Abstract
Voltage-gated sodium channels (VGSC) are transmembrane proteins that generate an action potential in excitable cells and play an essential role in neuronal signaling. Since VGSCs play a crucial role in nerve transmission they have become primary targets for a broad range of commercial insecticides. RNA interference (RNAi) is a valuable reverse genetics tool used in functional genomics, but recently, it has also shown promise as a novel agent that could be used to control agricultural insect pests. In this study, we targeted the VGSC (MpNav) gene in the peach-potato aphid Myzus persicae, by oral feeding of artificial diets mixed with dsRNAs. Knock-down of MpNav gene expression caused up to 65% mortality in 3rd instar nymphs. Moreover, significantly lower fecundity and longevity was observed in adult aphids that had been fed with dsMpNav solution at the nymphal stage. Analysis of gene expression by qRT-PCR indicated that the aphid mortality rates and the lowered fecundity and longevity were attributable to the down-regulation of MpNav by RNAi. Taken together, our results show that MpNav is a viable candidate target gene for the development of an RNAi-based bio-aphicide.
Similar content being viewed by others
Introduction
The peach-potato aphid Myzus persicae (Hemiptera: Aphididae) is a major sap-sucking insect pest that infest more than 400 species of plants belonging to 40 different families including Brassicaceae and Solanaceae. Owing to direct plant feeding and the transmission of more than 100 plant viruses, it is considered one of the most destructive agricultural pests worldwide1,2. Its broad host range, telescoping of generations and cyclical parthenogenesis make it a highly successful insect pest3. Synthetic chemical insecticides are considered the most effective way to combat M. persicae. However, it has developed resistance against several different insecticide classes4, resulting in significant control failures and losses in protected crops.
An appropriate functioning of the voltage-gated sodium channel (VGSC) is essential for the normal transmission of the nerve impulse in insects, including aphids, and disruption of the action potential by insecticides leads to paralysis and eventually death of the insect5. The insecticides that target the VGSC have broad spectrum effects, as the structure of the VGSC is highly conserved across the animal kingdom, so there is a consequential detrimental impact on non-target species including pollinators and other beneficial’s. There is thus a real need to develop alternative (species specific) control methods with lower environmental impacts6.
RNAi is a genetically conserved post-transcriptional gene silencing mechanism that facilitates down regulation of gene expression by small noncoding RNA molecules in almost all eukaryotes7,8. RNAi has over the past decade been developed as a molecular tool to knock-down target gene transcripts in insects9. Due to its high degree of specificity, RNAi is also being widely explored as a potential novel pest control strategy for a variety of pest species10, with the perceived benefits of increased species discrimination and decreased risk to the environment and non-target species11.
Unusually in M. persicae (and other aphids), the functioning VGSC is encoded by two genes (NCBI Accessions FN601405 and FN601406)12 with some unique properties that are not present in the VGSCs of other insects. Unlike the channels of other insects, the aphid has a unique heterodimeric channel (composed of two subunits, H1 and H2, encoding DI-II and DIII-IV of the VGSC respectively) with an atypical ion selectivity filter (similar that found in the mammalian sodium sensor channel Nax)13, which, atypically for insect VGSCs, is extremely insensitive to tetrodotoxin.
Abd El Halim et al.14 recently reported that RNAi-mediated knock-down of VGSC gene expression, through application of complementary dsRNA, caused significantly high larval mortalities and severe developmental arrest in the red flour beetle Tribolium castaneum, a coleopteran insect pest that has proved to be particularly amenable to RNAi15,16. In comparison levels of gene-knock down and systemic RNAi responses (following injection or ingestion of dsRNA) in many other insect classes are extremely variable17. In particular, RNAi outcomes in phloem sap feeding hemipteran species such as aphids, whitefly, psyllids and plant hoppers can be exceedingly disparate, ranging from no phenotype to significant mortality and from very low to complete gene knock-down18,19,20. Hemipteran species are therefore more challenging to work with as they appear to be somewhat intractable to RNAi manipulations21. In this study, we demonstrate that oral delivery of MpNav dsRNA to the peach-potato aphid M. persicae successfully down-regulates the expression of the aphids VGSC and causes significant nymphal mortalities. Moreover, lower fecundity and longevity was also observed in dsRNA treated insects. These results suggest that RNAi targeting the VGSC could be a promising novel bio-pesticide against this hemipteran pest.
Materials and Methods
Insect culture
Myzus persicae were collected from a cabbage field in Mardan, Khyber Pakhtunkhwa, Pakistan, and established as a laboratory colony that was maintained on cabbage plants (variety Golden Acre) under standard laboratory conditions (25 ± 2 °C, 70 ± 10% relative humidity, 12:12 (light: dark) photoperiod).
Total RNA extraction and cDNA synthesis
Total RNAs were isolated from 1st, 2nd, 3rd, 4th instar nymphs and adults of M. persicae using Wizol™ Reagent (Wizbiosolutions Inc., Korea) following the manufacturer’s recommended protocol. RNA integrity was analyzed on 1.5% (w/v) agarose gels as described in Sambrook and Russell22. cDNA’s were synthesized using 1 μg total RNA with WizScript™ First Strand cDNA synthesis kit (Wizbiosolutions Inc. Korea), according to the manufacturer’s recommended protocol.
Synthesis of dsRNA molecules
For RNAi experiments, a 289 bp fragment of the M. persicae MpNav (heteromer H1) gene (NCBI Accession FN601405) and a 329 bp fragment of the Aequorea victoria green-fluorescent protein (GFP) gene (NCBI Accession M62653) were amplified from cDNA obtained from adult aphids using PCR. The T7 promoter sequence 5′GGATCCTAATACGACTCACTATAGGA3′ was added in front of the forward and reverse PCR primers as required for subsequent dsRNA synthesis (Table 1). Primers were chosen based on the results of GeneScripts primer design software (https://www.genscript.com/tools/pcr-primers-designer). The reaction mixture for PCR contained 500 ng of the cDNA template, 0.5 µm of forward and reverse primers, 1.75 mM Mg2+, 0.25 mM of each dNTP (Takara Bio, Japan), 1X PCR buffer, 5U/µl Taq DNA polymerase (Takara Bio) and double-distilled H2O to make a 25 µl total reaction volume. The PCR conditions were 95 °C for 5 min, followed by 35 cycles of 95 °C for 30 s, 55–60 °C for 30 s and 72 °C for 30 s, and an additional final polymerization step of 72 °C for 5 min. PCR products were purified using TIANgel Midi Purification Kit (Tiangen, Beijing, China). These purified products were then used to synthesize dsRNA using the T7 RiboMAX™ Express RNAi System (Promega, US), following the manufacturer’s protocol. The dsRNA quality was monitored on agarose gel electrophoresis, and the concentration was determined by spectrophotometry (Nano Drop 1000, Thermo Scientific, US). dsRNA products were stored at −80 °C prior to further use.
Dietary delivery of the double-stranded RNA to nymphs
The artificial feeding diet for M. persicae was prepared as described by Pan et al.23, albeit with a 20% lower sucrose content24. In the RNAi diet, dsMpNav or dsGFP RNA, or DEPC water was mixed in and fed to 3rd instar nymphs. To rear aphids on this artificial diet, 6 well culture plates were used with small ventilation holes incorporated in the bottom of the plates. Using a fine paintbrush, one hundred and fifty 3rd instar nymphs were carefully collected from cabbage leaves and transferred to each well of the plate, and the plate sealed with Parafilm®M. 0.75 mg/ml25 dsMpNav, or dsGFP RNA, or DEPC water was incorporated into the artificial diet as required, and the mixture loaded on the parafilm membrane above the wells; feeding sachets were created with 2 cm2 pieces of parafilm, cleaned with ethanol and DEPC water. Employing this strategy, the aphids could puncture the inner layer of parafilm membrane and feed on the mixture of diet sandwiched between the two layers of parafilm. The plates were kept under laboratory conditions (25 ± 2 °C, 70 ± 10% RH, and 12:12 (light: dark) photoperiod) and the diet mixtures renewed every day for 7 days. Mortality of the aphids was recorded daily. All bioassay treatments were repeated in triplicate.
MpNa v expression analysis by Quantitative Real-Time PCR
Quantitative reverse transcription PCRs (RT-qPCR) was performed to analyze the expression level of MpNav (heteromer H1). Primers for the MpNav H1, GFP and Actin genes were designed online using Primer-BLAST26 (https://www.ncbi.nlm.nih.gov/tools/primer-blast/). qRT-PCR reactions were performed using iTaq™ Universal SYBR Green Supermix (Bio-Rad) in a Bio-Rad iCycler (Bio-Rad, Hercules, CA, USA), following the manufacturer’s instructions. PCR conditions were 95 °C for 10 min, 40 cycles of 95 °C for 15 s, 60 °C for 30 s and 72 °C for 30 s. All the analyses were repeated in triplicate. Relative expression levels were calculated using 2−△△Ct method27. Actin was used as the internal control28. All the primers used in this study (Table 1) were designed to avoid the homologous ORF region used for the synthesis of dsRNA.
Longevity and fecundity analysis
After continuous feeding of dsRNA for 7 days, each survivor aphid was transferred to a cabbage leaf placed on a freshly prepared agar plate. Plates were kept under laboratory condition as described above. The number of new born aphid nymphs from each plate was recorded daily until the aphid died. The nymphs produced by the aphids were removed from the dishes after counting. The cabbage leaf was replaced every alternate day with a fresh leaf.
Statistical analysis
The data were statistically analyzed with GraphPad prism 5.0. The results were analyzed using One-way analysis of variance (ANOVA) with subsequent Tukey Kramer multiple comparison. For all tests, P < 0.05 was considered significant.
Results
Temporal expression of MpNa v gene
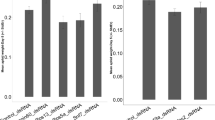
The relative abundance of MpNav H1 mRNA at specific developmental stages of M. persicae was estimated by qRT-PCR. Transcripts were detected in all life stages investigated. However, MpNav H1 was most abundantly expressed (P < 0.005) in 3rd and 4th instar nymphs and adults (Fig. 1). There was no significant difference in the level of MpNav gene expression in the 3rd instar, 4th instar, and adult aphids. Based on this, 3rd instar nymphs were selected as a suitable developmental stage for RNAi studies.
Stage-specific expression pattern of the M. persicae gene MpNav H1 in the whole insect body analyzed by qRT-PCR. The letters above the bars show significant differences (least significant difference in one-way analysis of variance, P < 0.05) in MpNav expression. Means with the same letter are not significantly different. Mean ± SEM of three independent experiments are shown. RNA was normalized with Actin as the internal control.
Bioinformatic analysis of targeted sequence
Homology of the selected MpNav dsRNA fragment to other insect Nav sequences, including representatives from important pollinator and beneficial species, was investigated using the NCBI BLASTn algorithm. The BLASTn search indicated a high degree of similarity between the MpNav dsRNA fragment and the Nav coding sequence in other aphid species (overall homology scores ranging between 95% identity, e-value 5e-125 to the pea aphid Acyrthosiphon pisum to 84% identity, e-value 3e-83 to the sugar-cane aphid Sipha flava). For important pollinating and beneficial insects, Blastn scores ranged from 74% identity, e-value 2e-16 identity for the common eastern bumble bee, Bombus impatiens, 73% identity, e-value 1e-13 for the common honey bee, Apis mellifera, and 71% identity, e-value 1e-12 for the parasitic wasp Trichogramma pretiosum.
MpNa v expression after dsRNA feeding
The silencing efficiency of dsMpNav was examined using real-time quantitative PCR. We analyzed expression of MpNav at daily intervals following continuous oral delivery of dsRNA. A direct correlation was observed between the amounts of dsRNA ingested and a consequent decrease in the abundance of MpNav mRNA transcript. At the 2nd day of ingestion, MpNav dsRNA caused significant (P < 0.01) reduction in MpNav mRNA abundance. During the 3rd to 7th day of dsMpNav feeding the MpNav transcript level showed very significant differences (P < 0.001, 2.5-fold decrease) compared with the DEPC water and dsGFP control treatments, indicating a substantial gene knockdown (Fig. 2).
Dietary delivery of MpNav dsRNA alters MpNav expression in M. persicae. Relative abundances of MpNav H1 gene transcripts, as determined by qRT-PCR of cDNA made from total RNA isolated from whole body 1 to 7 days after the indicated dietary treatments of M. persicae. Data represent the mean ± SEM of three independent experiments. MpNav mRNA was normalized with Actin as an endogenous control. Treatment was compared with controls using ANOVA (Tukey Kramer multiple comparison, p < 0.05). Symbols ns, ** and *** indicates non-significant, P < 0.01 and P < 0.005 respectively.
Effects of dsRNA on aphid development and mortality
No significant aphid mortality was observed for the DEPC water, dsGFP and dsMpNav treatments after the first day of feeding. The mortality differences between treatment become more and more obvious from the 2nd day of feeding onwards. The average mortality for dsMpNav treatments reached 34.7%, 43.6%, 58.2%, 60.0%, 63.6% and 65.7% respectively after the second, third, fourth, fifth, sixth and seventh day of continuous feeding, whereas the mortality of the DEPC water treated group was 5.5%, 4.3%, 6.2%, 5.0%, 8.1% and 8.4% respectively, whilst the mortality for dsGFP oral feeding treatments was 3.8%, 6.2%, 4.8%, 5.7%, 7.6% and 8.1% respectively (Fig. 3). Relative to the -ve (DEPC water) and +ve (dsGFP) control diets, which exhibited only a small decrease in survival over an assay period of 7 days, the dsMpNav containing diet resulted in a significant (p < 0.005) impact on aphid mortality between days 3 and 7. At day 7, any surviving insects were transferred to cabbage leaves and their longevity and fecundity evaluated. Prior dietary delivery of dsMpNav significantly decreased (p < 0.01, p < 0.05) the longevity and fecundity of M. persicae survivors compared to control treatments (Figs 4 and 5).
Mortality rate of M. persicae fed artificial diets. Mortality rates by feeding artificial diets containing DEPC water, dsGFP and dsMpNav over time. Mean ± SEM of three replications (n = 150) are shown. Different treatments were compared using ANOVA (Tukey Kramer multiple comparison, p < 0.05). Symbols ns and *** indicates non-significant and P < 0.005 respectively.
Effect of MpNav silencing on adult longevity of M. persicae. Life-spans of insects fed on artificial diets containing DEPC water, dsGFP and dsMpNav at their nymphal stage. Mean ± SEM of three replications (n = 20, 18, or 17) are shown. Treatments are compared using ANOVA (Tukey Kramer multiple comparison, p < 0.05). Symbol ** indicates P < 0.01.
Effect of MpNav knock-down on fecundity of M. persicae. Fecundity of insects previously fed on artificial diets containing DEPC water, dsGFP and dsMpNav at their nymphal stage. Mean ± SEM of three replications (n = 18) are shown. Treatments are compared using ANOVA (Tukey Kramer multiple comparions, p < 0.05). Symbol ** indicates P < 0.01.
Discussion
For electrical signaling in eukaryotes, VGSCs are essential and ubiquitous29. Insect VGSCs are normally encoded by a single gene (para in Drosophila melanogaster and its equivalent orthologs in other insect species), that can generate a large number of transcriptional editing and splice variants that are differentially expressed at various developmental stages within the life cycle and in specific cell types30. Unusually in M. persicae (and other aphids), the functioning VGSC is encoded by two genes (designated heteromer H1 and H2) with some unique channel properties13. These two heteromer’s are hypothesized to have arisen due to a gene inversion event having occurred at some time during the evolution of aphids, resulting in a novel two-subunit channel.
Abd El Halim et al.14 recently demonstrated that RNAi-mediated knockdown of a VGSC gene (TcNav) by oral feeding causes up to 51.34% larval mortality and developmental arrest in the red flour beetle T. castaneum. This dose dependent larval mortality was observed after the beetles were continuously fed dsRNA for 6 days. Moreover, adult emergence was also significantly reduced in insects fed with dsRNA. Our results for the peach-potato aphid channel MpNav are on a par with this study, with up to 63.6% mortality observed for aphid nymphs by day 6 of exposure to dsRNA targeted against the H1 subunit. In our experiments, knock-down of the MpNav gene also significantly reduced adult longevity (by up to 7 days) and the fecundity (a decline of approximately 45%) of survivors. A previous study in Drosophila melanogaster has reported that decreased levels of the para VGSC in mlenapts (no-action potential temperature sensitive mutation of the maleless (mle) gene) mutant flies resulted in decreased longevity and fecundity31. mlenapts is a recessive gain-of-function mutation of mle that results in a splicing defect of the Na+ channel transcript and a severe reduction of VGSC RNA levels and channel activity. Another recent RNAi study targeting the VGSC in the bird cherry-oat aphid Rhopalosiphum padi32, a global pest of wheat, indicated significant suppression of the transcript levels of heteromers H1 and H2, in this case by direct injection (rather than oral feeding) of their respective dsRNA, and a significant cross-suppressions in the transcript levels between H1 and H2 subunit genes. Although in our investigations we did not analyse expression of the M. persicae H2 subunit in response to application of H1 dsRNA, the likelihood is that significant cross-suppression will also have occurred.
If we compare the regions targeted for RNAi suppression in these three individual studies, whereas in T. castaneum a 239 nt dsRNA covering the membrane spanning regions S1 and S2 in Domain I of the channel (GenBank accession NM_001165908.1) was used, the two separate aphid studies (M. persicae and R, padi) employed dsRNAs encompassing the DI-DII linker region of heteromer H1. For R. padi H1 (GenBank accession KJ872633) a larger 485 bp Rpvgsc1 fragment was amplified. However, in both aphid studies a common region of 71 amino acids (214 nucleotides) was covered in the amplified fragments. In both cases similar levels of suppression were reported, suggesting that this region is a good target for RNAi suppression technology. In the R. padi study a 358 bp dsRNA fragment encompassing the DIV S4-S5 and S5-S6 linker region (Rpvgsc2 GenBank accession no. KP966088) of heteromer H2 (which is the more evolutionarily divergent gene when compared to classical monomeric VGSCs) was also selected for suppression studies. It is not, however, clear from these experiments which region of the gene (coding region 3′ or 5′ end) is ideal for dsRNA design, and in fact it may not really be that important e.g. in the pea aphid Acyrthosiphon pisum, no difference in mortality was observed in groups of insects fed with dsRNA matching either the 5′ or 3′ end of the hunchback (hb) gene25.
Unintentional gene silencing in non-target species33 is the primary risk posed by pesticidal RNAi. Bioinformatic analyses that compares the pesticidal RNAs to non-target genomes has been recognised as an useful initial screen that can help to predict potential non-target risks posed by RNAi33,34. In the present study, we used in-silico searches to determine whether our 289 bp dsRNA shared prohibitive sequence similarities with the genes from key insect pollinators and beneficial insects. We obtained somewhat similar homology scores in the range 73–74% for the bee species represented in the NCBI nucleotide database and the highest score for a beneficial insect (predatory wasp) was 71%. In comparison, other aphid species returned high homology scores of 84–95%, suggesting that in practice the dsRNA used in this study may suppress the VGSC expression of more than one aphid type. No significant hits were obtained when the off-target search algorithm dsCheck35 was used to identify exact and near nucleotide matches of the 289 bp M. persicae dsRNA to the genome of the model insect D. melanogaster. However, even considerable sequence divergence between an mRNA and the dsRNA does not rule-out unintentional gene silencing since, once ingested, the dsRNA is cleaved into numerous very short (19–23 nucleotides) small interfering RNAs (siRNA). These siRNAs most likely have abundant direct sequence matches throughout most eukaryotic and prokaryotic genomes, thereby increasing the chances for off-target (i.e. silencing of a gene with sufficient sequence similarity to the dsRNA) and non-targeted (silencing of the intended gene in an unintended organism) effects36.
E-RNAi37 analysis used for design of the dsRNA used in the T. castaneum study14 predicted over two hundred 19 nt siRNAs could be generated from the 239 bp dsRNA fragment (albeit with an average efficiency score of 54.16, and no off-target predictions); there is thus considerable scope for unforeseen off-target effects. Most studies report that dsRNA ranging from 140 to 500 nucleotides in length are required for successful RNAi in insects, although successful suppression with dsRNA of 50 bp has been reported38. This finding could be helpful to further optimize the dsRNA specificity by using shorter amplified dsRNA fragments, thereby decreasing the likelihood of non-target and off-target effects. The success of RNAi is also dependent on the molecules stability and an effective uptake of the dsRNA by the target species39. A rapid degradation of dsRNA by extracellular ribonucleases in the insect haemolymph and gut is increasingly recognised as a fundamental factor influencing RNAi efficiency in several insect orders40. This is exacerbated in hemipteran species since extra-oral salivary degradation of dsRNAs provides an additional block to cellular uptake17,21,41. To translate RNAi to field applicability, RNAi silencing elicitors will most likely need to be combined with biotic or abiotic systems that mediate both protection and uptake of the eliciting RNAi trigger39.
To conclude, our study demonstrates that silencing of a voltage-gated sodium channel gene (MpNav) via RNAi in M. persicae caused larval mortality. It provides evidence to propose that MpNav is a feasible candidate gene to target for the control of this insect pest using RNA interference technology and provides the foundation to design dsRNA molecules and associated delivery systems with a high degree of species specificity that could ultimately be used as commercial bio-aphicides.
Data Availability
All data generated or analysed during this study are included in this published article.
References
Blackman, R. L. & Eastop, V. F. Aphids on the world’s crops. An identification and information guide; John Wiley, Chichester, UK (1984).
Hogenhout, S. A., Ammar, E.-D., Whitfield, A. E. & Redinbaugh, M. G. Insect vector interactions with persistently transmitted viruses. Annual Review of Phytopathology 46, 327–359, https://doi.org/10.1146/annurev.phyto.022508.092135 (2008).
Dixon, A. F. G. Aphid ecology. An optimization approach. Springer, Netherlands (1985).
Bass, C. et al. The evolution of insecticide resistance in the peach potato aphid, Myzus persicae. Insect Biochem Mol Biol 51, 41–51, https://doi.org/10.1016/j.ibmb.2014.05.003 (2014).
Davies, T. G. E., Field, L. M., Usherwood, P. N. & Williamson, M. S. DDT, pyrethrins, pyrethroids and insect sodium channels. IUBMB Life 59, 151–162, https://doi.org/10.1080/15216540701352042 (2007).
Food 2030. Department for Environment, Food and Rural Affairs (2010)
Jinek, M. & Doudna, J. A. A three-dimensional view of the molecular machinery of RNA interference. Nature 457, 405–412, https://doi.org/10.1038/nature07755 (2009).
Whangbo, J. S. & Hunter, C. P. Environmental RNA interference. Trends Genet 24, 297–305, https://doi.org/10.1016/j.tig.2008.03.007 (2008).
Burand, J. P. & Hunter, W. B. RNAi: Future in insect management. Journal of Invertebrate Pathology 112, S68–S74, https://doi.org/10.1016/j.jip.2012.07.012 (2013).
Gu, L. Q. & Knipple, D. C. Recent advances in RNA interference research in insects: Implications for future insect pest management strategies. Crop Prot 45, 36–40, https://doi.org/10.1016/j.cropro.2012.10.004 (2013).
Scott, J. G. et al. Towards the elements of successful insect RNAi. J Insect Physiol 59, 1212–1221, https://doi.org/10.1016/j.jinsphys.2013.08.014 (2013).
Amey, J. S. et al. An evolutionarily-unique heterodimeric voltage-gated cation channel found in aphids. FEBS Lett 589, 598–607, https://doi.org/10.1016/j.febslet.2015.01.020 (2015).
Noda, M. & Hiyama, T. Y. Sodium sensing in the brain. Pflügers Archiv - European Journal of Physiology 467, 465–474, https://doi.org/10.1007/s00424-014-1662-4 (2015).
Halim, A. E. H. M. et al. RNAi-mediated knockdown of the voltage-gated sodium ion channel TcNav causes mortality in Tribolium castaneum. Sci Rep 6, 29301, https://doi.org/10.1038/srep29301 (2016).
Knorr, E. et al. Gene silencing in Tribolium castaneum as a tool for the targeted identification of candidate RNAi targets in crop pests. Sci Rep 8, 2061, https://doi.org/10.1038/s41598-018-20416-y (2018).
Baum, J. A. et al. Control of coleopteran insect pests through RNA interference. Nat Biotechnol 25, 1322–1326, https://doi.org/10.1038/nbt1359 (2007).
Singh, I. K., Singh, S., Mogilicherla, K., Shukla, J. N. & Palli, S. R. Comparative analysis of double-stranded RNA degradation and processing in insects. Sci Rep 7, 17059, https://doi.org/10.1038/s41598-017-17134-2 (2017).
Christiaens, O. & Smagghe, G. The challenge of RNAi-mediated control of hemipterans. Curr Opin Insect Sci 6, 15–21, https://doi.org/10.1016/j.cois.2014.09.012 (2014).
Luo, Y. et al. Towards an understanding of the molecular basis of effective RNAi against a global insect pest, the whitefly Bemisia tabaci. Insect Biochem Mol Biol 88, 21–29, https://doi.org/10.1016/j.ibmb.2017.07.005 (2017).
Tzin, V. et al. RNA interference against gut osmoregulatory genes in phloem-feeding insects. J Insect Physiol 79, 105–112, https://doi.org/10.1016/j.jinsphys.2015.06.006 (2015).
Cao, M., Gatehouse, J. A. & Fitches, E. C. A systematic study of RNAi effects and dsRNA stability in Tribolium castaneum and Acyrthosiphon pisum, following injection and ingestion of analogous dsRNAs. Int J Mol Sci 19, https://doi.org/10.3390/ijms19041079 (2018).
Sambrook, J. & Russell, D. W. Molecular cloning: a laboratory manual. 3rd edn, Cold Spring Harbor Laboratory Press (2001).
Pan, K., Huang, B.-Q. & Hou, X.-W. A modified practical method of rearing Aphis craccivora with meridic liquid mutrients. Chinese Bulletin of Entomology 43, 728–730 (2006).
Rahbé, Y. & Febvay, G. Protein toxicity to aphids: an in vitro test on Acyrthosiphon pisum. Entomologia Experimentalis et Applicata 67, 149–160, https://doi.org/10.1111/j.1570-7458.1993.tb01663.x (1993).
Mao, J., Zeng, F. & Feeding-based, R. N. A. interference of a gap gene is lethal to the pea aphid, Acyrthosiphon pisum. PLoS One 7, e48718, https://doi.org/10.1371/journal.pone.0048718 (2012).
Ye, J. et al. Primer-BLAST: A tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 13, 134, https://doi.org/10.1186/1471-2105-13-134 (2012).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408, https://doi.org/10.1006/meth.2001.1262 (2001).
Zhi‐Wei, K. et al. Evaluation of the reference genes for expression analysis using quantitative real‐time polymerase chain reaction in the green peach aphid, Myzus persicae. Insect Science 24, 222–234, https://doi.org/10.1111/1744-7917.12310 (2017).
Hille, B. Ion channels of excitable membranes 3rd ed.; Sinauer Associates, Ltd: Sunderland, M.A (2001).
Soderlund, D. M. Sodium channels. In: Comprehensive molecular insect science; Gilbert, L. I., Iatrou, K., Gill, S. S., Eds.; Elsevier: New York, Vol. 5, pp 1−24 (2005).
Garber, G., Smith, L. A., Reenan, R. A. & Rogina, B. Effect of sodium channel abundance on Drosophila development, reproductive capacity and aging. Fly (Austin) 6, 57–67, https://doi.org/10.4161/fly.18570 (2012).
Zuo, Y. et al. Expression patterns, mutation detection and RNA interference of Rhopalosiphum padi voltage-gated sodium channel genes. Scientific reports 6, 30166, https://doi.org/10.1038/srep301661 (2016).
Mogren, C. L. & Lundgren, J. G. In-silico identification of off-target pesticidal dsRNA binding in honey bees (Apis mellifera). PeerJ 5, e4131, https://doi.org/10.7717/peerj.4131 (2017).
Zotti, M. J. & Smagghe, G. RNAi technology for insect management and protection of beneficial insects from diseases: lessons, challenge sand risk assessments. Neotrop. Entomol 44, 197–213, https://doi.org/10.1007/s13744-015-0291-8 (2015).
Naito, Y. et al. dsCheck: highly sensitive off-target search software for double-stranded RNA-mediated RNA interference. Nucleic Acids Research 33, W589–W591, https://doi.org/10.1093/nar/gki419 (2005).
Lundgren, J. G. & Duan, J. J. RNAi-based insecticidal crops: potential effects on nontarget species. BioScience 63, 657–665, https://doi.org/10.1525/bio.2013.63.8.8 (2013).
Horn, T. & Boutros, M. E-RNAi: a web application for the multi-species design of RNAi reagents—2010 update. Nucleic Acids Research 38, W332–W339, https://doi.org/10.1093/nar/gkq317 (2010).
Jing, Y. & Zhao-jun, H. Optimisation of RNA interference-mediated gene silencing in Helicoverpa armigera. Australian Entomology 53, 83–88, https://doi.org/10.1111/aen.12052 (2014).
Joga, M. R., Zotti, M. J., Smagghe, G. & Christiaens, O. RNAi efficiency, systemic properties, and novel delivery methods for pest insect control: What we know so far. Front Physiol 7, https://doi.org/10.3389/fphys.2016.00553 (2016).
Wang, K. X. et al. Variation in RNAi efficacy among insect species is attributable to dsRNA degradation in-vivo. Insect Biochem Molec Biol 77, 1–9, https://doi.org/10.1016/j.ibmb.2016.07.007 (2016).
Christiaens, O., Swevers, L. & Smagghe, G. DsRNA degradation in the pea aphid (Acyrthosiphon pisum) associated with lack of response in RNAi feeding and injection assay. Peptides 53, 307–314, https://doi.org/10.1016/j.peptides.2013.12.014 (2014).
Acknowledgements
This work was funded by Higher Education Commission of Pakistan (Grant No. 1594/SRGP/R&D/HEC/2017). The Smart Crop Protection (SCP) strategic programme (BBS/OS/CP/000001) at Rothamsted Research is funded through the Biotechnology and Biological Sciences Research Council’s Industrial Strategy Challenge Fund.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: K.T. and A.A. Performed the experiments: K.T., E.N. and L.N. Analyzed the data: K.T., S.S., M.H. and F.U. Contributed reagents/materials: K.T., E.N. L.N. and M.H. Performed the bio-informatic analysis: T.G.E.D. Wrote the paper: K.T. and T.G.E.D.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tariq, K., Ali, A., Davies, T.G.E. et al. RNA interference-mediated knockdown of voltage-gated sodium channel (MpNav) gene causes mortality in peach-potato aphid, Myzus persicae. Sci Rep 9, 5291 (2019). https://doi.org/10.1038/s41598-019-41832-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-41832-8
This article is cited by
-
A first greenhouse application of bacteria-expressed and nanocarrier-delivered RNA pesticide for Myzus persicae control
Journal of Pest Science (2023)
-
Flock house virus as a vehicle for aphid Virus-induced gene silencing and a model for aphid biocontrol approaches
Journal of Pest Science (2023)
-
RNA interference-mediated knockdown of odorant-binding protein 2 and MP58 gene causes mortality in Myzus persicae
International Journal of Tropical Insect Science (2022)
-
Intron-containing hairpin RNA interference vector for OBP8 show promising mortality in peach potato aphid
Plant Cell, Tissue and Organ Culture (PCTOC) (2022)
-
Efficacy of various natural plant extracts and the synthetic insecticide cypermethrin 25EC against Leucinodes orbonalis and their impact on natural enemies in brinjal crop
International Journal of Tropical Insect Science (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.