR, with Pbgd as a control (n = 4–7). Results represent the mean ± SEM (*p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.005; ****p ≤ 0.001).
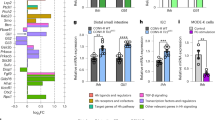
Having demonstrated specific and similar alterations between Hdac1- and Hdac2-deficient enteroids, we then determined global patterns of RNA expression by RNA-Seq, selecting log2 > 1 and log2 < 1 Hdac1 or Hdac2 specific genes with DESeq adjusted p-value ≤ 0.05. Based on this selection, RNA-Seq analysis revealed 1491 and 1119 genes respectively increased in Hdac1- and Hdac2-deficient enteroids, as well as 694 and 569 genes separately decreased (Fig. 5A, Supplementary Tables S5, S6). Increased or decreased expression of respectively 713 and 274 genes overlapped with both Hdac1- and Hdac2-deficient enteroids. FXR/RXR activation was a common top canonical pathway while lipopolysaccharide represented a common top upstream regulator, as determined by IPA analysis (Fig. 5B, Supplementary Tables S1, S2). Bioinformatics analysis revealed other specific top canonical pathways (LXR/RXR activation, Nicotine degradation II and III, LPS/IL-1 mediated inhibition of RXR function) in Hdac1-depleted enteroids. Additional specific predicted top upstream regulators were either inhibited and related to Ppara (ACOX1, ciprofibrate), or activated and related to Rara signaling (tretinoin) (Supplementary Table S1). Likewise, other top canonical pathways for Hdac2-depleted enteroids included retinoate biosynthesis I and retinol biosynthesis, with TNF as an activated top upstream regulator (Fig. 5B, Supplementary Table S2).
Transcript changes in Hdac1- or Hdac2-deficient enteroids, and top canonical pathways. (A) Venn diagram illustrating the specificity and the overlap of increased or decreased gene expression patterns between Hdac1-deficient and Hdac2-deficient enteroids (P-value < 0.05; log2 > 1 and log2 < 1), as assessed by RNA-Seq analysis. Top canonical pathways for gene expression identified by IPA analysis are shown for Hdac1-deficient enteroids (B-1) and Hdac2-deleted enteroids (B-2).
Biological processes identified by Gene Ontology analysis with DAVID identified common up-regulated processes, including inflammatory response, response to lipopolysaccharide, extracellular matrix organization, oxido-reduction process, lipid transport and retinol metabolic process (P-value between 3.80E-06 and 1.70E-03) (Supplementary Fig. S5). Common down-regulated processes included multicellular organism development and angiogenesis (P-value between 2.40E-08 and 1.20E-03). Specific increased biological processes included cholesterol homeostasis, superoxide and lipoprotein metabolic process for Hdac1-depleted enteroids (P-value between 8.10E-05 and 1.10E-03), while negative regulation of cell proliferation and negative regulation of cell migration were decreased (P-value of 9.8E-05 and 2.10E-03). Likewise, specific increased biological processes included response and defense response to virus for Hdac2-depleted enteroids (P-value between 1.90E-05 and 1.70E-04), while response to interferon-gamma, positive regulation of gene expression and of DNA-templated transcription were decreased (P-value between 6,10E-04 and 3.10E-03).
Changes in global protein expression between wild type and mutant enteroid cells were then quantified by quantitative mass spectrometry after SILAC labeling, selecting >1.5-fold and <1.5-fold specific proteins identified by at least two peptides, with an FDR 5%. Proteomic analysis revealed 141 and 78 proteins respectively increased in Hdac1- and Hdac2-deficient enteroids, as well as 132 and 90 proteins separately decreased in Hdac1- and Hdac2-deleted enteroids (Fig. 6A, Supplementary Table S7). Increased or decreased expression of respectively 52 and 58 proteins overlapped in both mutated enteroids.
Protein changes in Hdac1- or Hdac2-deficient enteroids, and top canonical pathways. (A) Venn diagram illustrating the specificity and the overlap of increased or decreased protein expression patterns between Hdac1-deficient and Hdac2-deficient enteroids (P-value < 0.05; log2 > 1 and log2 < 1), as assessed by SILAC proteomics analysis. Top canonical pathways for protein expression identified by IPA analysis are shown for Hdac1-deficient enteroids (B-1) and Hdac2-deleted enteroids (B-2).
LPS/IL-1 mediated inhibition of RXR function was a common top canonical pathway while CFTR represented a common top upstream regulator, as determined by IPA analysis (Fig. 6B). Further specific top canonical pathways were revealed in Hdac1- (Fatty acid oxidation I, Ethanol degradation) and Hdac2-depleted enteroids (Aryl hydrocarbon receptor signaling, Estrogen biosynthesis, Xenobiotic metabolism signaling, PXR/RXR activation) (Fig. 6B), in addition to predicted top upstream regulators associated with Hdac1- (PPARA, CFTR, PPARG, Essra, IL10RA) and Hdac2-deficient enteroids (CFTR, NFE2L2, IL10RA) (Fig. 6B, Supplementary Tables S3, S4). While not predicted as activated, top upstream regulators identified in Hdac2-deficient enteroid proteomes included steroid receptors such as NR1I3 and PXR ligand-PXR-retinoic acid-RXR.
Biological processes identified by Gene Ontology analysis with DAVID included common up-regulated Hdac1- and Hdac2-dependent processes, such as oxidation-reduction and lipid metabolic process (P-value between 2.40E-15 and 3.1E-03) (Supplementary Fig. S6). Common down-regulated processes included glutathione metabolic process, metabolic process, cell-cell adhesion and oxido-reduction process (P-value between 8.20E-05 and 3.10E-03). Specific increased biological processes included fatty acid, retinoid, carbohydrate, cholesterol, lipoprotein and steroid metabolic processes, as well as lipid and cholesterol transport for Hdac1-depleted enteroids (P-value between 2.90E-07 and 4.30E-04), while ATP-dependent chromatin remodeling, RNA splicing, mRNA processing and retinoic acid metabolic processes were decreased (P-value between 7.10E-04 and 4.90-E03). Likewise, specific increased biological processes included response to drug, nutrient, and hormone, among others, for Hdac2-depleted enteroids (P-value between 1.40-05 and 1.10E-03), while responses to different stimuli were decreased (P-value between 9.60E-04 and 3.70E-02). Oxido-reduction process was the GO term shared by the transcriptome and the proteome of both Hdac1- and Hdac2-depleted enteroids. Thus, HDAC1 and HDAC2 may both regulate, to different extent, metabolic and oxido-reduction processes, as well as cell responses to endogenous and exogenous environmental metabolites, in part through steroid receptor signaling.
Proteome and transcriptome analysis revealed common activation of STAT signaling in Hdac1- and Hdac2-deficient enteroids. Indeed, STAT1 was the top transcriptional regulator identified by RNA-Seq analysis while IL10RA was one of the top upstream regulators identified by proteome analysis. Since an increased Stat3 phosphorylation can be observed in dual Hdac1 and Hdac2 IEC-specific deleted mice32, Stat3 phosphorylation levels were verified in mutated enteroids. The data show an increase in Stat3 phosphorylated forms in Hdac1- as well as Hdac2-deleted enteroids (Fig. 7A).
Hdac1 or Hdac2 depletion alters the expression of specific metabolic and inflammatory related genes. Total protein extracts from control, Hdac1- or Hdac2-deficient enteroids were separated on SDS-PAGE gels for Western blot analysis, and selected proteins were revealed with specific antibodies against phosphorylated and total STAT3 (A), against HMGCS2 (C), and against β-ACTIN as a loading control (A,C) (n = 3). Densitometric analysis of Phospho-STAT3 compared to STAT3 is indicated (**p ≤ 0.01; ****p ≤ 0.001). Cropped images for Phospho-STAT3, STAT3 and β-ACTIN are from immunoblotting experiments on the same membrane. Cropped images for HMGCS2 and β-ACTIN are from immunoblotting experiments on another membrane. Samples for (A,C) derive from the same experiments. Full-length blots are presented in Supplementary Fig. S8. (A). Total RNAs were isolated from 5-day cultured control, Hdac1- or Hdac2-deficient enteroids. Expression levels of Hmgcs2 and Creb3l1 (B), Nox1 and Xdh (D), St3gal4, Bcl2l15 and Il18 (E) were determined by qPCR, with Pbgd as a control (n = 4–7). Results represent the mean ± SEM (*p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.005; ****p ≤ 0.001).
Based on common and specific biological processes revealed by RNA-Seq and proteomic analysis, we selected a subset of genes associated with metabolism, stress and environmental responses, for further verification of gene expression patterns by qPCR. Expression of Hmgcs2, a mitochondrial enzyme involved in the ketogenesis pathway and regulated in part by PPARA39, was decreased at both mRNA and protein levels (Fig. 7B,C). Expression of Creb3l1, an unfolded protein response regulator40, was decreased in Hdac1- or Hdac2-deficient organoids, as assessed by qPCR (Fig. 7B). Increased expression of Nox1, a NADPH oxidase41, Xdh, involved in purine oxidative metabolism42, Il18, a proinflammatory cytokine induced by inflammasome signaling43, and Bcl2l15, an apoptosis regulator expressed at high levels in the intestine44, was observed (Fig. 7D,E). Other genes encoding Nfkbiz, a regulator of NF-κB activity45, Nos2, a nitric oxide synthase46, and Apoa4, an apolipoprotein47, displayed increased expression in both Hdac1- and Hdac2-deficient enteroids while expression of Dusp4, a MAP kinase phosphatase48, was decreased (Supplementary Fig. S7). Expression of St3gal4, a glycosyltransferase49, was increased specifically in Hdac1-deficient enteroids (Fig. 7E). These results indicate common and specific patterns of gene regulation by Hdac1 or Hdac2 deficiency in enteroids, which affect cell-intrinsic metabolic as well as cell-extrinsic environmental responses.
Discussion
In order to determine the IEC intrinsic role of histone deacetylase HDAC1 and HDAC2, we have performed a global phenotypic, transcriptomic and proteomic analysis of Hdac1- and Hdac2-deficient enteroids. Our results show that Hdac1 or Hdac2 deletion in enteroids leads to increased production of secretory Paneth and goblet cells, suggesting a contribution of HDAC1 and HDAC2 in the regulation of Notch signaling. Indeed, active Notch signaling prevents secretory cell differentiation12,50. We have also observed increases in the number of intermediate cells, considered as precursors of both Paneth and goblet cell lineages in the small intestine37,51. Notch signaling is also involved in the production of intermediate cells. Indeed, intermediate as well as secretory cell numbers are increased in the small intestine of mice treated with the Notch pathway inhibitor DBZ38. Intestinal epithelial-specific deletion of the Adam10 sheddase gene in mice leads to increased numbers of Paneth/goblet cell intermediates as well as enteroendocrine cells through Notch signaling regulation52.
In vivo, we have previously observed that IEC-specific Hdac1- or Hdac2-deficient mice do not display intestinal architectural defects. However, in contrast to Hdac1+/−; Hdac2−/− IEC-deficient mice, IEC-deleted Hdac1−/−; Hdac2−/− or Hdac1−/−; Hdac2+/− are characterized by IEC loss of polarity and barrier function, resulting in a chronic inflammation, concomitant with increased crypt proliferation, decreased secretory cell numbers and Notch pathway activation32,35. These data suggest that different HDAC1 and HDAC2 levels may alter the balance between stem and progenitor cell proliferation, and differentiation, in part by regulating Notch pathway activity. In vivo, some additional contributions from the intestinal environment may differentially regulate intestinal homeostasis. Further studies to compare Hdac1+/−; Hdac2−/− or Hdac1−/−; Hdac2+/− enteroids to single Hdac1- or Hdac2-deleted organoids, and to assess their responses to mucosal environment factors, including cytokines, could be performed. Nevertheless, our data indicate that HDAC1 and HDAC2 may regulate intestinal epithelial cell fate, in part through Notch pathway regulation, as well as the relative production and maturation of differentiated cells from progenitors.
An increase in the number of phosphorylated γ-H2AX foci is observed after Hdac1 or Hdac2 deletion. HDAC1 and HDAC2 play important roles in DNA repair and the DNA damage response to double-strand breaks36,53, major inducers of γ-H2AX phosphorylation54. HDAC1 and HDAC2 are also recruited to replication forks55 to regulate DNA replication56. It is thus possible that Hdac1 or Hdac2 deletion renders enteroids more sensitive to DNA-strand breaks or replication stresses, leading to increased phosphorylated γ-H2AX foci. Recent data suggest that changes in chromatin structure per se may also result in γ-H2AX phosphorylation. For example, hypotonic treatment of different cell lines leads to increased γ-H2AX foci formation, in the relative absence of DNA strand-breaks57. Thus, Hdac1 or Hdac2 deletion could induce the formation of γ-H2AX foci by affecting chromatin structure, in part through histone acetylation.
Our data indicate that HDAC1 or HDAC2 homodimers, while sharing some targets, are not functionally equivalent to HDAC1/HDAC2 heterodimers in HDAC1/HDAC2 co-repressor complexes25. Hdac1 or Hdac2 deletion could lead to global effects on histone acetylation patterns, resulting in transcriptional changes. It has been proposed that HDAC associated with actively transcribed gene promoters could be important in resetting active promoters through deacetylation, for further rounds of transcription58,59. While increased histone acetylation correlates with active gene expression, it was shown that HDAC inhibition with HDAC pharmacological inhibitors, which results in increased histone acetylation in gene bodies and intragenic domains, may hinder transcriptional elongation and eRNA-associated enhancer transcription, suggestive of a positive role in transcriptional elongation60. Whether specific Hdac1 or Hdac2 deletion in enteroids could affect the expression of a subset of genes at the level of elongation or enhancer/promoter usage remains to be determined.
Our transcriptomic and proteomic analysis has uncovered various HDAC1- or HDAC2-dependent pathways acting as internal or external signaling sensors or transmitters. For example, HDAC1 and HDAC2 may play a role in the production of retinoic acid (RA), as suggested by increased Retinol metabolic processes in mutant enteroids. IEC-derived RA is considered as an important mucosal signal to mediate immune responses61. Of note, IEC-derived RA production is increased by the bacterial-derived short-chain fatty acid butyrate, a HDAC inhibitor, as well as by HDAC pharmacological inhibitors62,63.
IPA and GO term analysis of transcriptome and proteome data have revealed shared pathways regulated by Hdac1 or Hdac2. Hdac1 and Hdac2 deletion alters inflammatory signaling pathways. For example, STAT1 and RELA are in the top transcription factor upstream regulator, as well as IL10RA, suggesting the implication of HDAC1 and HDAC2 in respectively IFN signaling64, in cytokine-mediated NF-κB signaling65 and in STAT3 activation by cytokines66. HDAC may directly regulate the acetylation levels of these transcription factors, and their activity. For example, HDAC1 and HDAC2 negatively regulate NF-κB65. Class I HDAC, including HDAC1 and HDAC2, regulate STAT1 as well as STAT3 acetylation and transcriptional activity64. We have shown an increase in STAT3 phosphorylation in Hdac1- and Hdac2-deleted enteroids. This induced phosphorylation could be indirect, through autocrine production of cytokines, or direct as STAT3 acetylation may promote STAT3 phosphorylation, leading to increased transcriptional activity64,67. STAT3 is an important downstream effector of many cytokine-induced pathways which regulate IEC barrier function and mucosal wound healing68. For example, IL-22-dependent induction of STAT3 is essential for IEC barrier integrity and epithelial regeneration after injury, both in vivo and in organoid cultures69. Thus, HDAC1 and HDAC2 regulation of STAT3 activity may play an important role to insure intestinal homeostasis.
IPA and GO term analysis of transcriptome and proteome data have also revealed specific pathways regulated by Hdac1 or Hdac2. Hdac1 deletion leads to PPARα pathway inhibition, as a top upstream regulator, and to alterations in many metabolic pathways related to lipid and fatty acid metabolic processes. PPARα activation by fatty acids ligands, among others, regulates fatty acid metabolism, through peroxisomal and mitochondrial β-oxidation, inflammation and ketogenesis39,70. For example, expression of PPARα-regulated mitochondrial Hmgcs2, the rate-limiting enzyme involved in the production of ketone bodies39,71, including β-hydroxybutyrate, a known endogenous HDAC inhibitor17,18, is decreased in mutant enteroids. Thus, Hdac1 or Hdac2 enteroid deficiency may alter directly or indirectly the production of endogenous inhibitors, which could maintain HDAC activity.
Hdac2-deleted enteroids display AhR and PXR/CAR xenobiotic signaling as top canonical pathway and top upstream regulator, respectively. AhR is a transcription factor regulated by many endogenous and exogenous diet- and microbiota-derived ligands. AhR play important roles in the intestinal mucosal immune response and the regulation of the intestinal barrier function, as well as in antioxidant and xenobiotic responses72,73. AhR signaling is increased after HDAC inhibitor treatment of cultured murine and human IEC74. In addition to their role as xenobiotic sensors, recent data have uncovered roles for CAR and PXR in energy homeostasis75, as well as a role for PXR in intestinal epithelial repair responses76.
In addition to metabolic processes, oxido-reduction is one of the top processes affected in Hdac1- and Hdac2-deleted enteroids. This suggests that lack of HDAC1 or HDAC2 may affect enteroid metabolism. Acetyl-CoA levels vary according to the cellular metabolic state, with high levels associated with cell growth and survival, and global histone acetylation77. Acetyl-CoA is generated in various cellular compartments, including the nucleus, the cytoplasm and mitochondria. In the nucleus, the acetyl group linked to lysines on histones may supply acetate, in order to restore nuclear acetyl-CoA levels. Changes in HDAC activity could affect acetate availability, and thus influence enteroid homeostasis through metabolic pathway alterations, as proposed before78. Indeed, we have previously found that Hdac1 depletion in the intestinal epithelial cell line IEC-6 alters metabolic processes79. Whether metabolic process modifications observed in Hdac1- and Hdac2-deleted enteroids are indeed a compensatory response to maintain proper acetyl-CoA levels remains to be determined.
The extensive transcriptomic and proteomic changes observed upon Hdac1 or Hdac2 deletion in enteroids may be direct, through lysine acetylated mark alterations in regulatory regions, or indirect. This could occur either through acetylation-dependent alterations of the function of non-histone proteins, including transcription factors such as STAT380, or by metabolite changes that could result from decreased acetate availability. To identify more specific targets, chromatin immunoprecipitation experiments for specific acetylated marks, HDAC1 and HDAC2 could be performed with, for example, inducible villin-CreER Hdac1- or Hdac2-deleted enteroids. Mass spectrometry experiments could be carried out in order to assess the changes in the acetylome of Hdac1- or Hdac2-depleted enteroids.
We have identified here, homeostatic pathways dependent on HDAC1 and HDAC2 activity in enteroids. Our data suggest that HDAC1 and HDAC2 regulate differentiation, barrier function and stress responses in enteroids. Furthermore, HDAC1 and HDAC2 in part regulate common pathways related to oxydo-reduction and inflammatory processes in enteroids. Specific pathways include lipid metabolism for HDAC1 and response to environmental signals for HDAC2. Nuclear HDAC activity may well be important to control cellular acetyl-CoA levels necessary to insure enteroid homeostasis. Thus, IEC-intrinsic modifications of HDAC1 and/or HDAC2 activity may mediate the intestinal mucosal responses observed in IEC-specific Hdac1- and/or Hdac2-deficient mice32,33,35. As specific HDAC inhibitors are developed for use in cancer and in various immune and neurological diseases81,82, analysis of their impact on global and specific effects of HDAC1 and HDAC2 should be considered.
Methods
Enteroid culture
Hdac1 and Hdac2 floxed mice29 were crossed with villin-Cre transgenic mice83 in a C57BL/6 J X 129SV X CD1 background. All experiments were performed in accordance with relevant guidelines and regulations, and were approved by the Institutional Animal Research Review Committee of the Université de Sherbrooke (protocol 360-14B)33,35. Genomic DNA was isolated with the Spin Doctor genomic DNA kit (Gerard Biotech) to determine the genotypes33,35. Jejunal crypts from one wild-type, Hdac1 or Hdac2 mutant mouse were isolated by EDTA fractionation and cultured according to previous publications84,85, and as we have done before86. Matrigel (Growth Factor Reduced, BD Corning) embedded enteroids were grown in ENR medium containing 70% Advanced DMEM/F-12 Flex medium (Thermo Fisher Scientific), 1.25 mM N-acetylcysteine (Sigma), 50 ng/mL EGF (Life Technologies), B27 supplement 1X (Life Technologies), N2 supplement 1X (Life Technologies), 10% Advanced DMEM/F-12 Noggin conditioned medium and 20% Advanced DMEM/F-12 R-Spondin 1 conditioned medium. Penicillin (100 U/mL) and streptomycin (100 µg/mL) were also added to the medium. Enteroids were passaged every 5 days by mechanical disruption, and embedded 1:3 in 20 µL Matrigel (about 10 enteroids per µL).
Enteroid characterization
Wild-type, Hdac1- and Hdac2-deficient enteroid surface areas were measured after 5 days in culture (n = 3; 3 to 6 wells for each). Perimeters to measure enteroid areas were determined from horizontal cross-sections with the ZEISS ZEN Microscope Software. Budding efficiencies were measured under light microscopy by scoring for the number of buds per enteroid after 5 days: no budding (0), one bud (1), two buds (2) and three or more buds (3+). Imaging was performed with Cell Discoverer 7 microscope (magnification: 10X) (Zeiss, Toronto, ON, Canada).
HDAC activity measurement
Matrigel was dissolved in ice-cold Cell Recovery solution (Corning), and nuclear protein extracts were prepared from recovered wild-type, Hdac1- and Hdac2-deficient enteroids grown for 5 days, by using Abcam nuclear extraction kit (ab113474). 7.5 µg of nuclear extracts were used to measure nuclear HDAC activity with the colorimetric Epigenase HDAC activity/inhibition direct assay kit (Epigentek), on a Versamax ELISA microplate reader at 450 nm (Molecular Devices), according to the manufacturer’s protocol (n = 3; 2–3 wells for each). As a control, nuclear extracts were incubated with the HDAC pan-inhibitor Trichostatin A (1 μM). Results are expressed as the mean ± SD. Statistical significance was determined by Student’s t-test.
Histological analysis and immunofluorescence
Five-day wild-type, Hdac1- or Hdac2-deleted enteroid cultures were used. Enteroids recovered from Matrigel were separated by mechanical disruption, centrifuged for 3 min at 7000 rpm, and suspended in 4% paraformaldehyde for 1 h at 4oC. Fixed enteroids were centrifuged at 7000 rpm for 3 min, suspended in 70% ethanol for 1 h at 4oC and centrifuged at 7000 rpm for 3 min. Enteroids were mixed with Histogel (Thermo Fisher Scientific) before paraffin inclusion32,33. Sections were stained with hematoxylin and eosin for histological analysis, with Alcian blue for goblet cell mucins and with Best’s Carmine for Paneth cells. Goblet, Paneth and intermediate cell numbers in Hdac1- or Hdac2-depleted enteroids relative to control enteroids (n = 3, 25–35 enteroids per experiment) were counted in a blinded fashion, by two independent investigators. For immunofluorescence experiments, paraffin-embedded enteroid sections were rehydrated with graded ethanol series and boiled for 6 min in 10 mM citric acid. Treated sections were then blocked in PBS supplemented with 0.1% BSA and 0.2% Triton for 45 min86. Proliferation was assessed by fluorescein-conjugated mouse anti-bromodeoxyuridine (BrdU) (1:50, BMC 9318, Roche Diagnostics) staining of sections obtained from enteroid cultures incubated for 90 min with 10 µM BrdU. BrdU positive nuclei were counted with Cell Profiler 3.1587. Other primary antibodies include rabbit anti-HDAC1 (ab7028) (1:500, Abcam), rabbit anti-HDAC2 (ab7029) (1:500, Abcam), rabbit anti-phosphohistone γH2A.X (sc-101696) (1:500, Santa Cruz Biotechnology). γH2A.X positive stained nuclei were counted with Cell Profiler 3.15 (surface area for each experiment: 7.8 mm2, n = 2). Secondary donkey F(ab’)2 Anti-Rabbit IgG H&L (Alexa Fluor 568) preadsorbed antibodies (Abcam) were incubated at room temperature for 2 h (n = 3). Additional information about antibodies is included in Supplementary Table S8.
Protein isolation and Western blotting
Five-day wild-type, Hdac1- or Hdac2-deleted enteroid cultures were used, unless otherwise stated. Enteroids were recovered and directly lysed in 1 X Laemmli buffer 1 (62.5 mM Tris-HCl pH 6.8, 2% SDS, 10% glycerol) supplemented with protease and phosphatase inhibitors. Whole enteroid protein content was measured with the Pierce BCA Protein Assay Kit (Thermo Fisher Scientific). Fifteen µg of total protein extracts were added on 4–12% SDS-polyacrylamide gels, and transferred on PVDF membranes (Roche Molecular Biochemicals). Western blotting was performed as described previously79. Membranes were incubated 1 h at room temperature or overnight at 4 °C with the following primary antibodies: rabbit anti-HDAC1, rabbit anti-HDAC2 (Abcam); rabbit anti-HMGCS2, rabbit anti-phosphoSTAT3, rabbit anti-STAT3 (Cell Signaling); mouse anti-ACTIN (EMD Millipore). Secondary antibodies included goat anti-mouse and goat anti-rabbit (Invitrogen). Immune complexes were revealed with Amersham ECLTM Western blotting detection reagents (GE Healthcare) (n = 3). Additional information about antibodies is included in Supplementary Table S8.
RNA isolation, qPCR and semi-quantitative RT-PCR
Total RNAs from wild-type, Hdac1- or Hdac2-deleted enteroids were purified with the RNeasy mini kit (Qiagen), and quantified with a NanoDrop ND-1000 spectrophotometer (Thermo Fisher Scientific). cDNAs were synthetized from 1 µg of RNA with oligodT and Superscript II reverse transcriptase (Life Technologies). For semi-quantitative RT-PCR, cDNA amplification for Hdac1, Hdac2 and Gapdh with the Taq PCR Master Mix Kit (Qiagen, Mississauga, ON, Canada) was performed by a first 94 °C cycle for 5 min, followed by 29 cycles of 1 min at 94 °C, 45 sec starting at 62 °C and decreasing in increments of 0.3 °C every cycle, 1 min at 72 °C, and a final cycle of 1 min at 94 °C and 10 min at 72 °C (Supplementary Table S9 for PCR primer sequences). For qPCR analysis, 10 ng of cDNAs were amplified with the Brilliant III Ultra-fast SYBR Green qPCR Master Mix (Agilent Technologies) and specific gene upstream and downstream oligonucleotides (Supplementary Table S9), with a 10 min cycle at 95 °C, followed by 40 cycles of 10 sec at 95 °C, 10 sec at 60 °C and 20 sec at 72 °C. qPCR was performed with a Corbett RotorGene (Qiagen/Corbett Research). Relative RNA amounts were determined by using porphobilinogen deaminase (Pbgd) amplification (n = 4–7).
RNA-Seq analysis
Total RNAs from wild-type, Hdac1- or Hdac2-deleted enteroids were purified with the RNeasy mini kit (Qiagen) (n = 4), and quantified. RNA integrity was evaluated with a 2100 Bioanalyzer (Agilent Technologies). Only samples with RNA Integrity Number (RIN) > 6.5 were selected. cDNA library preparation and transcriptome analysis with the Illumina HiSeq. 4000 PE100 sequencing system (Illumina) were performed at the McGill University and Génome Québec Innovation Center. Sequence alignment was performed with the Star 2.4.0.1 software package based on genome reference Mus_musculus:GRCm38. Differentially expressed genes were identified with DESeq adjusted p-value ≤ 0.0588. Only transcripts increased or decreased more than 2-fold between wild-type, Hdac1- or Hdac2-deleted enteroids were selected for further bioinformatics analysis. RNA-Seq data have been deposited in the Gene Expression Omnibus database (GSE124146).
Proteome analysis
Wild-type, Hdac1- or Hdac2-deleted enteroids were grown for 6 passages in SILAC ENR medium supplemented with arginine and lysine, either the normal light isotopes of carbon, hydrogen and nitrogen (12C14N) (L), the medium L-arginine-13C614N4 and L-lysine-2H4 isotopes (M) or the heavy L-arginine- 13C6-15N4 and L-lysine-13C6-15N2 isotopes (H), as done before86. SILAC-labeled enteroids were recovered after six days in culture. Enteroid pellets were suspended in a 10 mM HEPES pH 7.0–7.6, 8 M urea solution, and proteins were processed for HPLC-MS/MS analysis (n = 2)86. Briefly, proteins were reduced in 3.24 mM dithiothreitol (DTT) before being alkylated in 13.5 mM iodoacetamide. After BCA protein quantitation, pools of 15 µg of light-, medium- or heavy-isotope labeled samples were digested by trypsin. Trypsin digested peptides were separated with an Ultimate U3000 nanoflow LC-system (Dionex Corporation). The HPLC system was coupled to an OrbiTrap Q Exactive, via an EasySpray source. Spray voltage was set to 1.5 kV, with a 40 °C column temperature. Full scan MS survey spectra (m/z 350–1800) in profile mode were acquired in the Orbitrap with a resolution of 70,000 after accumulation of 1,000,000 ions. The ten most intense peptide ions were fragmented by collision-induced dissociation (normalized collision energy 35% with a resolution of 17,500) after the accumulation of 50,000 ions. Maximal filling times were 250 ms for the full scans and 60 ms for the MS/MS scans. Data were acquired using the XCalibur software.
SILAC-labeled proteins were identified and quantified with MaxQuant version 1.5.2.8 software89 with the Uniprot mouse protein database containing 89,422 proteins, to which 175 commonly observed contaminants and all the reversed sequences had been added. To achieve reliable identification, proteins with the number of forward hits in the database at least 100-fold higher than the number of reverse database hits, were accepted, thus resulting in less than 5% false discovery rate (FDR). Quantification was performed with a minimum of 2 peptides for each protein, and only in the presence of 3 ratio counts. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD011869 (http://proteomecentral.proteomexchange.org).
Bioinformatics analysis
For both transcriptome and proteome, classification of genes according to Gene ontology was performed with the Database for Annotation, Visualization and Integrated Discovery software (DAVID 2.0)90 (p < 0.05), and with the Ingenuity Pathway Analysis software (IPA, Qiagen). These classifications identified altered biological processes and variations between samples, in order to predict pathway activation (URA, z-score)91.
Statistical analysis
Data were expressed as means ± SEM, or SD for HDAC activity measurement. Groups were compared by Student’s t-test (unpaired), one-way ANOVA with tukey multiple comparison test.
References
Clevers, H. The intestinal crypt, a prototype stem cell compartment. Cell 154, 274–284, https://doi.org/10.1016/j.cell.2013.07.004 (2013).
Yousefi, M., Li, L. & Lengner, C. J. Hierarchy and Plasticity in the Intestinal Stem Cell Compartment. Trends in cell biology 27, 753–764, https://doi.org/10.1016/j.tcb.2017.06.006 (2017).
Asselin, C. & Gendron, F. P. Shuttling of information between the mucosal and luminal environment drives intestinal homeostasis. FEBS letters 588, 4148–4157, https://doi.org/10.1016/j.febslet.2014.02.049 (2014).
Turner, J. R. Intestinal mucosal barrier function in health and disease. Nature reviews. Immunology 9, 799–809, https://doi.org/10.1038/nri2653 (2009).
Peterson, L. W. & Artis, D. Intestinal epithelial cells: regulators of barrier function and immune homeostasis. Nature reviews. Immunology 14, 141–153, https://doi.org/10.1038/nri3608 (2014).
Kurashima, Y. & Kiyono, H. Mucosal Ecological Network of Epithelium and Immune Cells for Gut Homeostasis and Tissue Healing. Annual review of immunology 35, 119–147, https://doi.org/10.1146/annurev-immunol-051116-052424 (2017).
Miron, N. & Cristea, V. Enterocytes: active cells in tolerance to food and microbial antigens in the gut. Clinical and experimental immunology 167, 405–412, https://doi.org/10.1111/j.1365-2249.2011.04523.x (2012).
Johansson, M. E., Sjovall, H. & Hansson, G. C. The gastrointestinal mucus system in health and disease. Nature reviews. Gastroenterology & hepatology 10, 352–361, https://doi.org/10.1038/nrgastro.2013.35 (2013).
Clevers, H. C. & Bevins, C. L. Paneth cells: maestros of the small intestinal crypts. Annual review of physiology 75, 289–311, https://doi.org/10.1146/annurev-physiol-030212-183744 (2013).
Gassler, N. Paneth cells in intestinal physiology and pathophysiology. World journal of gastrointestinal pathophysiology 8, 150–160, https://doi.org/10.4291/wjgp.v8.i4.150 (2017).
Clevers, H., Loh, K. M. & Nusse, R. Stem cell signaling. An integral program for tissue renewal and regeneration: Wnt signaling and stem cell control. Science 346, 1248012, https://doi.org/10.1126/science.1248012 (2014).
Sancho, R., Cremona, C. A. & Behrens, A. Stem cell and progenitor fate in the mammalian intestine: Notch and lateral inhibition in homeostasis and disease. EMBO reports 16, 571–581, https://doi.org/10.15252/embr.201540188 (2015).
Fan, J., Krautkramer, K. A., Feldman, J. L. & Denu, J. M. Metabolic regulation of histone post-translational modifications. ACS chemical biology 10, 95–108, https://doi.org/10.1021/cb500846u (2015).
Filippakopoulos, P. & Knapp, S. Targeting bromodomains: epigenetic readers of lysine acetylation. Nature reviews. Drug discovery 13, 337–356, https://doi.org/10.1038/nrd4286 (2014).
Pietrocola, F., Galluzzi, L., Bravo-San Pedro, J. M., Madeo, F. & Kroemer, G. Acetyl coenzyme A: a central metabolite and second messenger. Cell metabolism 21, 805–821, https://doi.org/10.1016/j.cmet.2015.05.014 (2015).
Shi, L. & Tu, B. P. Acetyl-CoA and the regulation of metabolism: mechanisms and consequences. Current opinion in cell biology 33, 125–131, https://doi.org/10.1016/j.ceb.2015.02.003 (2015).
Shimazu, T. et al. Suppression of oxidative stress by beta-hydroxybutyrate, an endogenous histone deacetylase inhibitor. Science 339, 211–214, https://doi.org/10.1126/science.1227166 (2013).
Newman, J. C. & Verdin, E. beta-Hydroxybutyrate: A Signaling Metabolite. Annual review of nutrition 37, 51–76, https://doi.org/10.1146/annurev-nutr-071816-064916 (2017).
Huang, H. et al. L-carnitine is an endogenous HDAC inhibitor selectively inhibiting cancer cell growth in vivo and in vitro. PloS one 7, e49062, https://doi.org/10.1371/journal.pone.0049062 (2012).
Hait, N. C. et al. Regulation of histone acetylation in the nucleus by sphingosine-1-phosphate. Science 325, 1254–1257, https://doi.org/10.1126/science.1176709 (2009).
Delcuve, G. P., Khan, D. H. & Davie, J. R. Roles of histone deacetylases in epigenetic regulation: emerging paradigms from studies with inhibitors. Clinical epigenetics 4, 5, https://doi.org/10.1186/1868-7083-4-5 (2012).
Krautkramer, K. A., Rey, F. E. & Denu, J. M. Chemical signaling between gut microbiota and host chromatin: What is your gut really saying? The Journal of biological chemistry 292, 8582–8593, https://doi.org/10.1074/jbc.R116.761577 (2017).
Bultman, S. J. Interplay between diet, gut microbiota, epigenetic events, and colorectal cancer. Molecular nutrition & food research 61, https://doi.org/10.1002/mnfr.201500902 (2017).
Ji, J. et al. Microbial metabolite butyrate facilitates M2 macrophage polarization and function. Scientific reports 6, 24838, https://doi.org/10.1038/srep24838 (2016).
Kelly, R. D. & Cowley, S. M. The physiological roles of histone deacetylase (HDAC) 1 and 2: complex co-stars with multiple leading parts. Biochemical Society transactions 41, 741–749, https://doi.org/10.1042/BST20130010 (2013).
Laugesen, A. & Helin, K. Chromatin repressive complexes in stem cells, development, and cancer. Cell stem cell 14, 735–751, https://doi.org/10.1016/j.stem.2014.05.006 (2014).
Moser, M. A., Hagelkruys, A. & Seiser, C. Transcription and beyond: the role of mammalian class I lysine deacetylases. Chromosoma 123, 67–78, https://doi.org/10.1007/s00412-013-0441-x (2014).
Lagger, G. et al. Essential function of histone deacetylase 1 in proliferation control and CDK inhibitor repression. The EMBO journal 21, 2672–2681, https://doi.org/10.1093/emboj/21.11.2672 (2002).
Montgomery, R. L. et al. Histone deacetylases 1 and 2 redundantly regulate cardiac morphogenesis, growth, and contractility. Genes & development 21, 1790–1802, https://doi.org/10.1101/gad.1563807 (2007).
Hagelkruys, A. et al. A single allele of Hdac2 but not Hdac1 is sufficient for normal mouse brain development in the absence of its paralog. Development 141, 604–616, https://doi.org/10.1242/dev.100487 (2014).
Winter, M. et al. Divergent roles of HDAC1 and HDAC2 in the regulation of epidermal development and tumorigenesis. The EMBO journal 32, 3176–3191, https://doi.org/10.1038/emboj.2013.243 (2013).
Turgeon, N. et al. HDAC1 and HDAC2 restrain the intestinal inflammatory response by regulating intestinal epithelial cell differentiation. PloS one 8, e73785, https://doi.org/10.1371/journal.pone.0073785 (2013).
Turgeon, N. et al. The acetylome regulators Hdac1 and Hdac2 differently modulate intestinal epithelial cell dependent homeostatic responses in experimental colitis. American journal of physiology. Gastrointestinal and liver physiology 306, G594–605, https://doi.org/10.1152/ajpgi.00393.2013 (2014).
Zimberlin, C. D. et al. HDAC1 and HDAC2 collectively regulate intestinal stem cell homeostasis. FASEB journal: official publication of the Federation of American Societies for Experimental Biology 29, 2070–2080, https://doi.org/10.1096/fj.14-257931 (2015).
Gonneaud, A. et al. Distinct Roles for Intestinal Epithelial Cell-Specific Hdac1 and Hdac2 in the Regulation of Murine Intestinal Homeostasis. Journal of cellular physiology 231, 436–448, https://doi.org/10.1002/jcp.25090 (2016).
Roos, W. P. & Krumm, A. The multifaceted influence of histone deacetylases on DNA damage signalling and DNA repair. Nucleic acids research 44, 10017–10030, https://doi.org/10.1093/nar/gkw922 (2016).
Calvert, R., Bordeleau, G., Grondin, G., Vezina, A. & Ferrari, J. On the presence of intermediate cells in the small intestine. The Anatomical record 220, 291–295, https://doi.org/10.1002/ar.1092200310 (1988).
VanDussen, K. L. et al. Notch signaling modulates proliferation and differentiation of intestinal crypt base columnar stem cells. Development 139, 488–497, https://doi.org/10.1242/dev.070763 (2012).
Grabacka, M., Pierzchalska, M., Dean, M. & Reiss, K. Regulation of Ketone Body Metabolism and the Role of PPARalpha. International journal of molecular sciences 17, https://doi.org/10.3390/ijms17122093 (2016).
Kondo, S., Saito, A., Asada, R., Kanemoto, S. & Imaizumi, K. Physiological unfolded protein response regulated by OASIS family members, transmembrane bZIP transcription factors. IUBMB life 63, 233–239, https://doi.org/10.1002/iub.433 (2011).
Perez, S., Talens-Visconti, R., Rius-Perez, S., Finamor, I. & Sastre, J. Redox signaling in the gastrointestinal tract. Free radical biology & medicine 104, 75–103, https://doi.org/10.1016/j.freeradbiomed.2016.12.048 (2017).
Battelli, M. G., Polito, L., Bortolotti, M. & Bolognesi, A. Xanthine Oxidoreductase-Derived Reactive Species: Physiological and Pathological Effects. Oxidative medicine and cellular longevity 2016, 3527579, https://doi.org/10.1155/2016/3527579 (2016).
Ranson, N., Kunde, D. & Eri, R. Regulation and Sensing of Inflammasomes and Their Impact on Intestinal Health. International journal of molecular sciences 18, https://doi.org/10.3390/ijms18112379 (2017).
Coultas, L. et al. Bfk: a novel weakly proapoptotic member of the Bcl-2 protein family with a BH3 and a BH2 region. Cell death and differentiation 10, 185–192, https://doi.org/10.1038/sj.cdd.4401204 (2003).
Willems, M., Dubois, N., Musumeci, L., Bours, V. & Robe, P. A. IkappaBzeta: an emerging player in cancer. Oncotarget 7, 66310–66322, https://doi.org/10.18632/oncotarget.11624 (2016).
Bogdan, C. Nitric oxide synthase in innate and adaptive immunity: an update. Trends in immunology 36, 161–178, https://doi.org/10.1016/j.it.2015.01.003 (2015).
Wang, F. et al. Apolipoprotein A-IV: a protein intimately involved in metabolism. Journal of lipid research 56, 1403–1418, https://doi.org/10.1194/jlr.R052753 (2015).
Low, H. B. & Zhang, Y. Regulatory Roles of MAPK Phosphatases in Cancer. Immune network 16, 85–98, https://doi.org/10.4110/in.2016.16.2.85 (2016).
Li, Y. & Chen, X. Sialic acid metabolism and sialyltransferases: natural functions and applications. Applied microbiology and biotechnology 94, 887–905, https://doi.org/10.1007/s00253-012-4040-1 (2012).
Gehart, H. & Clevers, H. Tales from the crypt: new insights into intestinal stem cells. Nature reviews. Gastroenterology & hepatology, https://doi.org/10.1038/s41575-018-0081-y (2018).
Kamal, M. et al. Mucosal T cells regulate Paneth and intermediate cell numbers in the small intestine of T. spiralis-infected mice. Clinical and experimental immunology 126, 117–125 (2001).
Tsai, Y. H. et al. ADAM10 regulates Notch function in intestinal stem cells of mice. Gastroenterology 147, 822–834 e813, https://doi.org/10.1053/j.gastro.2014.07.003 (2014).
Miller, K. M. et al. Human HDAC1 and HDAC2 function in the DNA-damage response to promote DNA nonhomologous end-joining. Nature structural & molecular biology 17, 1144–1151, https://doi.org/10.1038/nsmb.1899 (2010).
Turinetto, V. & Giachino, C. Multiple facets of histone variant H2AX: a DNA double-strand-break marker with several biological functions. Nucleic acids research 43, 2489–2498, https://doi.org/10.1093/nar/gkv061 (2015).
Sirbu, B. M. et al. Analysis of protein dynamics at active, stalled, and collapsed replication forks. Genes & development 25, 1320–1327, https://doi.org/10.1101/gad.2053211 (2011).
Stengel, K. R. & Hiebert, S. W. Class I HDACs Affect DNA Replication, Repair, and Chromatin Structure: Implications for Cancer Therapy. Antioxidants & redox signaling 23, 51–65, https://doi.org/10.1089/ars.2014.5915 (2015).
Baure, J. et al. Histone H2AX phosphorylation in response to changes in chromatin structure induced by altered osmolarity. Mutagenesis 24, 161–167, https://doi.org/10.1093/mutage/gen064 (2009).
Wang, Z. et al. Genome-wide mapping of HATs and HDACs reveals distinct functions in active and inactive genes. Cell 138, 1019–1031, https://doi.org/10.1016/j.cell.2009.06.049 (2009).
Dovey, O. M., Foster, C. T. & Cowley, S. M. Emphasizing the positive: A role for histone deacetylases in transcriptional activation. Cell cycle 9, 2700–2701, https://doi.org/10.4161/cc.9.14.12626 (2010).
Greer, C. B. et al. Histone Deacetylases Positively Regulate Transcription through the Elongation Machinery. Cell reports 13, 1444–1455, https://doi.org/10.1016/j.celrep.2015.10.013 (2015).
Erkelens, M. N. & Mebius, R. E. Retinoic Acid and Immune Homeostasis: A Balancing Act. Trends in immunology 38, 168–180, https://doi.org/10.1016/j.it.2016.12.006 (2017).
Schilderink, R. et al. The SCFA butyrate stimulates the epithelial production of retinoic acid via inhibition of epithelial HDAC. American journal of physiology. Gastrointestinal and liver physiology 310, G1138–1146, https://doi.org/10.1152/ajpgi.00411.2015 (2016).
Goverse, G. et al. Diet-Derived Short Chain Fatty Acids Stimulate Intestinal Epithelial Cells To Induce Mucosal Tolerogenic Dendritic Cells. Journal of immunology 198, 2172–2181, https://doi.org/10.4049/jimmunol.1600165 (2017).
Zhuang, S. Regulation of STAT signaling by acetylation. Cellular signalling 25, 1924–1931, https://doi.org/10.1016/j.cellsig.2013.05.007 (2013).
Shakespear, M. R., Halili, M. A., Irvine, K. M., Fairlie, D. P. & Sweet, M. J. Histone deacetylases as regulators of inflammation and immunity. Trends in immunology 32, 335–343, https://doi.org/10.1016/j.it.2011.04.001 (2011).
Nguyen, P. M., Putoczki, T. L. & Ernst, M. STAT3-Activating Cytokines: A Therapeutic Opportunity for Inflammatory Bowel Disease? Journal of interferon & cytokine research: the official journal of the International Society for Interferon and Cytokine Research 35, 340–350, https://doi.org/10.1089/jir.2014.0225 (2015).
Avalle, L. & Poli, V. Nucleus, Mitochondrion, or Reticulum? STAT3 a La Carte. International journal of molecular sciences 19, https://doi.org/10.3390/ijms19092820 (2018).
Buzzelli, J. N. et al. Overexpression of IL-11 promotes premalignant gastric epithelial hyperplasia in isolation from germline gp130-JAK-STAT driver mutations. American journal of physiology. Gastrointestinal and liver physiology 316, G251–G262, https://doi.org/10.1152/ajpgi.00304.2018 (2019).
Lindemans, C. A. et al. Interleukin-22 promotes intestinal-stem-cell-mediated epithelial regeneration. Nature 528, 560–564, https://doi.org/10.1038/nature16460 (2015).
Pawlak, M., Lefebvre, P. & Staels, B. Molecular mechanism of PPARalpha action and its impact on lipid metabolism, inflammation and fibrosis in non-alcoholic fatty liver disease. Journal of hepatology 62, 720–733, https://doi.org/10.1016/j.jhep.2014.10.039 (2015).
Puchalska, P. & Crawford, P. A. Multi-dimensional Roles of Ketone Bodies in Fuel Metabolism, Signaling, and Therapeutics. Cell metabolism 25, 262–284, https://doi.org/10.1016/j.cmet.2016.12.022 (2017).
Roager, H. M. & Licht, T. R. Microbial tryptophan catabolites in health and disease. Nature communications 9, 3294, https://doi.org/10.1038/s41467-018-05470-4 (2018).
Lamas, B., Natividad, J. M. & Sokol, H. Aryl hydrocarbon receptor and intestinal immunity. Mucosal immunology 11, 1024–1038, https://doi.org/10.1038/s41385-018-0019-2 (2018).
Jin, U. H. et al. Short Chain Fatty Acids Enhance Aryl Hydrocarbon (Ah) Responsiveness in Mouse Colonocytes and Caco-2 Human Colon Cancer Cells. Scientific reports 7, 10163, https://doi.org/10.1038/s41598-017-10824-x (2017).
Mackowiak, B., Hodge, J., Stern, S. & Wang, H. The Roles of Xenobiotic Receptors: Beyond Chemical Disposition. Drug metabolism and disposition: the biological fate of chemicals 46, 1361–1371, https://doi.org/10.1124/dmd.118.081042 (2018).
Terc, J., Hansen, A., Alston, L. & Hirota, S. A. Pregnane X receptor agonists enhance intestinal epithelial wound healing and repair of the intestinal barrier following the induction of experimental colitis. European journal of pharmaceutical sciences: official journal of the European Federation for Pharmaceutical Sciences 55, 12–19, https://doi.org/10.1016/j.ejps.2014.01.007 (2014).
Ye, C. & Tu, B. P. Sink into the Epigenome: Histones as Repositories That Influence Cellular Metabolism. Trends in endocrinology and metabolism: TEM 29, 626–637, https://doi.org/10.1016/j.tem.2018.06.002 (2018).
Martinez-Pastor, B., Cosentino, C. & Mostoslavsky, R. A tale of metabolites: the cross-talk between chromatin and energy metabolism. Cancer discovery 3, 497–501, https://doi.org/10.1158/2159-8290.CD-13-0059 (2013).
Gonneaud, A., Turgeon, N., Boisvert, F. M., Boudreau, F. & Asselin, C. Loss of histone deacetylase Hdac1 disrupts metabolic processes in intestinal epithelial cells. FEBS letters 589, 2776–2783, https://doi.org/10.1016/j.febslet.2015.08.009 (2015).
Narita, T., Weinert, B. T. & Choudhary, C. Functions and mechanisms of non-histone protein acetylation. Nature reviews. Molecular cell biology 20, 156–174, https://doi.org/10.1038/s41580-018-0081-3 (2019).
Falkenberg, K. J. & Johnstone, R. W. Histone deacetylases and their inhibitors in cancer, neurological diseases and immune disorders. Nature reviews. Drug discovery 13, 673–691, https://doi.org/10.1038/nrd4360 (2014).
Felice, C., Lewis, A., Armuzzi, A., Lindsay, J. O. & Silver, A. Review article: selective histone deacetylase isoforms as potential therapeutic targets in inflammatory bowel diseases. Alimentary pharmacology & therapeutics 41, 26–38, https://doi.org/10.1111/apt.13008 (2015).
Madison, B. B. et al. Cis elements of the villin gene control expression in restricted domains of the vertical (crypt) and horizontal (duodenum, cecum) axes of the intestine. The Journal of biological chemistry 277, 33275–33283, https://doi.org/10.1074/jbc.M204935200 (2002).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265, https://doi.org/10.1038/nature07935 (2009).
Sato, T. & Clevers, H. Growing self-organizing mini-guts from a single intestinal stem cell: mechanism and applications. Science 340, 1190–1194, https://doi.org/10.1126/science.1234852 (2013).
Gonneaud, A. et al. A SILAC-Based Method for Quantitative Proteomic Analysis of Intestinal Organoids. Scientific reports 6, 38195, https://doi.org/10.1038/srep38195 (2016).
Carpenter, A. E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome biology 7, R100, https://doi.org/10.1186/gb-2006-7-10-r100 (2006).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome biology 11, R106, https://doi.org/10.1186/gb-2010-11-10-r106 (2010).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nature biotechnology 26, 1367–1372, https://doi.org/10.1038/nbt.1511 (2008).
Dennis, G. Jr. et al. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome biology 4, P3 (2003).
Kramer, A., Green, J., Pollard, J. Jr. & Tugendreich, S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 30, 523–530, https://doi.org/10.1093/bioinformatics/btt703 (2014).
Acknowledgements
This work was supported by a grant from Crohn and Colitis Canada to Claude Asselin. Claude Asselin, François Boudreau and François-Michel Boisvert are members of the Fonds de recherche du Québec-Santé-funded CR-CHUS. We thank Dr E.N. Olson for providing Hdac1 and Hdac2 conditional knockout mice. The authors thank Dr C. Kuo (Stanford University, USA) for providing the R-spondin 1 producing cell line and Dr G.R. van den Brink (Hubrecht Institute, The Netherlands) for providing the Noggin producing cell line.
Author information
Authors and Affiliations
Contributions
A.G. performed all the experiments. N.T. was involved in the qPCR experiments. C.A. and A.G. wrote the main manuscript text. C.J. and C.C. contributed to enteroid cultures. A.G. prepared Figures 1–7 with N.T., as well as Supplementary Tables and Figures, except Supplementary Figure S6 (N.T.). F.M.B. and D.L. provided technical assistance for MS experiments. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gonneaud, A., Turgeon, N., Jones, C. et al. HDAC1 and HDAC2 independently regulate common and specific intrinsic responses in murine enteroids. Sci Rep 9, 5363 (2019). https://doi.org/10.1038/s41598-019-41842-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-41842-6
This article is cited by
-
Hypoxia-induced LncRNA DACT3-AS1 upregulates PKM2 to promote metastasis in hepatocellular carcinoma through the HDAC2/FOXA3 pathway
Experimental & Molecular Medicine (2022)
-
Matrix metalloproteinases, purinergic signaling, and epigenetics: hubs in the spinal neuroglial network following peripheral nerve injury
Histochemistry and Cell Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.