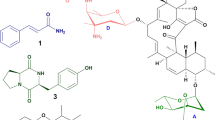
Abstract
Actinobacteria, a group of gram-positive bacteria, can produce plenty of valuable bioactive secondary metabolites, especially antibiotics. Hence, in order to search for new actinobacteria, actinobacterial isolates were obtained from rhizosphere soil collected from the Futian mangrove ecosystem in Shenzhen, China. According to 16S rRNA sequences, 14 actinobacterial strains of the genus Streptomyces, Rhodococcus, Microbacterium, Micromonospora, Actinoplanes and Mycobacterium were isolated and identified. Among these, strain Mycobacterium sp.13 was described as a potential new species belonging to the genus Mycobacterium within the class of actinobacteria according to the genomic analysis. The genome-based 16S rRNA sequences had 98.48% sequence similarity with Mycobacterium moriokaense DSM 44221T. Meanwhile, the genome sequences of Mycobacterium sp.13 showed an average nucleotide identity (ANI) with the Mycobacterium mageritense DSM 44476, Mycobacterium smegmatis MKD8 and Mycobacterium goodii strain X7B of only 74.79%, 76.12% and 76.42%, respectively. Furthermore, genome-mining results showed that Mycobacterium sp.13 contained 105 gene clusters encoding to the secondary metabolite biosynthesis, where many kinds of terpene, bacteriocin, T1pks, Nrps, saccharide, fatty acid, butyrolactone, ectoine and resorcinol were included. Finally, through LC-MS and HR-MS, analyzing the small molecules from ethyl acetate extract of this strain, asukamycin C and apramycin were for the first time found present to be in Mycobacterium moriokaense strain. Our study provides evidence in support of the potential new Mycobacterium sp.13 isolated from the mangrove environment as a possible novel source of natural products.
Similar content being viewed by others
Introduction
Natural products, including drugs, are clearly considered to occupy larger and distinct chemical spaces compared to combinatorial chemicals1. Natural products are derived from microorganisms, plants or animals2,3. In all known small molecule natural products producers, the microbial community represent a major source of biologically active secondary metabolites4. As we know, many important natural products isolated from bacteria that becoming potential leads for the human diseases treatment have been identified5. To date, about 20,000 secondary metabolites produced from microorganisms have been discovered6. Moreover, recent reports of new chemical entities and first-in-class drug candidates were frequently derived from actinobacterial strains7,8. As we know, Streptomyces is by far the most prolific genus, which produced about 80% of all described antibiotics9. However, the rare actinobacterial genus of Mycobacterium are little known to produce secondary metabolites10. Several mycobacteria, namely Mycobacterium tuberculosis and Mycoabcterium smegmatis, are only known to produce some sulfated compounds11. It is therefore, deducing the secondary metabolites producing ability by rare actinobacterial genus will require a substantial effort.
Increasing attention has already been placed on new or extreme environments for the exploration of novel natural compounds, such as marine12, desert13 and mangrove environments14. Mangroves are an ecosystem with high moisture, high salinity and tolerance to oxygen15. Many new types of microorganisms have been discovered from mangrove ecosystems, which have always been a natural source of secondary metabolites with biologically active. For example, A novel actinobacterial strain was isolated by Ruan et al. (2015) from mangrove soil in Thailand belonging to the genus Streptomyces16. A novel Jiangella strain was obtained by Suksaard et al. (2015) from a mangrove soil in 201517. As we know, microbial community are valuable and rich sources of structurally peerless bioactive natural products18. In recent years, studies have obtained compounds from mangrove actinobacterial strains with a unique structure and potential medicinal applications. Such as, a novel analogue of 1-isoquinolone, described as marinamide (A), and its methyl ester (B), were isolated from two endophytic fungi19. More recently, two novel indolocarbazoles isolated from a mangrove-derived Streptomyces showed unprecedented structure of cyclic N-glycosidic bond between 1,3-carbon atoms within the glycosyl moiety and two indole nitrogen atoms in the indolocarbazole with enhanced anticancer activity20. Therefore, the mangrove ecosystem can be regarded as a new and diverse source of natural products that remains untapped.
The recent surge in genome-wide sequencing projects has once again yielded insight into the production of secondary metabolites, where the genetic data showed that the metabolic potential of these natural resources was seriously underestimated21. Thus, a promising method for drug discovery is based on combining genome mining analysis in genomic sequences to reveal the silent pathway for natural products22. The most widely studied model strain of Streptomyces coelicolor had been shown to contain more than 20 gene clusters involved with potential secondary metabolites genes in the genome23. With the intensification of genome sequencing and metabolite analysis efforts, the metabolic function of other actinobacterial taxa is gradually being explored. Recent progress in several rare actinobacterial genus had suggested that it had the same potential of secondary metabolites production as Streptomyces24. The complete genome of a marine actinobacterial species, Salinispora tropica, was released during 2007, which identified 15 secondary metabolite gene clusters, including the Salinosporamide A gene cluster, a potent anticancer agent25. McyLeod et al. (2016) found the genome of Rhodococcus sp RHA1 contained 24 non ribosomal peptide synthase genes and 7 polyketide synthase genes, in which, 6 genes exceed 25k bp, provided evidence that this strain harbored an extensive pathway of secondary metabolism26. All of these findings showed that the biosynthetic capability of natural products from microorganisms had been greatly underestimated by traditional methods of bioassay-guided natural product discovery27. Thus, a better understanding of the whole bacterial genome is the key to successfully identifying new natural products.
Accordingly, the aims of this research were (1) to obtain novel actinobacterial isolates from mangrove ecosystem in Futian National Reserve, Shenzhen by a plate culture method, and to characterize the isolates based on the 16S rRNA gene sequences; (2) to identify the biosynthetic gene cluster involved with antibiotic production by whole-genome sequencing; and (3) to determine small molecule metabolites in fermentation products of the isolated strain by LC-MS and HR-MS. The results of this study should provide reference data informing the discovery of major antibiotics producing strain for further research, which can also bring benefit to the pharmaceuticals industry.
Materials and Methods
Materials
All chemical reagents were purchased from Sigma Aldrich (USA). TIANamp Bacteria DNA Kit was purchased from TIANGEN Biotech (Beijing) Co., Ltd.
Isolation medias were used in this study including ISP media 2 (yeast extract, 4.0 g; dextrose, 4.0 g; malt extract, 10.0 g; agar, 20.0 g; distilled water 1 L, pH 7.2)28, ISP media 4 (starch, 10.0 g; NaCl, 1.0 g; K2PHO4, 1.0 g; (NH4)2SO4, 2.0 g; FeSO4.7H2O, 0.001 g; CaCO3, 2.0 g; MnCl2.4H2O, 0.001 g; ZnSO4.7H2O, 0.001 g; agar, 20.0 g; distilled water 1 L, pH 7.2)29, Gauze No. 1 (soluble starch, 20.0 g; ferrous sulfate, 0.01 g; sodium chloride, 0.5 g; potassium nitrate, 1.0 g; magnesium sulfate, 0.5 g; dipotassium hydrogen phosphate, 0.5 g; agar, 15.0 g; distilled water 1 L, pH 7.2)30, nutrient agar (peptone, 10.0 g; sodium chloride, 5.0 g; beef extract, 3.0 g; agar, 15.0 g; distilled water 1 L, pH 7.2)31, Czapek’s medium (sodium nitrate, 3.0 g; magnesium sulfate, 0.5 g; dipotassium hydrogen phosphate, 1.0 g; potassium chloride, 0.5 g; sucrose, 30.0 g; ferrous sulfate, 0.01 g; agar, 15.0 g; distilled water 1 L, pH 7.2)32 and halothiobacillus HL2 medium (glucose, 10.0 g; tryptone, 3.0 g; peptone, 5.0 g; NaCl, 5.0 g; agar, 20.0 g; distilled water 1 L, pH 7.2)33.
Environmental sampling
The Futian National Nature Reserve, is situated at 22°32′ N and 114°05′ E and covers an area of about 369 ha, Shenzhen, the People’s Republic of China, Vegetation types include semi-mangroves, mangroves and seashore plants, with Kandelia candel and Aegiceras corniculatum being dominant in the Nature Reserve34. Samples were collected from rhizosphere soil, related to Aegiceras corniculatum and Kandelia candel, in August 2017. Collected soil samples were immediately placed in a sterile plastic bags and then transported them to the laboratory.
Selective isolation procedures and medium
Air dried soil samples were firstly sieved to exclude organic matter particles and large mineral, then grounded in a mortar and pestle. Selective soil samples pretreatment method including dry heat (120 °C, 60 min) and phenol (1.5%, 30 min at 30 °C)35. The pretreated soil samples were serially diluted with steriziled water down to 10–4. Then, 100 μL amounts of the 10−4 suspensions were spread onto selected isolation medium. The culture-plate procedure was designed to isolate pure cultures of bacteria and conducted according to previous publication by Hu et al.36. Briefly, suspensions of each sample were diluted and spreaded onto six different kinds of isolation medium: ISP media 2, ISP media 4, Gauze No. 1, nutrient agar, Czapek’s medium and halothiobacillus HL2 medium. All isolation media were cultured and stored for future use.
Extraction of DNA and PCR amplification
DNA was extracted directly from single colony to identify the species. The procedures of DNA extraction and PCR amplification were followed previous publication by Hu et al.36. DNA was extracted from the each single colony and used as the DNA template to do the PCR amplification37. The detailed PCR reaction for 16S rRNA amplification using the universal bacteria primer pair 27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492R (5′-TACGGCTACCTTGTTACGACTT-3′)38. The final products of PCR amplification were subsequently assessed in agarose gel by gel electrophoresis. Sanger sequencing platform was applied to sequence the products. The 16S rRNA sequences were compared with EzBioCloud database to determine the species (https://www.ezbiocloud.net/).
Genome sequencing, assembly and annotation
The procedures of genomic DNA extraction, whole-genome sequencing and subsequently analysis were performed according to previous publication by Hu et al.36. Firstly, genomic DNA was extracted to construct the genomic DNA library, which was sequenced on the Illumina NovaSeq HiSeq. 4000. Then, genome assembly, annotation and gene prediction were performed by SOAPdenovo, idba and gmhmmp, respectively. Transfer RNAs (tRNAs) and ribosomal RNAs (rRNAs) were predicted by tRNAscan-SE and rnammer. All the protein-encoding genes were annotated by searching against the non-redundant protein sequence (NR) database of the National Center for Biotechnology Information (NCBI) using BLAST. Finally, the database of Kyoto Encyclopedia of Genes and Genomes (KEGG) was provided to assign the functional categories of the query sequences.
Phylogenetic analysis and genome mining
The evolutionary relationships among different actinobacterial species were analyzed by Mega 7.0. 16S rRNA sequences of Mycobacterium sp.13 was retrieved from the genomic annotation results. In our study, we selected eight relevant mycobacteria species to construct the phylogenetic tree, whose sequences were downloaded from the NCBI database. An additional strain, Nocardia abscessus strain IMMIB D-1592 (Biosample accession: NR 025059.1) was used as the phylogenetic root39.
These species are Mycobacterium moriokaense strain CIP 105393 (Biosample accession: NR115331.1), Mycobacterium goodii strain ATCC 700504 (Biosample accession: AY457079.1), Mycobacterium smegmatis strain ATCC 19420 (Biosample accession: AY457078.1), Mycobacterium mageritense strain DSM 44476 (Biosample accession: AY457076.1), Mycobacterium avium strain JCM 15429 (Biosample accession: LC020093.1), Mycobacterium ulcerans strain ATCC 19423 (Biosample accession: NR113138.1), Mycobacterium bovis strain GTC 602 (Biosample accession: AB292583.1) and Mycobacterium tuberculosis strain UKR100 (Biosample accession: MG995565.1).
The online software of anti-SMASH was applied to mine the secondary metabolite biosynthesis gene clusters and predict the small molecule compounds production (https://antismash.secondarymetabolites.org/#!/about), which followed previous publication by Hu et al.36.
Preparation of crude extract and LC-MS analysis
The procedures of crude extract preparation and subsequent MS analysis according to the previous work by Hu et al.36. Firstly, the purified culture was processed the fermentation culture following by the extraction with ethyl acetate. Then, MS/MS analysis was performed on a 4000 Q TRAP LC/MS/MS system. Furthermore, a high accurate mass spectrometric analysis was performed on a LTQ-Orbitrap mass spectrometer XL MS equipped with an ESI mode and operated in the positive ion. The mass accuracy of the instrument lower than 3 ppm. Full scan data was acquired from 500 to 600 m/z under the same condition.
Nucleotide sequence accession numbers
The sequences of Mycobacterium sp.13 generated in this study have been deposited with the GenBank database under the accession number QQBJ00000000 for bacterial whole genome genes.
Results
Diversity of actinobacterial strains by 16S rRNA analysis
In total, 39 bacterial isolates were obtained from soil samples, 14 of which were related to the filamentous bacteria belonging to the class of actinobacteria (Supplementary Table S1 and Table 1). Isolates were identified by PCR and sequencing of 16S rRNA gene sequences, it turned out that they were classified into six genus in the class of actinobacteria, composing of Micromonospora, Streptomyces, Mycobacterium, Microbacterium, Rhodococcus and Actinoallomurus. Actinobacterial species were mainly isolated from five types of isolation media, e.i. ISP2, Nutrient agar, ISP4, HL2 and Gauze No. 1. Due to low 16S sequence identity with the nearest type strain (97.00%), one actinobacterial strain of Mycobacterium sp.13 was selected for further genomic analysis.
Genome mining and annotation
One of the isolated strains, Mycobacterium sp.13, the genome-based 16S rRNA gene isolated from strain, had 98.48% similarity to the sequence of Mycobacterium moriokaense DSM 44221T. Whole-genome sequencing of Mycobacterium sp.13 generated a total of 12,385,360 sequence reads, yielding approximately 268 scaffolds (Table 2). SOAPdenovo was used to do the de novo assembly, resulted in a total of 7.2 Mb (66.95% G+C content) distributed within one main scaffold having an average length of 26,880 bp. The assembled genome sequences were compared with the KEGG and NR databases to process the procedure of annotation. Within the Mycobacterium sp.13, a total of 7,144 protein-encoding genes were contained in the genome, 4 rRNA and 50 tRNA were predicted, the coding density was about 92.07% and the average CDS length was 928 bp. Strain Mycobacterium sp.13 also presented with low sequence similarity to other Mycobacterium strains, but these strains do not have genomic data within the NCBI database except for Mycobacterium mageritense DSM 44476, Mycobacterium smegmatis MKD8 and Mycobacterium goodii strain X7B. Therefore, these three strains were selected to calculate the average nucleotide identity (ANI), the value of which with the Mycobacterium mageritense DSM 44476, Mycobacterium smegmatis MKD8 and Mycobacterium goodii strain X7B was only 74.79%, 76.12% and 76.42%, respectively. The ANI result indicated that the isolate was a potential novel strain within the genus Mycobacterium, based on the genomic data.
Phylogenetic analysis
A phylogenetic analysis was performed to assess the evolutionary relationship among different bacterial strains. In the tree, eight 16S sequences from the genus of Mycobacterium were compared. Phylogenetic analysis of Mycobacterium sp.13 based on 16S rRNA sequences showed that this potential novel species occupied a unique position at the genus level (Fig. 1). It was on a single branch, which was relatively close to Mycobacterium goodii strain ATCC 700504 and Mycobacterium smegmatis strain ATCC 19420. This suggested that Mycobacterium sp.13 had a close genetic relationship with them. The isolated Mycobacterium sp.13 was also close to Mycobacterium mageritense strain DSM 44476 and Mycobacterium moriokaense strain CIP 105393, but was relatively distant from Mycobacterium bovis strain GTC 602, Mycobacterium avium strain JCM 15429, Mycobacterium ulcerans strain ATCC 19423 and Mycobacterium tuberculosis strain UKR100.
Comparison of eight 16S rRNA sequences from the genus of Mycobacterium with other orthologous sequences. Complete 16S rRNA sequence of the Mycobacterium sp.13 was extracted from the genome data. The tree is rooted with Nocardia abscessus strain IMMIBD-1592. The neighbor-joining method was used to construct the phylogenetic tree. The number of bootstrap replications was set to 1000.
Detection of gene clusters involved in antibiotics and secondary metabolites
A total of 105 biosynthetic gene clusters related to the secondary metabolite production were identified in the genome of Mycobacterium sp.13 (Supplementary Table S2), which were predicted for T1pks, terpene, Nrps, fatty acid, bacteriocin, ectoine, butyrolactone, saccharide, resorcinol and other products, in which 6 gene clusters showed similarity exceeding 20%. The analysis showed that at least 10 kinds of PKS and NRPS gene clusters can be observed in the Mycobacterium sp.13 genome, 3 of which had no similarity with any known gene cluster and the core structures of these clusters were identified (Table 3). In addition, seven of the biosynthetic gene clusters were encoded to antibiotics compounds, namely sch47554/sch47555, azinomycin B, apramycin, maklamicin, u-68204, nosiheptode and caprazamycin (Table 4).
The antibiotic of the maklamicin biosynthesis gene cluster of Mycobacterium sp.13, contained 20 ORFs, and three catalytic domains related to the polyketide synthase (Supplementary Fig. 1). In the genome of Mycobacterium sp.13, core secondary biosynthetic genes were related to AMP-dependent synthase and ligase, phosphopantetheine-binding domain-containing protein and thioesterase, additional biosynthetic genes were annotated to aminotransferase class V, GCN5-related N-acetyltransferase, pyruvate oxidase/decarboxylase and monooxygenase FAD-binding; some regulatory and transport-related genes can also be observed in the Mycobacterium sp.13 genome. In addition, maklamicin biosynthesis clusters of Mycobacterium sp.13 had 4% homology to the existing cluster of Micromonospora sp GMKU326. This may play an important role in the synthesis of the maklamicin-derived primer unit, which meant that these clusters may be derived from donor microorganisms via horizontal gene transfer. Another PKS-type biosynthetic gene cluster contained 42 ORFs and four domains related to antibiotic sch47554/sch47555 production (Supplementary Fig. 2). In sch47554/sch47555-like gene cluster, core and additional biosynthetic genes were encoded to rhodanese domain-containing protein, AMP-dependent synthetase and ligase, MMPL domain-containing transport protein, beta-ketoacyl synthase, phosphopantetheine-binding domain-containing protein, acyl carrier protein, 3-oxoacyl, o-methyltransferase, monooxygenase FAD-binding, phosphoglycerate mutase, isochorismate synthase, argininosuccinate lyase/adenylosuccinate lyase and alpha/beta hydrolase fold protein. Sch47554/sch47555 biosynthetic gene cluster was also constituted of considerable proteins with unknown function. Therefore, Mycobacterium sp.13 had great potential for producing an analogue of sch47554/sch47555.
In addition to the analysis of PKS type cluster, the fatty acid type gene clusters responsible for antibiotic synthase were found in the genome of Mycobacterium sp.13. The first fatty acid gene cluster related to caprazamycin-like compound production was consisted of 30 ORFs, 3 of which encoding key biosynthetic genes including 3-oxoacyl and NAD-dependent epimerase/dehydratase (Supplementary Fig. 3). This cluster of Mycobacterium sp.13 had different biosynthetic enzymes compared with the most-similar known cluster in the genome of Streptomyces sp MK730-62F2, which indicated that an unknown compound or caprazamycin analog might can be discovered from the new strain isolated from mangrove environment. Another fatty acid pathway was annotated to synthesis of U-68204 (Supplementary Fig. 4). It is worth noting that numerous genes in the U-68204-like gene cluster cannot be annotated. The core biosynthetic enzymes were related to malonyl CoA-acyl carrier protein transacylase, beta-ketoacyl synthase and putative acyl carrier protein.
Besides the fatty acid and PKS type gene clusters, other putative gene clusters can be found in the genome of Mycobacterium sp.13, which were also responsible for antibiotic production including of nosiheptode, apramycin and azinomycin B. The nosiheptode biosynthesis gene cluster of Mycobacterium sp.13 contained 24 ORFs (Supplementary Fig. 5). However, this cluster of Mycobacterium sp.13 had more annotated functional known biosynthetic genes involved with the antibiotics production than the Streptomyces actuosus cluster. In the genome of Mycobacterium sp.13, core biosynthetic genes were related to the putative esterase and glycosyltransferase. Furthermore, biosynthetic gene clusters of apramycin and azinomycin B were discovered in Mycobacterium sp.13 (Supplementary Figs 6 and 7). In the azinomycin biosynthetic gene cluster, the core genes were related to the synthase of haloalkane dehalogenase, oxidoreductase and short-chain dehydrogenase/reductase SDR. No core biosynthetic genes were observed in the genome responsible for apramycin biosynthesis.
Detection of secondary metabolites produced by Mycobacterium sp.13
To test the results of genome-guided secondary metabolites prediction, a small-molecule profile was established. Mass spectral analysis were performed in positive mode. According to the mass-to-charge ratio of molecular ions, the metabolic substances in the fermentation broth were identified from the UPLC-MS profiles. After that, obtained profiles of secondary metabolites by mass spectrometry were compared with the genome mining data. As thus, two predicted secondary metabolites were detected in the fermentation product of Mycobacterium sp.13: asukamycin C (cluster 25, m/z = 521.5[M + H]+)40 and apramycin (cluster 39, m/z = 540.4 [M + H]+)41 (Fig. 2A). Corresponding fractions were also observed, in-source fragments at m/z 217.1 correspond to apramycin (Fig. 2B) and m/z 163.2 correspond to asukamycin C (Fig. 2C) were identified. Furthermore, the accurate identification of the compounds was conducted by HR-MS with high mass accuracy. The theoretical exact mass in positive mode was calculated by ChemDraw used as the reference in our study. The theoretical exact mass of MH+ (m/z) of asukamycin C and apramycin was 521.22878 (C29H33N2O7+) and 540.28809 (C21H42N5O11+), respectively. The measured mass using HR-MS was 521.25092 and 540.22180, which was shown in Fig. 3.
LC-MS analysis of ethyl acetate (EA) extract of asukamycin C and apramycin in the fermentation broth of Mycobacterium sp.13. The apramycin ([M + H]+ at m/z 540.4 (A), in-source fragment at m/z 217.1 (B)) and asukamycin C ([M + H]+ at m/z 521.5 (A), in-source fragment at m/z 163.2 (C)) were identified.
Discussion
Currently, tremendous research interest is concentrated on the isolation and identification of new actinobacterial species from unique and extreme environmental surroundings, as the screening of these microorganisms has increased the prospect of exploring novel natural products that could be exploited as biotechnological resources42,43. Such as, four hundred and forty eight actinobacterial strains were isolated by Suksaard et al. (2017) from water and sediment samples of mangroves and assessed the plant growth promoting potential of them44. Ruttanasutja et al. (2015) designed different isolation method to isolate diverse actinobacterial strains, including five pretreatments process, three enrichment media and fifteen selective media45. For this purpose, a mangrove forest was selected as the sampling site for this study, which was located in the regions along estuaries with tropical and subtropical climate and proved to be the driving force of bacterial diversity46. In our study, the soil used in the experiment were collected from Futian mangrove ecosystem in Shenzhen, China, which located in a tidal swamp of tropical delta. The 16S rRNA sequences were amplified by universal primers and then sequenced via Sanger sequencing platform to identify the species47. The isolation results indicated that 39 bacterial isolates were obtained from rhizosphere soil samples, 14 isolates of which were identified as the actinobacterial strains belonging to six genera: Streptomyces, Micromonospora, Mycobacterium, Microbacterium, Actinoallomurus and Rhodococcus. Among them, one Mycobacterium sp.13 only presented 97.00% 16S sequence similarity with Mycobacterium moriokaense. The 16S rRNA gene can be used for species classification as it is a highly conserved gene; the application of the 98.65% threshold could strictly classify bacterial community into different species depending on the 16S rRNA gene copy analyzed48. Thus, Mycobacterium sp.13 was selected to perform the whole genome sequencing. The complete 16S rRNA sequences obtained from the genome had 98.48% sequence similarity with Mycobacterium moriokaense DSM 44221T. Therefore, this isolate could be regarded as a potential new actinobacterial strain and was subjected to further genome mining. The genome sequence of the potential new actinobacterial strain showed an ANI with the Mycobacterium mageritense DSM 44476, Mycobacterium smegmatis MKD8 and Mycobacterium goodii strain X7B of only 74.79%, 76.12% and 76.42%, respectively. Among the available indices of genome-relatedness, the ANI value between two bacterial genome sequences is one of the most robust measurements of genome correlations and has great potential as a compensation for the DNA–DNA hybridization (DDH) technique in the classification of bacteria and archaea49. The ANI threshold range for species delineation (95–96%) has been determined based on a comparative study between the DDH and ANI values50. On account of genome-based results, Mycobacterium sp.13 can be classified as a potential novel species within the genus of Mycobacterium.
Previous efforts in genomic analysis have proven that actinobacterial strain isolated from mangrove environment could have a greater gene clusters involved in multiple secondary metabolic pathway compared with normal strains36. This observation has led us to explore this potential novel species isolated from mangrove which should offer a higher probability of discovering novel compounds than previously screened strains. The results of genome mining showed that Mycobacterium sp.13 had the capacity to produce such diverse antibiotics compounds, such as sch47554/sch47555, azinomycin B, apramycin, maklamicin, u-68204, nosiheptode and caprazamycin. The identification of biosynthetic gene cluster in Mycobacterium sp.13 genome revealed that a total of 105 gene clusters were related to the secondary metabolites production, which was significantly higher than other actinobacterial model strain. To date, roughly more than 2,000 actinobacterial strains genomes have been completed and annotated; these have been deposited in the database of NCBI. Homology searching for genes encoding the known or putative gene clusters responsible for secondary metabolite production revealed that Streptomyces coelicolor A3(2) predicted a further 17 gene clusters related to secondary metabolism23; Streptomyces avermitilis MA-4680 harbored 37 secondary metabolic gene clusters51, and the genome of Saccharopolyspora erythraea NRRL2338 suggested at least 27 biosynthesis genes involved with a various of secondary metabolites of mostly unknown composition of chemical compounds52. As noted before, the best-known Mycobacterium species in previous reports are pathogenic; there have been almost no reports of extensive antibiotics being derived from it, even belonging to the actinobacterial taxon53,54,55. In our study, the Mycobacterium sp.13 genome had rich genetic potential for secondary metabolite production; at least seven different kinds of antibiotic compound were found by the biosynthetic gene cluster prediction. In addition, there were many putative gene clusters in the genome involving secondary metabolites, and many predicted secondary metabolites were previously found to be isolated from the non-Mycobacterium strain. For example, the gene organization of polyketide azinomycin B was quite similar to that in Streptomyces sahachiroi56. The other predicted antibiotics of sch47554/sch47555, apramycin, maklamicin, u-68204, nosiheptode and caprazamycin were commonly found in Streptomyces sp SCC 2136, Streptoalloteichus hindustanus DSM 44523T, Micromonospora sp GMKU326, Streptomyces sp MG11, Streptomyces actuosus and Streptomyces sp MK730-62F2, respectively57,58,59,60,61. These observations indicated that this new strain had great potential for biosynthesis of diverse secondary metabolites, likely via different synthetic pathways compared to other known Mycobacterium species. All these observations proved that a complex and diverse secondary metabolome existed within the Mycobacterium strain genome. Genome mining efforts in our study indicated that the capability of Mycobacterium to produce secondary metabolites had been underestimated.
With the availability of the genome sequences of Mycobacterium sp.13, we are now in a position to address the possible secondary metabolites. Based on this, analysis of fermentation product of the Mycobacterium sp.13 can led to identification of the secondary metabolites asukamycin C and apramycin, which will further validate the genome mining prediction results. Asukamycin was isolated from the fermentation culture broth of a Streptomyces designated as Streptomyces nodosus subsp. Asukaensis in 197662, which was a member of a potential antitumor agent within the manumycin family of metabolites63. In addition, apramycin, an aminocyclitol antibiotic complex produced by Streptomyces tenebrarius, was first reported in 196764. In vitro experiment proved that apramycin can act against multidrug-resistant Pseudomonas aeruginosa and Acinetobacter baumannii65. To our knowledge, this study represented a first record of asukamycin C and apramycin being produced from the Mycobacterium moriokaense strain. In fact, few secondary metabolites have been described as produced by strains belonging to the genus of Mycobacterium, because the cell wall of them is shown to be an effective permeability barrier to hydrophilic compounds66. On the other hand, the expression of secondary metabolites gene clusters is regulated by many different protein families, and led to a range of secondary metabolites with varying concentrations67. Thus, a step in this direction is to develop a method to identify the secondary metabolites that share common structural features68. Although our study does not provide the absolute information on identifying the complete structure of the metabolites, some of fragments of metabolites can be observed by MS/MS spectrum. Therefore, an integrative genome- and LC-MS-based metabolomics strategy could facilitate the screening of natural products from actinobacterial strain.
References
Bauer, A. & Bronstrup, M. Industrial natural product chemistry for drug discovery and development. Nat Prod Rep 31, 35–60, https://doi.org/10.1039/c3np70058e (2014).
Zhang, A., Sun, H. & Wang, X. Recent advances in natural products from plants for treatment of liver diseases. Eur J Med Chem 63, 570–577, https://doi.org/10.1016/j.ejmech.2012.12.062 (2013).
Ngo, L. T., Okogun, J. I. & Folk, W. R. 21st century natural product research and drug development and traditional medicines. Nat Prod Rep 30, 584–592, https://doi.org/10.1039/c3np20120a (2013).
Xiong, Z. Q., Wang, J. F., Hao, Y. Y. & Wang, Y. Recent advances in the discovery and development of marine microbial natural products. Mar Drugs 11, 700–717, https://doi.org/10.3390/md11030700 (2013).
Shen, B. A new golden age of natural products drug discovery. Cell 163, 1297–1300, https://doi.org/10.1016/j.cell.2015.11.031 (2015).
Choi, S.-S. et al. Genome engineering for microbial natural product discovery. Curr Opin Microbiol 45, 53–60 (2018).
Fu, P. & MacMillan, J. B. Spithioneines A and B, two new bohemamine derivatives possessing ergothioneine moiety from a marine-derived Streptomyces spinoverrucosus. Org Lett 17, 3046–3049, https://doi.org/10.1021/acs.orglett.5b01328 (2015).
Fu, P. & MacMillan, J. B. Thiasporines A-C, thiazine and thiazole derivatives from a marine-derived Actinomycetospora chlora. J Nat Prod 78, 548–551, https://doi.org/10.1021/np500929z (2015).
Toumatia, O. et al. Antifungal properties of an actinomycin D‐producing strain, Streptomyces sp IA1, isolated from a Saharan soil. J Basic Microbiol 55, 221–228 (2015).
Lau, S. K. et al. Identification of specific metabolites in culture supernatant of Mycobacterium tuberculosis using metabolomics: exploration of potential biomarkers. Emerg Microbes Infect 4, e6 (2015).
Walsh, C. T. & Wencewicz, T. A. Prospects for new antibiotics: a molecule-centered perspective. J. Antibiot 67, 7–22, https://doi.org/10.1038/ja.2013.49 (2014).
Kiuru, P. et al. Exploring marine resources for bioactive compounds. Planta Med 80, 1234–1246, https://doi.org/10.1055/s-0034-1383001 (2014).
Mohammadipanah, F. & Wink, J. Actinobacteria from arid and desert habitats: diversity and biological activity. Front Microbiol 6, https://doi.org/10.3389/Fmicb.2015.01541 (2016).
Mohamed, H. et al. Isolation and characterization of actinobacteria from Algerian Sahara Soils with antimicrobial activities. Int J Mol Cell Med 6, 109–120, https://doi.org/10.22088/acadpub.BUMS.6.2.5 (2017).
Friess, D. A. Mangrove forests. Curr Biol 26, R746–748, https://doi.org/10.1016/j.cub.2016.04.004 (2016).
Ruan, C.-y et al. Streptomyces ferrugineus sp nov., isolated from mangrove soil in Thailand. Antonie van Leeuwenhoek 107, 39–45 (2015).
Suksaard, P. et al. Jiangella mangrovi sp nov., isolated from mangrove soil. Int J Syst Ecol Microbiol 65, 2569–2573 (2015).
Rashad, F. M., Fathy, H. M., El-Zayat, A. S. & Elghonaimy, A. M. Isolation and characterization of multifunctional Streptomyces species with antimicrobial, nematicidal and phytohormone activities from marine environments in Egypt. Microbiol Res 175, 34–47, https://doi.org/10.1016/j.micres.2015.03.002 (2015).
Zhu, F. & Lin, Y. C. Marinamide, a novel alkaloid and its methyl ester produced by the application of mixed fermentation technique to two mangrove endophytic fungi from the South China Sea. Chinese Sci Bull 51, 1426–1430, https://doi.org/10.1007/s11434-006-1426-4 (2006).
Fu, P. et al. Streptocarbazoles A and B, two novel indolocarbazoles from the marine-derived actinomycete strain Streptomyces sp FMA. Org Lett 14, 2422–2425, https://doi.org/10.1021/ol3008638 (2012).
Hidalgo, P. I. et al. Molecular characterization of the PR-toxin gene cluster in Penicillium roqueforti and Penicillium chrysogenum: cross talk of secondary metabolite pathways. Fungal Genet Biol 62, 11–24, https://doi.org/10.1016/j.fgb.2013.10.009 (2014).
Wu, C., Du, C., Ichinose, K., Choi, Y. H. & van Wezel, G. P. Discovery of C-Glycosylpyranonaphthoquinones in Streptomyces sp MBT76 by a combined NMR-Based metabolomics and bioinformatics workflow. J Nat Prod 80, 269–277, https://doi.org/10.1021/acs.jnatprod.6b00478 (2017).
Bentley, S. D. et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417, 141–147, https://doi.org/10.1038/417141a (2002).
Tiwari, K. & Gupta, R. K. Rare actinomycetes: a potential storehouse for novel antibiotics. Crit Rev Biotechnol 32, 108–132, https://doi.org/10.3109/07388551.2011.562482 (2012).
Udwary, D. W. et al. Genome sequencing reveals complex secondary metabolome in the marine actinomycete Salinispora tropica. P Natl Acad Sci USA 104, 10376–10381, https://doi.org/10.1073/pnas.0700962104 (2007).
McLeod, M. P. et al. The complete genome of Rhodococcus sp RHA1 provides insights into a catabolic powerhouse. P Natl Acad Sci USA 103, 15582–15587, https://doi.org/10.1073/pnas.0607048103 (2006).
Kalaitzis, J. A., Ingrey, S. D., Chau, R., Simon, Y. & Neilan, B. A. Genome-guided discovery of natural products and biosynthetic pathways from Australia’s untapped microbial megadiversity. Aust J Chem 69, 129–135, https://doi.org/10.1071/CH15601 (2016).
Tokuyama, S., Hatano, K. & Takahashi, T. Discovery of a novel enzyme, N-Acylamino acid racemase in an actinomycete - screening, isolation, and identification. Biosci Biotech Biochem 58, 24–27, https://doi.org/10.1271/Bbb.58.24 (1994).
Carpenterboggs, L., Loynachan, T. E. & Stahl, P. D. Spore germination of gigaspora-margarita stimulated by volatiles of soil-isolated actinomycetes. Soil Biol Biochem 27, 1445–1451, https://doi.org/10.1016/0038-0717(95)00075-P (1995).
Zakharova, O. S., Zenova, G. M. & Zvyagintsev, D. G. Some approaches to the selective isolation of actinomycetes of the genus Actinomadura from soil. Microbiology 72, 110–113, https://doi.org/10.1023/A:1022294526830 (2003).
Hu, R. M., Cheng, L. & Wei, G. Z. Saccharomonospora-Cyanea Sp-Nov. Int J Syst Bacteriol 38, 444–446 (1988).
Porter, J. N., Wilhelm, J. J. & Tresner, H. D. Method for the preferential isolation of actinomycetes from soils. J Appl Microbiol 8, 174–178 (1960).
Kurtboke, D. I. Actinophages as indicators of actinomycete taxa in marine environments. Antonie van Leeuwenhoek 87, 19–28, https://doi.org/10.1007/s10482-004-6535-y (2005).
Yang, Q. et al. Potential use of mangroves as constructed wetland for municipal, sewage treatment in Futian, Shenzhen, China. Mar Pollut Bull 57, 735–743, https://doi.org/10.1016/j.marpolbul.2008.01.037 (2008).
Pisano, M. A., Sommer, M. J. & Lopez, M. M. Application of pretreatments for the isolation of bioactive actinomycetes from marine-sediments. Appl Microbiol Biotechnol 25, 285–288 (1986).
Hu, D. et al. Genome guided investigation of antibiotics producing actinomycetales strain isolated from a Macau mangrove ecosystem. Scientific reports 8, 14271 (2018).
Walsh, P. S., Metzger, D. A. & Higuchi, R. Chelex 100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. Biotechniques 54, 134–139 (2013).
Mao, D. P., Zhou, Q., Chen, C. Y. & Quan, Z. X. Coverage evaluation of universal bacterial primers using the metagenomic datasets. BMC Microbiol 12, https://doi.org/10.1186/1471-2180-12-66 (2012).
Devulder, G., De Montclos, M. P. & Flandrois, J. A multigene approach to phylogenetic analysis using the genus Mycobacterium as a model. Int. J.S. Syst. Evol. Microbiol 55, 293–302 (2005).
Pospisil, S. et al. Effect of starter unit availability on the spectrum of manumycin-type metabolites produced by Streptomyces nodosus ssp. asukaensis. J Appl Microbiol 111, 1116–1128, https://doi.org/10.1111/j.1365-2672.2011.05132.x (2011).
Zhu, Z., Liu, G. H., Wang, F., Sasanya, J. J. & Cannavan, A. Development of a liquid chromatography tandem mass spectrometric method for simultaneous determination of 15 aminoglycoside residues in porcine tissues. Food Anal Method 9, 2587–2599, https://doi.org/10.1007/s12161-016-0446-1 (2016).
Dionisi, H. M., Lozada, M. & Olivera, N. L. Bioprospection of marine microorganisms: biotechnological applications and methods. Rev Argent Microbiol 44, 49–60 (2012).
Cheeptham, N. & Saiz-Jimenez, C. New sources of antibiotics: caves. Antibiotics: Current Innovations and Future Trends, 213–227 (2015).
Suksaard, P., Pathom-aree, W. & Duangmal, K. Diversity and plant growth promoting activities of actinomycetes from mangroves. Chiang Mai J Sci 44, 1210–1223 (2017).
Ruttanasutja, P. & Pathom-aree, W. Selective isolation of cultivable actinomycetes from thai coastal marine sediment. Chiang Mai J Sci 42, 88–103 (2015).
Wu, P. et al. Bacterial communities in the rhizospheres of three mangrove tree species from Beilun Estuary, China. Plos One 11, https://doi.org/10.1371/journal.pone.0164082 (2016).
Lo, J. R., Lang, J. M., Darling, A. E., Eisen, J. A. & Coil, D. A. Draft genome sequence of an actinobacterium, Brachybacterium muris strain UCD-AY4. Genome announc 1, e00086–00013 (2013).
Kim, M., Oh, H.-S., Park, S.-C. & Chun, J. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int. J. Syst. Evol. Microbiol 64, 346–351, https://doi.org/10.1099/ijs.0.059774-0 (2014).
Kim, M., Oh, H. S., Park, S. C. & Chun, J. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int. J. Syst. Evol. Microbiol 64, 1825–1825, https://doi.org/10.1099/ijs.0.064931-0 (2014).
Ramasamy, D. et al. A polyphasic strategy incorporating genomic data for the taxonomic description of novel bacterial species. Int. J. Syst. Evol. Microbiol 64, 384–391, https://doi.org/10.1099/ijs.0.057091-0 (2014).
Komatsu, M. et al. Engineered Streptomyces avermitilis host for heterologous expression of biosynthetic gene cluster for secondary metabolites. ACS Synth Biol 2, 384–396, https://doi.org/10.1021/sb3001003 (2013).
Oliynyk, M. et al. Complete genome sequence of the erythromycin-producing bacterium Saccharopolyspora erythraea NRRL23338. Nat Biotechnol 25, 447–453, https://doi.org/10.1038/nbt1297 (2007).
Liu, Z. et al. Analysis of miRNA expression profiling in human macrophages responding to Mycobacterium infection: induction of the immune regulator miR-146a. J Infection 68, 553–561, https://doi.org/10.1016/j.jinf.2013.12.017 (2014).
Stragier, P., Hermans, K., Stinear, T. & Portaels, F. First report of a mycolactone-producing Mycobacterium infection in fish agriculture in Belgium. Fems Microbiol Lett 286, 93–95, https://doi.org/10.1111/j.1574-6968.2008.01264.x (2008).
Yotsu, R. R., Nakanaga, K., Hoshino, Y., Suzuki, K. & Ishii, N. Buruli ulcer and current situation in Japan: a new emerging cutaneous Mycobacterium infection. J Dermatol 39, 587–593, https://doi.org/10.1111/j.1346-8138.2012.01543.x (2012).
Corre, C. & Lowden, P. A. S. The first biosynthetic studies of the azinomycins: acetate incorporation into azinomycin B. Chem Commun, 990–991, https://doi.org/10.1039/b400093e (2004).
Flatt, P. M. & Mahmud, T. Biosynthesis of aminocyclitol-aminoglycoside antibiotics and related compounds. Nat Prod Rep 24, 358–392, https://doi.org/10.1039/b603816f (2007).
Igarashi, Y. et al. Maklamicin, an antibacterial polyketide from an endophytic Micromonospora sp. J Nat Prod 74, 670–674, https://doi.org/10.1021/np100727h (2011).
Tao, W. et al. A genomics-led approach to deciphering the mechanism of thiotetronate antibiotic biosynthesis. Chem Sci 7, 376–385, https://doi.org/10.1039/c5sc03059e (2016).
Mocek, U. et al. Biosynthesis of the modified peptide antibiotic nosiheptide in. Streptomyces-Actuosus. J Am Chem Soc 115, 7557–7568, https://doi.org/10.1021/Ja00070a001 (1993).
Igarahi, M. et al. Caprazamycin B, a novel anti-tuberculosis antibiotic, from Streptomyces sp. J Antibiot 56, 580–583, https://doi.org/10.7164/antibiotics.56.580 (2003).
Omura, S., Kitao, C., Tanaka, H., Oiwa, R. & Takahashi, Y. A new antibiotic,, asukamycin, produced by. Streptomyces. J Antibiot 29, 876–881 (1976).
Rui, Z. et al. Biochemical and genetic insights into asukamycin biosynthesis. J Biol Chem 285, 24915–24924, https://doi.org/10.1074/jbc.M110.128850 (2010).
O’Connor, S., Lam, L. K., Jones, N. D. & Chaney, M. O. Apramycin, a unique aminocyclitol antibiotic. J Org 41, 2087–2092 (1976).
Kang, A. D. et al. In vitro apramycin activity against multidrug-resistant Acinetobacter baumannii and Pseudomonas aeruginosa. Diagn Microbiol Infect Dis 88, 188–191, https://doi.org/10.1016/j.diagmicrobio.2017.03.006 (2017).
Minnikin, D. E. et al. Pathophysiological implications of cell envelope structure in Mycobacterium tuberculosis and related taxa. In Tuberculosis, ed. W Ribón chap. 7, https://doi.org/10.5772/59585 (Intech, 2015).
Zarins-Tutt, J. S. et al. Prospecting for new bacterial metabolites: a glossary of approaches for inducing, activating and upregulating the biosynthesis of bacterial cryptic or silent natural products. Nat Prod Rep 33, 54–72 (2016).
Medema, M. H. & Fischbach, M. A. Computational approaches to natural product discovery. Nat Chem Biol 11, 639, https://doi.org/10.1038/nchembio.1884 (2015).
Heidelberg, T. & Martin, O. R. Synthesis of the glycopeptidolipid of Mycobacterium avium serovar 4: first example of a fully synthetic C-Mycoside GPL. The Journal of organic chemistry 69, 2290–2301 (2004).
Xie, P., Sheng, Y., Ito, T. & Mahmud, T. Transcriptional regulation and increased production of asukamycin in engineered Streptomyces nodosus subsp asukaensis strains. Appl Microbiol Biotechol 96, 451–460 (2012).
Fidan, O. et al. New insights into the glycosylation steps in the biosynthesis of Sch47554 and Sch47555. Chembiochem (2018).
Alcaro, S., Ortuso, F. & Coleman, R. S. DNA cross-linking by azinomycin B: Monte Carlo simulations in the evaluation of sequence selectivity. J Med Chem 45, 861–870 (2002).
Walton, J. Apramycin, a new aminocyclitol antibiotic: I. In vitro microbiological studies. J Antimicrob Chemoth 4, 309–313 (1978).
Yu, Y. et al. NosA catalyzing carboxyl-terminal amide formation in nosiheptide maturation via an enamine dealkylation on the serine-extended precursor peptide. J Am Chem Soc 132, 16324–16326 (2010).
Igarashi, M. et al. Caprazamycins, novel lipo-nucleoside antibiotics, from Streptomyces sp. J Antibiot 58, 327 (2005).
Acknowledgements
This study was supported by The Macau Science and Technology Development Fund (FDCT) and the Ministry of Science and Technology of China (MOST) joint funding scheme (Ref. No. FDCT 017/2015/AMJ), The International S&T Cooperation (2016YFE0122000) and Research Committee, University of Macau (MYRG2016-00056-FST, MYRG2015-00182-ICMS-QRCM, MYRG139(Y1-L4)-ICMS12-LMY, and MYRG2016-00129-ICMS-QRCM).
Author information
Authors and Affiliations
Contributions
Simon Ming-Yuen Lee, Dini Hu and Chenghang Sun collected samples. Dini Hu, Cheng Gao, Tao Jin and Guangyi Fan conducted the genomic analysis. Simon Ming-Yuen Lee, Dini Hu and Kai Meng Mok wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hu, D., Gao, C., Sun, C. et al. Genome-guided and mass spectrometry investigation of natural products produced by a potential new actinobacterial strain isolated from a mangrove ecosystem in Futian, Shenzhen, China. Sci Rep 9, 823 (2019). https://doi.org/10.1038/s41598-018-37475-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-37475-w
This article is cited by
-
Metabarcoding expands knowledge on diversity and ecology of rare actinobacteria in the Brazilian Cerrado
Folia Microbiologica (2024)
-
Novel approach in whole genome mining and transcriptome analysis reveal conserved RiPPs in Trichoderma spp
BMC Genomics (2020)
-
In silico genomic mining reveals unexplored bioactive potential of rare actinobacteria isolated from Egyptian soil
Bulletin of the National Research Centre (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.