Abstract
Peptidoglycan (PG) biosynthesis and assembly are needed for bacterial cell wall formation. Lipid II is the precursor in the PG biosynthetic pathway and carries a nascent PG unit that is processed by glycosyltransferases. Despite its immense therapeutic value as a target of several classes of antibiotics, the conformational ensemble of lipid II in bacterial membranes and its interactions with membrane-anchored enzymes remain elusive. In this work, lipid II and its elongated forms (lipid VI and lipid XII) were modeled and simulated in bilayers of POPE (palmitoyl-oleoyl-phosphatidyl-ethanolamine) and POPG (palmitoyl-oleoyl-phosphatidyl-glycerol) that mimic the prototypical composition of Gram-negative cytoplasmic membranes. In addition, penicillin-binding protein 1b (PBP1b) from Escherichia coli was modeled and simulated in the presence of a nascent PG to investigate their interactions. Trajectory analysis reveals that as the glycan chain grows, the non-reducing end of the nascent PG displays much greater fluctuation along the membrane normal and minimally interacts with the membrane surface. In addition, dihedral angles within the pyrophosphate moiety are determined by the length of the PG moiety and its surrounding environment. When a nascent PG is bound to PBP1b, the stem peptide remains in close contact with PBP1b by structural rearrangement of the glycan chain. Most importantly, the number of nascent PG units required to reach the transpeptidase domain are determined to be 7 or 8. Our findings complement experimental results to further understand how the structure of nascent PG can dictate the assembly of the PG scaffold.
Similar content being viewed by others
Introduction
Bacterial cell wall biogenesis is critically important to the viability of bacteria and underpin bacterial pathogenesis and human microbiome interactions1. Considering that peptidoglycan (PG) is the major component of most bacteria, the cellular machinery that processes building blocks related to the cell wall is tightly linked to PG structural integrity and cellular maintenance2. Moreover, molecules that disrupt PG assembly are some of the most efficacious antibiotics discovered to date3,4. Gaining deeper understanding into PG biosynthesis may reveal potential targets for drug discovery and development.
Lipid II is the lipid-bound precursor of PG, a primary component of bacterial cell walls. PG consists of cross-linked layers that contribute to cell shape and cell wall strength5,6,7. Lipid II consists of β-(1→4)-N-acetylglucosamine (GlcNAc)-N-acetylmuramic acid (MurNAc) disaccharide unit linked to a pentapeptide stem peptide via MurNAc. The disaccharide unit is connected to an undecaprenyl tail by pyrophosphate and the tail anchors lipid II onto the membrane. While the PG pentapeptide sequence can vary depending on the bacterial species, the canonical sequence for the majority of Gram-negative bacteria is l-Ala-γ-d-Glu-m-DAP-d-Ala-d-Ala, where m-DAP is meso-diaminopimelic acid2.
Lipid II molecules are processed by a number of membrane-anchored enzymes4,8,9,10. Among them, penicillin-binding protein 1b (PBP1b) is a member of the bifunctional class A that carries out both glycosyltransferase (GTase) and transpeptidase (TPase) activities11. GTase domains processes lipid II by polymerizing the glycan chains, whereas TPase domains perform covalent cross-links between neighboring stem peptides12. Given the importance of GTase activity for PG biosynthesis, it is important to understand how glycan chain elongation can control substrate access and priming. Towards these goals, the crystal structure of GTase from Staphylococcus aureus was previously solved with the antibiotic moenomycin to gain structural insight into the mechanism of action of moenomycin13. Moreover, the full-length Escherichia coli (E. coli) PBP1b structure containing the transmembrane helix subdomain was recently published14. Based on these works, we have herein set out to model nascent PG in complex with PBP1b with the ultimate goal of gaining structural insight into PBP-PG interactions.
Given the importance of lipid-bound PG precursors in PG biosynthesis and drug-discovery4,15, there are unanswered fundamental questions about how PG elongation affects lipid II processing by PBPs. Elegant experimental studies have described the elongation of nascent PG building blocks into extended lipid-bound precursors by PBPs16,17,18,19. It was established that during glycan elongation by GTase, glycan chains are stitched together unidirectionally before being loaded onto the existing PG scaffold. Yet, these studies did not provide atomistic details about the dynamics of lipid-bound PG precursors in a membrane environment and within PBPs. How do elongated lipid-anchored PG intermediates behave within bacterial membranes? How does the size of the nascent PG control association with PBPs? Most critically, how many elongation steps are needed for the nascent PG to reach the TPase domain? Towards answering some aspects of these fundamental questions, we first modeled and simulated Gram-negative lipid-anchored PG precursors such as lipid II, lipid VI, and lipid XII in membranes that are representative of Gram-negative bacteria (Figs 1 and 2). Next, E. coli PBP1b was modeled and simulated in the presence of nascent PG (PBP1b-lipidXX complex) to gain atomistic insight into the dynamics of nascent PG in association with PBP1b (Fig. 1).
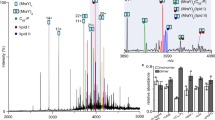
Schematic representation of the E. coli cell envelope with the structures of (A) AcrAB-TolC, (B) lipid II, (C) lipid VI, (D) lipid XII, and (E) the most tilted and (F) the least tilted PBP1b-lipidXX complex. The AcrAB-TolC structure is used to approximately measure the vertical distance between the phosphate of the upper leaflet of the inner membrane (IM) to the phosphate of the lower leaflet of the outer membrane (OM). The periplasmic length (~170 Å) is measured by the length of AcrAB-TolC39, and the distance between the PG and the inner membrane is ~90 Å40.
(A) Chemical structure of lipid II composed of MurNAc (green), GlcNAc (blue), pyrophosphate (red), undecaprenyl tail (black), and pentapeptide (black). Schematic representations of (B) lipid II, (C) lipid VI, and (D) lipid XII; pyrophosphate and undecaprenyl moieties are abbreviated as PP and UND, respectively.
Results and Discussion
Isolated lipid II, lipid VI, and lipid XII in a Gram-negative Bacterial Cytoplasmic Membrane
To characterize the conformational dynamics of isolated lipid II, VI, and XII (Fig. 2B–D) embedded in a Gram-negative cytoplasmic membrane composition (POPE:POPG = 3:1), we performed 1-μs simulation for each system (Fig. S1; Table S1; see Methods for details). We calculated the average Z-position of lipid II/VI/XII components relative to the phosphate groups of the membrane, as well as the fraction of interaction time between each residue and the membrane surface. Figure 3 shows the average Z-position (dot) for lipid II, VI, and XII with standard deviations over the entire trajectory. The fraction of interaction time (Fig. 3, shaded box) was defined as the fraction of the simulation time that each residue interacts with membrane head groups. As the length of the nascent PG increases, the non-reducing (distal) end of the glycan strand samples a larger physical space (particularly along the membrane normal). In turn, such sampling facilitates the glycan moiety to interact with lipid head groups even in lipid XII, albeit the fraction time is relatively short. During lipid II polymerization, transglycosylation occurs on the outer leaflet of the cytoplasmic membrane18. As shown in the following section, the GTase domain needs to bind two nascent PGs to elongate the lipid-anchored PG precursor: one in the donor site and the other in the acceptor site20. Thus, the non-reducing end of acceptor strand needs to access the reducing end of donor strand for the completion of the transglycosylation step. Our simulation results show that for the elongated forms of lipid II (e.g., lipid VI and XII), the non-reducing end has limited time to access to the membrane surface, which may disfavor its ability to act as the intermolecular nucleophile onto the reducing end of a nascent PG.
For (A) lipid II, (B) lipid VI, and (C) lipid XII systems, the average Z-distances (dot) from the lipid phosphate of the upper leaflet of a Gram-negative cytoplasmic membrane with the standard deviations (error bar; n = 4167 for the lipid II system and n = 8335 for the other systems), and the interaction fraction time (box) of the pyrophosphate (PP, red), glycan chain (MG, blue), and pentapeptide stem (black): l-Ala (A), γ-d-Glu (e), m-DAP (m), and d-Ala (a).
Prior studies have suggested that the pyrophosphate moiety plays an important role in the binding and stabilization of lipid II within positively charged cages of various proteins that process lipid II molecules21,22. Likewise, several residues from GTase domains of PBPs have been observed to coordinate the pyrophosphate moiety of the lipid II-mimetic donor chain20. In our simulation, lipid II, VI, and XII show distinct patterns of pyrophosphate dihedral angles (Fig. 4A–C; see Fig. S2 for the characterized Newman projection of each hot spot). The distribution of dihedral angles of lipid II pyrophosphate is restricted in range (Fig. 4A). Moreover, the PG moieties such as glycan and peptide (head) and the undecaprenyl moiety (tail) of lipid II are preferably found to be on the same side (syn-configuration) (Figs 4E; S2). In contrast, the pyrophosphate moieties in lipid VI and lipid XII become frequently distorted (Fig. 4B,C) in that the head and tail moieties are preferably placed on different sides (anti-configuration) (Fig. 4F). The implication of having syn configurations during transglycosylation is discussed in the next section. It is known that the number of saccharide units within the lipid-anchored PG precursor plays a prominent role in GTase activity23. The catalytic efficiency of lipid II processing is 20–40 times greater than lipid IV, and elongated nascent PGs (longer than lipid IV) are poor GTase substrates. Our results suggest that elongated nascent PGs frequently form unfavorable pyrophosphate angles that cause anti-configuration relative to lipid II within the positively charged cage in GTase domains. In turn, altered angles can potentially have a significant impact on the processing efficiency of elongated nascent PGs.
(A) Two-dimensional pyrophosphate dihedral angle plots of (A) lipid II, (B) lipid VI, and (C) lipid XII. (D) The two dihedral angles of pyrophosphate are defined as α (O11UND-P1-O12-P2) and β (P1-O12-P2-O22MurNAc), and (E) a syn-configuration (purple arrow in A) and (F) an anti-configuration (red arrow in C) of the pyrophosphate moiety.
Dynamics of PBP1b associated with lipid-anchored nascent PG
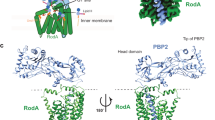
Transglycosylation of PG precursors are proposed to be carried out when two lipid-anchored PG precursors are properly placed in the donor and acceptor sites (Fig. 5A)13. The product of this reaction is the formation of an extended lipid-anchored PG precursor (Fig. 5B). Next, the growing nascent PG is translocated from the acceptor site to the donor site for further transglycosylation reaction(s). In this study, a PBP1b-nascent PG complex was built to represent a final state after translocation of nascent PG, which is an appropriate model to investigate the overall function of PBP1b in the elongation stage. For this purpose, antibiotics (moenomycin and aztreonam) that were originally embedded with PBP1b in the crystal structure (PDB:5HLB) were removed, and lipid XX (20 monosaccharides) was placed in the donor site (Fig. 5A, blue sphere). The placement of the nascent PG was based on the position of moenomycin in the X-ray structure. The PBP1b-lipidXX complex was embedded into a Gram-negative cytoplasmic membrane and the entire system was simulated for 2 μs (Fig. 6; Table S1; see Methods for details).
A proposed mechanism for transglycosylation of lipid II by the GTase domain in PBP1b (white). (A) Unbound PBP1b, active sites of two domains (yellow), and the donor (blue) and acceptor sites (orange) in the GTase domain. (B) PBP1b-lipidII-lipidII ternary complex, PBP1b with turn-over product after glycosylation, and a nascent PG in the donor site after translocation.
A representative snapshot of PBP1b-lipidXX complex embedded in a Gram-negative cytoplasmic membrane (PE:PG = 3:1): white surface for PBP1b, magenta surface for GTase, green surface for TPase, blue sticks for pentapeptide stem, orange sticks for undecaprenyl pyrophosphate, yellow sticks for glycan chain, violet sticks for POPE, and pink sticks for POPG. Water molecules and KCl ions are not shown for clarity.
In the crystal structure of PBP1b, moenomycin is bound to the GTase donor site, and two positive residues in the cationic cage (Lys274 and Arg286) make salt bridges with the moenomycin phosphate moiety (Fig. 7A)14. These results provide critical insight into the mechanism of action of moenomycin by showing that it is actively competing against the GTase substrate. In our study, another lysine residue (Lys287) that was not initially coordinated the pyrophosphate unit (Fig. 7B) became accessible in the course of the simulation (Fig. 7C). The significance of our findings is that Lys287 is relatively more mobile than Lys274 or Arg286, suggesting that Lys287 also plays a role in substrate orientation and accessibility. Consistent with these results, it has been previously determined that mutation of Lys287 to Ala decreases GTase activity by ~40% but does not abrogate it24. Moreover, Lys274 and Arg286 residues have been shown to be critical for enzymatic activity as their mutation to Ala leads to the loss of GTase activity24. The three positive residues together form a positive cage in the donor site, which holds the pyrophosphate dihedral angles preferably in syn configurations (Fig. 4E) during the simulation (Fig. 7D). More specifically, a population of syn configurations is about 74% in the PBP1b-lipidXX complex (Fig. S3). These findings imply that the cationic cage in GTase is suitable for preferably maintaining the pyrophosphate moiety as syn-configurations, which is speculated to be appropriate configuration for the transglycosylation reaction and the favorable configuration for the accessibility of lipid-anchored PG-precursor.
Next, we set out to explore the conformational dynamics and interactions of the PBP1b-lipidXX complex in a Gram-negative cytoplasmic membrane. During the simulation, a tilting motion of the entire protein axis is observed (Fig. 8A); in other words, the tilt angle of PBP1b with respect to the Z-axis (the membrane normal) changes from 12° to 60° frequently as a function of simulation time (Fig. 8B1,2). This tilting motion results in the effective samples of the TPase domain across 30 Å along the Z-axis (Fig. 8C). Such a large spatial sampling potentially allows for the TPase domain of PBP to interact with the existing PG for the transpeptidation reaction onto the growing lipid-anchored nascent PG (Fig. 1F). The GTase site is near the base of PBP. Thus, for the nascent PG to be stitched onto the existing PG, the backbones of the sugar units have to be connected. The tilting motion may provide the accessibility necessary for the large existing PG to reach or dock onto the base of PBP to reach the GTase active site.
(A) Tilt angle with respect to the Z axis measured by the vector connecting the centers of mass of the GTase and TPase domains. (B) Time-series of tilt angle of PBP1b; 1 and 2 correspond to Fig. 1E and F. (C) Time-series of Z-position of the TPase domain’s center of mass from the upper phosphate of the upper leaflet of a Gram-negative cytoplasmic membrane.
While the GTase domain has two separate substrate binding sites, both donor and acceptor sites are connected continuously through a glycan binding groove for the transglycosylation enzymatic activity. Thus, two lipid II molecules need to properly dock into the donor and acceptor sites. In our simulation, the Z-position of bound nascent PG in the donor site (Fig. 9A,C, blue circle) is raised approximately 10 Å relative to isolated (unbound) lipid II (Fig. 9B) with the assumption of a lipid II molecule at the acceptor site (Fig. 9C, red circle). We speculate that the glycan binding groove coordinates the Z-positioning of two substrates, and bind to all glycan strands collinearly. Most importantly, our findings suggest that the association of PBP1b with lipid II serves to properly align the nucleophilic attack of the non-reducing end at the acceptor on the reducing end at the donor (Fig. 9D).
Average Z-distances with standard deviations (error bar) of pyrophosphate (red), MurNAc (green), and GlcNAc (blue) of (A) the bound nascent PG (in the PBP1b-lipidXX complex) and (B) the isolated (unbound) lipid II from the lipid phosphate Z-position. (C) A schematic representation of a nascent PG in the donor site (blue circle), and lipid II in the acceptor site (red circle) of PBP1b with the Z-position of phosphate head groups (red dotted line). (D) A proposed mechanism for lipid II polymerization by PBP1b.
At the initial stage of the transglycosylation, the length of the growing nascent PG is too short to reach the TPase domain. Therefore, several transglycosylation reactions would need to occur prior to processing by the TPase domain. In our simulation, the twisted glycan strand at the beginning of the simulation becomes more stretched out and makes extensive contact with the PBP1b surface as the simulation progresses (Fig. 10A,B), leading to a higher frequency of interactions between the pentapeptide stems and TPase site (Fig. 10C). During the simulation, the 7th and 8th pentapeptide stems (green and purple in Fig. 10B,C) migrated and bound to the TPase site (Fig. 10B, blue Arg555). Presumably electrostatic forces are driving the interaction between the negatively charged nascent PG (Fig. S4A) and the positively charged surface of PBP1b (Fig. S4B). Based on our results, it was determined that approximately 7 or 8 repetitive transglycosylation reactions are required for a distal pentapeptide stem to reach the TPase site.
A series of (A) glycan chain snapshots and (B) snapshots of PBP1b-lipidXX complex as the simulation progresses (cyan for glycan chain; white for PBP1b; blue for Arg555; pink for phosphate in the membrane; purple, green, orange, and red for 7th–10th pentapeptide chains). (C) Interaction patterns of 7th–10th peptide stems with the TPase domain residues from 547 to 556.
Conclusions
We have investigated the structure and dynamics of isolated lipid II, its elongated forms (lipid VI and XII), and the PBP1b-nascent PG complex in Gram-negative bacterial-mimetic membranes to understand how nascent PG structures control PG elongation and processing. Lipid II needs to be polymerized by the GTase domains of PBPs for the proper assembly of PG. The GTase domain of PBP1b is located near the membrane surface, which can impose some structural constraints on the substrate structure. More specifically, the non-reducing end of lipid II in the acceptor site must also be present near the membrane surface for a productive transglycosylation reaction with another nascent PG in the donor site.
As glycan chains of lipid-anchored PG precursors are extended, while fluctuations of the non-reducing end (particular along the membrane normal) appear to increase, the non-reducing end remains distant from the membrane surface and would likely reduce the probability that its elongated nascent PG units are used as an acceptor strand. Glycan chain length also alters the dihedral angle distributions of the pyrophosphate moiety, which could be critical during glycan transfer. During our 2-μs simulation, PBP1b-nascent PG complex shows a prominent tilting motion. As a result, the Z-position of the TPase domain is varied considerably, which can contribute to the binding of the nascent PG with the large existing PG or any of the stem peptides within the growing nascent PG. The non-reducing end of the bound nascent PG is found to interact with the TPase domain for the duration of the simulation, and 7 or 8 elongation steps are needed for a pentapeptide stem to reach the TPase site in PBP1b.
Methods
A lipid II molecule was assembled (Fig. 2A,B) to represent the prototypical Gram-negative PG precursor with m-DAP in the 3rd residue25. In addition, two elongated forms (lipid VI and lipid XII) were modeled (Fig. 2C,D). All saccharides in lipid II/VI/XII were linked with β-(1→4) glycosidic bond, and our naming scheme is based on the number of monosaccharide units26. POPE/POPG mixed bilayer were built to mimic the prototypical Gram-negative cytoplasmic membrane composition (a ratio of PE:PG = 3:1)27, and each lipid II/VI/XII was embedded in the upper leaflet of the bilayer (Fig. S1; Table S1).
A previously published X-ray structure of PBP1b from E. coli (PDB:5HLB) was used to model a lipid anchored nascent PG with PBP1b14. For our modeling, the PBP1b-nascent PG complex was built to represent a final state after translocation of nascent PG. Antibiotics (moenomycin and aztreonam) in the X-ray structure were removed, and lipid XX was placed in the donor site (Fig. 5A, blue sphere) as a nascent PG based on the position of moenomycin in the X-ray structure. The PBP1b-lipidXX complex was embedded into a Gram-negative cytoplasmic membrane (Fig. 6; Table S1).
CHARMM-GUI Membrane Builder protocols were used to build all systems and to prepare the necessary input files for equilibration28,29,30. In all simulations performed in this study, the CHARMM36 force field was used for proteins31, carbohydrates32,33, and lipids34. Also, SHAKE algorithm35 was applied to all bonds containing hydrogen atoms, and temperature and pressure were held at 310.15 K and 1 bar, respectively. For each system, the well-validated Membrane Builder’s six-step equilibrations were initially performed for 375 ps by gradually reducing dihedral (saccharide and undecaprenoid) and planar (phospholipid and water) harmonic restraints using CHARMM36. During the gradual equilibration steps, Langevin dynamics was used for 75 ps with NVT (constant particle number, volume, and temperature) and it was subsequently changed to the following 300 ps with NPT (constant particle number, pressure, and temperature) maintained by Hoover thermostat and Langevin piston. The additional 30-ns NPT equilibrations were performed using NAMD37 to further equilibrate the systems before 1-μs (for lipid II system) and 2-μs (for the other systems) ANTON38 simulations for the data presented in this paper. For Anton simulations, the Multigrator integrator was used for NPT ensemble, which controls the pressure and temperature using the Nosé-Hoover method. A 2-fs time step was used, and simulation frames were saved every 240 ps.
References
Silhavy, T. J., Kahne, D. & Walker, S. The bacterial cell envelope. Cold Spring Harb. Perspect. Biol. 2, a000414, https://doi.org/10.1101/cshperspect.a000414 (2010).
Vollmer, W., Blanot, D. & de Pedro, M. A. Peptidoglycan structure and architecture. FEMS Microbiol. Rev. 32, 149–167, https://doi.org/10.1111/j.1574-6976.2007.00094.x (2008).
Silver, L. L. Multi-targeting by monotherapeutic antibacterials. Nat. Rev. Drug Discov. 6, 41–55, https://doi.org/10.1038/nrd2202 (2007).
Breukink, E. & de Kruijff, B. Lipid II as a target for antibiotics. Nat. Rev. Drug Discov. 5, 321–332, https://doi.org/10.1038/nrd2004 (2006).
Holtje, J. V. Growth of the stress-bearing and shape-maintaining murein sacculus of Escherichia coli. Microbiol. Mol. Biol. Rev. 62, 181–203 (1998).
Middelberg, A. P. & O’Neill, B. K. A correlation for the effective strength of Escherichia coli during homogenization. Biotechnol. Prog. 9, 109–112, https://doi.org/10.1021/bp00019a017 (1993).
Vollmer, W. & Holtje, J. V. The architecture of the murein (peptidoglycan) in gram-negative bacteria: vertical scaffold or horizontal layer(s)? J. Bacteriol. 186, 5978–5987, https://doi.org/10.1128/JB.186.18.5978-5987.2004 (2004).
van Heijenoort, J. Lipid intermediates in the biosynthesis of bacterial peptidoglycan. Microbiol. Mol. Biol. Rev. 71, 620–635, https://doi.org/10.1128/MMBR.00016-07 (2007).
Mengin-Lecreulx, D. & van Heijenoort, J. Effect of growth conditions on peptidoglycan content and cytoplasmic steps of its biosynthesis in Escherichia coli. J. Bacteriol. 163, 208–212 (1985).
Perlstein, D. L., Wang, T. S., Doud, E. H., Kahne, D. & Walker, S. The role of the substrate lipid in processive glycan polymerization by the peptidoglycan glycosyltransferases. J. Am. Chem. Soc. 132, 48–49, https://doi.org/10.1021/ja909325m (2010).
Goffin, C. & Ghuysen, J. M. Multimodular penicillin-binding proteins: an enigmatic family of orthologs and paralogs. Microbiol. Mol. Biol. Rev. 62, 1079–1093 (1998).
Vollmer, W. & Bertsche, U. Murein (peptidoglycan) structure, architecture and biosynthesis in Escherichia coli. Biochim. Biophys. Acta 1778, 1714–1734, https://doi.org/10.1016/j.bbamem.2007.06.007 (2008).
Lovering, A. L., de Castro, L. H., Lim, D. & Strynadka, N. C. Structural insight into the transglycosylation step of bacterial cell-wall biosynthesis. Science 315, 1402–1405, https://doi.org/10.1126/science.1136611 (2007).
King, D. T., Wasney, G. A., Nosella, M., Fong, A. & Strynadka, N. C. J. Structural Insights into Inhibition of Escherichia coli Penicillin-binding Protein 1B. J. Biol. Chem. 292, 979–993, https://doi.org/10.1074/jbc.M116.718403 (2017).
Sarkar, S., Libby, E. A., Pidgeon, S. E., Dworkin, J. & Pires, M. M. In Vivo Probe of Lipid II-Interacting Proteins. Angew. Chem. Int. Ed. Engl. 55, 8401–8404, https://doi.org/10.1002/anie.201603441 (2016).
Schwartz, B., Markwalder, J. A. & Wang, Y. Lipid II: Total synthesis of the bacterial cell wall precursor and utilization as a substrate for glycosyltransfer and transpeptidation by penicillin binding protein (PBP) 1b of Eschericia coli. J. Am. Chem. Soc. 123, 11638–11643, https://doi.org/10.1021/ja0166848 (2001).
VanNieuwenhze, M. S. et al. The first total synthesis of lipid II: The final monomeric intermediate in bacterial cell wall biosynthesis. J. Am. Chem. Soc. 124, 3656–3660, https://doi.org/10.1021/ja017386d (2002).
Ye, X.-Y. et al. Better Substrates for Bacterial Transglycosylases. J. Am. Chem. Soc. 123, 3155–3156, https://doi.org/10.1021/ja010028q (2001).
Zhang, Y. et al. Synthesis of heptaprenyl-lipid IV to analyze peptidoglycan glycosyltransferases. J. Am. Chem. Soc. 129, 3080–3081, https://doi.org/10.1021/ja069060g (2007).
Huang, C. Y. et al. Crystal structure of Staphylococcus aureus transglycosylase in complex with a lipid II analog and elucidation of peptidoglycan synthesis mechanism. Proc. Natl. Acad. Sci. USA 109, 6496–6501, https://doi.org/10.1073/pnas.1203900109 (2012).
Hu, Y. et al. Crystal structure of the MurG:UDP-GlcNAc complex reveals common structural principles of a superfamily of glycosyltransferases. Proc. Natl. Acad. Sci. USA 100, 845–849, https://doi.org/10.1073/pnas.0235749100 (2003).
Hsu, S. T. et al. The nisin-lipid II complex reveals a pyrophosphate cage that provides a blueprint for novel antibiotics. Nat. Struct. Mol. Biol. 11, 963–967, https://doi.org/10.1038/nsmb830 (2004).
Barrett, D. et al. Analysis of glycan polymers produced by peptidoglycan glycosyltransferases. J. Biol. Chem. 282, 31964–31971, https://doi.org/10.1074/jbc.M705440200 (2007).
Terrak, M. et al. Importance of the conserved residues in the peptidoglycan glycosyltransferase module of the class A penicillin-binding protein 1b of Escherichia coli. J. Biol. Chem. 283, 28464–28470, https://doi.org/10.1074/jbc.M803223200 (2008).
Lebar, M. D. et al. Forming Cross-Linked Peptidoglycan from Synthetic Gram-Negative Lipid II. J. Am. Chem. Soc. 135, 4632–4635, https://doi.org/10.1021/ja312510m (2013).
Wang, T.-S. A., Manning, S. A., Walker, S. & Kahne, D. Isolated Peptidoglycan Glycosyltransferases from Different Organisms Produce Different Glycan Chain Lengths. J. Am. Chem. Soc. 130, 14068–14069, https://doi.org/10.1021/ja806016y (2008).
Morein, S., Andersson, A.-S., Rilfors, L. & Lindblom, G. Wild-type Escherichia coli Cells Regulate the Membrane Lipid Composition in a Window between Gel and Non-lamellar Structures. J. Biol. Chem. 271, 6801–6809, https://doi.org/10.1074/jbc.271.12.6801 (1996).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865, https://doi.org/10.1002/jcc.20945 (2008).
Jo, S., Lim, J. B., Klauda, J. B. & Im, W. CHARMM-GUI Membrane Builder for mixed bilayers and its application to yeast membranes. Biophys. J. 97, 50–58, https://doi.org/10.1016/j.bpj.2009.04.013 (2009).
Wu, E. L. et al. CHARMM-GUI Membrane Builder toward realistic biological membrane simulations. J. Comput. Chem. 35, 1997–2004, https://doi.org/10.1002/jcc.23702 (2014).
MacKerell, A. D. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616, https://doi.org/10.1021/jp973084f (1998).
Guvench, O. et al. Additive empirical force field for hexopyranose monosaccharides. J. Comput. Chem. 29, 2543–2564, https://doi.org/10.1002/jcc.21004 (2008).
Guvench, O., Hatcher, E. R., Venable, R. M., Pastor, R. W. & Mackerell, A. D. CHARMM Additive All-Atom Force Field for Glycosidic Linkages between Hexopyranoses. J. Chem. Theory Comput. 5, 2353–2370, https://doi.org/10.1021/ct900242e (2009).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843, https://doi.org/10.1021/jp101759q (2010).
Ryckaert, J. P., Ciccotti, G. & Berendsen, H. J. C. Numerical-Integration of Cartesian Equations of Motion of a System with Constraints - Molecular-Dynamics of N-Alkanes. J. Comput. Phys. 23, 327–341, https://doi.org/10.1016/0021-9991(77)90098-5 (1977).
Brooks, B. R. et al. CHARMM: the biomolecular simulation program. J. Comput. Chem. 30, 1545–1614, https://doi.org/10.1002/jcc.21287 (2009).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802, https://doi.org/10.1002/jcc.20289 (2005).
Shaw, D. E. et al. Anton, a special-purpose machine for molecular dynamics simulation. Commun. ACM 51, 91–97, https://doi.org/10.1145/1364782.1364802 (2008).
Murakami, S., Nakashima, R., Yamashita, E. & Yamaguchi, A. Crystal structure of bacterial multidrug efflux transporter AcrB. Nature 419, 587–593, https://doi.org/10.1038/nature01050 (2002).
Matias, V. R., Al-Amoudi, A., Dubochet, J. & Beveridge, T. J. Cryo-transmission electron microscopy of frozen-hydrated sections of Escherichia coli and Pseudomonas aeruginosa. J. Bacteriol. 185, 6112–6118 (2003).
Acknowledgements
This work was supported by the NSF MCB-1810695, DBI- 1707207, XSEDE MCB070009 (WI), and GM124893-01 (MMP). Anton 2 computer time was provided by the Pittsburgh Supercomputing Center (PSC) through Grant R01GM116961 from the National Institutes of Health. The Anton 2 machine at PSC was generously made available by D.E. Shaw Research.
Author information
Authors and Affiliations
Contributions
S.K. and W.I. conceived and designed the modeling and simulation. S.K. performed the MD simulations, and S.K., M.M.P. and W.I. analyzed the MD simulations. S.K., M.M.P. and W.I. wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, S., Pires, M.M. & Im, W. Insight into Elongation Stages of Peptidoglycan Processing in Bacterial Cytoplasmic Membranes. Sci Rep 8, 17704 (2018). https://doi.org/10.1038/s41598-018-36075-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-36075-y
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.