Abstract
Spliced leader trans-splicing (SLTS) plays a part in the maturation of pre-mRNAs in select species across multiple phyla but is particularly prevalent in Nematoda. The role of spliced leaders (SL) within the cell is unclear and an accurate assessment of SL occurrence within an organism is possible only after extensive sequencing data are available, which is not currently the case for many nematode species. SL discovery is further complicated by an absence of SL sequences from high-throughput sequencing results due to incomplete sequencing of the 5’-ends of transcripts during RNA-seq library preparation, known as 5′-bias. Existing datasets and novel methodology were used to identify both conserved SLs and unique hypervariable SLs within Heterodera glycines, the soybean cyst nematode. In H. glycines, twenty-one distinct SL sequences were found on 2,532 unique H. glycines transcripts. The SL sequences identified on the H. glycines transcripts demonstrated a high level of promiscuity, meaning that some transcripts produced as many as nine different individual SL-transcript combinations. Most uniquely, transcriptome analysis revealed that H. glycines is the first nematode to demonstrate a higher SL trans-splicing rate using a species-specific SL over well-conserved Caenorhabditis elegans SL-like sequences.
Similar content being viewed by others
Introduction
Pre-mRNA splicing is a vital mechanism associated with the expression and regulation of eukaryotic genes. The most widely deployed splicing mechanism is cis-splicing, which enables the removal of intron sequences from mRNA molecules. Trans-splicing is less widespread and results in the fusion of RNA molecules that are transcribed from different genomic loci. The most prevalent form of trans-splicing involves the addition of a short, spliced leader (SL) sequence to the 5′ end of mRNA transcripts, referred to as spliced leader trans-splicing (SLTS). SLTS has evolved independently in a diverse set of phyla including Nematoda, Platyhelminthes, Trypanosoma, Cnidaria, Rotifera, Chordata, Arthropoda and Dinoflagellata1,2,3,4,5,6,7,8.
SLs originate from SL RNA genes, whose transcripts are divided into two parts by a donor splice site: a 5′ exon-like SL region and a 3′ intron-like region9,10,11. SL RNAs maintain a conserved secondary structure comprised of hairpins and a single-stranded Sm binding site (5′-purine-AU4–6G-purine-3), which allows the SL RNA to interact with proteins that are required for SLTS3,12,13.
It is evident that SLTS has a role in resolving polycistonic mRNAs in Caenorhabditis elegans, acting as a prerequisite for subsequent translation14. In C. elegans, approximately 70% of transcripts are trans-spliced to a 22nt SL: SL1 or SL23,15,16,17. However, operon resolution is not the sole function of SLTS in C. elegans, as only 17% of C. elegans transcripts originate from operons15,18. It has been hypothesized that SLTS is involved in many translational regulation mechanisms, including the replacement of deleterious sequences in the 5′-untranslated region, addition of translational motifs from within the SL sequence, or by replacing a transcript’s 5′-monomethylated cap with a 5′-hypermodified cap structure18,19,20,21,22,23,24.
Sequence data indicate that all nematode species studied to date utilize SL trans-splicing. In all nematodes, SLs with similarity to SL1 and/or SL2 are found, with an exception of Trichinella spiralis, which uses non-canonical spliced leaders25,26,27,28,29,30,31,32. Interestingly, sequence analysis of the potato cyst nematodes Globodera rostochiensis and G. pallida, identified multiple hypervariable SL sequences in addition to SL1 and SL226,33. The diversity of SL sequences found in Globodera spp. and the dearth of information regarding their functionality highlights a need to improve our understanding through the investigation of nematode genomic and transcriptomic data. Previous studies have identified SL1 in the soybean cyst nematode, Heterodera glycines, a highly damaging plant parasite closely related to Globodera spp.34. Subsequently, the SL1 sequence has been used to successfully generate H. glycines cDNA libraries (LIBEST_005577; unpublished McCarter, J., Clifton, S., Chiapelli, B., Pape, D., Martin, J., Wylie, T., Dante, M., Marra, M., Hillier, L., Kucaba, T. et al.).
In this current study, we utilize the recently assembled H. glycines genome35 and the RNA-seq reads from an early-life stages transcriptome36 to extensively characterize SLs and their usage in H. glycines. Serendipitous observation of variation in the 5′-end of two previously sequenced H. glycines transcripts led to the discovery of a novel SL. Through subsequent bioinformatic approaches utilizing both H. glycines genomic and transcriptomic data, this report shows that H. glycines possesses at least twenty-one SLs, found on a total of 2,532 H. glycines transcripts, which account for approximately one-third of H. glycines genes. Functional analysis of the H. glycines SL trans-spliced transcripts reveals involvement in a variety of biological processes. Interestingly, around 45% of the transcripts are promiscuously trans-spliced by SLs suggesting that there is functional redundancy amongst SL RNA molecules. Furthermore, H. glycines is the first nematode to show a transcriptome-wide preference for a species-specific SL sequence over the well-conserved C. elegans SL-like sequences.
Results
Discovery of a novel spliced leader in H. glycines
Exploring the available H. glycines expressed sequence tags on NCBI revealed two transcripts coding for chorismate mutase proteins (AY16022537 & MH119144), which are important enzymes for parasitism in multiple plant-parasitic nematodes37,38,39,40,41,42. Alignment of the 5′ end of MH119144 and the SL1 primer sequence used to clone AY160225 revealed divergent 5′ ends, suggesting the presence of a novel SL sequence (Fig. 1A).
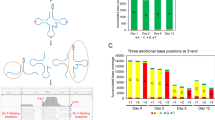
MH119144 is SL trans-spliced by a novel SL sequence. (A) Pairwise sequence alignment of C. elegans SL1 (CeSL1) and the first 22 nucleotides of MH119144 using EMBOSS needle with default settings. (|) = matching nucleotides, (.) = mismatched nucleotides, (−) = gap. (B) Secondary structural prediction of the putative MH119144 SL RNA consensus sequence using RNAfold shows a characteristic 3-hairpin SL RNA structure. (C) RT-PCR using H. glycines gDNA (g), cDNA (c) or no template DNA (NT). All lanes use a gene-specific reverse primer with either a gene-specific (GSP) forward primer or a SL forward primer.
To investigate the putative MH119144 SL sequence, the entire transcript was mapped to the H. glycines genome with BLASTn. All but the first fifteen nucleotides of MH119144 mapped to scaffold_282 (Supplemental Fig. S1). To locate the 5′-end of MH119144 in the H. glycines genome, the twenty-two-nucleotide putative SL was mapped to the genome with BLASTn. The putative SL had four exact hits in the H. glycines genome, all of which mapped within a 2.5 Kb region on scaffold_362, confirming that MH119144 is comprised of two sequences that are located in distinctly separate regions of the genome (Supplemental Fig. S1).
In order for a SL to be functional, transcription must create a distinct non-coding hairpin SL RNA structure with a single-stranded Sm motif3,12,13. To identify the presence of these features, the ninety-eight nucleotides downstream of the four putative SL hits were extracted from the genome. All sequences had 99% sequence identify and displayed the typical secondary structure of functional SL RNAs (Fig. 1B).
The validity of the putative SL was tested further using RT-PCR to search for the putative SL chorismate mutase sequence in H. glycines gDNA and cDNA (Fig. 1C). Using the putative SL sequence as a forward primer and a gene-specific reverse primer, a visible band was produced when using a cDNA template, but not gDNA (Fig. 1C). Genic structure predictions performed on chorismate mutase indicate that the absence of a band within the gDNA reaction is not due to the primers being located on intron/exon borders. Furthermore, a control PCR amplification with cDNA and gDNA templates was performed using a gene-specific primer pair to verify the presence of the chorismate mutase gene in both DNA samples (Fig. 1C).
Collectively, three tiers of evidence support the legitimacy of this novel SL, including: mapping of the putative SL and the remainder of the transcript to separate locations within the genome, the similarity of the putative SL RNA sequence to known SL RNAs, and the absence of a SL chorismate mutase PCR product when using gDNA. This novel SL will subsequently be named Heterodera glycines spliced leader 3 (HgSL3) to distinguish it from C. elegans SL1-like and SL2-like sequences in other nematode species.
The H. glycines genome contains multiple novel SL sequences
To investigate the existence of previously identified SL sequences in H. glycines, all known SLs from C. elegans and Globodera spp. were mapped to the H. glycines genome. SL1 mapped to 180 loci in the H. glycines genome, twenty-two sequences of which were located within close proximity to a Sm motif13. The only Globodera spp. SL variant present in the H. glycines genome was SL1b, which lacked a proximal Sm motif (Supplemental Table 1).
To search for novel HgSL RNA genes, HgSL3 RNA was queried with BLAST against the H. glycines genome. A total of twenty sequences were identified that also contained single-stranded Sm-binding sites flanked by hairpins (Supplemental Table 1). Alignment of the first twenty-two nucleotides of the putative HgSL RNA sequences yielded ten additional unique HgSLs, numbered HgSL4–13 (Fig. 2).
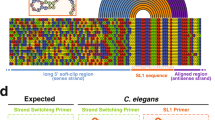
Sequence alignment of novel hypervariable H. glycines SLs. Multiple sequence alignment of H. glycines HgSL3 RNA blast results with SL RNA-like secondary structural predictions. The alignment was performed using T-coffee with default settings. (*) indicates a consensus nucleotide position and (−) denotes a sequence gap. Pink, yellow and green shading indicates good, average and bad nucleotide matches respectively.
Splice leaders are promiscuously present on multiple H. glycines transcripts
To assess SLTS in H. glycines, known SLs were truncated to the 3′ most 11nt yielding a total of twenty-six unique sequences (7 from C. elegans, 5 from H. glycines, and 14 from Globodera spp.). The use of truncated SLs has been demonstrated to circumvent the low availability of complete 5′-ends in RNA-seq data25,26.
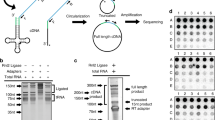
The truncated SLs were used as query sequences for three separate BLAST analyses. In the first approach, SLs were queried against the NCBI EST database, in the second approach SLs were queried against a H. glycines transcriptome36. A two-tiered third approach that involved SL queries to trimmed Illumina reads with subsequent mapping to the 5′ end of transcripts (Fig. 3). This third approach circumvents RNA-seq 5′ bias, which may result in the misassembly at the 5′ end of transcripts43.
BLAST searches to ESTs and transcripts yielded 187 and 2,215 SL trans-spliced transcripts respectively, with a 40% (74/187) rediscovery rate of ESTs within the transcriptome (Table 1 and Fig. 4). After removing the SLs from the sequences, all ESTs were unique, while only 2076/2,215 transcripts were unique, revealing that in some cases transcripts are not uniquely spliced to one SL (Table 1 and Fig. 4). Using the read-based approach, 85,876 of ~11.4 million reads had a terminal SL, the legitimacy of which is supported by SL BLAST hits preferentially locating to the 5′-read ends (Fig. 5). Subsequent mapping of the SL-reads to the H. glycines transcriptome revealed a false positive rate of SL-reads at 88.4%, with 9,927/85,876 reads mapping to the 5′ end of 1,635 unique SL trans-spliced transcripts. Again, a portion of the transcripts appeared to be the target of more than one SL RNA molecule, resulting in 6,350 SL-transcript combinations (Table 1 and Fig. 4). Collectively, these analyses identified in 2,532 unique SL trans-spliced transcripts and 21 functional SLs (Table 1 and Fig. 4). Interestingly, when combining all three analyses, HgSL3 is present on 30.9% of SL trans-spliced transcripts making it the most abundantly used SL, a finding unique to H. glycines. Furthermore, 45.5% of the 2,532 SL trans-spliced transcripts were spliced by two or more SLs, with trans-splicing of five or more different SLs onto 6.8% of these transcripts (Fig. 6).
Summary of the H. glycines SL trans-spliced transcript analyses. The Venn diagram represents the overlapping and unique SL trans-spliced transcripts identified through the three-part approach using BLASTn against the NCBI ESTs (EST BLAST), the Trinity transcriptome (Direct BLAST) and the unassembled reads (read-to-transcript).
HgSLs are preferentially located at the 5′ end of H. glycines reads. All truncated HgSLs were queried with BLASTn against the H. glycines raw reads, which were also used for the Trinity assembly. The read nucleotide start positions were plotted to show the strong preference for HgSLs to be located at the 5′ end of the read.
Genomic features of transcripts that possess spliced leaders
To functionally characterize the genes that give rise to SL trans-spliced transcripts, all SL trans-spliced transcripts were mapped to the H. glycines genome using GMAP44. Exonic overlap between H. glycines genes and SL trans-spliced transcripts accounted for approximately one-third of the genes in the genome (9,042/29,959). It is interesting that of the 9,042 SL trans-spliced genes, approximately one-third (3,013) co-align with annotated repeats in the H. glycines genome. The ten most abundant repeats comprised 27.7% of the 3,013 trans-spliced genes. The most abundant functionally annotated repeat is associated with a LINE/CR1 retrotransposon (3.4%), suggesting that a significant portion of SL trans-spliced transcripts are associated with transposons (Table 2).
To assess the positioning of SL trans-spliced genes within the H. glycines genome, the genome was partitioned into 50 kb bins. Analysis of the 50 kb genomic segments showed that SL trans-spliced genes were dispersed throughout the genome. However, clustering of SL trans-spliced genes was also evident, as 40 of the 2,640 50 kb bins had 14 or more consecutively arranged SL trans-spliced genes (Table 3).
Functional analysis of SL trans-spliced transcripts reveals involvement in a variety of biological processes
In order to gain functional insight into the role of SL trans-splicing in H. glycines, the SL trans-spliced transcripts were annotated with Blast2go45 (Supplemental Table 2). Over half (52%) of the annotated transcripts were involved in metabolic and developmental processes (Fig. 7A), with the top two biological processes involved in ‘Embryo development ending or egg hatching’ and ‘Nematode larval development’ (Fig. 7B). A complementary GO enrichment analysis was performed on the corresponding genomic genes, revealing a similar profile of functions involved in metabolic processes (Fig. 7C, Supplemental Table 3).
Gene Ontology (GO) biological processes for SL trans-spliced transcripts and genes. (A) SL trans-spliced transcripts. (B) Specific child terms for SL trans-spliced transcripts. (C) SL trans-spliced genes. Bars represent the number of proteins represented for each GO biological process that was identified using Blast2go (transcripts) or Ontologizer (genes).
Effector transcripts are SL trans-spliced and display an all-or-none relationship with multi-gene copy effectors
To investigate whether spliced leaders could be involved in parasitism, we searched for exon-exon overlap between SL trans-spliced transcripts and effector genes in the genome. Effector genes produce proteins that are secreted by H. glycines during parasitism and are thought to play a major role in altering host cell structure and function. Reviewed by46,47,48,49. Within the H. glycines genome there are 80 known bona fide effector proteins, 28 of which originate from multiple gene copies and 51 are single gene effectors, to make a total of 121 currently confirmed effector genes. SL trans-spliced transcripts overlapped with the first exon of 29/121 effector genes, indicating that approximately 24% of the currently known bona fide effector genes are subject to SL trans-splicing (Table 4). Interestingly 23/28 multi-gene copy effectors display an all-or-none relationship with SL trans-splicing. For example, 5/5 genes corresponding to 11A06 are SL trans-spliced, while 0/5 genes are SL trans-spliced for 4D06 and 32E03 (Table 4, Supplemental Table 4).
Discussion
This study identified and functionally characterized SLs and SL trans-spliced transcripts of the plant-parasitic soybean cyst nematode Heterodera glycines. The recent availability of both the H. glycines genome and transcriptome has provided an opportunity to extensively characterize SL use and function in a parasitic nematode.
This study was prompted by the discovery of HgSL3 at the 5′-end of a chorismate mutase effector cDNA, leading to the identification of a unique set of hypervariable HgSLs. Novel hypervariable SLs have previously been discovered in the potato cyst nematode G. rostochiensis and the animal-parasitic nematode T. spiralis25,26. Interestingly, despite the high volume of SLs that have been discovered in these three species, genomic data suggest a low interspecies conservation of SLs. Given the parasitic nature of all three species, as well as the perceived link between SLs and translational regulation, it is possible that the hyper-variation of SLs is a response to parasitism of different hosts. This study investigated a possible link between SLs and known parasitic molecules, referred to as effectors, and found that 24% (29/121) of bona fide effector genes are subject to SL trans-splicing. Previous hypotheses indicate that species use SL trans-splicing as a form of translational control to respond to changing environments, particularly in response to nutrient availability20. The existence of two subsets of effector transcripts, one SL trans-spliced and one not, may provide H. glycines with a way to mitigate host defense responses through differentially regulating the two subsets of effectors.
To identify H. glycines SL trans-spliced transcripts, SLs were first truncated at the 5′-ends before being queried using BLAST against H. glycines sequences. The use of truncated SLs was previously utilized in G pallida26. Before truncating the SLs in H. glycines, we first verified that this approach was necessary by using the full-length SLs as query sequences against the H. glycines ESTs and transcriptome. Only fifteen sequences, none of which were SL1, were identified across both databases (available in GitHub). The failure to recover full-length SL1 supports the lack of 5′-ends within the H. glycines datasets, as SL1 is present in H. glycines and other related nematodes26,27. The read-based approach further verified the lack of complete 5′-ends within the H. glycines transcriptome by showing that the truncated SLs were predominantly located at the first nucleotide of raw reads that were underrepresented in mature transcripts. To further complicate transcriptome assembly in SLTS organisms, this study revealed that 45.5% of SL trans-spliced transcripts do not have one unique SL-transcript combination. The promiscuous nature of SLs on otherwise identical transcripts may cause high ambiguity in the assembly step, resulting in 5′ truncations or the assembly of a transcript that reflects only the most abundant SL-transcript while discarding lower expressed SL-transcripts.
Analysis of the available H. glycines ESTs, transcriptome and raw reads used in this study concluded that HgSL3 is the most prevalent SL in H. glycines, with 30.9% of the SL trans-spliced transcripts being trans-spliced by HgSL3. The predominant use of a non-SL1 sequence is unique to H. glycines and contrasts with findings in C. elegans and the animal-parasitic nematode Ascaris suum, as well as G. pallida where SL1 and SL1 variants were identified on >90% of the SL-containing G. pallida reads26.
C. elegans operon genes, which are resolved into monocistronic transcripts using SL trans-splicing, are upregulated during recovery from growth-arrested states14,50. Operon arrangement is believed to be advantageous in C. elegans during times of limited resources as there are less promoters competing for transcriptional resources50. In the case of H. glycines, SL trans-spliced transcripts were found to be involved in ‘Embryo development ending or egg hatching’ and ‘Nematode larval development,’ suggesting that SL trans-splicing may also be involved in initiating developmental changes in H. glycines. Operon arrangement has not yet been defined in H. glycines, however the clustering of SL trans-spliced transcripts in the genome suggests the presence of operon-like structures.
To both adapt and improve upon existing SL identification pipelines18,51,52, we developed a SL identification pipeline that utilizes generic RNA-seq, assembled transcripts, and ESTs, rather than requiring SL trapping prior to sequencing53,54. This method provides an alternative to existing pipelines by utilizing the propensity for SLs to be trans-spliced at 5′ ends and avoiding the requirement of unmapped reads having dual genomic mapping18,51. Additionally, using both pre-existing and predicted SL sequences that follow canonical SL RNA structures, we allow for the identification of novel SLs. A drawback of this approach may lie in the requirement of SLs to reside at 5′ ends, which is reliant on accurate adaptor trimming and prior knowledge of the anticipated SL length.
In summary, H. glycines possesses a unique set of hypervariable SLs, which are promiscuously trans-spliced to the 5′ end of >2,000 H. glycines transcripts, equivalent to approximately one-third of H. glycines genes. A robust identification of SLs was possible through novel methodology and the availability of H. glycines genome and transcriptome sequences. As more data becomes available for H. glycines and other parasitic and non-parasitic nematodes, the functional significance of SLTS may become more apparent and potentially lead to novel control measures.
Materials and Methods
Identification and structure prediction of putative SLRNAs
All G. rostochiensis26, C. elegans (PRJNA13758) SL sequences and the 22nt SL sequence from MH119144 were queried to the H. glycines genome with BLASTn V2.4.0 + (E-value 1.0e-3)55. SL hits and the adjacent 3′ 98 nucleotides were extracted using Samtools V1.456. Secondary structure was predicted using RNAfold V2.1.9 with unpaired bases participating in at most one dangling end. All extracted sequences were analyzed for a downstream Sm motif (5′-purine-AU4–6G-purine-3′)57.
DNA extraction and amplification
To confirm the functionality of putative SL on transcript MH119144 OP50 H. glycines was propagated on Williams 82 soybean. To isolate mixed-stage nematodes, root tissue was macerated with a blender, sieved and separated with a sucrose gradient58. Nematodes were ground in liquid nitrogen and total RNA was extracted using a RNeasy Mini Kit (Qiagen, Valencia, CA, USA). One μg of total RNA was treated with DNase I (Thermo Fisher Scientific, Waltham, MA, USA) and cDNA was synthesized using qScript cDNA SuperMix (Quantabio, Beverly, MA, USA). Genomic DNA was also extracted from ground nematode tissue using QIAamp DNA Mini Kit (Qiagen). RT-PCR was performed on a Bio-Rad S1000TM thermal cycler with reactions containing 1X PCR buffer, 1.5 mM MgCl2, 0.2 mM dNTP, and 1 unit of Taq DNA Polymerase (ThermoFisher Scientific). Thermocycler conditions were: 94 °C for 3 min, 35 cycles of 95 °C for 45 s, 55 °C for 30 s and 72 °C for 1 min, followed by 10 min at 72 °C.
Identification of SL trans-spliced transcripts
All putative SL sequences were queried with BLASTn to the H. glycines NCBI EST database, referred to as EST BLAST55,59 and a H. glycines de novo Trinity transcriptome assembled from NCBI SRA accession SRP122521, referred to as Direct BLAST36. BLAST hits were filtered to within the first thirteen nucleotides of the transcript, and with a 10nt minimum alignment length. ESTs and trinity transcripts were mapped to the genome using GMAP 20170317, and transcript to gene relationships were identified using exon to exon overlaps with Bedtools intersect V2.26.044,60. Genes were clustered by location using custom bash scripts.
Read Analysis
In the method referred to as Read-to-transcript, SLs were truncated to the 11 3′ nucleotides and queried with BLASTn V2.4.0+ to Sickle-trimmed (default)61, paired-end reads (word_size 5, -dust no, -task blastn-short) used in generating a H. glycines trinity transcriptome55. The subject start position for hits was graphed using GraphPad Prism 4. BLAST output was filtered by a 10 bp minimum alignment length and hits within 12 bp of the appropriate read end. Putative SL-containing reads were queried with BLASTn V2.4.0+ to the transcriptome and filtered by 80 bp min alignment length, and within 12 bp of the 5′ transcript end55.
Functional Analysis
Functional annotation was performed using Blast2go V4.1. All transcript sequences were searched against the NCBI NR database using Blastx (e-value 1.0–5. Interpro scan was performed using all default applications, and sequences were annotated with an annotation cutoff of 55 and a GO weight of 545,62. GO enrichment for trans-spliced genes was performed using Ontologizer V2.0 with gene functions from the H. glycines genome63.
Trans-splicing in Effector and Repetitive Genes
Bedtools intersect V2.26.0 and custom bash scripts were used to identify trans-spliced repetitive genes from a Repeatmodeler V1.0.8 tracks of the genome60,64. Effector genes were mapped to the genome using GMAP 2017031744, and were subjected to bedtools intersect V2.26.0 and custom bash scripts to identify trans-splicing effectors60.
Data Availability
The H. glycines expressed sequence tags analyzed during the current study are available through the National Center for Biotechnology Information website. The H. glycines transcriptome data is from the publication https://doi.org/10.1038/s41598-018-20536-5. The H. glycines genome is publically available on the SCNbase website (https://www.scnbase.org). All scripts used for bioinformatic analysis are available at (https://github.com/ISUgenomics/Heterodera-glycines-Spliced-Leaders/blob/master/StaceyBarnesRestart.md).
References
Douris, V., Telford, M. J. & Averof, M. Evidence for Multiple Independent Origins of trans-Splicing in Metazoa. Molecular Biology and Evolution 27, 684–693, https://doi.org/10.1093/molbev/msp286 (2010).
Ganot, P., Kallesoe, T., Reinhardt, R., Chourrout, D. & Thompson, E. M. Spliced-leader RNA trans splicing in a chordate, Oikopleura dioica, with a compact genomet. Molecular and Cellular Biology 24, 7795–7805, https://doi.org/10.1128/mcb.24.17.7795-7805.2004 (2004).
Krause, M. & Hirsh, D. A trans-spliced leader sequence on actin messenger-RNA in C-elegans. Cell 49, 753–761, https://doi.org/10.1016/0092-8674(87)90613-1 (1987).
Pouchkina-Stantcheva, N. N. & Tunnacliffe, A. Spliced leader RNA-mediated trans-splicing in phylum Rotifera. Molecular Biology and Evolution 22, 1482–1489, https://doi.org/10.1093/molbev/msi139 (2005).
Rajkovic, A., Davis, R. E., Simonsen, J. N. & Rottman, F. M. A Spliced Leader is Present on a Subset of mRNAs from the Human Parasite Schistosoma mansoni. Proceedings of the National Academy of Sciences of the United States of America 87, 8879–8883, https://doi.org/10.1073/pnas.87.22.8879 (1990).
Stover, N. A. & Steele, R. E. Trans-spliced leader addition to mRNAs in a cnidarian. Proceedings of the National Academy of Sciences of the United States of America 98, 5693–5698, https://doi.org/10.1073/pnas.101049998 (2001).
Vandenberghe, A. E., Meedel, T. H. & Hastings, K. E. M. mRNA 5 ′-leader trans-splicing in the chordates. Genes & Development 15, 294–303, https://doi.org/10.1101/gad.865401 (2001).
Zhang, H. et al. Spliced leader RNA trans-splicing in dinoflagellates. Proceedings of the National Academy of Sciences of the United States of America 104, 4618–4623, https://doi.org/10.1073/pnas.0700258104 (2007).
Bruzik, J. P., Vandoren, K., Hirsh, D. & Steitz, J. A. Trans splicing involves a novel form of small nuclear ribonuceloprotein-particles. Nature 335, 559–562, https://doi.org/10.1038/335559a0 (1988).
Hannon, G. J., Maroney, P. A., Yu, Y. T., Hannon, G. E. & Nilsen, T. W. Interaction of U6 snrna with a sequence required for function of the nematode SL RNA in transsplicing. Science 258, 1775–1780, https://doi.org/10.1126/science.1465612 (1992).
Sharp, P. A. Trans splicing: variation on a familiar theme? Cell 50, 2 (1987).
Thomas, J., Lea, K., Zuckeraprison, E. & Blumenthal, T. The spliceosomal snrnas of caenorhabditis-elegans. Nucleic Acids Research 18, 2633–2642, https://doi.org/10.1093/nar/18.9.2633 (1990).
Riedel, N., Wolin, S. & Guthrie, C. A subset of yeast snrnas contains functional binding-sites for the highly conserved sm antigen. Science 235, 328–331, https://doi.org/10.1126/science.2948278 (1987).
Spieth, J., Brooke, G., Kuersten, S., Lea, K. & Blumenthal, T. Operons in C-elegans - polycistronic messenger-RNA precursors are processed by transsplicing of SL2 to downstream coding regions. Cell 73, 521–532, https://doi.org/10.1016/0092-8674(93)90139-h (1993).
Allen, M. A., Hillier, L. W., Waterston, R. H. & Blumenthal, T. A global analysis of C. elegans trans-splicing. Genome Research 21, 255–264, https://doi.org/10.1101/gr.113811.110 (2011).
Huang, X. Y. & Hirsh, D. A. 2nd Trans-spliced RNA leader sequence in the nematode caenorhabditis-elegans. Proceedings of the National Academy of Sciences of the United States of America 86, 8640–8644, https://doi.org/10.1073/pnas.86.22.8640 (1989).
Zorio, D. A. R., Cheng, N. S. N., Blumenthal, T. & Spieth, J. Operons as a common form of chromosomal organization in C-elegans. Nature 372, 270–272, https://doi.org/10.1038/372270a0 (1994).
Tourasse, N. J., Millet, J. R. M. & Dupuy, D. Quantitative RNA-seq meta-analysis of alternative exon usage in C. elegans. Genome Research 27, 2120–2128, https://doi.org/10.1101/gr.224626.117 (2017).
Hastings, K. E. M. SL trans-splicing: easy come or easy go? Trends in Genetics 21, 240–247, https://doi.org/10.1016/j.tig.2005.02.005 (2005).
Danks, G. B. et al. Trans-Splicing and Operons in Metazoans: Translational Control in Maternally Regulated Development and Recovery from Growth Arrest. Molecular Biology and Evolution 32, 585–599, https://doi.org/10.1093/molbev/msu336 (2015).
Lall, S. et al. Contribution of trans-splicing, 5 ′-leader length, cap-poly(A) synergism, and initiation factors to nematode translation in an Ascaris suum embryo cell-free system. Journal of Biological Chemistry 279, 45573–45585, https://doi.org/10.1074/jbc.M407475200 (2004).
Maroney, P. A., Denker, J. A., Darzynkiewicz, E., Laneve, R. & Nilsen, T. W. Most messenger-rnas in the nematode Ascaris-limbricoides are trans-spliced - a role for spliced leader addition in translational efficiency. Rna-a Publication of the Rna Society 1, 714–723 (1995).
Wallace, A. et al. The Nematode Eukaryotic Translation Initiation Factor 4E/G Complex Works with a trans-Spliced Leader Stem-Loop To Enable Efficient Translation of Trimethylguanosine-Capped RNAs. Molecular and Cellular Biology 30, 1958–1970, https://doi.org/10.1128/mcb.01437-09 (2010).
Yang, Y. F. et al. Trans-splicing enhances translational efficiency in C. elegans. Genome Research 27, 1525–1535, https://doi.org/10.1101/gr.202150.115 (2017).
Pettitt, J., Mueller, B., Stansfield, I. & Connolly, B. Spliced leader trans-splicing in the nematode Trichinella spiralis uses highly polymorphic, noncanonical spliced leaders. Rna-a Publication of the Rna Society 14, 760–770, https://doi.org/10.1261/rna.948008 (2008).
Cotton, J. A. et al. The genome and life-stage specific transcriptomes of Globodera pallida elucidate key aspects of plant parasitism by a cyst nematode. Genome Biology 15, https://doi.org/10.1186/gb-2014-15-3-r43 (2014).
Mitreva, M. et al. A survey of SL1-spliced transcipts from the root-lesion nematode Pratylenchus penetrans. Molecular Genetics and Genomics 272, 138–148, https://doi.org/10.1007/s00438-004-1054-0 (2004).
Harrison, N., Kalbfleisch, A., Connolly, B., Pettitt, J. & Mueller, B. SL2-like spliced leader RNAs in the basal nematode Prionchulus punctatus: New insight into the evolution of nematode SL2 RNAs. Rna-a Publication of the Rna Society 16, 1500–1507, https://doi.org/10.1261/rna.2155010 (2010).
Ray, C., Abbott, A. G. & Hussey, R. S. Trans-splicing of a Meloidogyne incognita mRNA encoding a putative esophageal gland protein. Molecular and biochemical parasitology 68, 9, https://doi.org/10.1016/0166-6851(94)00153-7 (1994).
Takacs, A. M., Denker, J. A., Perrine, K. G., Maroney, P. A. & Nilsen, T. W. A 22-nucleotide spliced leader sequence in the human parasitic nematode Brugia-malayi is identical to the trans-spliced leader exon in Caenorhabditis-elegans. Proceedings of the National Academy of Sciences of the United States of America 85, 7932–7936, https://doi.org/10.1073/pnas.85.21.7932 (1988).
Nilsen, T. W. et al. Characterization and expression of a spliced leader RNA in the parasitic nematode Ascaris-lumbricoides var suum. Molecular and Cellular Biology 9, 3543–3547 (1989).
Goyal, K. B., Browne, J. A., Burnell, A. M. & Tunnacliffe, A. Dehydration-induced tps gene transcripts from an anhydrobiotic nematode contain novel spliced leaders and encode atypical GT-20 family proteins. Biochimie 87, 10, https://doi.org/10.1016/j.biochi.2005.01.010 (2005).
Bers, N. E. M. v. Characterization of genes coding for small hypervariable peptides in Globodera rostochiensis. Characterization of genes coding for small hypervariable peptides in Globodera rostochiensis, 228 pp (2008).
Fosu-Nyarko, J., Nicol, P., Naz, F., Gill, R. & Jones, M. G. K. Analysis of the Transcriptome of the Infective Stage of the Beet Cyst Nematode, H-schachtii. Plos One 11, https://doi.org/10.1371/journal.pone.0147511 (2016).
Masonbrink, R. E. et al. The genome of the soybean cyst nematode (Heterodera glycines) reveals complex patterns of duplications involved in the evolution of parasitism genes. https://doi.org/10.1101/391276 (2018).
Gardner, M. et al. Novel global effector mining from the transcriptome of early life stages of the soybean cyst nematode Heterodera glycines. Scientific Reports 8, https://doi.org/10.1038/s41598-018-20536-5 (2018).
Bekal, S., Niblack, T. L. & Lambert, K. N. A chorismate mutase from the soybean cyst nematode Heterodera glycines shows polymorphisms that correlate with virulence. Molecular Plant-Microbe Interactions 16, 439–446, https://doi.org/10.1094/mpmi.2003.16.5.439 (2003).
Lambert, K. N., Allen, K. D. & Sussex, I. M. Cloning and characterization of an esophageal-gland-specific chorismate mutase from the phytoparasitic nematode Meloidogyne javanica. Molecular Plant-Microbe Interactions 12, 328–336, https://doi.org/10.1094/mpmi.1999.12.4.328 (1999).
Jones, J. T. et al. Characterization of a chorismate mutase from the potato cyst nematode Globodera pallida. Molecular Plant Pathology 4, 43–50, https://doi.org/10.1046/j.1364-3703.2003.00140.x (2003).
Doyle, E. A. & Lambert, K. N. Meloidogyne javanica chorismate mutase 1 alters plant cell development. Molecular Plant-Microbe Interactions 16, 123–131, https://doi.org/10.1094/mpmi.2003.16.2.123 (2003).
Vanholme, B. et al. Structural and functional investigation of a secreted chorismate mutase from the plant-parasitic nematode Heterodera schachtii in the context of related enzymes from diverse origins. Molecular Plant Pathology 10, 189–200, https://doi.org/10.1111/j.1364-3703.2008.00521.x (2009).
Gao, B. L. et al. The parasitome of the phytonematode Heterodera glycines. Molecular Plant-Microbe Interactions 16, 720–726, https://doi.org/10.1094/mpmi.2003.16.8.720 (2003).
Lahens, N. F. et al. IVT-seq reveals extreme bias in RNA sequencing. Genome Biology 15, https://doi.org/10.1186/gb-2014-15-6-r86 (2014).
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875, https://doi.org/10.1093/bioinformatics/bti310 (2005).
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676, https://doi.org/10.1093/bioinformatics/bti610 (2005).
Mitchum, M. G. et al. Nematode effector proteins: an emerging paradigm of parasitism. New Phytologist 199, 879–894, https://doi.org/10.1111/nph.12323 (2013).
Hewezi, T. & Baum, T. J. Manipulation of Plant Cells by Cyst and Root-Knot Nematode Effectors. Molecular Plant-Microbe Interactions 26, 9–16, https://doi.org/10.1094/mpmi-05-12-0106-fi (2013).
Davis, E. L., Hussey, R. S., Mitchum, M. G. & Baum, T. J. Parasitism proteins in nematode-plant interactions. Current Opinion in Plant Biology 11, 360–366, https://doi.org/10.1016/j.pbi.2008.04.003 (2008).
Juvale, P. S. & Baum, T. J. “Cyst-ained” research into Heterodera parasitism. PLOS Pathogens 14, doi:10.1371 (2018).
Zaslaver, A., Baugh, L. R. & Sternberg, P. W. Metazoan Operons Accelerate Recovery from Growth-Arrested States. Cell 145, 981–992, https://doi.org/10.1016/j.cell.2011.05.013 (2011).
Yague-Sanz, C. & Hermand, D. SL-quant: a fast and flexible pipeline to quantify spliced leader trans-splicing events from RNA-seq data. Gigascience 7, https://doi.org/10.1093/gigascience/giy084 (2018).
Guo, Y., McK Bird, D. & Nielsen, D. M. Improved structural annotation of protein-coding genes in the Meloidogyne hapla genome using RNA-Seq. Worm 3, https://doi.org/10.4161/worm.29158 (2014).
Nilsson, D. et al. Spliced Leader Trapping Reveals Widespread Alternative Splicing Patterns in the Highly Dynamic Transcriptome of Trypanosoma brucei. Plos Pathogens 6, https://doi.org/10.1371/journal.ppat.1001037 (2010).
Boroni, M. et al. Landscape of the spliced leader trans-splicing mechanism in Schistosoma mansoni. Scientific Reports 8, https://doi.org/10.1038/s41598-018-22093-3 (2018).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. Journal of Molecular Biology 215, 403–410, https://doi.org/10.1006/jmbi.1990.9999 (1990).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079, https://doi.org/10.1093/bioinformatics/btp352 (2009).
Thomas, J. D., Conrad, R. C. & Blumenthal, T. The C-elegans trans-spliced leader RNA is bound to sm and has a trimethylguanosine cap. Cell 54, 533–539, https://doi.org/10.1016/0092-8674(88)90075-x (1988).
de Boer, J. M. et al. Production and characterization of monoclonal antibodies to antigens from second stage juveniles of the potato cyst nematode, Globodera rostochiensis. Fundamental and Applied Nematology 19, 545–554 (1996).
NCBI. (Expressed Sequence Tags [Internet] Bethesda (MD): National Library of Medicine (US), National Center for Biotechnology Information; https://www.ncbi.nlm.nih.gov/nucest/?term = heterodera + glycines).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842, https://doi.org/10.1093/bioinformatics/btq033 (2010).
N. A. & J. N. F. Sickle: A sliding-window, adaptive, quality-based trimming tool for FastQ files (Version 1.33) [Software], http://github.com/najosh/sickle.
Conesa, A. & Gotz, S. Blast2GO: A comprehensive suite for functional analysis in plant genomics. International journal of plant genomics 2008, 619832–619832, https://doi.org/10.1155/2008/619832 (2008).
Bauer, S., Grossmann, S., Vingron, M. & Robinson, P. N. Ontologizer 2.0 - a multifunctional tool for GO term enrichment analysis and data exploration. Bioinformatics 24, 1650–1651, https://doi.org/10.1093/bioinformatics/btn250 (2008).
Smit, A. & Huxley, R. RepeatModeler Open-1.0. 2008–2015 http://www.repeatmasker.org.
Acknowledgements
This is a publication of the Iowa Agriculture and Home Economics Experiment Station, Ames, IA, supported by Hatch Act and State of Iowa funds. This work was supported by funds from the Iowa Soybean Association and the United States Department of Agriculture NIFA award 2015-67013-2351.
Author information
Authors and Affiliations
Contributions
S.B., R.M., T.M. and T.B. conceptualized this work. S.B., R.M., T.M. Arun. S. and Anoop. S. designed and performed the research. All authors contributed to writing and reviewing the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Barnes, S.N., Masonbrink, R.E., Maier, T.R. et al. Heterodera glycines utilizes promiscuous spliced leaders and demonstrates a unique preference for a species-specific spliced leader over C. elegans SL1. Sci Rep 9, 1356 (2019). https://doi.org/10.1038/s41598-018-37857-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-37857-0
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.