Abstract
Gene therapy has always been a promising therapeutic approach for Cystic Fibrosis (CF). However, numerous trials using DNA or viral vectors encoding the correct protein resulted in a general low efficacy. In the last years, chemically modified messenger RNA (cmRNA) has been proven to be a highly potent, pulmonary drug. Consequently, we first explored the expression, function and immunogenicity of human (h)CFTR encoded by cmRNAhCFTR in vitro and ex vivo, quantified the expression by flow cytometry, determined its function using a YFP based assay and checked the immune response in human whole blood. Similarly, we examined the function of cmRNAhCFTR in vivo after intratracheal (i.t.) or intravenous (i.v.) injection of the assembled cmRNAhCFTR together with Chitosan-coated PLGA (poly-D, L-lactide-co-glycolide 75:25 (Resomer RG 752 H)) nanoparticles (NPs) by FlexiVent. The amount of expression of human hCFTR encoded by cmRNAhCFTR was quantified by hCFTR ELISA, and cmRNAhCFTR values were assessed by RT-qPCR. Thereby, we observed a significant improvement of lung function, especially in regards to FEV0.1, suggesting NP-cmRNAhCFTR as promising therapeutic option for CF patients independent of their CFTR genotype.
Similar content being viewed by others
Introduction
Cystic fibrosis (CF), the most common life-limiting autosomal-recessive disease in the Caucasian population (1/2,500 newborns), affects more than 80,000 people worldwide1. It is caused by different mutations within the gene encoding for the CF transmembrane conductance regulator (CFTR). Those mutations result in impaired anion secretion and hyper-absorption of sodium ions across epithelia2,3. Chronic lung disease and slow lung degradation are the major contributing factors to both mortality and strongly reduced quality of life4,5. With currently available therapies, the mean survival is between 35 and 45 years6,7. Since the CFTR gene was first cloned in 1989, many efforts have been made to deal with the mutations at a cellular and genetic level8,9. Gene therapy approaches made it quickly to the clinic aiming to deliver viral CFTR-encoding vectors (such as adenoviruses (Ad) or adeno-associated viruses (AAV)) to CF patients10. However, none of the clinical studies and current treatments seem to provide sufficient human (h)CFTR expression to prevent the ultimately lethal CF symptoms in the respiratory tract of CF patients. Furthermore, repeated administration of viral vectors or DNA may lead to the development of unwanted immune reactions, mainly due to viral capsids and vector-encoded proteins10,11,12.
Newly designed viral vectors circumvent those problems and can be administered repeatedly, but from a clinical perspective, the field is still in need of a therapeutic tool that combines efficient expression in lungs and other (affected) organs and cells while avoiding immunogenicity and genotoxicity completely13,14,15. The non-viral CFTR-encoding plasmid–liposome complex pGM169/GL67A has been one of the most promising therapeutical approach used in clinical trial by Alton’s group. Beside some encouraging results, the trial only managed to modestly improve forced expiratory volume in 1 s (FEV1) after repeated administration along with no improvement in patient’s quality of life16,17. Recently, in vitro transcribed (IVT) chemically modified messenger RNA (cmRNA) came into focus, which has the potential to combine striking advantages in a single-stranded molecule18,19. Chemically modified mRNA has been tested for repeated administration, without developing immune responses or losing efficacy, presenting cmRNAhCFTR complexed with biodegradable chitosan-coated PLGA nanoparticles (NPs) as a promising therapeutic for the treatment of CF patients19,20,21. Versatile delivery options of mRNA ensure the unique possibility to utilize NP-cmRNAhCFTR in early infants as well as adults, independent of the underlying CFTR mutation. To the best of our knowledge, we provide the first in vivo studies delivering cmRNAhCFTR to the lungs of CFTR deficient mice (Cftr−/−) by intravenous (i.v.) and intratracheal (i.t.) administration, complexed with NPs. We provide a proof of concept of NP- cmRNAhCFTR mediated, ELISA quantified, hCFTR expression in the lungs of Cftr−/− mice, leading to significantly reduced chloride secretion and, more importantly, restored criticial lung function parameters, including the most important parameter to evaluate mortality and morbidity of CF patients, the forced expiratory volume (FEV) in 1s or 0.1s in small animals, respectively22,23,24.
Materials and Methods
mRNA production
hCFTR was PCR amplified from pcDNA3.hCFTR with primers adding NheI (Fwd: 5′-TTAGCTAGATGCAGAGGTCGCCTC-3′) and KpnI (Rev: 5′-GCGGGTACCTATCTTGCATCTCTTCT -3′) restriction sites to each end. The PCR product was cloned into a poly(A)-120 containing pVAX (pVAX.A120, www.lifetechnologies.com) by sticky-end ligation using the mentioned restriction sites. pVAX.A120 containing hCFTR is referred as pDNAhCFTR throughout this study. For control experiments, DsRed reporter protein was sub-cloned into pVAX.A120 vector from its original vector pDsRed (www.clontech.com). For in vitro transcription (IVT), the plasmids were linearized downstream of the poly(A) tail with XhoI (www.neb.com). IVT reaction was carried out using MEGAscript T7 Transcription kit (www.ambion.com) with an anti-reverse CAP analog (ARCA) at the 5′ end (www.trilink.com). To produce chemically modified mRNA, the following chemically modified nucleosides were added to the IVT reaction in the indicated ratios: uridine-tri-phosphate (UTP) and cytidine-tri-phosphate (CTP) were fully replaced by N1-Methylpseudo-UTP and 5-Methyl-CTP, abbreviated to \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{{\rm{m5C}}}_{1.0}}^{{\rm{h}}{CFTR}}\) and partly replaced by the incorporation of 25% 2-Thio-UTP and 25% 5-Methyl-CTP, respectively, abbreviated to \({{\rm{c}}{\rm{m}}{\rm{R}}{\rm{N}}{\rm{A}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}{/{\rm{m}}5{\rm{C}}}_{0.25}}^{{\rm{h}}CFTR}\) (www.trilink.com). The cmRNAhCFTR and cmRNADsRed were purified using the MEGAclear kit (www.ambion.com) and analyzed for size and concentration using an RNA NanoChip 6000 for Agilent 2100 Bioanalyzer (Supplement, Fig. S1) (ww.agilent.com).
Cell culture and Transfection
CFBE41o− and 16HBE14o- cells (from Gruenert’s lab) were incubated at 37 °C in a humidified atmosphere containing 5% CO2 until they reached 80–90% confluency. Cell lines were washed with cold, sterile PBS and detached by Trypsin-EDTA. Trypsinization was stopped by adding minimum essential medium (MEM; www.thermofisher.com) containing 10% fetal calf serum. Cells were collected and spun down at 500 × g for 5 minutes before resuspension in fresh MEM. One day before transfection, 250,000 cells/well/1 ml were plated in 12-well plates and grown overnight in MEM without antibiotics. At confluence of 70–90%, cells were then transfected with 1000 ng (c)mRNAhCFTR or equivalent (in nmol) pDNAhCFTR using Lipofectamine 2000 (www.invitrogen.com) following the manufacturer’s instructions and after changing the media to the reduced serum media, Opti-MEM (www.thermofisher.com). After 5 hours, the complexes were removed by replacement with fresh culture medium. Cells were kept for 24 h and 72 h before further analyses.
Flow cytometry analyses
All flow cytometry analyses were performed using a BD LSR Fortessa X-20 SORP (www.bdbioscience.com). For detection of hCFTR protein in 16HBE14o- and CFBE41o− cell lines, cells were transfected as described above and subsequently prepared for intracellular staining using a Fixation/Permeabilization Solution Kit as directed in the manufacturer’s instruction (www.bdbioscience.com). As primary antibody mouse anti-human hCFTR clone 596 (1:500, kindly provided by the cystic fibrosis foundation therapeutics Inc.) has been used. As secondary antibody served Alexa Fluor 488 goat anti-mouse IgG (1:1,000, www.lifetechnologies.com). At least 20,000 gated cells per tube were counted. Data were analyzed with FlowJo software, version 10.
Western blot analysis
Protein isolated from cell lines was separated on Bolt NuPAGE 4–12% Bis-Tris Plus gels and a Bolt Mini Gel Tank (all from www.lifetechnologies.com). Immunoblotting for hCFTR was performed by standard procedures according to the manufacturer’s instructions using the XCell II Mini-Cell and blot modules (www.lifetechnologies.com). After blocking for 1 h in 5% dry milk at room temperature, primary antibody against hCFTR clone 596 (1:500, kindly provided by the cystic fibrosis foundation therapeutics Inc.) or anti-GAPDH (1:1000) (www.scbt.com) was incubated overnight, horseradish peroxidase–conjugated secondary antibodies (anti-mouse from www.dianova.com) were incubated for 1 h at room temperature. All blots were processed by using ECL Prime Western Blot Detection Reagents for 30 min exposure time (www.gelifesciences.com). Semiquantitative analysis was performed using the ImageJ software and overexposure has been avoided as per as digital image and integrity policies.
Immunofluorescence
CFBE41o− and 16HBE14o- were plated on a cell culture insert (0.75 × 106 cells per insert) containing a PET membrane (0.4 μm pore size) (www.corning.com) to provide an air-liquid interface. Cells were transfected 12 h after plating with 5000 ng cmRNAhCFTR or equivalent (in nmol) pDNAhCFTR using Lipofectamine 2000 (www.invitrogen.com) according to manufacturer’s instructions. Membranes were cut out from the insert 24 h after transfection, fixed with 4% PFA, blocked with 0.1% BSA and Fc blocker. Blocking was followed by overnight incubation with hCFTR clone 596 (1:250, kindly provided by the cystic fibrosis foundation therapeutics Inc.). As secondary antibody served Alexa Fluor 594 goat anti-mouse IgG (1:250, www.abcam.com, (ab150116)). Membranes were mounted on a coverslip and images were acquired by Zeiss Confocal Laser Scanning Microscope (CLSM) 710 NLO with Zen software.
YFP-based functional assay
CFTR activity following transient transfection of (c)mRNAhCFTR in A549 and CFBE41o− cells was determined using the halide-sensitive yellow fluorescent protein YFP-H148Q/I152L25. CFTR deficient A549 or CFBE14o- cells stably expressing the YFP were plated in 96-well microplates (50,000 cells/well) in 100 µl of antibiotic-free culture medium and, after 6 h, transfected (in case of CFBE14o- reverse transfected) with either plasmids carrying the coding sequence for CFTR or different (c)mRNAhCFTR. For each well, 0.25 µg of mRNA or plasmid DNA and 0.25 µl of Lipofectamine 2000 were pre-mixed in 10 µl of Opti-MEM (www.invitrogen.com) to generate transfection complexes that were then added to the cells. After 24 hours, the complexes were removed by replacement with fresh culture medium. The CFTR functional assay was carried out 24, 48 or 72 h after transfection. For this purpose, the cells were washed with PBS and incubated for 20–30 min with 60 µl PBS containing forskolin (20 µM). After incubation, cells were transferred to a microplate reader (FluoStar Galaxy; www.bmg.labtech.com) for CFTR activity determination. The plate reader was equipped with high-quality excitation (HQ500/20X: 500 ± 10 nm) and emission (HQ535/30M: 535 ± 15 nm) filters for yellow fluorescent protein (www.chroma.com). Each assay consisted of a continuous 14-s fluorescence reading (5 points per second) with 2 s before and 12 s after injection of 165 µl of a modified PBS containing 137 mM NaI instead of NaCl, resulting in a final 100 mM NaI concentration in the well). To determine iodide (I−) influx rate, the final 11 s of the data for each well were fitted with an exponential function to extrapolate initial slope. After background subtraction, cell fluorescence recordings were normalized for the initial average value measured before addition of I−. Maximum slopes were converted to rates of variation of intracellular I− concentration (in mM/s) using the equation: d[I−]/dt = KI[d(F/F0)]/dt, where KI is the affinity constant of YFP for I−, and F/F0 is the ratio of the cell fluorescence at a given time vs. initial fluorescence25.
Whole blood assay
Ethical approval for using whole blood from healthy donor was obtained from Ethics Commission University Clinic of Tuebingen, Germany (349/2013BO2) and experiments were conducted in accordance with relevant guidelines and regulations. Informed consent form (following WHO guideline) was signed by each volunteer (healthy donor) and kept safely by principal investigator for privacy requirement. Blood samples from three healthy donors were collected in EDTA collection tubes (www.sarstedt.com). For each treatment group, 2 ml of EDTA-blood was transferred into 12-well plates and treated accordingly. R848 (Resiquimod, www.sigmaaldrich.com) was added at a concentration of 1 mg/ml to the respective blood positive controls. cmRNAhCFTR and pDNAhCFTR (7 picomol each) were complexed to NPs at a ratio of 1:10. The samples were kept at 37 °C incubator maintaining 5% CO2. At 6 h and 24 h, 1 ml of whole blood was transferred into microtubes containing serum gel (www.sarstedt.com) and spun down at 10,000 × g for 5 min to obtain serum. Sera were stored at −20 °C for further cytokine measurement analyses. Serum was used to conduct IFN-α, TNF-α and IL-8 ELISA at manufacturer’s instruction (www.thermofisher.com).
Animal experiments
All animal experiments were approved by Regierungspräsidium Tübingen, Baden-Württemberg and carried out according to the guidelines of the German Law for the Protection of Animals (File Number: 35/9185.81-2/K3/16). Cftr−/− mice (CFTRtm1Unc) were purchased from Jackson Laboratory (www.jax.org) at the age of 6 to 8 weeks and were maintained under standardized specific pathogen-free conditions on a 12 h light-dark cycle. Food, water as well as nesting material were provided ad libitum. Prior to i.t. spray applications, mice were anesthetized intraperitoneally (i.p.) with a mixture of medetomidine (0.5 mg/kg), midazolam (5 mg/kg) and fentanyl (50 μg/kg). Cftr−/− mice received 40 μg or 80 µg of cmRNAhCFTR or an equivalent amount (calculated using nmols) pDNAhCFTR complexed with chitosan-coated PLGA nanoparticles [Chitosan (83% deacetylated (Protasan UP CL 113) coated PLGA (poly-D,L-lactide-co-glycolide 75:25 (Resomer RG 752 H)) nanoparticles; short: NPs] (Full details of nanoparticles are provided in Supplement Table T1 and Fig. S5) by intratracheal (i.t.) spraying (n = 4), and intravenous (i.v.) injection (n = 4–7) into the tail vein. Mock-treated control Cftr−/− mice received 40 μg or 80 μg cmRNADsRed complexed to NPs (n = 5) by i.v. or i.t administration, respectively, or just 200 μl NPs by both i.v. and i.t. delivery. An antidote with a mixture of naloxone (1.2 mg/kg), flumazenil (0.5 mg/kg) and atipamezol (2.5 mg/kg) was used against anesthetizing reagents. For both interventions, NP-cmRNA and NP-pDNA complexes were administered in a total volume of 200 μl, twice at an interval of 3 days (day 0 and day 3). After 6 days, mice were sacrificed for endpoint analyses. A detailed description of the i.t. procedures are explained in previously published study26.
Pulmonary mechanics
Lung function for each group was evaluated using a FlexiVent® equipped with FX1 module and NPFE extension and was operated by the flexiWare v7.2 software (www.scireq.com). Prior to tracheostomy, mice were anesthetized intraperitoneally as described above. After anesthesia, a 0.5 cm incision was performed in rostral to caudal direction. A flap of skin was retracted, the connective tissue was dissected, and the trachea was exposed. The trachea was then cannulated between the second and third cartilage ring with a blunt-end stub adapter. The mouse was connected to the FlexiVent® system and quasi-sinusoidally ventilated27 with a tidal volume of 10 ml/kg. A breathing frequency of 150 breaths per min was maintained with an inspiratory to expiratory ratio of 2:3.
Airway resistance (Rn), which is dominated by the resistance of the large conducting airways was considered in this study when the coefficient of determination of the model fit was ≥0.9. Compliance (Cst) was calculated straight from deflating arm of the pressure volume (PV) loops and ramp style pressure-driven maneuver (PVr-P). For obtaining FEV0.1 data a NPFE maneuver was performed which results in FV loops and FE-related parameters. The mice lung was inflated by a pressure of +30 cmH2O over 1.2 s and rapidly deflated to a negative pressure of −55 cmH2O to generate an imposed negative expiratory pressure gradient.
Salivary assay
Prior to tracheostomy, anesthetized mice were injected with 50 µl of 1 mM acetylcholine (ACh) in the cheek to stimulate the production of saliva. The fluid was collected via glass capillaries and a chloride assay was performed using the Chloride (Cl−) Assay Kit according to the manufacturer’s protocol (www.sigmaaldrich.com). Briefly, saliva was diluted at a ratio of 1:100 with water in a total volume of 50 µl and subsequently 150 µl chloride reagent was added. After 15 min incubation at room temperature in the dark, absorbance was measured at 620 nm using an Ensight Multimode plate reader (www.perkinelmer.com).
Enzyme-linked immunosorbent assays (ELISAs)
To detect protein levels of hCFTR after i.t. or i.v. injection of differently modified cmRNAhCFTR in Cftr−/− mice (CFTRtm1Unc), the lungs were isolated at day 6 (experimental endpoint), homogenized and lysed in 600 µl RIPA-buffer and 5 µl protease inhibitor cocktail with tubes of the Precellys Ceramic Kit 1.4/2.8 mm at 6,500 rpm for 10 s for a total of three cycles, each interrupted by a 15 s break in a Precellys Evolution Homogenizer for protein isolation (all from www.peqlab.com). Subsequently, supernatants were kept on ice and additionally homogenized 10 times with a 20G needle and incubated for 20 min (www.bdbioscience.com). Lysates were spun down for 20 min at 13,000 × g and 4 °C. The supernatant was collected and stored at −20 °C for further use. Prior to hCFTR ELISA detection, protein concentration was measured using the Pierce BCA protein assay kit (www.thermofisher.com). For each sample, an equal amount of 15 µg whole protein lysate was used. A human CFTR ELISA kit (www.elabscience.com) was used for hCFTR detection according to manufacturer’s instructions.
Real-time RT-PCR
After i.t. or i.v. injection of cmRNAhCFTR the lungs were isolated at day 6 (experimental endpoint), homogenized and lysed with tubes of the Precellys Ceramic Kit 1.4/2.8 mm at 5,000 rpm for 20 s in a Precellys Evolution Homogenizer for subsequent RNA-isolation (all from www.peqlab.com). Reverse transcription of 200 ng RNA was carried out using an iScript cDNA synthesis kit (www.bio-rad.com) and 1:20 dilution of the cDNA product had been used for further experiment. Detection of mRNAhCFTR was performed by SYBR-Green based quantitative Real-time PCR in 15 μl reactions on a ViiA7 (www.lifetechnologies.com). In all involved procedures, we strictly followed the MIQE protocols for RealTime experiments28. Pre- and post-reaction rooms were strictly separated. Reactions were incubated for 10 min at 95 °C, followed by 40 cycles of 15 s at 95 °C and 2 min at 50 °C (annealing and extension), followed by standard melting curve analysis. The following primer pairs were used:
hCFTR fwd 5′-GAGATGCTCCTGTCTCCTGG-3′, rev 5′-CCTCTCCCTGCTCAGAATCT-3′; 18S rRNA fwd 5′-GGGAGCCTGAGAAACGGC-3′, rev 5′-GACTTGCCCTCCAATGGATCC-3′. Differences in mRNA expression between groups were analyzed by pair-wise fixed reallocation randomization tests with REST 2009 software after collection of the data from Viia7.
Immune response in vivo
To assess immune responses to (c)mRNAhCFTR and pDNAhCFTR, C57BL/6 (Jackson Laboratory (www.jax.org)) mice (n = 4 per group) were treated as described for Cftr−/− mice. As positive controls a group of mice received two administrations of E. coli mRNA-NPs (20 µg) i.v. or i.t. C57BL/6 mice received two injections of 20 µg cmRNAhCFTR complexed to NPs i.v. or i.t. After 6 h, 24 h, and 72 h of second injection mice were sacrificed and blood was collected. For cytokine measurement, blood from mice was used to obtain serum using a serum separator (www.sarstedt.com) and tested for IFN-α and TNF-α production as directed in the manufacturer’s instructions (www.thermofisher.com).
Statistics
All analyses were performed using the Kruskal-Wallis test with GraphPad Prism Version 6 (www.graphpad.com). Most of the data are represented as mean ± SD; box plot data are represented as a mean ± minimum to maximum values. P ≤ 0.05 was considered statistically significant.
Results
(c)mRNAhCFTR and hCFTR protein quantification in vitro
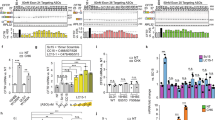
To evaluate the influence of chemical nucleoside modification, we first conducted a set of in vitro analyses to characterize the expression and functionality of hCFTR protein. First, we compared the expression profile of plasmid-encoded hCFTR (pDNAhCFTR), unmodified hCFTR mRNA (mRNAhCFTR) and two well-defined nucleoside modifications (\({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) and \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\)) which have been described to exert state-of-the-art stability/expression in vitro or lung-specific cell contexts in vivo21,29,30,31. Flow cytometry analyses 24 h after transfection of human cystic fibrosis bronchial epithelial (CFBE41o−) cells with pDNAhCFTR, mRNAhCFTR, \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) and \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\), showed hCFTR positive cells (marked as black dots) ranging from 15.8% (pDNAhCFTR) to 49.6% (\({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\)) (P ≤ 0.01; Fig. 1A, lower panel). At 24 h, hCFTR positive cells and hCFTR median fluorescence intensities (MFIs, marked as columns) of (c)mRNAhCFTR were significantly higher compared to pDNAhCFTR (P ≤ 0.05; Fig. 1A, lower panel). At 24 h the total hCFTR expression, defined as median fluorescent intensity (MFI) multiplied by the transfection efficiency, of \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\) was significantly higher compared to pDNAhCFTR and mRNAhCFTR (P ≤ 0.01; Fig. 1A, upper panel). In contrast, after 72 h (c)mRNAhCFTR expressed significantly lower compared to pDNAhCFTR transfected cells, reflected in the percentage of positive cells, MFI and in total hCFTR expression (P ≤ 0.05; Fig. 1B).
(c)mRNAhCFTR and pDNAhCFTR mediated expression of hCFTR in vitro (A) Total expression of hCFTR (calculated by multiplying positive cells (dots) and MFI (bars)) 24 h after transfection with 1 µg (c)mRNAhCFTR and equivalent nmols of pDNAhCFTR detected by flow cytometry. (B) Total expression of hCFTR 72 h after transfection with 1 µg (c)mRNAhCFTR and equivalent nmols of pDNAhCFTR detected by flow cytometry. (C) Western Blots, semi-quantifying human CFTR in transfected CFBE41o- cells, normalized to GAPDH and put relative to CFTR levels in 16HBE14o- cells. Blot section cropped from different blots are delineated with clear dividing lines (black) and full blot of same exposure time (30 mins) are depicted in Supplement Fig. S4. All bar graph data are depicted as means ± SDs while box plots data are depicted as the means ± minimum to maximum values. *P ≤ 0.05 versus unmodified mRNAhCFTR; §P ≤ 0.05 and §§P ≤ 0.01 vs. pDNAhCFTR.
To confirm and substantiate those findings, we performed Western blot analyses of protein lysates taken from transfected CFBE41o- cells at 24 h and 72 h post treatment (Fig. 1C). As a positive control served protein lysate from untransfected 16HBE14o- cells, and GAPDH was used to normalize band intensities. At 24 h pDNAhCFTR transfected CFBE41o- cells showed an average of 22.8% of the protein expression compared to hCFTR observed in 16HBE14o- cells, which increased 4.1-fold to 94.0% at 72 h (P ≤ 0.05; Fig. 1C). This drastic increase of hCFTR expression after pDNA transfection goes well in line with the observations in flow cytometry. As well as the quick onset of hCFTR expression after (c)mRNAhCFTR transfection at 24 h (P ≤ 0.05; Fig. 1C). However, relative to the 24 h time-point, hCFTR expression after 72 h either remained nearly static (mRNAhCFTR resulted in 33.8% and 34.7% expression at 24 h and 72 h, respectively), decreased (\({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) resulted in 45% and dropped to 29.3% hCFTR expression at 24 h and 72 h, respectively) or increased (\({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\), 46.4% at 24 h and raised to 63.3% at 72 h). Ultimately, the expression of hCFTR mRNA in vitro was strongly dependent on its chemical modification, with \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\) resulting in the most robust hCFTR expression among all (c)mRNA transfections (All the blots are separately provided in Supplement Fig. S4).
All in vitro results are also underlined by the conducted immunofluorescence imaging. All tested samples show a higher amount of hCFTR positive cells compared to the negative control (CFBE41o- cells; Fig. 2A). Additionally, transfection with unmodified mRNAhCFTR produced a lower amount of hCFTR positive cells compared to both pDNAhCFTR and cmRNAhCFTR with the highest amount of hCFTR positive cells in the samples transfected with \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\) (Fig. 2A). Looking at the fluorescence image itself transfection of pDNAhCFTR shows a quite dispersed appearance of hCFTR within the cells compared to cmRNAhCFTR transfection seeming to have a higher abundance of hCFTR towards the cell membrane (Fig. 2A, left panel). In general, the Immunofluorescence imaging confirms that transfection with pDNAhCFTR as well as (c)mRNAhCFTR leads to increased levels of hCFTR protein within the transfected cells.
(c)mRNAhCFTR and pDNAhCFTR mediated expression of hCFTR by immunofluorescence and functional hCFTR in vitro and immunogenicity in human whole blood. (A) Detection of hCFTR protein by immunofluorescence (after 24 h), percent of hCFTR expression in pDNAhCFTR or (c)mRNAhCFTR transfected CFBE41o- cells compare to untransfected CFBE41o- and 16HBE14o- cells. Image J has been used for calculating means ± SDs of hCFTR positive cells; (B) Quenching efficacy of pDNAhCFTR or (c)mRNAhCFTR transfected CFBE41o- and CFTR null A549 cells relative to un-transfected controls was measured at 24 h, 48 h and 72 h post-transfection. *P ≤ 0.05 versus un-transfected controls; (C) 2 ml whole blood, each from three different healthy human donors, were incubated with either R848 (1 mg/ml) or 7 pmol pDNAhCFTR or 7 pmol (c)mRNAhCFTR (providing the same total number of nucleic acid molecules) and NPs at a 1:10 ratio; after 6 h and 24 h the immune response was determined by ELISA in the sera; The blue area represents the variance of the negative controls which are biological replicates. n.d., not detectable and red dotted lines mark the detection limit as specified in the respective ELISA kit. All bar graph data are depicted as means ± SDs while box plots data are depicted as the means ± minimum to maximum values. *and §P ≤ 0.05 (§§P ≤ 0.01) versus control at 6 h and 24 h, respectively.
hCFTR (c)mRNA functionality test in vitro
For functional analysis of the (c)mRNAhCFTR -encoded CFTR channel, we performed a YFP-based functional assay using CFTR null A549 cells or ΔF508 CFBE41o- cells which stably express halide-sensitive YFP-H148Q/I152L30. Quenching of the YFP signal induced by hCFTR channel-mediated I− influx is reciprocally proportional to hCFTR channel function25,32. Figure 2B shows the quenching efficacy after transfection of 250 ng (c)mRNAhCFTR, for three different time points, normalized to mock-transfected cells. In pDNAhCFTR transfected cells, the quenching efficacy was significantly higher after 48 h and stayed high even after 72 h (P ≤ 0.05), while mRNAhCFTR as well as modified cmRNAhCFTR transfected cells revealed a single peak quenching at 48 h (P ≤ 0.05), which was undetectable at 72 h in A549 cells. In CFBE41o- cells mRNAhCFTR could not provide any detectable quenching but \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\) produced significant quenching at all the time points (P ≤ 0.05) and \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) showed very significant quenching at 48 h (P ≤ 0.001), which is in line with expression patterns seen in Fig. 1A,B.
(c)mRNAhCFTR immunogenicity ex vivo by an adapted human whole blood assay
Due to lack of a reliable method to detect immune responses that therapeutic mRNAs may trigger in a living organism, we focused on an innovative approach to using whole blood from humans. Blood was collected from three healthy donors and used fresh to conduct whole blood assays. Interestingly, the negative control groups (blood only and NP only) did not raise IFN-α values above the detection limit (Fig. 2C, red dotted lines), while TNF-α and IL-8 were already measurable in human blood untreated or treated only with NPs. That is the reason why we adapted the graphical presentation, using a blue colored area that represents the variance of the negative controls, which are biological replicates. The positive control (R848) lead to a strong and significant production of IFN-α (6 h and 24 h, respectively; P ≤ 0.05), IL-8 (6 h and 24 h, respectively; P ≤ 0.01) and TNF-α (6 h and 24 h, respectively; P ≤ 0.05) (Fig. 2C). All cmRNAhCFTR showed a very similar result in cytokine expression as observed for negative controls: the IFN-α levels did not reach the detection limit of the ELISA; IL-8 and TNF-α responses were not statistically significant at 6 h and 24 h, respectively (Fig. 2C). Unmodified mRNAhCFTR resulted in a significant increase of IFN-α at 6 h and 24 (P ≤ 0.05), only significant increase in IL-8 at 24 hours (P ≤ 0.05) and the TNF-α levels were in line with the negative control. While pDNAhCFTR triggered high TNF-α responses at 6 h (P ≤ 0.05), significant and detectable IFN-α and IL-8 responses after 6 h and 24 h (P ≤ 0.05). Due to both, significantly lower expression of mRNAhCFTR in vitro (Fig. 1) and unwanted higher immune responses of mRNAhCFTR, we focused on cmRNAhCFTR and pDNAhCFTR in the following therapeutic studies.
Therapeutic effect of cmRNAhCFTR in vivo in mice after i.t. and i.v. application
All in vivo experiments were performed with nanoparticles if not stated otherwise. Therapeutic potential of cmRNAhCFTR was investigated in a mouse model of Cystic Fibrosis. Cftr−/− and Cftr+/+ mice have been used in several experimental settings that are explained and color-coded in Fig. 3A. To assess the impact of cmRNAhCFTR on lung function, we evaluated clinically relevant parameters using the FlexiVent® lung function measurement system. We observed significant differences between Mock controls, Cftr−/− and healthy wild-type mice for all parameters measured (P ≤ 0.05; Figs 3 and 4B, P ≤ 0.01; Figs 3 and 4C and P ≤ 0.001 Figs 3 and 4D). I.v. administration of \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) significantly increased the compliance from 0.02 ± 0.01 ml/cmH2O (Cftr−/− mice) to 0.03 ± 0.01 ml/cmH2O (P ≤ 0.05), reaching equivalent values to those measured in Cftr+/+ mice (Fig. 3B). In contrast, the i.v. application of 40 µg \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\) or pDNAhCFTR did not alter compliance significantly. Applying 40 µg of \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) or \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\) i.v. significantly lowered the resistance (P ≤ 0.01 and P ≤ 0.05 respectively, Fig. 3C) but pDNAhCFTR did not alter the resistance at a significant level. FEV0.1 (human equivalent of FEV1) of Cftr+/+ mice defined as projecting 100% forced expiratory volume is pointedly different compared to the FEV0.1 value of Cftr−/− mice of only 66% of the wild-type (P ≤ 0.001). I.v. injection of 40 µg \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) significantly improved the FEV0.1 by 23 percentage points (P ≤ 0.01) and i.v. injection of 40 µg \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\) provided a significant FEV0.1 improvement of 14 percentage points compared to FEV0.1 value of untreated Cftr−/− mice (P ≤ 0.05; Fig. 3D). However, i.v. administration of pDNAhCFTR showed no significant improvement of FEV0.1. I.v. injected mock cmRNADsRED or nanoparticles alone encouragingly aligned with untreated groups in all determined lung function parameters. Taken together, these results demonstrate significant lung function improvement in all relevant lung function parameters of Cftr−/− mice intravenously (i.v.) treated with cmRNAhCFTR.
In vivo lung function measurements in cmRNAhCFTR and pDNAhCFTR treated Cftr−/− mice by i.v. route. All mouse groups utilized in (B–D) are color-coded for their treatment schemes (A), including dosage and application routes. (B–D) Precision in vivo lung function measurements covering all relevant outcome parameters on in Cftr−/− mice treated twice via i.v. route and measured 72 hours after the 2nd instillment; n = 4–7 mice per group. The blue area represents the variance of the negative controls which are biological replicates. Data represent the means ± SD on compliance, resistance and Forced Expiratory Volume in 0.1 seconds (FEV0.1). *P ≤ 0.05; **P ≤ 0.01 and ***P ≤ 0.001 versus untreated Cftr−/− mice.
In vivo lung function measurements in cmRNAhCFTR and pDNAhCFTR treated Cftr−/− mice by i.t. route. All mouse groups utilized in (B–D) are color-coded for their treatment schemes (A), including dosage and application routes. (B–D) Precision in vivo lung function measurements covering all relevant outcome parameters on Cftr−/− mice treated twice via i.t route and measured 72 hours after the 2nd instillment; n = 4–7 mice per group. The blue area represents the variance of the negative controls which are biological replicate. Data represent the means ± SD on compliance, resistance and Forced Expiratory Volume in 0.1 seconds (FEV0.1). *P ≤ 0.05; **P ≤ 0.01 and ***P ≤ 0.001 versus untreated Cftr−/− mice.
In the i.t. treated groups, a substantial improvement in compliance and resistance could be detected when the \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) dose was increased to 80 µg (0.04 ± 0.01 ml/cmH2O and 0.86 ± 0.18 cmH2O.s/ml respectively; P ≤ 0.05; Fig. 4B,C). However, 80 µg of \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\) i.t. lowered the resistance but did not improve the compliance as effectively as \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) (P ≤ 0.05; Fig. 4B,C). pDNAhCFTR (80 µg) i.t. treated mice also produced significant improvements of resistance and compliance (P ≤ 0.05, Fig. 4B,C). In terms of FEV0.1, i.t. application of 80 µg \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) was improved by 19 percentage points and i.t. application of 80 µg \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\) improved the FEV0.1 by 12 percentage points with respect to untreated Cftr−/− mice (P ≤ 0.05, Fig. 4D). I.t. administration of pDNAhCFTR showed no significant improvement of FEV0.1. Taken together, these results demonstrate significant lung function improvement in all relevant lung function parameters of Cftr−/− mice treated intratracheally with cmRNAhCFTR.
A well-established functional test, measuring the mouse saliva chloride concentration33 was conducted to complement the functional results observed using FlexiVent. The saliva chloride concentration detected in Cftr−/− mice (4084 ± 236.8 ng/µl) was significantly higher compared to Cftr+/+ mice (748.8 ± 96.9 ng/µl, P ≤ 0.001; Fig. 5A,B). The treatment of \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) i.v. significantly lowered the chloride concentrations in the saliva of Cftr−/− mice by more than 52 percentage points (P ≤ 0.01; Fig. 5A) underlining the FlexiVent results. However, \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\) and pDNAhCFTR treated mice (i.v.) only provided about 20 percentage points reduction. The treatment with \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) i.t. (80 µg) significantly lowered the chloride concentrations in the saliva of Cftr−/− mice by 36 percentage points (P ≤ 0.01; Fig. 5B). \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\) treated mice (i.t.) abridged the chloride concentration not significantly in saliva of Cftr−/− mice but pDNAhCFTR treated Cftr−/− provided a significant reduction (P ≤ 0.01; Fig. 4B) but not as proficiently as \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\). The mock mRNA treated group and just nanoparticle treated group failed to decrease the chloride concentration.
In vivo saliva chloride concentration measurement of cmRNAhCFTR and pDNAhCFTR treated Cftr−/− mice by i.v./ i.t. route (A,B) Functional test of reconstituted CFTR channel and reduced chloride concentration after i.v. (A) or i.t. (B) treatment of Cftr−/− mice compared to untreated Cftr−/− (black), positive controls (violet), and percentages relative to the positive control; n = 4 mice per group; two mock controls were included (white); boxes represent the means ± minimum and maximum values. The blue area represents the variance of the negative controls which are biological replicates. *P ≤ 0.05; **P ≤ 0.01 versus untreated Cftr−/− mice.
cmRNAhCFTR and hCFTR protein quantification in lungs after application in vivo
All in vivo experiments were performed with nanoparticles if not stated otherwise. We tested for the localization of cmRNAhCFTR complexed with nanoparticle in the lungs after i.t. or i.v. application via RT-qPCR, quantified the hCFTR protein expression with hCFTR ELISA and then evaluated its immunogenicity depending on modification. In contrast to the in vitro data, when 40 µg \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) was i.v. injected into the mice, this resulted in a higher accumulation of that mRNA in the lung as compared to 40 µg \({{\rm{cmRNA}}}_{{\rm{N}}1{{\rm{\Psi }}}_{1.0}/{\rm{m}}5{{\rm{C}}}_{1.0}}^{{\rm{h}}{CFTR}}\) and pDNAhCFTR (P ≤ 0.01, Fig. 6C). More importantly, we wanted to analyze if there is a significant increase in hCFTR protein levels in the lungs of treated mice by hCFTR ELISA (Fig. 6B,E). These analyses confirmed that mice treated with 40 µg \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) i.v. had a highly significant increase of hCFTR protein in the lungs of treated mice vs. control mice (P ≤ 0.01; Fig. 6B). Besides, we tested the effects of an increased amount (80 µg) of cmRNAs and pDNAhCFTR with i.t. instillation, \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) and pDNAhCFTR showed a clear and significant increase of hCFTR protein compared to control mice (Fig. 6E) (P ≤ 0.01). All the mock controls used in hCFTR ELISA have proven to be not significantly different from the negative control.
Expression of hCFTR protein in mouse lungs and delivery of cmRNAhCFTR and pDNAhCFTR in lungs. (A,D) All mouse groups, particles and particle combinations depicted in the study plan are color-coded for their treatment schemes, including dosage and application routes. (B,E) hCFTR ELISA, detecting specifically human CFTR, was performed on lung preparations at day 6 from Cftr−/− mice treated twice via i.v. (B) or i.t. (E) route and measured 72 hours after the 2nd instillment (endpoint); the same n = 4–7 mice per group were used. (C,F) Relative amounts of differently modified hCFTR mRNAs in the lungs applied i.v. or i.t., then determined by RT-quantitative PCR, compared to untreated Cftr−/− mice (*P ≤ 0.05); n = 4–7 mice per group. All bar graph data are depicted as means ± SDs while box plots data are depicted as the means ± minimum to maximum values. The blue area represents the variance of the negative controls which are biological replicates. *P ≤ 0.05; **P ≤ 0.01 and ***P ≤ 0.001 versus untreated Cftr−/− mice.
cmRNAhCFTR immunogenicity in vivo in mice after i.v. application
All in vivo experiments were performed with nanoparticles if not stated otherwise. First, we applied different compounds such as nanoparticles, E. coli extract total RNA (positive control), cmRNAhCFTR and pDNAhCFTR i.v. or i.t. to mice and monitored their immune reaction at three different time points. Applying 40 µg cmRNAhCFTR (with any modifications used) or pDNAhCFTR i.v. or i.t. did not lead to detectable responses of key cytokines IFN-α or TNF-α (detected by ELISA) at all three-time points (Fig. 7A)34,35. Nanoparticles alone (used in all in vivo experiments) showed no immune response over the detection limit. However, as expected the positive control (E. coli extract total RNA) i.v. and i.t. resulted in a significant increase of IFN-α and TNF-α at 6 h and a trend increase of IFN-α at 24 h, while an effect at 72 h was not detectable (Fig. 7A). No immune response had been observed apart from positive control in groups treated intratracheally (i.t) (Fig. 7B).
(c)mRNAhCFTR and pDNAhCFTR mediated immunogenicity in vivo Mice were i.v. or i.t. injected with a mix of (c)mRNA and NPs at a 1:10 ratio and ELISAs were performed post-i.v./i.t.-injection at three different time points. n.d., not detectable. The red dotted lines in (A,B) mark the detection limit as specified in the respective ELISA kit. All bar graph data are depicted as the means ± SD and box plots data are represented as the means ± minimum to maximum values.
Discussion
Although much progress has been achieved since the discovery of the CFTR gene 25 years ago, there is still a substantial need to restore robust CFTR function in patients suffering from cystic fibrosis8. With the recent approvals of the small molecule agents ivacaftor and lumacaftor, science has paved a possible way to overcome the hurdles caused by the disease-conferring gene. Those treatments can be more or less effectively applied to patients bearing CFTR mutations delF508 (Lumacaftor-ivacaftor/Orkambi) and G551D (ivacaftor)36,37,38,39. However, lung function, as one of the main outcome parameters probably having the most significant influence on life quality of CF patients, is rarely tested in preclinical models. In fact, actual effects of (modern) existing drugs on lung function, with forced expiratory volume in one second (FEV1) as a key parameter, are quite low40. Here, by using cmRNAhCFTR, we are presenting a proof of concept for a viable and potent therapeutic alternative. We have vigorously tested mRNA therapy with focus on in vivo lung function normalization while avoiding any possible, unwanted immune responses for a possibility of repeated dosing. The unique formulation utilized can be used both topically (intratracheally) and systemically (via i.v. injection), having in both cases a profound effect on normalizing the lung function parameters, including compliance, resistance and FEV0.1 of treated Cftr−/− mice to values obtained from Cftr+/+ mice.
In vitro, using cmRNAhCFTR, CFTR protein expression in CFBE41o− cells was increased up to 5.5-fold compared to mRNAhCFTR, which is consistent with previous studies obtained by us and others18,31,41. Incorporation of naturally occurring chemically modified nucleosides has been shown to suppress inhibitory effects on translation by avoiding detection by pattern recognition receptors (PRRs) such as Toll-like receptors (TLRs) TLR3, TLR7, and TLR834,35. Those receptors play a crucial role in the detection, processing, and degradation of mRNA. Interestingly, depending on the mRNA modification, kinetics of hCFTR expression varies upon the different nucleosides used. In fact, after 72 h we only observe an increased quenching of Yellow fluorescent protein (YFP) in YFP assay in CFTR null A549 and CFBE41o− cells by pDNAhCFTR which would corroborate our findings from our flow cytometry and western blot analyses in CFBE41o− cells. In contrast there is a significant increase in I− influx by functional hCFTR channels and quenching of YFP at 48 h post transfection by cmRNAhCFTR. Consequently, we assume that upon different cell lines, kinetics by which the hCFTR protein is expressed varies. Earlier studies support our notion that differently modified mRNAs can have an impact on the translational effect between distinct cell lines31,35.
To better mimic in vivo human conditions, we performed an ex vivo whole blood assay (WBA) which offers a more complex environment to test for immune responses. This assay has already been used in a number of preclinical settings, and Coch and colleagues could demonstrate that it has the potential to reflect broad aspects of the in vivo cytokine release caused by oligonucleotides42. Indeed, we could show that the small molecule Resiquimod (serving as a positive control by activating TLR7 and TLR8) lead to a substantial release of IFN-α, TNF-α and IL-8. pDNAhCFTR, as well as unmodified mRNAhCFTR, also showed elevated cytokine levels probably due to the activation of innate immune receptors34,35. In contrast, incorporation of modified nucleosides into hCFTR mRNA (cmRNAhCFTR) abolished such responses, with no detectable amounts of IFN-α. This is in concert with previously published data, demonstrating cmRNA’s limited immune responses, mainly by evading detection from receptors such as TLRs, RIG-1, MDA-5 or PKR34,41. Interestingly, even though TNF-α or IL-8 could be detected, it rather shows donor-dependency than effects deriving from NPs and/or cmRNAhCFTR with cytokine levels being all within the variance of negative controls. Although it mirrors only the blood compartment and does not reflect the more complex in vivo situation, the WBA can give a prediction of how cytokines are released in the human system in response to systemically applied (c)mRNA prior to clinical testing.
To determine the clinical potential of CFTR-encoded cmRNA we compared not only different modifications in vivo but also two different routes of administration. I.t. application has been chosen for this study on the base of our previous findings of applying cmRNA i.t. in a surfactant protein-B deficient mouse model leading to significantly prolonged survival26. Given the fact that in patients suffering from CF one of the key barriers is the airway mucus layer in which inhaled particles are more likely to get trapped and removed, we sought to apply cmRNAhCFTR/pDNAhCFTR complexed to NPs by i.v. injection as an alternative administration route. Systemic delivery via lipid-modified polymeric nanoparticles have been already shown to target the lungs efficiently43.
To support our notion of improved CFTR activity, we performed extensive lung function measurements using state-of-the-art technology to provide detailed in vivo information on different lung function parameters. There are doubts about Cftr−/− mice as a proper model for cystic fibrosis as it does not reflect the typical lung phenotype seen in CF patients44. However, the reason behind that seems to be in how deeply lungs or other affected organs had been investigated. A layer of material can be observed with characteristics of an acid mucopolysaccharide on the bronchiolar surface and is also evident in alveoli by using scanning electron microscopy in Cftr−/− mice, which is not evident in Cftr+/+ mice45. It has also been reported Cftr−/− mice shows similar effect of CF patients like, age-dependent pulmonary inflammation, death of respiratory epithelial cells and high vulnerability to severe Pseudomonas aeruginosa infection46. Recent studies could demonstrate reduced airway compliance and increased resistance in comparison to wild-type mice47,48. Indeed, we observed significantly higher and lower levels regarding resistance and compliance, respectively, in Cftr−/− controls and mock-treated Cftr−/− mice compared to homozygous wild-type mice (Cftr+/+) mice and demonstrated that treatment with cmRNAhCFTR-NPs improved compliance and resistance significantly equal to those seen in healthy Cftr+/+ mice. FEV1 percentage (for mouse or small animal FEV0.1) is related to survival in CF and a most important physiological parameter for CF patients. A previous study demonstrated that patients with a %FEV1 of <30 compared to healthy individuals had a 50% chance of mortality within 2 years and hence are regularly examined in clinical setup49. A strong variance amid Cftr−/− controls and mock-treated Cftr−/− mice compared to homozygous wild-type mice (Cftr+/+) mice has been perceived in the case of FEV0.1. Our study provides a significant improvement of FEV0.1 due to treatment with NP-cmRNAhCFTR. Interestingly, NP-pDNAhCFTR when administered via i.t. route improved parameters of lung function measurements including FEV0.1, but not as significant as cmRNAhCFTR. We also observed i.v. or i.t. administration of \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) to positively compensate most of lung function parameters. Overall, we could demonstrate that certain protocols, applying cmRNAhCFTR either i.v. or i.t. efficiently restored lung function values equal to those of wild-type. Suggesting a more even distribution through arteries and the bronchial circulation by i.v. injection, this route and formulation could lead to a very potent therapy especially for newborns and young infants. By providing functional CFTR early in life, the lungs could be protected from irreversible damage. Nevertheless, when applied intratracheally, which mimics deep inhalation of a spray or powder formulation (primary application route in adults), an adjustment in dose and/or formulation (e.g. \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) increased to 80 µg) might easily abrogate any negative effect of the Cftr−/− genetic background on lung function.
Eventually, we determined the impact of cmRNAhCFTR and pDNAhCFTR on another relevant physiological outcome such as the saliva chloride concentration to evaluate therapeutic effect and complement the lung function results. Sweat chloride concentration has become an accepted method as a diagnostic readout to assess treatment effects of CF patients50. As an analog, chloride concentration of β-adrenergic stimulated salivary glands of Cftr−/− mice can be investigated as it complies with findings in CF patients33. In this study, we could show a substantial difference in salivary Cl− content of cmRNAhCFTR and pDNAhCFTR treated mice – both, i.v. and i.t. – compared to their untreated counterpart. With end point-analysis, a significant decrease in Cl− to nearly 50% was observed, indicating a restoration of CFTR in the duct compartment of salivary glands and thus leading to an improved Cl− absorption. Previous studies estimated that a restoration of CFTR activity to 50% could lead to sweat chloride levels to near normal levels in CF patients. Given that, it is possible that cmRNAhCFTR treatment has the potential to improve CFTR activity to levels that are at least similar to those in patients with a mild CF phenotype51.
In this study, by applying cmRNAhCFTR consecutively, both modifications were successfully delivered to the lungs with the i.v. route being more efficient at doses of 40 µg (2 mg/kg body weight) per treatment. Intriguingly, in contrast to the results obtained in vitro, \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) showed a significantly higher CFTR protein expression with higher accumulation of hCFTR mRNA in lung cells. Assuming differences of cmRNA-encoded transgene expression between distinct cell lines, it is plausible to consider such differences between in vitro versus in vivo applications, which is by far more complex. In this respect, the higher amount of \({{\rm{cmRNA}}}_{{\rm{s}}2{{\rm{U}}}_{0.25}/{{\rm{m5C}}}_{0.25}}^{{\rm{h}}{CFTR}}\) found in lung cells after i.v. injection, might be due to the fact that its nucleoside composition is more favorable to evade PRRs, thus being less degraded. However, regardless of cmRNA kinetics we also observed differences in the delivery route of cmRNAhCFTR/pDNAhCFTR -NPs. Our data suggest i.v. injection to be more efficient in delivering such complexes to the lung than topical administration. Tests of cmRNAhCFTR -NP’s capacity of mucus penetration are in planning phase including detection of cmRNAhCFTR and CFTR protein (glycosylated) in a Cftr-deficient mouse model especially at the apical side of the bronchial epithelium. The upper airways are lined with mucus and mucociliary movements clear foreign particles immediately. In addition, the main barriers in the deeper areas are the alveolar lining, scavenger transporters and alveolar macrophages52,53. We, therefore, concluded that the original dosing by which cmRNA-NPs were delivered i.t. was not as efficient as using the i.v. route. Indeed, increasing the amount by doubling the dose (to 80 µg) for each treatment showed a hCFTR expression close to levels seen using the i.v. route.
To exclude immune reactions caused by either NPs or the cmRNAhCFTR itself, we conducted extensive immune assay tests in vivo. Except for the positive control (E. coli total mRNA), we could not detect any immunostimulatory effect in vivo that could arise from NPs or the cmRNAhCFTR. These results confirm our previous studies in which we showed that NPs, as well as modified mRNA, could be administered safely to the lungs without any substantial increase in cytokines, or inflammatory-related cells such as macrophages or neutrophils26. Systemic delivery has also been reported to have no impact on proinflammatory cytokine secretion29.
Taken together, this study is the first proof of concept of efficient application of NP-cmRNAhCFTR in vivo to restore lung function in a Cftr-deficient mouse model. Importantly, we could neither detect immune responses in vivo nor in a more defined setting ex vivo. Applying cmRNAhCFTR to Cftr−/− mice could efficiently restore lung function close to levels of healthy control mice. In addition, our study compared - apart from two well-known mRNA modifications and pDNAhCFTR - also two different delivery routes, demonstrating that systemic administration of cmRNA targets lung cells more efficiently at lower dosages. This study provides a proof of concept for alternative treatment of patients suffering from CF. cmRNAhCFTR transcript supplementation may be broadly applicable for most CFTR mutations, not only in adults but already in the postnatal state, thereby protecting the lungs from exacerbations from the very beginning of life.
References
Brown, S. D., White, R. & Tobin, P. Keep them breathing: Cystic fibrosis pathophysiology, diagnosis, and treatment. JAAPA: official journal of the American Academy of Physician Assistants 30, 23–27, https://doi.org/10.1097/01.jaa.0000515540.36581.92 (2017).
Cutting, G. R. Cystic fibrosis genetics: from molecular understanding to clinical application. Nat Rev Genet 16, 45–56, https://doi.org/10.1038/nrg3849 (2015).
Mall, M. A. & Galietta, L. J. V. Targeting ion channels in cystic fibrosis. Journal of Cystic Fibrosis 14, 561–570, https://doi.org/10.1016/j.jcf.2015.06.002 (2015).
Havermans, T., Colpaert, K., Vanharen, L. & Dupont, L. J. Health related quality of life in cystic fibrosis: To work or not to work? J Cyst Fibros 8, 218–223, https://doi.org/10.1016/j.jcf.2009.03.002 (2009).
Cohen, M. A., Ribeiro, M. A., Ribeiro, A. F., Ribeiro, J. D. & Morcillo, A. M. Quality of life assessment in patients with cystic fibrosis by means of the Cystic Fibrosis Questionnaire. J Bras Pneumol 37, 184–192 (2011).
MacKenzie, T. et al. Longevity of patients with cystic fibrosis in 2000 to 2010 and beyond: survival analysis of the Cystic Fibrosis Foundation patient registry. Ann Intern Med 161, 233–241, https://doi.org/10.7326/M13-0636 (2014).
Keogh, R. H., Szczesniak, R., Taylor-Robinson, D. & Bilton, D. Up-to-date and projected estimates of survival for people with cystic fibrosis using baseline characteristics: A longitudinal study using UK patient registry data. J Cyst Fibros 17, 218–227, https://doi.org/10.1016/j.jcf.2017.11.019 (2018).
Kerem, B. et al. Identification of the cystic fibrosis gene: genetic analysis. Science (New York, N.Y.) 245, 1073–1080 (1989).
Riordan, J. R. et al. Identification of the cystic fibrosis gene: cloning and characterization of complementary DNA. Science (New York, N.Y.) 245, 1066–1073 (1989).
Bellon, G. et al. Aerosol administration of a recombinant adenovirus expressing CFTR to cystic fibrosis patients: a phase I clinical trial. Human gene therapy 8, 15–25, https://doi.org/10.1089/hum.1997.8.1-15 (1997).
Zuckerman, J. B. et al. A phase I study of adenovirus-mediated transfer of the human cystic fibrosis transmembrane conductance regulator gene to a lung segment of individuals with cystic fibrosis. Human gene therapy 10, 2973–2985, https://doi.org/10.1089/10430349950016384 (1999).
Thaci, B., Ulasov, I. V., Wainwright, D. A. & Lesniak, M. S. The Challenge for Gene Therapy: Innate Immune Response to Adenoviruses. Oncotarget 2, 113–121 (2011).
Takaoka, A. et al. DAI (DLM-1/ZBP1) is a cytosolic DNA sensor and an activator of innate immune response. Nature 448, 501–505, https://doi.org/10.1038/nature06013 (2007).
Anson, D. S. The use of retroviral vectors for gene therapy-what are the risks? A review of retroviral pathogenesis and its relevance to retroviral vector-mediated gene delivery. Genetic vaccines and therapy 2, 9, https://doi.org/10.1186/1479-0556-2-9 (2004).
Griesenbach, U., Geddes, D. M. & Alton, E. W. Gene therapy for cystic fibrosis: an example for lung gene therapy. Gene Ther 11(Suppl 1), S43–50, https://doi.org/10.1038/sj.gt.3302368 (2004).
Alton, E. et al. Repeated nebulisation of non-viral CFTR gene therapy in patients with cystic fibrosis: a randomised, double-blind, placebo-controlled, phase 2b trial. The Lancet. Respiratory medicine 3, 684–691, https://doi.org/10.1016/s2213-2600(15)00245-3 (2015).
Alton, E. et al. In A randomised, double-blind, placebo-controlled trial of repeated nebulisation of non-viral cystic fibrosis transmembrane conductance regulator (CFTR) gene therapy in patients with cystic fibrosis (NIHR Journals Library Copyright (c) Queen’s Printer and Controller of HMSO 2016. This work was produced by Alton et al. under the terms of a commissioning contract issued by the Secretary of State for Health. This issue may be freely reproduced for the purposes of private research and study and extracts (or indeed, the full report) may be included in professional journals provided that suitable acknowledgement is made and the reproduction is not associated with any form of advertising. Applications for commercial reproduction should be addressed to: NIHR Journals Library, National Institute for Health Research, Evaluation, Trials and Studies Coordinating Centre, Alpha House, University of Southampton Science Park, Southampton SO16 7NS, UK., 2016).
Mays, L. E. et al. Modified Foxp3 mRNA protects against asthma through an IL-10-dependent mechanism. J Clin Invest 123, 1216–1228, https://doi.org/10.1172/JCI65351 (2013).
Antony, J. S., Dewerth, A., Haque, A., Handgretinger, R. & Kormann, M. S. Modified mRNA as a new therapeutic option for pediatric respiratory diseases and hemoglobinopathies. Mol Cell Pediatr 2, 11, https://doi.org/10.1186/s40348-015-0022-6 (2015).
Zangi, L. et al. Modified mRNA directs the fate of heart progenitor cells and induces vascular regeneration after myocardial infarction. Nature biotechnology 31, 898–907, https://doi.org/10.1038/nbt.2682 (2013).
Kormann, M. S. et al. Expression of therapeutic proteins after delivery of chemically modified mRNA in mice. Nature biotechnology 29, 154–157, https://doi.org/10.1038/nbt.1733 (2011).
Corey, M. & Farewell, V. Determinants of mortality from cystic fibrosis in Canada, 1970-1989. American journal of epidemiology 143, 1007–1017 (1996).
Zemanick, E. T. et al. Measuring and improving respiratory outcomes in cystic fibrosis lung disease: opportunities and challenges to therapy. Journal of cystic fibrosis: official journal of the European Cystic Fibrosis Society 9, 1–16, https://doi.org/10.1016/j.jcf.2009.09.003 (2010).
Szczesniak, R., Heltshe, S. L., Stanojevic, S. & Mayer-Hamblett, N. Use of FEV(1) in Cystic Fibrosis Epidemiologic Studies and Clinical Trials: A Statistical Perspective for the Clinical Researcher. Journal of cystic fibrosis: official journal of the European Cystic Fibrosis Society 16, 318–326, https://doi.org/10.1016/j.jcf.2017.01.002 (2017).
Galietta, L. V., Jayaraman, S. & Verkman, A. S. Cell-based assay for high-throughput quantitative screening of CFTR chloride transport agonists. American journal of physiology. Cell physiology 281, C1734–1742 (2001).
Mahiny, A. J. et al. In vivo genome editing using nuclease-encoding mRNA corrects SP-B deficiency. Nature biotechnology 33, 584–586, https://doi.org/10.1038/nbt.3241 (2015).
Devos, F. C. et al. Forced expiration measurements in mouse models of obstructive and restrictive lung diseases. Respir Res 18, 123, https://doi.org/10.1186/s12931-017-0610-1 (2017).
Bustin, S. A. et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clinical chemistry 55, 611–622, https://doi.org/10.1373/clinchem.2008.112797 (2009).
Pardi, N. et al. Administration of nucleoside-modified mRNA encoding broadly neutralizing antibody protects humanized mice from HIV-1 challenge. Nature Communications 8, 14630, https://doi.org/10.1038/ncomms14630, https://www.nature.com/articles/ncomms14630#supplementary-information (2017).
Pardi, N. et al. Expression kinetics of nucleoside-modified mRNA delivered in lipid nanoparticles to mice by various routes. Journal of controlled release: official journal of the Controlled Release Society 217, 345–351, https://doi.org/10.1016/j.jconrel.2015.08.007 (2015).
Andries, O. et al. N(1)-methylpseudouridine-incorporated mRNA outperforms pseudouridine-incorporated mRNA by providing enhanced protein expression and reduced immunogenicity in mammalian cell lines and mice. Journal of controlled release: official journal of the Controlled Release Society 217, 337–344, https://doi.org/10.1016/j.jconrel.2015.08.051 (2015).
Goda, K. & Szaloki, G. A sensitive tool to measure CFTR channel activity. Cytometry. Part A: the journal of the International Society for Analytical Cytology 83, 528–529, https://doi.org/10.1002/cyto.a.22290 (2013).
Droebner, K. & Sandner, P. Modification of the salivary secretion assay in F508del mice–the murine equivalent of the human sweat test. J Cyst Fibros 12, 630–637, https://doi.org/10.1016/j.jcf.2013.05.001 (2013).
Kariko, K., Buckstein, M., Ni, H. & Weissman, D. Suppression of RNA recognition by Toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA. Immunity 23, 165–175, https://doi.org/10.1016/j.immuni.2005.06.008 (2005).
Kariko, K. et al. Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability. Mol Ther 16, 1833–1840, https://doi.org/10.1038/mt.2008.200 (2008).
Cholon, D. M. et al. Potentiator ivacaftor abrogates pharmacological correction of DeltaF508 CFTR in cystic fibrosis. Science translational medicine 6, 246ra296, https://doi.org/10.1126/scitranslmed.3008680 (2014).
Van Goor, F. et al. Rescue of CF airway epithelial cell function in vitro by a CFTR potentiator, VX-770. Proceedings of the National Academy of Sciences of the United States of America 106, 18825–18830, https://doi.org/10.1073/pnas.0904709106 (2009).
Clancy, J. P. et al. Results of a phase IIa study of VX-809, an investigational CFTR corrector compound, in subjects with cystic fibrosis homozygous for the F508del-CFTR mutation. Thorax 67, 12–18, https://doi.org/10.1136/thoraxjnl-2011-200393 (2012).
Ramsey, B. W. et al. A CFTR potentiator in patients with cystic fibrosis and the G551D mutation. The New England journal of medicine 365, 1663–1672, https://doi.org/10.1056/NEJMoa1105185 (2011).
Cholon, D. M., Esther, C. R. Jr. & Gentzsch, M. Efficacy of lumacaftor-ivacaftor for the treatment of cystic fibrosis patients homozygous for the F508del-CFTR mutation. Expert review of precision medicine and drug development 1, 235–243, https://doi.org/10.1080/23808993.2016.1175299 (2016).
Anderson, B. R. et al. Incorporation of pseudouridine into mRNA enhances translation by diminishing PKR activation. Nucleic acids research 38, 5884–5892, https://doi.org/10.1093/nar/gkq347 (2010).
Coch, C. et al. A Human In Vitro Whole Blood Assay to Predict the Systemic Cytokine Response to Therapeutic Oligonucleotides Including siRNA. PLOS ONE 8, e71057, https://doi.org/10.1371/journal.pone.0071057 (2013).
Kaczmarek, J. C. et al. Polymer-Lipid Nanoparticles for Systemic Delivery of mRNA to the Lungs. Angewandte Chemie (International ed. in English) 55, 13808–13812, https://doi.org/10.1002/anie.201608450 (2016).
Cohen, T. S. & Prince, A. Cystic fibrosis: a mucosal immunodeficiency syndrome. Nature medicine 18, 509–519, https://doi.org/10.1038/nm.2715 (2012).
Kent, G. et al. Lung disease in mice with cystic fibrosis. J Clin Invest 100, 3060–3069, https://doi.org/10.1172/jci119861 (1997).
Teichgraber, V. et al. Ceramide accumulation mediates inflammation, cell death and infection susceptibility in cystic fibrosis. Nature medicine 14, 382–391, https://doi.org/10.1038/nm1748 (2008).
Geiser, M., Zimmermann, B., Baumann, M. & Cruz-Orive, L. M. Does lack of Cftr gene lead to developmental abnormalities in the lung? Experimental lung research 26, 551–564 (2000).
Craig Cohen, J., Lundblad, L. K., Bates, J. H., Levitzky, M. & Larson, J. E. The “Goldilocks Effect” in Cystic Fibrosis: identification of a lung phenotype in the cftr knockout and heterozygous mouse. BMC Genetics 5, 21, https://doi.org/10.1186/1471-2156-5-21 (2004).
Kerem, E., Reisman, J., Corey, M., Canny, G. J. & Levison, H. Prediction of mortality in patients with cystic fibrosis. The New England journal of medicine 326, 1187–1191, https://doi.org/10.1056/nejm199204303261804 (1992).
Gonzalo-Ruiz, J. et al. Early determination of cystic fibrosis by electrochemical chloride quantification in sweat. Biosensors & bioelectronics 24, 1788–1791, https://doi.org/10.1016/j.bios.2008.07.051 (2009).
Rowe, S. M., Accurso, F. & Clancy, J. P. Detection of Cystic Fibrosis Transmembrane Conductance Regulator Activity in Early-Phase Clinical Trials. Proceedings of the American Thoracic Society 4, 387–398, https://doi.org/10.1513/pats.200703-043BR (2007).
Paranjpe, M. & Muller-Goymann, C. C. Nanoparticle-mediated pulmonary drug delivery: a review. International journal of molecular sciences 15, 5852–5873, https://doi.org/10.3390/ijms15045852 (2014).
Patton, J. S. et al. The particle has landed–characterizing the fate of inhaled pharmaceuticals. Journal of aerosol medicine and pulmonary drug delivery 23(Suppl 2), S71–87, https://doi.org/10.1089/jamp.2010.0836 (2010).
Acknowledgements
We thank Dr. Dominik Hartl for experimental guidance on CFTR KO mice; Dr. Sandra Beer-Hammer and Dr. Franz Iglauer for helping to draft the respective animal proposal; late Dr. Joachim Riethmüller for the numerous and fruitful discussions on translation of hCFTR mRNA therapy into the clinic; Dr. Markus Mezger for human whole blood assay; Katrin Ganzenberg for mRNA isolation from various organs of Cftr mice. This work was supported by the European Research Council (ERC Starting Grant to M.S.D.K., 637752 “BREATHE”), by the Deutsche Forschungsgemeinschaft (DFG KO 4258/2-1, to M.S.D.K. and Lauren Mays), Fortüne Grant (No. 1980-0-0, to M.S.D.K.) by the European Respiratory Society (Maurizio Vignola Award, to M.S.D.K.), in part by the Mukoviszidose e.V. (S03/12 to M.S.D.K.) and by the Italian Cystic Fibrosis Foundation (Grant FFC No. 2/2015 to N.P.).
Author information
Authors and Affiliations
Contributions
A.H., A.D., J.S.A. and M.S.D.K. designed the experiments. A.H., A.D., J.S.A., G.R.S., P.W., N.L., N.P., E.S., A.R. and J.L. performed experiments. A.H., A.D. and B.W. analyzed the data. P.S., C.S., H.Y., B.L., C.M.L. and R.H. provided material. A.H., A.D., J.R. and M.S.D.K. interpreted the data. A.H. A.D., J.R. and M.S.D.K. drafted the paper, and A.H. and M.S.D.K. wrote the final version of the paper.
Corresponding author
Ethics declarations
Competing Interests
M.S.D.K. holds a patent on RNA modification (EP2459231B1). M.S.D.K., A.H., A.D. and J.S.A., hold a European patent on delivery of cmRNAhCFTR complexed with nanoparticles (17169561.2-1401).
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Haque, A.K.M.A., Dewerth, A., Antony, J.S. et al. Chemically modified hCFTR mRNAs recuperate lung function in a mouse model of cystic fibrosis. Sci Rep 8, 16776 (2018). https://doi.org/10.1038/s41598-018-34960-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-34960-0
Keywords
This article is cited by
-
Assessment of respiratory mechanics and X-ray velocimetry functional imaging in two cystic fibrosis rat models
Scientific Reports (2024)
-
Therapeutic nucleic acids in regenerative medicine and tissue repair
Nano Research (2024)
-
mRNA biotherapeutics landscape for rare genetic disorders
Journal of Biosciences (2024)
-
mRNA nanodelivery systems: targeting strategies and administration routes
Biomaterials Research (2023)
-
Species-agnostic polymeric formulations for inhalable messenger RNA delivery to the lung
Nature Materials (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.