Abstract
Pancreatic ductal adenocarcinoma (PDAC) is one of the most deadly malignancies with <5% five-year survival rate due to late diagnosis, limited treatment options and chemoresistance. There is thus an urgent unmet clinical need to develop effective anticancer drugs to treat pancreatic cancer. Here, we study the potential of repurposing monensin as an anticancer drug for chemo-resistant pancreatic cancer. Using the two commonly-used chemo-resistant pancreatic cancer cell lines PANC-1 and MiaPaCa-2, we show that monensin suppresses cell proliferation and migration, and cell cycle progression, while solicits apoptosis in pancreatic cancer lines at a low micromole range. Moreover, monensin functions synergistically with gemcitabine or EGFR inhibitor erlotinib in suppressing cell growth and inducing cell death of pancreatic cancer cells. Mechanistically, monensin suppresses numerous cancer-associated pathways, such as E2F/DP1, STAT1/2, NFkB, AP-1, Elk-1/SRF, and represses EGFR expression in pancreatic cancer lines. Furthermore, the in vivo study shows that monensin blunts PDAC xenograft tumor growth by suppressing cell proliferation via targeting EGFR pathway. Therefore, our findings demonstrate that monensin can be repurposed as an effective anti-pancreatic cancer drug even though more investigations are needed to validate its safety and anticancer efficacy in pre-clinical and clinical models.
Similar content being viewed by others
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is one of the most deadly diseases and one of the leading causes of cancer-related deaths in United States1,2,3,4. Most PDAC patients remain asymptomatic until the disease reaches an advanced stage4. In fact, only less than 20% of patients are present with localized, potentially resectable tumors5. As lifespan is being improved in general population, it is conceivable that the absolute case numbers of pancreatic cancer are likely to rise, especially in China, India and other Asian regions with large populations6. For example, in 2015 there were about 90,000 new cases diagnosed with PDAC and nearly 80,000 deaths due to this disease in China7. While multiple factors may contribute to the dismal prognosis for patients with pancreatic cancer, two notable clinical features of this disease may share the blame, late diagnosis and resistance to the already limited treatment options2,8. Despite decades of efforts, the overall five-year survival rate for pancreatic cancer remains at only ~5%3,6,9.
Even though the detailed tumorigenic mechanism behind PDAC remains to be fully elucidated, most pancreatic cancers arise from microscopic non-invasive epithelial proliferations within the pancreatic ducts4. Alterations of the four driver genes KRAS, CDKN2A, TP53, and SMAD4 are thought critical to the development of pancreatic cancer, in which KRAS mutation and alterations in CDKN2A are considered early events in pancreatic tumorigenesis4. A recent integrated genomic analysis of 456 PDAC samples has identified 32 recurrently mutated genes that aggregate into 10 pathways, including KRAS, TGF-β, WNT, NOTCH, ROBO/SLIT signaling, G1/S transition, SWI-SNF, chromatin modification, DNA repair and RNA processing10. Furthermore, transcriptomic analysis classified PDAC into 4 subtypes: squamous tumors, pancreatic progenitor tumors, immunogenic tumors, and aberrantly differentiated endocrine exocrine (ADEX) tumors, which correlate well with PDAC histopathological characteristics10. It is conceivable that such integrative genomic analysis of the molecular evolution of pancreatic cancer subtypes should identify potential targets for therapeutic development in the near future.
The past two decades have witnessed numerous progresses in the development of new and effective targeted cancer therapeutics. However, limited progress has been made in the drug development for pancreatic cancer due to its heterogeneity and drug resistance9,10,11. In most cases, surgical resection remains as the only potentially curative treatment, followed by post-operative adjuvant chemotherapy with gemcitabine or S-1, an oral fluoropyrimidine derivative4. FOLFIRINOX (fluorouracil, folinic acid, irinotecan, and oxaliplatin) and gemcitabine plus nanoparticle albumin-bound paclitaxel (nab-paclitaxel) are the treatments of choice for those who do not have surgery indications4. The use of gemcitabine in patients with advanced pancreatic cancer is associated with a significant, though marginal, survival extension of approximately one month12. Gemcitabine has been the cornerstone of PDAC treatment in all stages of the disease for the last two decades, but gemcitabine resistance develops within weeks of chemotherapy initiation9. The epithelial growth factor receptor (EGFR) inhibitor erlotinib is one of a few targeted agents that show promise in combination with gemcitabine although only achieving a marginal survival benefit in unselected patients13. Thus, it is urgent to develop effective anticancer drugs to treat pancreatic cancer.
The unmet need for more effective anticancer drugs has sparked a growing interest for drug repurposing, which involves in using drugs already approved for other indications to treat cancer14. Drug repurposing can also be a cost-effective alternative strategy to identify new small molecule-based therapies and may significantly influences the discovery of therapeutics although successful drug repurposing is challenging and subject to particular limitations14,15. Nonetheless, several of such repurposed anticancer drugs are currently in clinical trials14,15,16,17.
Here, we study the anticancer activity of an antibiotic, monensin, in human pancreatic cancer. As a polyether innophore antibiotic secreted by the bacteria Streptomyce cinnamonensis18,19, monensin has recently been shown to exhibit anti-proliferative effect on several cancer types. However, it is not clear whether monensin has similar anticancer effect on pancreatic cancer, particularly on gemcitabine-resistant pancreatic cancer. Using two human pancreatic cancer cell lines Panc-1 and MiaPaCa-2, we show that monensin suppresses cell proliferation and migration, and cell cycle progression, and induces cell death of gemcitabine-resistant human pancreatic cancer cells. Furthermore, monensin functions synergistically with gemcitabine or erlotinib to suppress cell growth and induce cell death of gemcitabine-resistant human pancreatic cancer cells. Mechanistic studies indicate that monensin targets several cancer-related signaling pathways and effectively inhibits EGFR expression in pancreatic cancer cell, which is elevated in PDAC samples. Lastly, monensin effectively blunts tumor growth in vivo and inhibits cell proliferation and EGFR expression in the xenograft tumors of gemcitabine-resistant PDAC cells. Collectively, these results demonstrate that monensin can be repurposed to treat pancreatic cancer, in particular for chemo-resistant PDAC, although further studies are required to validate its safety and anticancer efficacy in various pre-clinical and clinical models.
Materials and Methods
Cell culture and chemicals
Human pancreatic cancer cell line Panc-1 and MiaPaCa-2 were kindly provided by Dr. Keping Xie of the University of Texas MD Anderson Cancer Center. All cells were cultured in complete DMEM containing 10% fetal bovine serum (FBS, ThermoFisher, Waltham, MA) with 100 units/mL penicillin and 100 µg/mL streptomycin at 37 °C in 5% CO2 as previously described20,21,22,23. Monensin, gemcitabine and erlotinib were obtained from Cayman Chemical (Ann Arbor, MI) and LC Laboratories (Woburn, MA), respectively. Unless indicated, all other reagents were purchased from Sigma-Aldrich (St. Louis, MO) or Thermos Fisher Scientific (Waltham, MA).
Crystal violet staining
Crystal violet staining was performed as previously described24,25,26,27. Briefly, subconfluent Panc-1 and MiaPaCa-2 were treated with the indicated concentrations of drugs (monensin, gemcitabine, erlotinib or drugs combinations). At 72 h post treatment, cells were washed, fixed and stained with 0.5% crystal violet/formalin for 30 min, followed by tape water rinse and air dry before taking macrographic images.
WST-1 assay
Cell proliferation was measured by using Premixed WST-1 Reagent (Takara Bio USA, Mountain View, CA) as previously described28,29,30,31. Briefly, subconfluent Panc-1 and MiaPaCa-2 cells were plated in 96-well plates and treated with different concentrations of drugs (gemcitabine, monensin, erlotinib or drugs combinations) for 48 h. The Premixed WST-1 substrate was added to the wells, and incubated at 37 °C for 30 min, followed byreading at 440 nm using a microplate reader. Each assay condition was performed in triplicate.
Cell wounding/migration assay
Cell wounding/migration experiments were carried out as previously described32,33,34,35,36. Specifically, cells were plated in 6-well culture plates to reach ~90% confluence. The monolayer cells were then scratched with pipette tips. At the indicated time points, wound healing status at the same locations was recorded. Each assay was set up in triplicate.
Transwell cell migration analysis
Transwell assay was carried out as previously described37,38,39. Briefly, resuspended Panc-1 or MiaPaCa-2 cells were place in the upper chamber containing a layer of the 8 µm pore Corning transwell membrane (Millipore-Sigma) and treated with 4 µM monensin or DMSO control, while the lower chamber was filled with culture medium. At 12 h post treatment, the cells that migrated through the membrane were fixed, stained, and counted (e.g., 10 high-power fields were counted to determine the average migrated cells).
Cell cycle analysis
Cell cycle analysis was carried out as previously described29,40,41,42,43. Specifically, Panc-1 and MiaPaCa-2 were plated in 6-well culture plates and treated with 2 µM monensin or solvent control. At 48 h, cells were harvested, fixed and stained with Magic Solution (10x stock: 0.5% NP-40, 3.4% formaldehyde, 10 ug/ml Hoechst 33258, in PBS) for 30 min, followed by flow cytometry using BD FACS Calibur-HTS. Data analysis was done with FlowJo v10.0 software. Each assay was performed in triplicate.
Apoptosis analysis
The apoptosis analysis was determined by using the Annexin V staining assay as described29,44,45,46,47. Briefly, Panc-1 and MiaPaCa-2 were plated in 6-well cell culture plates and treated with different concentrations of monensin or vehicle control. At 48 h, cells were collected, resuspended in Annexin V Binding Buffer at 106 cells/ml, and stained with Annexin V-FITC (BD Pharmingen, San Jose, CA) and propidium iodide (PI) for 15 min under a light-proof condition. The stained cells were subjected to flow cytometry using BD FACS Calibur-HTS. Data analysis was done with the FlowJo v10.0 Each assay was performed in triplicate.
Chou-Talalay drug combination index determination
The drug combination effects between gemcitabine or EGFR inhibitor (erlotinib) and monensin was analyzed by using Chou-Talalay method28,29,48,49. Dose-dependent effects of each drug alone and in combinations on cell proliferation were first determined by WST-1 assay. The acquired data were calculated by using CompuSyn software (ComboSyn, Inc.). The obtained combination index (CI) from Chou-Talalay method provides a quantitative definition for additive effect (CI = 1), synergism (CI < 1), and antagonism (CI > 1) in different drug combinations as previously described48,49.
Transfection and Gaussia luciferase assay
Gaussia luciferase (GLuc) reporter analysis was conducted as described50,51,52,53,54. A penal of cancer-associated signaling pathway reporters, such as E2F/DP1, Elk1/SRF, AP-1, NFκB, and STAT1/2 reporters, were homemade as previously described29,33,52. A constitutively active reporter pBG2Luc served as a control29,33. Briefly, Panc-1 cells were plated in 25 cm2 flasks at subconfluence and transfected with 3 µg/flask of different reporter plasmids using Lipofectamine according to the manufacturer’s instructions (Invitrogen). At 16 h post transfection, the transfected cells were reseeded into 24-well culture plates and treated with indicated concentrations of drug or vehicle control. At 24 h and 48 h post treatment, 50 µl of culture medium were taken for Gaussia luciferase assay using BioLux Gaussia Luciferase Assay Kit (New England Biolabs, NEB, Ipswich, MA). Each assay was carried out in triplicate.
Total RNA purification and Touchdown-quantitative real-time PCR (TqPCR)
Subconfluent pancreatic cancer cells were treated with different concentrations of monensin for 48 h. RNA was isolated by using TRIZOL Reagents (Invitrogen) for reverse transcription with hexamer and M-MuLV reverse transcriptase (NEB). The cDNA products were used qPCR with the primers of the genes of interest designed with Primer3 program (Supplementary Table S1)55. TqPCR was done by using SYBR Green-based qPCR on a CFX-Connect unit (Bio-Rad Laboratories, Hercules, CA) as previously described50,56,57,58. Each qPCR condition was done in triplicate. GAPDH was used to normalize gene expression levels.
Immunofluorescence staining
The immunofluorescence staining was performed as previously reported59,60,61. Briefly, Panc-1 and MiaPaCa-2 were exposed to various concentrations of drug or vehicle control. At 36 h, the cells were fixed and immunofluorescence stained with an anti–EGFR antibody (Santa Cruz Biotechnology, Santa Cruz, CA). Negative control was set up by incubating the cells with control IgG.
Xenograft tumors of human pancreatic cancer lines
Animal experiments were approved by the Institutional Animal Care and Use Committee at The University of Chicago. All experimental procedures were performed by following the approved protocol. Xenograft tumor model was established as previously described28,29,39,43,62,63,64,65. Briefly, Panc-1 cells were stably tagged with FLuc (firefly luciferase) (Panc-1/FLuc) using the piggyBac system30,41,60. Exponentially growing Panc-1/FLuc cells were harvested, prepared in sterile PBS at 107cells/ml and subcutaneously injected into the flanks of nude mice (ENVIGO, 6–8 week-old, male, 3.0 × 106 cells perinjection, 6 sites permouse, n = 5 per group). After 3 days, the mice were randomly divided into two groups: monensin (10 mg/kg, i.p., once every 36 h) or vehicle control (i.p., once every 36 h). Tumor masses were monitored by caliper measurement and whole body bioluminescence imaging with the Xenogen IVIS 200 Imaging System at 7, 11, 16, 26 days post treatment. All mice were euthanized after4 weeks. Tumor masses were harvested.
H & E staining and immunohistochemical (IHC) staining
The use of clinical pancreatic cancer samples and normal pancreatic samples was approved by the Institutional Ethic Committee of West China Hospital, Sichuan University. The archived pancreatic samples were delinked without the patients’ identifiers and used for IHC with the waived informed consent according to the U. S. NIH’s guidelines involving human subjects. A total of 8 cases of pancreatic cancer samples and 6 cases of non-tumor pancreatic samples were obtained from the Department of Pancreatic Surgery of West China Hospital at Sichuan University. The tumor samples recovered from the xenograft model were also subjected to H & E staining as previously described25,66,67.
The IHC staining was conducted as previously reported37,68,69,70. Briefly, paraffin-embedded tissue sections were processed with deparaffinization, rehydration, and antigen retrieval, followed by IHC staining with an anti-EGFR or anti-PCNA antibody (Santa Cruz Biotechnology). Negative controls were set up by incubating the slides with control IgGs.
Statistical analysis
All quantitative experiments were either repeated three times independently and/or performed in triplicate. The student’s t test and one-way analysis of variance were used to calculate the statistical significance, which was defined as p < 0.05.
Results
Monensin suppresses cell proliferation and migration of gemcitabine-resistant pancreatic cancer cells
We first tested the effect of gemcitabine on two commonly-used human pancreatic cancer cell lines Panc-1 and MiaPaCa-2. Gemcitabine is one of the first-line chemotherapy agents for pancreatic cancer. When subconfluent Panc-1 and MiaPaCa-2 cells were exposed to escalating concentrations of gemcitabine, crystal violet staining demonstrated that Panc-1 cells were resistant to gemcitabine and survived well at an even high concentration of 40 µM, although MiaPaCa-2 cells were slightly more sensitive to gemcitabine (Fig. 1A, panel a). However, under the same growth condition monensin was shown to effectively suppress cell proliferation of both lines at a concentration as low as 1 µM (Fig. 1A, panel b), suggesting that monensin may act as a potent anticancer agent for pancreatic cancer cells.
Monensin effectively inhibits cell proliferation and migration of gemcitabine-resistant pancreatic cancer cells. (A) Crystal violet staining assay. Subconfluent Panc-1 and MiaPaCa-2 cells were treated with gemcitabine (GEM) (a) or monensin (MON) (b) at the indicated concentrations. At 72 h post treatment, the cells were fixed and stained with crystal violet. Representative results are shown. (B) WST-1 assay. Panc-1 and MiaPaCa-2 cells were seeded in 96-well plates and treated with varied concentrations of gemcitabine (GEM) or monensin (MON). At 24 h (a) or 48 h (b) WST-1 reagent was added to plates and incubated for 30 min, and absorbance measurement was performed. All assay conditions were done in triplicate. “*”p < 0.05 and “**”p < 0.01, compared with that of the control groups. (C) Cell wound healing assay. Exponentially growing Panc-1 (a) and MiaPaCa-2 (b) cells were wounded with micro-pipette tips and treated with monensin at indicated concentrations. The gaps were recorded at 0 h, 12 h, 24 h, 36 h and 48 h after treatment. The dotted lines indicate the edge of the wound. Each assay condition was done in triplicate. Representative results are shown. (D) Transwell cell assay. Resuspended Panc-1 (a) and MiaPaCa-2 (b) cells were treated with 0 or 4 µM monensin, the cells migrated through the membrane were fixed, stained and quantitatively determined (c). Representative images are shown. “**”p < 0.01, compared with that of the control groups.
We further carried out the WST-1 cell proliferation assays and confirmed the above findings. Specifically, Panc-1 cells were inhibited ineffectively by gemcitabine and showed about only 20% inhibition even at 80 µM of gemcitabine, while MiaPaCa-2 cells were inhibited by 80% at 20 µM of gemcitabine (Fig. 1B, panel a). On the other hand, the WST-1 assay results showed that the cell proliferation was drastically suppressed by monensin at a concentration of as low as 0.5 µM monensin for both Panc-1 (p < 0.01) and MiaPaCa-2 (p < 0.01) (Fig. 1B, panel b). Taken together, the above results indicate that monensin can effectively suppress pancreatic cancer cell proliferation and overcome gemcitabine resistance in pancreatic cancer cells, particularly in Panc-1 cells.
Monensin suppresses cell wound healing and migration of pancreatic cancer cells
We next tested whether monensin treatment impacts wound healing and cell migration of pancreatic cancer cells. When confluent Panc-1 and MiaPaCa-2 monolayer cells were scratched and added with 0 or 4 uM monensin, we found that the rate of wound closure was significantly lower in monensin-treated Panc-1 (Fig. 1C, panel a) and MiaPaCa-2 (Fig. 1C, panel b) than that of the control group (or 0 µM monensin) at all examined the time points. For example, at 48 h, the wound gap was only ~20% and ~5% in Panc-1 and MiaPaCa-2 cells, respectively, compare to the starting time point (or 0 h) in the control groups. However, in the presence of 4 µM monensin the rate of wound closure in Panc-1 and MiaPaCa-2 was significantly reduced, and approximately 90% of the wound remained open in both cell lines (Fig. 1C, panels a vs. b).
We conducted the transwell cell migration assay to assess the effect of monensin on cell migration capability of pancreatic cells. While both Panc-1 and MiaPaCa-2 cells were shown to migrate through the transwell membrane rather effectively (Fig. 1D, panels a,b), the presence of monensin (at 4 µM) significantly reduced the average cell numbers migrated through the transwell membrane in both cell lines (p < 0.01) (Fig. 1D, panel c). Collectively, the above findings strongly suggest that, in addition to its ability to inhibit cell proliferation, monensin may significantly diminish the cell wound healing and migration capabilities of pancreatic cancer cells.
Monensin suppresses cell cycle progression and induces apoptosis in human pancreatic cancer cells
To uncover possible mechanism through which monensin inhibits pancreatic cancer cell proliferation, we analyzed the cell cycle profile in monensin-treated pancreatic cancer cells and found a drastic increase in the sub-G1 phase and a remarkable decrease in S phase of Panc-1 and MiaPaCa-2 cells, compared with that of the control groups (Fig. 2A, panels a,b), consistent with monensin’s ability to inhibit cell proliferation of pancreatic cancer cells. Notably, the G1 phase in both lines also significantly decreased, which may be caused by the marked increase in sub-G1 populations.
Monensin inhibits cell cycle progression and induces apoptosis in human pancreatic cancer cells. (A) Cell cycle analysis. Exponentially growing Panc-1 (a) and MiaPaCa-2 (b) cells were treated with monensin (2 µM) or vehicle control for 48 h. Cells were collected and stained, then subjected to FACS analysis. Percentages of cells in sub-G1 phase and S phase were graphed. “*” p < 0.05 and “**” p < 0.01, compared with that of the control groups. (B) Annexin-V apoptosis assay. Panc-1 (a) and MiaPaCa-2 (b) cells were treated with monensin (4 µM) or vehicle control. At 48 h post treatment, cells were collected and stained with Annexin V-FITC and propodium iodide, and subjected to flow cytometry. Average percentages of apoptotic cells (including early apoptosis and late apoptosis) were calculated and graphed. “**” p < 0.01, compared with that of the control groups.
Furthermore, Annexin-V based apoptosis analysis demonstrated that monensin was able to induce both early apoptosis and late apoptosis, compared with that of the control groups (Fig. 2B, panels a,b). For example, in Panc-1 cells, the percentage of early apoptotic cells and late apoptotic cells in the 4 µM monensin group were 12% and 20%, respectively, compared with 5.85% and 10.5% in the control group (p < 0.01) (Fig. 2B, panel a). Similarly, in MiaPaCa-2 cells the percentage of early apoptotic cells and late apoptotic cells in the 4 µM monensin group were 23% and 40.1%, respectively, compared with 1.95% and 5.77% in control group (p < 0.01) (Fig. 2B, panel b). We also assessed the nucleus morphologic evidence of monensin-induced apoptosis in pancreatic cancer cells through Hoechst 33258 staining and found that monensin (at 4 µM for 48 h) induced significantly higher % of nuclear condensation and DNA fragmentation in Panc-1 and MiaPaCa-2 cells, compared with that of the control groups (data not shown). Therefore, the above findings suggest monensin may suppress pancreatic cancer cell proliferation at least in part by inhibiting cell cycle progression and inducing apoptosis.
Monensin acts synergistically with gemcitabine or EGFR inhibitor erlotinib on suppressing cell growth and inducing cell death of human pancreatic cancer lines
We tested if monensin would synergize with and/or sensitize pancreatic cancer cells to the currently used first-line chemotherapeutic drugs, such as gemcitabine and EGFR targeted inhibitor erlotinib, to inhibit pancreatic cancer cell proliferation. Qualitative crystal violet staining assay showed that, when subconfluent Panc-1 cells were co-treated with gemcitabine or erlotinib and different concentrations of monensin, menonsin was shown to enhance the inhibitory effects of gemcitabine (Fig. 3A, panel a) or erlotinib (Fig. 3A, panel b) in a dose-dependent fashion. Similar results were obtained in MiaPaCa-2 cells (Fig. 3B, panels a,b). The above findings indicate that monensin can potentiate the inhibitory effects that gemcitabine or erlotinib exerts on pancreatic cancer cells.
Monensin acts synergistically with gemcitabine or erlotinib on inhibiting cell proliferation of human pancreatic cancer cells. (A,B) Crystal violet assay for drug combinations. Panc-1 (A) and MiaPaCa-2 (B) cells were treated with monensin/gemcitabine combination (a) or monensin/erlotinib combination (b) at the indicated concentrations. At 72 h post treatment, cells were fixed and stained with crystal violet. Representative results are shown. (C) WST-1 assay for drug combinations and Chou-Talalay drug combination index analysis. Panc-1 (a,b) and MiaPaCa-2 (c,d) cells were treated with monensin/gemcitabine combination or monensin/erlotinib combination at indicated concentrations (a,c). At 48 h post treatment, WST-1 reagent was added to each well and incubated for 30 min. WST-1 activities were measured and graphed. The WST-1 assay data were further calculated for the combination index (CI) using the Chou-Talalay method (b,d). CI < 1, synergistic effect; CI = 1, additive effect; and CI > 1, antagonistic effect.
We also carried out the quantitative WST-1 assays on drug combinations (Fig. 3C, panels a,c). Using the data acquired from the WST-1 assays, we analyzed the combination index (or CI) with the well-established Chou-Talalay method48,49. We found the CI values for both monensin/gemcitabine and monensin/erlotinib combinations in Panc-1 cells were <1, indicating that these combinations may exhibit synergistic effects (Fig. 3C, panel b). Similarly, in MiaPaCa-2 cells the CI values for both monensin/gemcitabine and monensin/erlotinib combinations were also <1, indicating that these combinations may exhibit synergistic effects (Fig. 3C, panel d).
Furthermore, we analyzed the effect of combinations of monensin and gemcitabine on inducing apoptosis in both Panc-1 and MiaPaCa-2 cells, and found that monensin significantly enhanced gemcitabine-induced early and late phases of apoptosis (Fig. 4A, panels a,b). Similarly, monensin significantly augmented erlotinib-induced early and late phases of apoptosis (Fig. 4B, panels a,b). Collectively, these findings strongly demonstrate that monensin can synergize with gemcitabine or erlotinib on inhibiting the cell growth and inducing cell death of pancreatic cancer cells.
Monensin potentiates gemcitabine or erlotinib-induced apoptosis of human pancreatic cancer cells. Monensin/gemcitabine combinations (A) or monensin/erlotinib combinations (B) at the indicated concentrations were carried out in Panc-1 (a) and MiaPaCa-2 (b). At 48 h post treatment, cells were collected, stained with Annexin V-FITC and propodium iodide, and subjected to flow cytometry. Average percentages of apoptotic cells were calculated and graphed. “*” p < 0.05 and “**” p < 0.01, compared with that of the control groups.
Monensin suppressed several cancer-associated pathways and effectively inhibits the expression of EGFR in pancreatic cancer cells
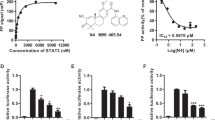
To explore the potential mechanisms through which monensin exerts anticancer effects on pancreatic cancer cells, we surveyed the effects of monensin on a panel of five well-characterized cancer-associated signal pathway reporters as we previously described28,29,33,45,52,71,72. When the Gaussia luciferase reporters were introduced into Panc-1 cells and treated with 0, 1 µM or 4 µM monensin for 24 h and 48 h, the Gaussia luciferase activities for the E2F/DP1, STAT1/2, NFκB, AP-1 and Elk-1/SRF reporters were significantly inhibited (p < 0.01) (Fig. 5A, panels a,b). It is noteworthy that we also analyzed the reporter activities for NFAT, HIF1A, RBP-JK, MYC/MAX, TCF/LEF, CREB, and TGFB/SMAD pathways and found their activities were not significantly affected by monensin.
Monensin inhibits multiple cancer-associated pathways and targets EGFR signaling in human pancreatic cancer cells. (A) Effect of monensin on several important cancer-associated pathways. Subconfluent Panc-1 cells were transfected with the indicated GLuc reporter plasmids and treated with monensin at the indicated concentrations. At 24 h (a) and 48 h (b) post treatment, culture media were collected for Gaussia luciferase activity assay. Each assay condition was done in triplicate. “**” p < 0.01, compared with that of the control groups. (B) Effect of monensin on EGFR signaling pathway. Subconfluent Panc-1 (a) and MiaPaCa-2 (b) cells were treated with the indicated concentrations of monensin for 48 h. Total RNA was isolated and subjected to TqPCR analysis of the expression of EGFR and related genes. Human GAPDH was used as the reference gene. “*” p < 0.05 and “**” p < 0.01, compared with that of the control groups. (C) Effect of monensin on EGFR protein level. Subconfluent Panc-1 (a) and MiaPaCa-2 (b) cells were treated with monensin at the indicated concentrations or vehicle control. At 36 h, cells were fixed and subjected to immunofluorescence staining with an EGFR antibody. The cell nuclei were counter-stained with DAPI. Control IgG was used as a negative control. Representative results are shown.
Based on the results from the reporter assays and the previous reports from ours and other labs19,29, we speculated that monensin might target EGFR pathway. Thus, we analyzed the expression levels of the EGFR and EGFR-regulated downstream genes following monensin treatment in Panc-1 and MiaPaCa-2 cells. Using TqPCR56, we found that the expression of EGFR and RAF1 was significantly repressed in both cell lines (Fig. 5B, panels a,b). Interestingly, KRAS and NRAS, and to a lesser extent MEK1, were shown to be significantly up-regulated by monensin, especially at 4 µM level (Fig. 5B, panels a,b). While we do not have any satisfactory explanations about such up-regulations, it is conceivable the up-regulation may be caused by negative feedback inhibitions upon monensin treatment.
We also examined the effect of monensin on EGFR expression at protein level. Subconfluent Panc-1 and MiaPaCa-2 were treated with varied concentrations of monensin. EGFR expression was examined by immunofluorescence analysis. We found that while the membrane-bound and/or whole cell expression of EGFR was readily detected in the control or vehicle treated cells, EGFR expression, especially at cell membrane, was significantly suppressed by monensin, although there was significant nuclear stainging upon monensin treatment (Fig. 5C, panel a,b). Nonetheless, the above results further indicate monensin can target EGFR signaling in pancreatic cancer.
We further analyzed EGFR expression in the clinical samples of pancreatic cancer. While EGFR expression in normal pancreatic samples was not apparently detectable, seven of the examined eight cases of pancreatic cancer samples exhibited strong EGFR staining in the cancerous ductal cells (Fig. 6A vs. B). The remaining samples had weaker but detectable EGFR expression in cancerous regions (data not shown). Taken together, these results demonstrate that EGFR may be targeted by monensin, which may at least in part explain how monensin exerts its effective anticancer activity against pancreatic cancer cells.
EGFR is highly expressed in human pancreatic cancer tissues. Two cases of normal pancreatic tissues (A) and four cases of representative pancreatic cancer samples (B) tissues were sectioned and subjected to immunohistochemical staining with an anti-EGFR antibody. Control IgG was used as a negative control. Representative images are shown.
Monensin effectively blunts the tumor growth and inhibits cell proliferation and EGFR expression in the xenograft model of human pancreatic cancer in vivo
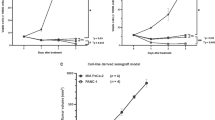
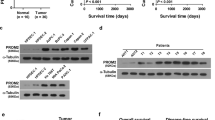
Lastly, we examined the in vivo anticancer activity of monensin in the xenograft tumor model of human pancreatic cancer. The firefly luciferase-tagged Panc-1 cells were first injected into the flanks of athymic mice. After 3 days, the mice were randomly divided into two groups and treated with monensin (10 mg/kg body weight) or vehicle control. Tumor progression was monitored through whole body Xenogen bioluminescence imaging (Fig. 7A panel a). Xenogen imaging data analysis indicates that monensin effectively suppressed tumor growth at as early as 11 days after treatment, compared with the control group (Fig. 7A, panel b). At 4 weeks after treatment, the tumor masses recovered from the control group are significantly larger, either in individual tumors or bulk tumor volumes, than that of the monensin treatment group (Fig. 7B, panels a–c). These findings further confirm that monensin can effectively suppress the tumor growth of gemcitabine-resistant pancreatic cancer cells in vivo.
Monensin blunts the tumor growth and inhibits cell proliferation and EGFR expression in the xenograft model of human pancreatic cancer cells in vivo. (A) Xenogen bioluminescence imaging of xenograft tumor growth. Firefly luciferase-labeled Panc-1 cells were subcutaneously injected into athymic nude mice and randomly divided into two groups. At 3 days post injection, the animals were treated with monensin (10 mg/kg) or vehicle control. The mice were imaged at 7, 11, 16, 26 days after treatment, and sacrificed at 4 weeks of injection. Representative images at day 26 are shown (a). The average signal for each group at different time points was calculated and graphed (b). (B) Retrieved tumor samples and average tumor volume. Representative gross image of the retrieved tumors (a) and accumulative tumor masses from each group (b) are shown. The average tumor volume for each group at the endpoint was also calculated (c). “**” p < 0.01, compared with that of the control group. (C) Histologic evaluation and immunohistochemical staining. The retrieved tumor samples from each group were fixed, paraffin-embedded and subjected to H & E staining (a) and immunohistochemical staining using anti-PCNA (b) or anti-EGFR antibody (c). Control IgGs were used as negative controls. Representative images are shown.
Nonetheless, histologic evaluation of the tumor masses retrieved from both groups did not show significant differences (Fig. 7C, panel a). However, IHC analysis indicated that the expression of the cell proliferation marker PCNA dramatically decreased in the tumor samples retrieved from the monensin treatment group, compared with that of the control group (Fig. 7C, panel b). Similarly, immunohistochemical staining indicated that the EGFR expression was significantly diminished in the tumor samples retrieved from the monensin treatment group, compared with that of the control group (Fig. 7C, panel c), consistent with the possibility that EGFR may be targeted by monensin in pancreatic cancer cells. Collectively, the in vitro and in vivo findings strongly demonstrate that monensin exerts a potent inhibitory effect on cell proliferation and tumor growth in drug-resistant pancreatic cancer cells, possibly through targeting the EGFR signaling pathway.
Discussion
Pancreatic cancer ranks fourth among cancer related deaths, and the disappointing five-year survival rate of below 5% results from drug resistance to all known therapies, as well as from disease presentation at a late stage when PDAC is already metastatic5,9. Most PDAC patients suffer from recurrence within 24 months and die of the progressively worsening treatment-resistant cancer. Recently, major treatment breakthroughs in many difficult-to-treat cancers, such as melanoma, have been facilitated by the identification of actionable mutant oncogenic driver genes73. While there has been an increasing understanding of the underlying biology of pancreatic cancer, unfortunately, until now, there are no actionable therapeutic targets for PDAC. Meanwhile, only modest improvement in effective systemic chemotherapy has been attained in pancreatic cancer. Therefore, pancreatic cancer is still one of the most lethal cancers with a dismal 5-year survival less than 5%5. Thus, there is an unmet clinical need to develop more effective and safe treatment for the clinical management of PDAC patients. The repurposing of existing non-cancer drugs represents a cost-effective alternative to develop new treatment options for cancer patients with high unmet medical needs. In this study, we demonstrate that monensin may be repurposed to treat chemo-resistant pancreatic cancer. Our results suggest the monensin may act synergistically with gemcitabine or erlotinib in combination chemotherapy for the treatment of drug-resistant pancreatic cancer.
Monensin was discovered as a polyether innophore antibiotic over half century ago18,19. It has a rather favorable biosafety profile as monensin is widely used in cattle and poultry feed18,19,29. It was reported that malignant cells were approximately 20 times more sensitive to monensin than normal cells74. We and others showed that monensin exhibits anti-proliferative effects on several other types of cancer, including renal cancer, lung cancer, colon cancer, myeloma, prostate cancer, and ovarian cancer cells19,29,74,75,76,77,78,79,80. In this study, we further demonstrate that monensin exerts potent anticancer activity in chemo-resistant pancreatic cancer cells by inhibiting its proliferation, cell cycle progression, and cell migration and by inducing apoptosis. Moreover, monensin acts synergistically with gemcitabine or erlotinib to inhibit cell proliferation and induce apoptosis in chemo-resistant pancreatic cancer cells. Our in vivo study in xenograft tumor model of PDAC cells further validates the biosafety and anticancer efficacy of monensin as a repurposed anti-PDAC agent.
Mechanistically, monensin may accomplish its anticancer effect by targeting multiple signaling pathways, particularly the EGFR signaling pathway. Given the fact that monensin exerts its anti-proliferation effect in chemo-resistant pancreatic cancer cells at very low micromole concentrations when compared with gemcitabine or erlotinib. While more mechanism-based studies are needed, it was reported that monensin can reduce the expression of cyclin A, CDK6, and cyclin D1while inducing programmed cell death-related genes, such as caspase-3, caspase-8, Bax, and mitochondria transmembrane potential in some types of human cancer lines76,77,78,79,80. It has been recently shown that monensin can suppress Wnt signaling in colorectal cancer cells81, and EGFR signaling in ovarian cancer cells29.
In this study, we also investigated the effect of monensin on multiple cancer-related pathways and found that monensin can inhibit E2F/DP1, STAT1/2, NFκB, AP-1 and Elk-1/SRF pathways. Furthermore, the expression of EGFR and its downstream genes, such as RAF1 and BRAF, is effectively suppressed by monensin. The above findings suggest that monensin may exert its potent proliferation suppression effect through the inhibition of multiple growth factor-induced signal pathways, especially EGFR, which is found overexpressed in the clinical samples of pancreatic cancer. Interestingly, it was reported that monensin may impact the endocytic recycling pathway of EGFR19,82. We further demonstrate that the EGFR expression level in the PDAC xenograft tumors was significantly inhibited by monensin. Therefore, our findings strongly demonstrate that monensin can exert its potent anticancer activity in chemo-resistant PDAC cells at least in part by targeting the EGFR signaling pathway. It is conceivable that, even though both target EGFR, the mode of action for monensin should be distinct from that of erlotinib’s since monensin can synergize with erlotinib in PDAC cells. Collectively, our findings suggest monensin can be repurposed to treat pancreatic cancer, although its safety and anticancer efficacy need to be further validated in preclinical and clinical studies.
References
Wolfgang, C. L. et al. Recent progress in pancreatic cancer. CA Cancer J Clin 63, 318–348, https://doi.org/10.3322/caac.21190 (2013).
Xie, D. & Xie, K. Pancreatic cancer stromal biology and therapy. Genes Dis 2, 133–143, https://doi.org/10.1016/j.gendis.2015.01.002 (2015).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2018. CA Cancer J Clin 68, 7–30, https://doi.org/10.3322/caac.21442 (2018).
Kamisawa, T., Wood, L. D., Itoi, T. & Takaori, K. Pancreatic cancer. Lancet 388, 73–85, https://doi.org/10.1016/S0140-6736(16)00141-0 (2016).
Hidalgo, M. Pancreatic cancer. N Engl J Med 362, 1605–1617, https://doi.org/10.1056/NEJMra0901557 (2010).
Raimondi, S., Maisonneuve, P. & Lowenfels, A. B. Epidemiology of pancreatic cancer: an overview. Nat Rev Gastroenterol Hepatol 6, 699–708, https://doi.org/10.1038/nrgastro.2009.177 (2009).
Chen, W. et al. Cancer statistics in China, 2015. CA Cancer J Clin 66, 115–132, https://doi.org/10.3322/caac.21338 (2016).
Rahib, L. et al. Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res 74, 2913–2921, https://doi.org/10.1158/0008-5472.CAN-14-0155 (2014).
Binenbaum, Y., Na’ara, S. & Gil, Z. Gemcitabine resistance in pancreatic ductal adenocarcinoma. Drug Resist Updat 23, 55–68, https://doi.org/10.1016/j.drup.2015.10.002 (2015).
Bailey, P. et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531, 47–52, https://doi.org/10.1038/nature16965 (2016).
Rajabpour, A., Rajaei, F. & Teimoori-Toolabi, L. Molecular alterations contributing to pancreatic cancer chemoresistance. Pancreatology 17, 310–320, https://doi.org/10.1016/j.pan.2016.12.013 (2017).
Burris, H. A. 3rd et al. Improvements in survival and clinical benefit with gemcitabine as first-line therapy for patients with advanced pancreas cancer: a randomized trial. J Clin Oncol 15, 2403–2413, https://doi.org/10.1200/JCO.1997.15.6.2403 (1997).
Moore, M. J. et al. Erlotinib plus gemcitabine compared with gemcitabine alone in patients with advanced pancreatic cancer: a phase III trial of the National Cancer Institute of Canada Clinical Trials Group. J Clin Oncol 25, 1960–1966, https://doi.org/10.1200/JCO.2006.07.9525 (2007).
Sleire, L. et al. Drug repurposing in cancer. Pharmacol Res 124, 74–91, https://doi.org/10.1016/j.phrs.2017.07.013 (2017).
Verbaanderd, C., Meheus, L., Huys, I. & Pantziarka, P. Repurposing Drugs in Oncology: Next Steps. Trends Cancer 3, 543–546, https://doi.org/10.1016/j.trecan.2017.06.007 (2017).
Gupta, S. C., Sung, B., Prasad, S., Webb, L. J. & Aggarwal, B. B. Cancer drug discovery by repurposing: teaching new tricks to old dogs. Trends Pharmacol Sci 34, 508–517, https://doi.org/10.1016/j.tips.2013.06.005 (2013).
Bhattarai, D. et al. An Insight into Drug Repositioning for the Development of Novel Anti-Cancer Drugs. Curr Top Med Chem 16, 2156–2168 (2016).
Pressman, B. C. Ionophorous antibiotics as models for biological transport. Fed Proc 27, 1283–1288 (1968).
Dayekh, K. et al. Monensin inhibits epidermal growth factor receptor trafficking and activation: synergistic cytotoxicity in combination with EGFR inhibitors. Mol Cancer Ther 13, 2559–2571, https://doi.org/10.1158/1535-7163.MCT-13-1086 (2014).
Zhu, G. H. et al. Activation of RXR and RAR signaling promotes myogenic differentiation of myoblastic C2C12 cells. Differentiation 78, 195–204, https://doi.org/10.1016/j.diff.2009.06.001 (2009).
Huang, E. et al. Conditionally immortalized mouse embryonic fibroblasts retain proliferative activity without compromising multipotent differentiation potential. PLoS One 7, e32428, https://doi.org/10.1371/journal.pone.0032428 (2012).
Huang, E. et al. Growth hormone synergizes with BMP9 in osteogenic differentiation by activating the JAK/STAT/IGF1 pathway in murine multilineage cells. J Bone Miner Res 27, 1566–1575, https://doi.org/10.1002/jbmr.1622 (2012).
Luo, X. et al. Characterization of gene expression regulated by American ginseng and ginsenoside Rg3 in human colorectal cancer cells. Int J Oncol 32, 975–983 (2008).
Lamplot, J. D. et al. Reversibly Immortalized Mouse Articular Chondrocytes Acquire Long-Term Proliferative Capability While Retaining Chondrogenic Phenotype. Cell Transplant 24, 1053–1066, https://doi.org/10.3727/096368914X681054 (2015).
Zhao, C. et al. Sox9 augments BMP2-induced chondrogenic differentiation by downregulating Smad7 in mesenchymal stem cells (MSCs). Genes Dis 4, 229–239, https://doi.org/10.1016/j.gendis.2017.10.004 (2017).
Su, Y. et al. Establishment and characterization of a new highly metastatic human osteosarcoma cell line. Clin Exp Metastasis 26, 599–610, https://doi.org/10.1007/s10585-009-9259-6 (2009).
Song, D. et al. BMP9 induces osteogenesis and adipogenesis in the immortalized human cranial suture progenitors from the patent sutures of craniosynostosis patients. J Cell Mol Med 21, 2782–2795, https://doi.org/10.1111/jcmm.13193 (2017).
Zhang, F. et al. Anthelmintic mebendazole enhances cisplatin’s effect on suppressing cell proliferation and promotes differentiation of head and neck squamous cell carcinoma (HNSCC). Oncotarget 8, 12968–12982, https://doi.org/10.18632/oncotarget.14673 (2017).
Deng, Y. et al. Antibiotic monensin synergizes with EGFR inhibitors and oxaliplatin to suppress the proliferation of human ovarian cancer cells. Sci Rep 5, 17523, https://doi.org/10.1038/srep17523 (2015).
Chen, X. et al. Sustained high level transgene expression in mammalian cells mediated by the optimized piggyBac transposon system. Genes Dis 2, 96–105, https://doi.org/10.1016/j.gendis.2014.12.001 (2015).
Li, M. et al. Establishment and characterization of the reversibly immortalized mouse fetal heart progenitors. Int J Med Sci 10, 1035–1046, https://doi.org/10.7150/ijms.6639 (2013).
Si, W. et al. CCN1/Cyr61 is regulated by the canonical Wnt signal and plays an important role in Wnt3A-induced osteoblast differentiation of mesenchymal stem cells. Mol Cell Biol 26, 2955–2964, https://doi.org/10.1128/MCB.26.8.2955-2964.2006 (2006).
Liao, Z. et al. The Anthelmintic Drug Niclosamide Inhibits the Proliferative Activity of Human Osteosarcoma Cells by Targeting Multiple Signal Pathways. Curr Cancer Drug Targets 15, 726–738 (2015).
Li, R. et al. Targeting BMP9-promoted human osteosarcoma growth by inactivation of notch signaling. Curr Cancer Drug Targets 14, 274–285 (2014).
Rastegar, F. et al. Lysophosphatidic acid acyltransferase beta (LPAATbeta) promotes the tumor growth of human osteosarcoma. PLoS One 5, e14182, https://doi.org/10.1371/journal.pone.0014182 (2010).
Luo, Q. et al. Connective tissue growth factor (CTGF) is regulated by Wnt and bone morphogenetic proteins signaling in osteoblast differentiation of mesenchymal stem cells. J Biol Chem 279, 55958–55968, https://doi.org/10.1074/jbc.M407810200 (2004).
Luu, H. H. et al. Increased expression of S100A6 is associated with decreased metastasis and inhibition of cell migration and anchorage independent growth in human osteosarcoma. Cancer Lett 229, 135–148, https://doi.org/10.1016/j.canlet.2005.02.015 (2005).
Luther, G. A. et al. IGFBP5 domains exert distinct inhibitory effects on the tumorigenicity and metastasis of human osteosarcoma. Cancer Lett 336, 222–230, https://doi.org/10.1016/j.canlet.2013.05.002 (2013).
Li, Y. et al. The Calcium-Binding Protein S100A6 Accelerates Human Osteosarcoma Growth by Promoting Cell Proliferation and Inhibiting Osteogenic Differentiation. Cell Physiol Biochem 37, 2375–2392, https://doi.org/10.1159/000438591 (2015).
Bi, Y. et al. Functional characteristics of reversibly immortalized hepatic progenitor cells derived from mouse embryonic liver. Cell Physiol Biochem 34, 1318–1338, https://doi.org/10.1159/000366340 (2014).
Wen, S. et al. Characterization of constitutive promoters for piggyBac transposon-mediated stable transgene expression in mesenchymal stem cells (MSCs). PLoS One 9, e94397, https://doi.org/10.1371/journal.pone.0094397 (2014).
Luo, J. et al. TGFbeta/BMP type I receptors ALK1 and ALK2 are essential for BMP9-induced osteogenic signaling in mesenchymal stem cells. J Biol Chem 285, 29588–29598, https://doi.org/10.1074/jbc.M110.130518 (2010).
Chen, X. et al. The E-F hand calcium-binding protein S100A4 regulates the proliferation, survival and differentiation potential of human osteosarcoma cells. Cell Physiol Biochem 32, 1083–1096, https://doi.org/10.1159/000354508 (2013).
He, B. C. et al. Synergistic antitumor effect of the activated PPARgamma and retinoid receptors on human osteosarcoma. Clin Cancer Res 16, 2235–2245, https://doi.org/10.1158/1078-0432.CCR-09-2499 (2010).
He, B. C. et al. Tetrandrine inhibits Wnt/beta-catenin signaling and suppresses tumor growth of human colorectal cancer. Mol Pharmacol 79, 211–219, https://doi.org/10.1124/mol.110.068668 (2011).
Shui, W. et al. Characterization of scaffold carriers for BMP9-transduced osteoblastic progenitor cells in bone regeneration. J Biomed Mater Res A 102, 3429–3438, https://doi.org/10.1002/jbma.35006 (2014).
Yang, K. et al. Conditional immortalization establishes a repertoire of mouse melanocyte progenitors with distinct melanogenic differentiation potential. J Invest Dermatol 132, 2479–2483, https://doi.org/10.1038/jid.2012.145 (2012).
Chou, T. C. Theoretical basis, experimental design, and computerized simulation of synergism and antagonism in drug combination studies. Pharmacol Rev 58, 621–681, https://doi.org/10.1124/pr.58.3.10 (2006).
Chou, T. C. Drug combination studies and their synergy quantification using the Chou-Talalay method. Cancer Res 70, 440–446, https://doi.org/10.1158/0008-5472.CAN-09-1947 (2010).
Fan, J. et al. Noncanonical Wnt signaling plays an important role in modulating canonical Wnt-regulated stemness, proliferation and terminal differentiation of hepatic progenitors. Oncotarget 8, 27105–27119, https://doi.org/10.18632/oncotarget.15637 (2017).
Zhang, H. et al. Canonical Wnt signaling acts synergistically on BMP9-induced osteo/odontoblastic differentiation of stem cells of dental apical papilla (SCAPs). Biomaterials 39, 145–154, https://doi.org/10.1016/j.biomaterials.2014.11.007 (2015).
Gao, J. L. et al. Ginseng saponin metabolite 20(S)-protopanaxadiol inhibits tumor growth by targeting multiple cancer signaling pathways. Oncol Rep 30, 292–298, https://doi.org/10.3892/or.2013.2438 (2013).
Ye, J. et al. A thermoresponsive polydiolcitrate-gelatin scaffold and delivery system mediates effective bone formation from BMP9-transduced mesenchymal stem cells. Biomed Mater 11, 025021, https://doi.org/10.1088/1748-6041/11/2/025021 (2016).
Zou, Y. et al. Gelatin-Derived Graphene-Silicate Hybrid Materials Are Biocompatible and Synergistically Promote BMP9-Induced Osteogenic Differentiation of Mesenchymal Stem Cells. ACS Appl Mater Interfaces 9, 15922–15932, https://doi.org/10.1021/acsami.7b00272 (2017).
Untergasser, A. et al. Primer3–new capabilities and interfaces. Nucleic Acids Res 40, e115, https://doi.org/10.1093/nar/gks596 (2012).
Zhang, Q. et al. TqPCR: A Touchdown qPCR Assay with Significantly Improved Detection Sensitivity and Amplification Efficiency of SYBR Green qPCR. PLoS One 10, e0132666, https://doi.org/10.1371/journal.pone.0132666 (2015).
Liao, J. et al. Characterization of retroviral infectivity and superinfection resistance during retrovirus-mediated transduction of mammalian cells. Gene Ther 24, 333–341, https://doi.org/10.1038/gt.2017.24 (2017).
Lu, S. et al. Bone morphogenetic protein 9 (BMP9) induces effective bone formation from reversibly immortalized multipotent adipose-derived (iMAD) mesenchymal stem cells. Am J Transl Res 8, 3710–3730 (2016).
Wang, N. et al. Adenovirus-mediated efficient gene transfer into cultured three-dimensional organoids. PLoS One 9, e93608, https://doi.org/10.1371/journal.pone.0093608 (2014).
Wang, N. et al. The piggyBac transposon-mediated expression of SV40 T antigen efficiently immortalizes mouse embryonic fibroblasts (MEFs). PLoS One 9, e97316, https://doi.org/10.1371/journal.pone.0097316 (2014).
Tang, N. et al. BMP-9-induced osteogenic differentiation of mesenchymal progenitors requires functional canonical Wnt/beta-catenin signalling. J Cell Mol Med 13, 2448–2464, https://doi.org/10.1111/j.1582-4934.2008.00569.x (2009).
Luu, H. H. et al. An orthotopic model of human osteosarcoma growth and spontaneous pulmonary metastasis. Clin Exp Metastasis 22, 319–329, https://doi.org/10.1007/s10585-005-0365-9 (2005).
Luo, X. et al. Osteogenic BMPs promote tumor growth of human osteosarcomas that harbor differentiation defects. Lab Invest 88, 1264–1277, https://doi.org/10.1038/labinvest.2008.98 (2008).
Luo, X., Sharff, K. A., Chen, J., He, T. C. & Luu, H. H. S100A6 expression and function in human osteosarcoma. Clin Orthop Relat Res 466, 2060–2070, https://doi.org/10.1007/s11999-008-0361-x (2008).
Yu, X. et al. Niclosamide Exhibits Potent Anticancer Activity and Synergizes with Sorafenib in Human Renal Cell Cancer Cells. Cell Physiol Biochem 47, 957–971, https://doi.org/10.1159/000490140 (2018).
Hu, X. et al. CRISPR/Cas9-mediated reversibly immortalized mouse bone marrow stromal stem cells (BMSCs) retain multipotent features of mesenchymal stem cells (MSCs). Oncotarget 8, 111847–111865, https://doi.org/10.18632/oncotarget.22915 (2017).
Wang, J. et al. NEL-Like Molecule-1 (Nell1) Is Regulated by Bone Morphogenetic Protein 9 (BMP9) and Potentiates BMP9-Induced Osteogenic Differentiation at the Expense of Adipogenesis in Mesenchymal Stem Cells. Cell Physiol Biochem 41, 484–500, https://doi.org/10.1159/000456885 (2017).
Hu, N. et al. BMP9-regulated angiogenic signaling plays an important role in the osteogenic differentiation of mesenchymal progenitor cells. J Cell Sci 126, 532–541, https://doi.org/10.1242/jcs.114231 (2013).
Haydon, R. C. et al. Cytoplasmic and/or nuclear accumulation of the beta-catenin protein is a frequent event in human osteosarcoma. Int J Cancer 102, 338–342, https://doi.org/10.1002/ijc.10719 (2002).
Liao, J. et al. Notch Signaling Augments BMP9-Induced Bone Formation by Promoting the Osteogenesis-Angiogenesis Coupling Process in Mesenchymal Stem Cells (MSCs). Cell Physiol Biochem 41, 1905–1923, https://doi.org/10.1159/000471945 (2017).
He, B. C. et al. Ginsenoside Rg3 inhibits colorectal tumor growth through the down-regulation of Wnt/ss-catenin signaling. Int J Oncol 38, 437–445, https://doi.org/10.3892/ijo.2010.858 (2011).
Deng, Y. et al. A Blockade of IGF Signaling Sensitizes Human Ovarian Cancer Cells to the Anthelmintic Niclosamide-Induced Anti-Proliferative and Anticancer Activities. Cell Physiol Biochem 39, 871–888, https://doi.org/10.1159/000447797 (2016).
Chapman, P. B. et al. Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N Engl J Med 364, 2507–2516, https://doi.org/10.1056/NEJMoa1103782 (2011).
Ketola, K., Vainio, P., Fey, V., Kallioniemi, O. & Iljin, K. Monensin is a potent inducer of oxidative stress and inhibitor of androgen signaling leading to apoptosis in prostate cancer cells. Mol Cancer Ther 9, 3175–3185, https://doi.org/10.1158/1535-7163.MCT-10-0368 (2010).
Choi, H. S. et al. Autophagy Inhibition with Monensin Enhances Cell Cycle Arrest and Apoptosis Induced by mTOR or Epidermal Growth Factor Receptor Inhibitors in Lung Cancer Cells. Tuberc Respir Dis (Seoul) 75, 9–17, https://doi.org/10.4046/trd.2013.75.1.9 (2013).
Park, W. H. et al. Monensin inhibits the growth of renal cell carcinoma cells via cell cycle arrest or apoptosis. Int J Oncol 22, 855–860 (2003).
Park, W. H., Kim, E. S., Jung, C. W., Kim, B. K. & Lee, Y. Y. Monensin-mediated growth inhibition of SNU-C1 colon cancer cells via cell cycle arrest and apoptosis. Int J Oncol 22, 377–382 (2003).
Park, W. H., Kim, E. S., Kim, B. K. & Lee, Y. Y. Monensin-mediated growth inhibition in NCI-H929 myeloma cells via cell cycle arrest and apoptosis. Int J Oncol 23, 197–204 (2003).
Park, W. H. et al. Monensin-mediated growth inhibition in acute myelogenous leukemia cells via cell cycle arrest and apoptosis. Int J Cancer 101, 235–242, https://doi.org/10.1002/ijc.10592 (2002).
Park, W. H. et al. Monensin-mediated growth inhibition in human lymphoma cells through cell cycle arrest and apoptosis. Br J Haematol 119, 400–407 (2002).
Tumova, L. et al. Monensin inhibits canonical Wnt signaling in human colorectal cancer cells and suppresses tumor growth in multiple intestinal neoplasia mice. Mol Cancer Ther 13, 812–822, https://doi.org/10.1158/1535-7163.MCT-13-0625 (2014).
Chung, B. M. et al. Aberrant trafficking of NSCLC-associated EGFR mutants through the endocytic recycling pathway promotes interaction with Src. BMC Cell Biol 10, 84, https://doi.org/10.1186/1471-2121-10-84 (2009).
Acknowledgements
A part of the reported work was orally presented at the American College of Surgeons Clinical Congress, October 22–26, 2017, San Diego, CA, USA. We wish to thank Dr. Keping Xie of the University of Texas MD Anderson Cancer Center, Houston, TX for the kind provision of human pancreatic cancer lines. The reported study was supported in part by research grants from the National Key Research and Development Program of China (2016YFC1000803 and 2011CB707906), the National Institutes of Health (CA226303 to TCH), the National Institutes of Health of China (W2017ZWS07 to XL and XW), and the 135 Research Project of West China Hospital of Sichuan University (XL and XW). XW was a recipient of pre-doctorate scholarship from China Scholarship Council. This project was also supported in part by The University of Chicago Cancer Center Support Grant (P30CA014599) and the National Center for Advancing Translational Sciences of the National Institutes of Health through Grant Number UL1 TR000430. TCH was also supported by the Mabel Green Myers Research Endowment Fund and The University of Chicago Orthopaedic Surgery Alumni Fund.
Author information
Authors and Affiliations
Contributions
T.C.H., X.L., Xin-W, B.P., H.H.L. and R.C.H. conceived and designed the project. Xin-W, X-Wu, Z-Zhang carried out most of the experiments. C.M., T.W., S.T., Z-Zeng, S.H., C.G., C.Y., L-Zhang, Y.F., B.H., W-Liu, B.Z., Y.S., W-Luo and Xi-W assisted in some experiments and helped on data collections and analyses. B.L., Y.L., Z.Y., L-Zhao, D.C., L.Y. and X.C. provided essential materials/reagents for the project and assisted data analyses. T.C.H., Xin-W, X.L., H.H.L. and R.C.H. drafted the manuscript. All authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, X., Wu, X., Zhang, Z. et al. Monensin inhibits cell proliferation and tumor growth of chemo-resistant pancreatic cancer cells by targeting the EGFR signaling pathway. Sci Rep 8, 17914 (2018). https://doi.org/10.1038/s41598-018-36214-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-36214-5
This article is cited by
-
Redefining bioactive small molecules from microbial metabolites as revolutionary anticancer agents
Cancer Gene Therapy (2024)
-
Endosomal recycling inhibitors downregulate the androgen receptor and synergise with enzalutamide
Investigational New Drugs (2024)
-
Combination of monensin and erlotinib synergistically inhibited the growth and cancer stem cell properties of triple-negative breast cancer by simultaneously inhibiting EGFR and PI3K signaling pathways
Breast Cancer Research and Treatment (2024)
-
Endosomal recycling inhibitors downregulate estrogen receptor-alpha and synergise with endocrine therapies
Breast Cancer Research and Treatment (2024)
-
MDHT-Net: Multi-scale Deformable U-Net with Cos-spatial and Channel Hybrid Transformer for pancreas segmentation
Applied Intelligence (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.