Abstract
Dynamic movements of the cardiac troponin complex are an important component of the cardiac cycle. Whether cardiac troponins are subjected to irreversible advanced glycation end-product (AGE) modification is unknown. This study interrogated human and rat cardiac troponin-C, troponin-I and troponin-T to identify endogenous AGE modifications using mass spectrometry (LC-MS/MS). AGE modifications were detected on two amino acid residues of human troponin-C (Lys6, Lys39), thirteen troponin-I residues (Lys36, Lys50, Lys58, Arg79, Lys117, Lys120, Lys131, Arg148, Arg162, Lys164, Lys183, Lys193, Arg204), and three troponin-T residues (Lys107, Lys125, Lys227). AGE modifications of three corresponding troponin-I residues (Lys58, Lys120, Lys194) and two corresponding troponin-T residues (Lys107, Lys227) were confirmed in cardiac tissue extracts from an experimental rodent diabetic model. Additionally, novel human troponin-I phosphorylation sites were detected (Thr119, Thr123). Accelerated AGE modification of troponin-C was evident in vitro with hexose sugar exposure. This study provides the first demonstration of the occurrence of cardiac troponin complex AGE-modifications. These irreversible AGE modifications are situated in regions of the troponin complex known to be important in myofilament relaxation, and may be of particular pathological importance in the pro-glycation environment of diabetic cardiomyopathy.
Similar content being viewed by others
Introduction
The cardiac troponin complex facilitates cardiomyocyte contraction via dynamic interaction between the troponin subunits and the thin myofilaments, actin and tropomyosin. Troponins control the positioning of tropomyosin on the actin filament to regulate myosin-actin cross-bridge formation in response to Ca2+. The trimeric complex comprises troponin-C, which binds Ca2+ in low affinity binding sites during systolic Ca2+ influx; troponin-I, which inhibits the actin-myosin interaction and is tightly regulated by phosphorylation; and troponin-T, which anchors the troponin complex into the thin filament structure1. Post-translational modification of the troponin subunits is an important mechanism for mediating modulation of cardiomyocyte contractility in health and disease2, and novel reports of citrullination3, O-GlcNAcylation4,5, S-nitrosylation6,7, and phosphorylation8,9,10,11 sites on troponins have emerged in recent years. Post-translational modification of cardiac proteins for biomarker evaluation is also gaining interest12,13,14. To date, only dynamic post-translational modifications have been investigated, as a mode of reversible signaling regulation of the troponin complex. Susceptibility of troponins to irreversible post-translational modifications such as advanced glycation end-product (AGE) formation, has not been previously evaluated.
Troponin mutations are detected in many inherited cardiomyopathies15, providing evidence that small changes in the troponin protein sequence may have major implications for cardiac function. Both Ca2+ sensitizing and Ca2+ desensitizing troponin mutations have been identified, associated with either a hypertrophic/restrictive cardiomyopathy or a dilated cardiomyopathy phenotype respectively15. Similarly, modified troponin 3-dimensional structure due to the presence of glycated amino acid residues would be expected to contribute to alterations in Ca2+ sensitivity and could underlie acquired cardiopathologies such as heart failure and/or diabetic cardiomyopathy. Surprisingly, no studies to date have investigated this proposition - perhaps reflecting a presumption that the relatively short half-life of troponins (3–5 days)16 would preclude a functional impact of a reputedly slow ‘permanent’ glycation modification (conventionally considered to be of importance in the context of more stabilized macroprotein entities). Yet emerging evidence suggests that intracellular cardiac proteins with normally high turnover rates are indeed modified by glycation. Mass spectrometry studies have shown that sarcoplasmic reticulum Ca2+ ATPase and ryanodine receptor proteins are present in AGE modified states in vivo in diabetic rat hearts17,18,19, despite half-lives of 5–8 days20. The mechanism and time-course of intracellular AGE-modification is not well established, and presence of glycated proteins considered to be relatively high turnover may reflect AGE-induced delayed protein degradation and/or an environment of accelerated AGE formation.
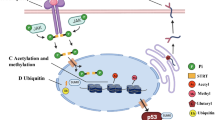
AGEs are formed by non-enzymatic free sugar attachment to proteins and these adducts are converted through oxidation into fixed moieties which have potential to confer protein deformation and rigidity21. A physiological role for AGEs is unknown. Conflicting hypotheses have been posed that AGEs may act as either signals for protein degradation22 or inhibitors of protein degradation23. A growing body of evidence suggests that AGEs are primarily a form of environment-induced pathological protein damage with no known enzymatic processes for addition or removal. Although extracellular AGE-crosslinking of myocardial collagen is well-described24, few studies have investigated the role of AGE formation on intracellular proteins.
The rationale for investigation of troponin-AGE involvement in diabetic cardiomyopathy etiology is compelling. Not only do diabetic cardiomyocytes exhibit the pro-glycation environment of increased AGE precursors and oxidative stress, but increased exposure to fructose sugar, a potent glycation agent, may also play a role. An understanding of cardiomyocyte fructose actions is emerging25,26, and high myocardial fructose levels (up to 60-fold elevated) in diabetic rat hearts has been reported, likely driven by upregulation of the sorbitol pathway27. In type 2 diabetic rats, myocardial levels of Nε-carboxymethyllysine (CML), a common AGE modification, are correlated with impaired left ventricular relaxation28. In particular AGE-modification of troponin proteins has potential to impair the trimeric complex operation and contribute to cardiomyocyte dysfunction in pathological settings.
The goal of this study was to investigate human cardiac troponins (TnC, TnI and TnT), seeking evidence of endogenous AGE modifications using mass spectrometry screening techniques. As an outcome of AGE discovery our aim was to link the amino acid residue modification sites to functional domains in the protein sequence, and thus obtain biological insight relating to potential cardiac functional impact of these irreversible post-translational modification types. Our studies provide the first evidence that all three troponin subunits exhibit endogenous AGE-modification and demonstrate that human cardiac troponin-I is particularly susceptible to glycation events, occurring at amino acid sites of functional importance. Two novel phosphorylation sites (Thr119 and Thr123) on human cardiac troponin-I have been identified. Additionally, susceptibility of human cardiac troponin-C to accelerated AGE formation under in vitro conditions of high glucose and fructose exposure was established.
Results
The investigative strategy firstly involved analyses of human-derived troponin subunits to identify occurrence of all glycated residues. This was followed by experimental confirmatory analyses of troponin glycation in cardiac tissue extract of control and diabetic rodents. Then an exploration of the positioning of glycation sites relative to phosphorylation sites detected was undertaken. Next, we evaluated the identified glycated residues in relation to known functional domains of each troponin. Finally, we performed experiments to assess residue susceptibility to glycation in a pro-glycation in vitro environment.
AGE modification of human cardiac troponins
To determine AGE modification sites on human cardiac troponins, liquid-chromatography tandem mass spectrometric analysis (LC-MS/MS) of trypsin-digested purified human cardiac troponin-C, troponin-I and troponin-T was performed. All AGE modifications detected above identity threshold are detailed in Table 1. Troponin-C exhibited only 2 modified residues, CML modification of Lys6, and Lys39. Troponin-I exhibited numerous glycated residues: 13 amino acid residues modified by AGEs were identified. Lys36, Lys50, Lys58, Lys117, Lys120, Lys131, Lys164, Lys183 and Lys193 were modified by CML, Lys120 was also modified by Nε-carboxyethyllysine (CEL), and Arg79, Arg148, Arg162 and Arg204 were modified by methylglyoxal-derived hydroimidazolone (MG-H). Exemplar spectra for two of the troponin-I peptides are shown in Fig. 1 each in unglycated and glycated forms – Lys193 (Fig. 1A,B) exhibited low detection frequency and Lys120 (Fig. 1C,D) exhibited high detection frequency as is evident in Table 1. An annotated expansion of Fig. 1A,B is provided in the Supplementary Information (Fig. S1) to allow for closer comparison and inspection of unglycated versus glycated peptide spectra.
Novel AGE modification sites on human cardiac troponin-I (TnI). (A) MS/MS spectrum of trypsin-digested TnI unglycated peptide 193–204 (m/z 645.828). (B) MS/MS spectrum of trypsin-digested TnI peptide 193–204 with CML modification (+29 m/z = +58 Da) of Lys193 (m/z 674.830), confirmed by 4 peptides. (C) MS/MS spectrum of trypsin-digested TnI unglycated peptide 118–131 (m/z 787.444). (D) MS/MS spectrum of trypsin-digested TnI peptide 118–131 with CML modification (+29 m/z = +58 Da) of Lys120 (m/z 816.446), confirmed by 23 peptides. (E) MS/MS spectrum of AspN-digested TnI unglycated peptide 113–126 (m/z 772.931). (F) MS/MS spectrum of AspN-digested TnI peptide 113–126 with CML modification (+29 m/z = +58 Da) of Lys120 (m/z 801.934), confirmed by 7 peptides. (G) MS/MS spectrum of trypsin-digested rat TnI unglycated peptide 119–132 (m/z 787.443) corresponding to human TnI unglycated peptide 118–131. (H) MS/MS spectrum of trypsin-digested rat TnI peptide 119–132 with CML modification (+29 m/z = +58 Da) of Lys121 (m/z 816.449) corresponding to human TnI Lys120. The b-ions denote N-terminal ions and y-ions denote C-terminal ions. CML, N ε-carboxymethyl-lysine.
Given the abundance of glycation sites on troponin-I, the endoproteinase AspN was used to digest troponin-I at aspartate and cysteic acid residues to confirm that AGE detection was not dependent on trypsin cleavage at Lys and Arg residues. As for trypsin digestion, AspN-digested troponin-I exhibited CML modification of Lys120 in both unglycated and glycated form (Fig. 1E,F). AspN digestion also yielded evidence of 2 additional CML modification sites, Lys117 and Lys183, which were not detected with trypsin-digestion (Table 1). Exploratory investigations were undertaken to establish evidence of AGE occurrence in an in vivo experimental setting. Crude tissue extract was prepared from myocardium of control and type1 diabetic rodents (8 weeks duration, streptozotocin treated). AGEs were detected on Lys59, Lys121, Lys194 of troponin-I derived peptides from diabetic but not control samples (Table S1). These sites represent the rodent equivalent of corresponding AGE modification sites identified on human troponin-I (Lys58, Lys120, Lys193). For these troponin-I AGE modifications, which could only be detected in the diabetic myocardium, selected exemplar spectra of unglycated and glycated troponin-I (Lys121) are provided in Fig. 1G,H. Human troponin-T exhibited 3 AGE modification sites with CML modification detected at Lys107, Lys125 and Lys227. Figure 2 shows spectra for those residues where multiple glycation detection events were observed. Thus, Lys227 and Lys125 in both unglycated and glycated form are shown in Fig. 2A–D. Three AGE modification sites were detected on troponin-T isolated from control and diabetic rat homogenate (Lys109, Lys202, Lys228) confirming two of the corresponding AGE-modification sites identified on human troponin-T (Lys107, Lys227). In addition, one new AGE-modification site was detected at Lys202 (Table S1).
Novel AGE modification sites on human cardiac troponin-T (TnT). (A) MS/MS spectrum of trypsin-digested TnT unglycated peptide 227–240 (m/z 832.461). (B) MS/MS spectrum of trypsin-digested TnT peptide 227–240 with CML modification (+29 m/z = +58 Da) of Lys227 (m/z 861.464), confirmed by 4 peptides. (C) MS/MS spectrum of trypsin-digested TnT unglycated peptide 124–134 (m/z 666.374). (D) MS/MS spectrum of trypsin-digested TnT peptide 124–134 with CML modification (+29 m/z = +58 Da) of Lys125 (m/z 695.377), confirmed by 3 peptides. The b-ions denote N-terminal ions and y-ions denote C-terminal ions. CML, N ε-carboxymethyl-lysine.
Identification of novel phosphorylation sites on human cardiac troponin-I using LC-MS/MS
LC-MS/MS analysis of trypsin-digested human cardiac troponin-I peptides identified 7 phosphorylation sites (Table 2 and Supplement Information). Analogous unmodified peptides were detected for all reported phosphorylation sites. Phosphorylation sites previously reported in the literature (Ser23/24, Thr51, Ser150, Ser166, Ser199) were confirmed in the purified human cardiac troponin-I samples in this study. The previously-reported dual Ser42/44 troponin-I phosphorylation events were not detected in these samples, likely due to the small size of the tryptic peptide containing these residues. Novel unreported troponin-I phosphorylation sites (Thr119 and Thr123) were also identified. Spectra of these troponin-I peptides in the presence and absence of phosphorylation at Thr119 and Thr123 are shown in Fig. 3A–D. Interestingly, these novel phosphorylation sites are in close proximity to the CML- and CEL-modification site Lys120 shown in Table 1 and Fig. 1D,F, suggesting that AGE modification at Lys120 has the potential to interfere with regulatory phosphorylation sites. All spectra where novel phosphorylation occurrence was detected on troponin-I (Thr119 and Thr123) are provided in the Supplementary Information.
Novel phosphorylation sites on human cardiac TnI. (A) MS/MS spectrum of trypsin-digested TnI unphosphorylated peptide 118–131 (m/z 787.444). (B) MS/MS spectrum of trypsin-digested TnI peptide 118–131 with phosphorylation (+40 m/z = +80 Da) of Thr119 (m/z 827.422), confirmed by 3 peptides. (C) MS/MS spectrum of trypsin-digested TnI unphosphorylated peptide 121–136 (m/z 945.521). (D) MS/MS spectrum of trypsin-digested TnI peptide 121–136 with phosphorylation (+40 m/z = +80 Da) of Thr123 (m/z 985.500). The b-ions denote N-terminal ions and y-ions denote C-terminal ions. Phos, phosphorylation.
Location of AGE- and phosphorylation-modification sites on the human cardiac troponin complex
As depicted in Fig. 4A, several AGE-modification sites identified by LC-MS/MS on the troponin subunits are located within protein-protein interaction regions of the 3-dimensional structure of the troponin complex. Figure 4B presents the identified phosphorylation sites on the 3-dimensional structure of the troponin complex. Figure 5A depicts the domain structure of the troponin subunits showing that CML-modified troponin-C amino acids, Lys6 and Lys39, are located within the N-terminal domain. Figure 5B illustrates that several glycated residues are situated adjacent to phosphorylated residues in the troponin-I sequence. Specifically, CML-modified Lys50 and Lys58 are in close proximity to the phosphorylation site Thr51. CEL/CML-modified Lys120 is located adjacent to the novel phosphorylation site Thr119 and near to Thr123. Glycation sites within the mobile domain (MG-H-Arg162, CML-Lys164, CML-Lys183, CML-Lys193, and MG-H-Arg204) may have important implications for movement of this domain in response to Ca2+ and for phosphorylation of Ser166 and Ser199. Figure 5C illustrates that the CML-modified Lys107, Lys125 and Lys227 are located within the central- and C-domain of troponin-T. Thus, glycated troponin residues may modify interaction of the troponin subunits and influence phosphorylation-mediated signaling regulation of the troponin complex.
Amino acid residue location of identified AGE and phosphorylation modification sites on human cardiac troponin complex proteins. (A,B) 3-dimensional structure of cardiac troponin-C (TnC, green), troponin-I (TnI, aqua) and troponin-T (TnT, mauve). Modification sites are shown in red. Location of Ca2+ binding pockets indicated by dark blue circles. TnI R204MG-H, TnI S23/24Phos, TnI S199Phos, TnT K107CML and TnT K125CML are not shown as the 3-dimensional structure is not resolved for the C-terminal domain of TnI and N-terminal of TnT. CML, N ε-carboxymethyl-lysine; CEL, N ε-carboxyethyl-lysine; MG-H, methylglyoxal-derived hydroimidazolone; Phos, Phosphorylation.
Domain structure of human troponin complex proteins indicating identified AGE and phosphorylation modification sites detected by LC-MS/MS. (A) Domain structure of human cardiac troponin-C. Ca2+ binding sites are depicted in green: dark green, high-affinity Ca2+ binding site; light green, low affinity Ca2+ binding sites. (B) Domain structure of human cardiac troponin-I. ID, inhibitory domain; RD, regulatory domain. Detected phosphorylation sites in blue, known but not detected site in grey. (C) Domain structure of human cardiac troponin-T. CML, N ε-carboxymethyl-lysine; CEL, N ε-carboxyethyl-lysine; MG-H, methylglyoxal-derived hydroimidazolone; P, phosphorylation.
In vitro high glucose or fructose incubation of human cardiac troponin-C accelerates AGE-formation
To experimentally examine whether glycation of human cardiac troponin could be promoted by hexose incubation in vitro, and to evaluate site-specific ‘glycation-vulnerability’, human purified cardiac troponin-C was incubated with 2 M glucose or 2 M fructose for 7 days at 37 C. Of the three purified troponin preparations, only troponin-C is soluble in the absence of urea. Thus troponin-C is suitable for in vitro investigations as urea-induced carbamylation of potential AGE sites (Lys and Arg) is avoided. Control (phosphate-buffered saline, PBS), glucose- and fructose-incubated troponin-C samples were trypsin-digested and processed by LC-MS/MS. Given that the conventional view is that the formation of AGEs can take weeks to months, we hypothesized that hexose attachments (AGE initiation) or Amadori products (AGE-precursors) might be evident. We anticipated that it would be unlikely that AGE formation (e.g. CML) would be observed within this time period. Surprisingly, CML-modification of troponin-C was observed after only 7-days of incubation with glucose or fructose (Fig. 6A–C). To identify the human cardiac troponin-C amino acid residues with the highest susceptibility to AGE formation, the LC-MS/MS fragmented peptide spectra were analyzed for 5–7 replicates per group (PBS control, 2 M glucose or 2 M fructose) and the percentage of replicates with AGE-modified amino acid residues are presented in Fig. 6A–C. The frequency of CML detection on amino acid residues of troponin-C was slightly higher in glucose-incubated samples, relative to PBS control (for residues Lys6, Lys17, Lys39, Lys92 and Lys106), although not all CML-modified residues reached the identity threshold of detection (Fig. 6A,B). Fructose-incubated troponin-C had the highest frequency of replicates with CML-modified amino acid residues, and detection of CML-modified Lys17, Lys92 and Lys106 was observed in 100% of replicates (Fig. 6C).
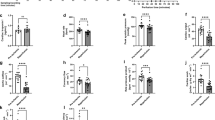
In vitro exposure to glucose and fructose increases AGE-modification of human cardiac troponin-C (TnC). (A–C) Percentage of technical replicates with detection of a CML modification at lysine residues of purified human cardiac TnC using LC-MS/MS (total 5–7 technical replicates per condition). Dark grey: modified, detection above identity threshold; light grey: modified, detection below identity threshold. CML, carboxymethyllysine. (D–F) Number of TnC MS/MS peptide spectra with hexose, CML, or oxidation modifications as a percentage of the total number of TnC MS/MS spectra detected for a given peptide (n = 5 replicates/group). Criteria for identifying a modification: above homology threshold and expect value < 0.5. Data presented as mean ± SEM. One-way ANOVA, Bonferroni post-hoc tests, *p < 0.05.
Spectral counting was performed to semi-quantify the hexose-, CML- and oxidation-modifications observed on troponin-C in response to glucose and fructose exposure. Troponin-C peptides containing the most frequently detected modified amino acid residues (Lys17, Lys92 and Lys106) were counted and presented as the number of modified peptides as a percentage of the total number of peptides (modified and unmodified). Comparisons were made between incubation groups for any single modification site. No differences in hexose modification were observed between PBS control, glucose or fructose incubated troponin-C for Lys17 and Lys92 sites (Fig. 6D). In contrast, hexose-modified Lys106 was significantly higher in the glucose-incubated troponin-C samples relative to fructose (44.4 ± 13.4% vs 8.0 ± 4.9%, p < 0.05, Fig. 6D). Considering all glycation sites, less than 2% of troponin-C peptides were modified by CML in the PBS control group. Glucose incubation increased CML modification to 10.5 ± 5.2% at the Lys92 site (p < 0.05, Fig. 6E). Fructose-incubated troponin-C exhibited the greatest extent of CML modification with 33.9 ± 5.8% of Lys92 and 71.3 ± 8.7% of Lys106 containing peptides detected with a CML-related mass shift (p < 0.05, Fig. 6E). Oxidation of methionine residues in troponin-C peptides was prominent in all groups. Relative to PBS incubated troponin-C, oxidation significantly increased in fructose incubated troponin-C derived Met103, and Met120/137 containing peptides (p < 0.05, Fig. 6F).
Collectively these findings suggest that glucose exposure may promote the initiation of AGE modification of troponin-C evidenced by increased hexose attachments, while fructose may accelerate AGE formation beyond the hexose initiation step evidenced by higher CML modification coincident with higher oxidation of methionine residues (and lower pre-AGE formation hexose attachments). These findings also confirm that AGE formation on troponin-C can occur within 7 days, a finding consistent with the observation that intracellular proteins of relatively short half-life exhibit AGE modification in vivo.
Discussion
This study provides the first evidence that all subunits of the human cardiac troponin complex are modified by endogenous AGE formation. Our data demonstrate that human cardiac troponin-I is particularly susceptible to AGE modification, with 13 glycation sites detected, many of which are located adjacent to phosphorylation sites or within protein-protein interaction regions. The findings show that some, but not all, troponin lysine and arginine residues exhibit AGE modification indicating that particular residues are more vulnerable to glycation than others. Analysis of troponins isolated from rodent hearts revealed troponin-I derived from diabetic but not control rat myocardium exhibited AGE modification. These studies have also revealed preliminary evidence for two novel phosphorylation sites on human cardiac troponin-I (Thr119 and Thr123). In vitro experiments with human cardiac troponin-C have demonstrated that AGE formation is promoted in a hexose-enhanced incubation setting, and that fructose in particular is an AGE accelerant. Collectively these findings provide new evidence that the human cardiac troponin complex is a potential target for glycation events at numerous sites, including locations of key importance for cardiomyocyte mechano-operation. Such molecular events may be of particular relevance in pathological settings, including diabetic cardiomyopathy, where hexose intracellular milieu is disturbed and reactive oxygen species levels are elevated.
During each cardiomyocyte activation cycle a complex sequence of molecular conformational changes involving the trimeric troponins underpins the conversion of an electrically-driven Ca2+ signal to a contractile outcome for the cardiac pump. Ca2+ binds to troponin-C and troponin-I changes conformation to reveal the myosin binding site on actin, thus allowing cross-bridge formation to occur. Dissociation of Ca2+ from troponin-C initiates contractile cycle relaxation by allowing troponin-I to reposition and re-instate the suppression of myosin-actin interaction. AGE-modification of troponins has considerable capacity to disturb the rapid cyclic dynamic movements of the troponin subunits, altering Ca2+-troponin-C association/dissociation kinetics, and disrupting protein-protein interactions. As apparent in Figs 4 and 5, the potential conformational impact of an AGE modified residue on troponin complex subunit positioning, and the spatial relationships between AGE- and phosphorylation responsive residues, provides for substantial pathophysiologic substrate.
All AGE modifications detected on troponin-I are located within protein binding sites, thus it is likely that binding of troponin-I to other troponins in the complex, and/or to myofilaments, would be affected. In the present study, multiple CML and MG-H sites were detected on troponin-I in the C-terminal tropomyosin positioning element. Mutations in this region elicit myofilament Ca2+ sensitizing effects and predispose to hypertrophic and/or restrictive cardiomyopathy phenotypes, where systolic function is maintained but diastolic function declines29,30,31,32. AGE modification within these regions may predispose for similar effects, consistent with increased Ca2+ sensitivity and diastolic dysfunction in diabetic cardiomyopathy. Based on the extensive literature relating to troponin-I mutation effects, the case for future studies designed to assess the potential modulation of contractile performance by troponin-I glycation is compelling.
CML is detected in the troponin-I binding site for the C-lobe of troponin-C, the secondary actin/tropomyosin binding site and the mobile domain of troponin-I. The positive charge of Lys and Arg residues is neutralized by AGE modification, potentially reducing hydrophilicity of the troponin-I domains33,34. Previous studies have identified charged amino acid residues as crucial elements of the troponin subunits for ensuring effective binding between all three subunits and for interaction with actin/tropomyosin35,36,37. AGE-induced neutralization of Lys and Arg in binding sites between the three subunits may decrease the structural stability of the complex and lead to uncoupling of Ca2+ binding and actomyosin cross-bridge formation. AGE-modification of the secondary actin/tropomyosin binding domain on troponin-I may also interfere with its role in Ca2+ dissociation and actin inhibition38. Neutralization of critical charged Lys and Arg residues by AGEs located in the mobile domain of troponin-I may be associated with slowed and/or incomplete myofilament relaxation and increased diastolic cardiomyocyte stiffness.
Several troponin-I domains are intrinsically disordered34,39,40, thus their conformation is highly responsive to the structural influence of post-translational modifications41,42. Key signaling kinases, protein kinase C, protein kinase A, protein kinase G, and p21 activated kinase 3, phosphorylate troponin-I resulting in modulation of myofilament Ca2+ sensitivity and contractility in response to various stimuli10,43. In the present study, five AGE-modified troponin-I residues are identified within two amino acids of a phosphorylation site, thus it could be expected that AGE-modification may interfere with signaling regulation of the troponin complex. In addition to confirmation of well-known and recently identified phosphorylation sites on troponin-I (Ser23/24, Thr51, Ser150, Ser166, Ser199), this study provides preliminary evidence for two novel phosphorylation sites (Thr119, Thr123). Interestingly, these sites span the identified robust CML and CEL modification of Lys120. Given that these residues are located within the troponin-I/troponin-T α-helical coiled coil1, phosphorylation and AGE-modification at these sites may be important for troponin complex structural backbone stability and tropomyosin anchoring. Further analysis of the Thr119 and Thr123 phosphorylation sites is now warranted, including identification of the relevant kinase(s) and characterization of functional implications.
Troponin-C and troponin-T exhibited fewer endogenous AGE-modification sites than troponin-I. CML-modification was detected on 2 troponin-C amino acid residues (Lys6 and Lys39) and 3 troponin-T amino acid residues (Lys107, Lys125 and Lys227). Spectral counting methods revealed that the overall percentage of troponin-C peptides with CML modification was very low (<2%) but increased after 1 week of in vitro hexose exposure (~70% for Lys106 exposed to fructose). Glucose incubation appeared to promote the initiation of AGE modification of troponin-C evidenced by increased hexose attachments on Lys106, while fructose accelerated AGE formation beyond the hexose initiation step, evidenced by lower hexose attachments but higher CML modification coincident with higher oxidation of methionine residues. Small changes in AGE modification of cardiac troponins would be expected to influence protein structure and function - a recent study demonstrated that a 6% change in Ser199 pseudo-phosphorylation status of troponin-I is sufficient to alter myofilament Ca2+ sensitivity8. These findings now prompt further investigation. As noted above, due to differing ionization efficiencies between peptides comparison could not be made between AGE modification sites. Development of site-specific quantitation methodology is required for further comparative analysis of in vitro and in vivo AGE accumulation. Further exploration of troponin-C AGE vulnerability is required, including more extensive proteolytic studies involving AspN. Additionally, systematically assessing the role of ambient hexose and oxidative stress conditions on AGE formation will yield important pathophysiologic insight. The in vitro hexose conditions utilized in this study are clearly artificial, contrived to achieve strongest evidence of AGE-site susceptibility. Similarly, while the atmospheric incubation environment utilized provides a starting point, a range of conditions should be investigated and the influence of subcellular compartmentalization, concentrating hexose and reactive oxygen species considered.
The 7 day in vitro incubation findings support the contention that AGE formation is more rapid than generally understood (previously estimated as weeks to months)44, further corroborated by the extensive observation of AGE modifications of troponin proteins considered to have short life times. AGE formation may delay protein turnover - in diabetic states where pronounced and sustained intracellular metabolic stress is evident, proteins and any accompanying AGE precursors (hexose, methylglyoxal, glyoxal, α- dicarbonyls) may persist longer in cytosolic locations due to dysfunction of intracellular degradative processes such as the glyoxalase45 and ubiquitin-proteasome system46 or autophagy pathways47. It could be expected that AGE modification of intracellular cardiac proteins may be promoted in the pro-glycation environment of altered hexose flux and oxidative stress in diabetic cardiomyocytes. Based on the ‘proof of concept’ data generated in this investigation, detailed evaluation of site-specific AGE modification accumulation and associated cardiomyocyte functional deficits in human and rodent diabetic heart tissues is warranted.
Limitations
While robust, the findings reported in this investigation involve aspects of inherent bias – both sampling and technical. As a ‘proof of concept’ study, the evaluation of purified proteins involved samples derived from a limited number of human subjects. More extensive studies are required to evaluate a range of tissue samples obtained from diabetic and non-diabetic patients. To maximize identification of AGE modification sites present in the samples evaluated in this study, technical replicates were utilized, and this may confer false positive risk. Conversely, a relatively conservative algorithm was employed to establish detection events of AGE-modified proteins, with a high threshold ‘identity’ requirement. This approach may produce an AGE ‘under-detection’ bias, the extent of which can only be determined by further investigation. These considerations highlight the imperative for further exploration of the role of troponin AGE-modification in myocardial pathophysiology.
Conclusions
This is the first study to demonstrate that all three human cardiac troponin subunits can exhibit AGE modification. Human cardiac troponin-I appears most susceptible to glycation, with thirteen modified amino acid residues detected. Given the importance of troponin-I in mediating the Ca2+-sensitive troponin complex conformational change with each cardiac cycle, it is likely that obstructive irreversible glycation modifications would elicit effects on cardiomyocyte function. Troponin-I AGE modifications were detected in regions of protein-protein interaction, adjacent to phosphorylation sites and in key regions of conformational change. The findings presented in this study generate new biological insight relating to potential cardiac functional impact of irreversible post-translational modification of cardiac troponins and provide the platform for new in vivo investigations to advance understanding of the functional outcomes of cardiac troponin AGE modification. In particular, investigations into the relative AGE abundance on cardiac troponins in a diabetic setting, where hexose flux disturbance and oxidative stress coincide to generate a pro-glycation environment, will be an important priority. Glycation of cardiac troponins may represent a novel mechanism underlying cardiomyocyte relaxation defects in diabetic cardiomyopathy, and offer an important new target for therapeutic intervention. Further studies of AGE occurrence in cardiac disease states are now warranted.
Methods
Animal model
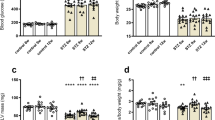
Sprague Dawley rats purchased from the Animal Resource Centre (WA, Australia) were housed in polypropylene cages in a temperature-controlled room at a 12:12 hour light and dark cycle at the Biomedical Sciences Animal Facility at the University of Melbourne and provided access to water and standard chow ad libitum for the duration of the experiment. Type 1 diabetes was induced in male rats at 8 weeks of age via a single 55 mg/kg tail vein injection of streptozotocin (STZ; Sigma) with weight and blood glucose monitored for 8 weeks. Hyperglycemia was present in STZ-treated animals from 2 days post injection and at 8 weeks post-injection, all STZ rats exhibited blood glucose >30 mmol/L vs. control animals 6.45 ± 0.16 mmol/L. At 8 weeks after diabetes induction, rats were euthanized by cervical dislocation (isoflurane anaesthesia), hearts were excised, and left ventricles were dissected and frozen at 80 °C. Frozen tissue was homogenized in 100 mM Tris·HCl, 5 mM EGTA, and 5 mM EDTA (Sigma-Aldrich) buffer containing protease and phosphatase inhibitors (Roche, Switzerland). STZ and Control hearts (N = 4 animals/group, n = 2 technical replicates/ heart) were used to generate in vivo rat TnI glycation data. All animals were cared for in accordance with the Australian Code of practice for the Care and Use of Animals for Scientific Purposes (NHMRC, 2013). All procedures conducted as part of this study were approved by the Animal Ethics Committee of The University of Melbourne (Approval Numbers:, 1613861).
Preparation of cardiac troponins for LC-MS/MS acquisition
Troponin-C, troponin-I and troponin-T were purified from human cardiac tissue harvested from non-myocardial infarction human donors by Life Diagnostics (PA, USA). Human troponin-I, C and T samples were each typically purified from a single donor heart. Each aliquot acquired from a troponin I (n = 10), C (n = 5) or T (n = 5) sample was considered a technical replicate for these studies. Troponin-I and C were purified by urea extraction of cardiac muscle, ion-exchange chromatography followed by hydrophobic affinity chromatography (TnC) or troponin-C affinity chromatography (TnI). Troponin-T was purified by salt extraction and fractionation of cardiac muscle followed by ion-exchange chromatography. The purity was confirmed by the manufacturer to be >98% for troponin-I and troponin-T, and >95% for troponin-C. For analysis of endogenous AGE- and phosphorylation-modifications, coomassie-stained troponin bands were excised from SDS-PAGE gels, diced into 1 mm3 cubes, destained for 2 hours with 50% acetonitrile in 50 mM triethylammonium bicarbonate buffer (TEAB), dehydrated with 100% acetonitrile for 30 minutes, reduced with 10 mM tris(2-carboxyethyl)phosphine (TCEP, Life Technologies, VIC, Australia) for 45 minutes at 55 °C and alkylated with 50 mM iodoacetamide (proteomics grade Life Technologies, Vic, Australia) for 30 minutes, shielded from light. After alkylation, gel pieces were washed 3 times in 50 mM TEAB for 10 minutes, dehydrated with 100% acetonitrile for 30 minutes and digested overnight in sequencing-grade trypsin (25 µg/mL, Sigma-Aldrich, MO USA) or sequencing-grade AspN (40 μg/mL; Promega, WI USA) in 25 mM TEAB at 37 °C with shaking. Samples were then acidified with 1% formic acid and centrifuged at 16,000 g for 10 minutes. The supernatant (20 μL) containing digested peptides was then transferred to Exigen vials, and stored at 4 oC pending LC-MS/MS acquisition48,49.
To determine whether particular lysine or arginine residues are more ‘glycation-vulnerable’ purified human cardiac troponin-C (0.137 µg/µL, Life Diagnostics, PA, USA) was incubated in PBS alone or containing 2 M glucose or 2 M fructose at 37 oC sealed under ambient atmospheric conditions for 7 days. Short incubation pilot experiments showed that hexose attachments formed within 2 days in 2 M glucose or 2 M fructose conditions so a longer 7 day incubation was selected to assess the formation of AGE-modifications. Following the incubation period, troponin-C (4 μg) was diluted with 50 mM TEAB, and reduced with 5 mM TCEP (Life Technologies, Vic, Australia) at 60 oC for 10 minutes. Samples were digested in sequencing-grade trypsin (5ug/ml; 1:50 protease: protein ratio; Sigma-Aldrich, MO, USA) at 37 oC overnight in TEAB with shaking followed by acidification in 1% v/v formic acid. Following centrifugation (16,000 g, 10 minutes), the supernatant (20 µL) containing digested peptides was transferred to Exigen vials, and stored at 4 oC pending LC-MS/MS acquisition48.
LC-MS/MS acquisition and analysis
Digested peptides were separated by liquid chromatography using a nanoflow-reversed-phase-HPLC (Ultimate 3000 RSLC; Dionex, CA, USA). The digested peptides were loaded onto an Acclaim Pepmap nano-trap column (C18, 100 Å, 75 μm × 2 cm; Dionex, CA, USA) with 3% acetonitrile/0.1% formic acid (5 μL/min, 6 min) and switched in-line to an Acclaim Pepmap RSLC analytical column (C18, 100 Å, 75 μm × 50 cm; Dionex, CA, USA) with solvent A, 0.1% formic acid and solvent B, 100% acetylnitrile/0.1% formic acid (gradient: 3–20% B for 95 min, 20-40% B for 10 min, 40–80% B for 5 min, and 80% B for 5 min). All spectra were acquired in positive mode with full scan MS with spectra range m/z 375–1400 at 70,000 resolution, automatic gain control target of 3e6 ions, maximum accumulation time of 50 ms and a lock mass of 445.120024 m/z. The 15 most intense peptide ions with charge states ≥2–5 were isolated (1.2 m/z window) and fragmented with normalized collision energy of 30 and spectra acquired in positive mode MS with 17,500 resolution, automatic gain control target of 1e5 ions, and maximum accumulation time of 100 ms. Mass spectra of peptide ion fragments were acquired using a Thermo Scientific OrbiTRAP Elite MS/MS, or a Thermo Scientific Q-Exactive plus mass spectrometer.
Mass spectra were collected and compiled using the MSILE2 platform (Bio21, University of Melbourne) and bioinformatics analysis performed using the MASCOT Pipeline (Matrix Science, London, UK)50. Peptide spectra were compared against theoretical in silico peptide signatures in the Mass Spectrometry protein sequence Database (MSDB; 53,659,159 sequences) to identify sequence coverage of troponin-C, troponin-I and troponin-T. MSDB database trawling was restricted to i) Homo sapien (purified human troponin-C, troponin-I and troponin-T samples), and ii) peptides were produced via trypsin proteolysis (C-term lysine and arginine cleavage only) or AspN proteolysis (N-terminal aspartic acid and cysteine cleavage only) with up to 2 missed cleavages per peptide permitted. Peptide scores were assigned based on the number of ions detected and matched with theoretical ions, and signal intensity relative to background noise. Peak match tolerance was 0.2 Da. An identity threshold score was assigned to represent a peptide score with 95% confidence. Criteria used to determine a protein match included: combined score >1000, 5 or more unique peptides detected, peptide scores greater than identity threshold scores and false discovery rate <5%. Peptide identity scores were determined using a ‘probability based scoring’ algorithm to evaluate the observed match between the experimental data and the database sequence50. The corresponding identity threshold calculation was based on the number of peptides that fell within the precursor mass tolerance window and the significance threshold that was selected (0.05). Probing for post-translational modification sites were limited to hexose, CML, CEL, MG-H, and phosphorylation (MASCOT bioinformatic searching platform). Methionine oxidation was selected as a variable modification. Carbamidomethyl cysteine was selected as a fixed modification as it is a result of iodoacetamide incubation during sample preparation (capping of cysteine residues to inhibit re-formation of disulphide bonds following TCEP reduction). Modifications were confirmed by manually matching theoretical ion masses to spectra with known mass shifts relating to hexose +162 Da, CML + 58 Da, CEL + 72 Da, and MG-H + 54 Da. Phosphorylation sites were identified as mass shifts of +80 Da, and/or by neutral loss scanning of phosphoric acid (−98 Da). Doubly charged peptides will present with half the mass shift e.g.(+29 for CML) and triply charged peptide ions will present a third of the expected mass shift e.g. (+19.3 for CML). Quantification of hexose, AGE- and oxidation-modification of troponin-C was performed by counting modified peptide spectra normalized to total (modified and unmodified) peptide counts (above homology threshold) for a particular peptide of interest. Amino acid residues were numbered as presented in the Uniprot database (UniProtKB - P63316 (TNNC1_HUMAN), P19429 (TNNI3_HUMAN), P45379 (TNNT2_HUMAN)) where full sequence data may be obtained.
Statistical analyses
Data are presented as mean ± sem. Statistical analyses were performed using GraphPad Prism V7.0 (GraphPad, CA, USA). Data were analyzed by one-way ANOVA with Bonferroni post-hoc tests where appropriate. A p-value of < 0.05 was considered statistically significant.
Data Availability
Full spectral dataset will be made available online (Scientific Data https://www.nature.com/sdata/).
References
Katrukha, I. A. Human cardiac troponin complex. Structure and functions. Biochemistry (Mosc) 78, 1447–1465, https://doi.org/10.1134/S0006297913130063 (2013).
Solaro, R. J., Henze, M. & Kobayashi, T. Integration of troponin I phosphorylation with cardiac regulatory networks. Circ Res 112, 355–366, https://doi.org/10.1161/CIRCRESAHA.112.268672 (2013).
Fert-Bober, J. et al. Citrullination of myofilament proteins in heart failure. Cardiovasc Res 108, 232–242, https://doi.org/10.1093/cvr/cvv185 (2015).
Dubois-Deruy, E. et al. Interplay between troponin T phosphorylation and O-N-acetylglucosaminylation in ischaemic heart failure. Cardiovasc Res 107, 56–65, https://doi.org/10.1093/cvr/cvv136 (2015).
Ramirez-Correa, G. A. et al. O-linked GlcNAc modification of cardiac myofilament proteins: a novel regulator of myocardial contractile function. Circ Res 103, 1354–1358, https://doi.org/10.1161/CIRCRESAHA.108.184978 (2008).
Figueiredo-Freitas, C. et al. S-Nitrosylation of sarcomeric proteins depresses myofilament Ca2+ sensitivity in intact cardiomyocytes. Antioxid Redox Signal 23, 1017–1034, https://doi.org/10.1089/ars.2015.6275 (2015).
Irie, T. et al. S-Nitrosylation of Calcium-Handling Proteins in Cardiac Adrenergic Signaling and Hypertrophy. Circ Res 117, 793–803, https://doi.org/10.1161/CIRCRESAHA.115.307157 (2015).
Wijnker, P. J. et al. A novel phosphorylation site, Serine 199, in the C-terminus of cardiac troponin I regulates calcium sensitivity and susceptibility to calpain-induced proteolysis. J Mol Cell Cardiol 82, 93–103, https://doi.org/10.1016/j.yjmcc.2015.03.006 (2015).
Zhang, P. et al. Multiple reaction monitoring to identify site-specific troponin I phosphorylated residues in the failing human heart. Circulation 126, 1828–1837, https://doi.org/10.1161/CIRCULATIONAHA.112.096388 (2012).
Kooij, V. et al. PKCalpha-specific phosphorylation of the troponin complex in human myocardium: a functional and proteomics analysis. PLoS One 8, e74847, https://doi.org/10.1371/journal.pone.0074847 (2013).
Solaro, R. J. & van der Velden, J. Why does troponin I have so many phosphorylation sites? Fact and fancy. J Mol Cell Cardiol 48, 810–816, https://doi.org/10.1016/j.yjmcc.2010.02.014 (2010).
Zhang, J. et al. Top-down quantitative proteomics identified phosphorylation of cardiac troponin I as a candidate biomarker for chronic heart failure. J Proteome Res 10, 4054–4065, https://doi.org/10.1021/pr200258m (2011).
Lindsey, M. L., Jung, M., Hall, M. E. & DeLeon-Pennell, K. Y. Proteomic analysis of the cardiac extracellular matrix: clinical research applications. Expert review of proteomics 15, 105–112, https://doi.org/10.1080/14789450.2018.1421947 (2018).
Soetkamp, D. et al. The continuing evolution of cardiac troponin I biomarker analysis: from protein to proteoform. Expert review of proteomics 14, 973–986, https://doi.org/10.1080/14789450.2017.1387054 (2017).
Lu, Q. W., Wu, X. Y. & Morimoto, S. Inherited cardiomyopathies caused by troponin mutations. J Geriatr Cardiol 10, 91–101, https://doi.org/10.3969/j.issn.1671-5411.2013.01.014 (2013).
Martin, A. F. Turnover of cardiac troponin subunits. Kinetic evidence for a precursor pool of troponin-I. J Biol Chem 256, 964–968 (1981).
Bidasee, K. R. et al. Chronic diabetes increases advanced glycation end products on cardiac ryanodine receptors/calcium-release channels. Diabetes 52, 1825–1836 (2003).
Bidasee, K. R. et al. Diabetes increases formation of advanced glycation end products on Sarco (endo) plasmic reticulum Ca2+ −ATPase. Diabetes 53, 463–473 (2004).
Shao, C. H. et al. Carbonylation of myosin heavy chains in rat heart during diabetes. Biochem Pharmacol 80, 205–217, https://doi.org/10.1016/j.bcp.2010.03.024 (2010).
Ferrington, D. A., Krainev, A. G. & Bigelow, D. J. Altered turnover of calcium regulatory proteins of the sarcoplasmic reticulum in aged skeletal muscle. J Biol Chem 273, 5885–5891 (1998).
Mellor, K. M., Brimble, M. A. & Delbridge, L. M. Glucose as an agent of post-translational modification in diabetes - New cardiac epigenetic insights. Life Sci 129, 48–53, https://doi.org/10.1016/j.lfs.2014.03.020 (2015).
Ahmed, N., Babaei-Jadidi, R., Howell, S. K., Beisswenger, P. J. & Thornalley, P. J. Degradation products of proteins damaged by glycation, oxidation and nitration in clinical type 1 diabetes. Diabetologia 48, 1590–1603, https://doi.org/10.1007/s00125-005-1810-7 (2005).
Uchiki, T. et al. Glycation-altered proteolysis as a pathobiologic mechanism that links dietary glycemic index, aging, and age-related disease (in nondiabetics). Aging Cell 11, 1–13, https://doi.org/10.1111/j.1474-9726.2011.00752.x (2012).
Cooper, M. E. Importance of advanced glycation end products in diabetes-associated cardiovascular and renal disease. Am J Hypertens 17, 31S–38S, https://doi.org/10.1016/j.amjhyper.2004.08.021 (2004).
Delbridge, L. M., Benson, V. L., Ritchie, R. H. & Mellor, K. M. Diabetic Cardiomyopathy: The Case for a Role of Fructose in Disease Etiology. Diabetes 65, 3521–3528, https://doi.org/10.2337/db16-0682 (2016).
Mellor, K. M. et al. Fructose modulates cardiomyocyte excitation-contraction coupling and Ca2+ handling in vitro. PLoS ONE 6, e25204, https://doi.org/10.1371/journal.pone.0025204 PONE-D-11-09982 (2011).
Kashiwagi, A. et al. Increase in cardiac muscle fructose content in streptozotocin-induced diabetic rats. Metabolism: clinical and experimental 41, 1041–1046 (1992).
Schafer, S. et al. Impaired left ventricular relaxation in type 2 diabetic rats is related to myocardial accumulation of N(epsilon)-(carboxymethyl) lysine. European journal of heart failure 8, 2–6, https://doi.org/10.1016/j.ejheart.2005.04.011 (2006).
Gomes, A. V., Liang, J. & Potter, J. D. Mutations in human cardiac troponin I that are associated with restrictive cardiomyopathy affect basal ATPase activity and the calcium sensitivity of force development. J Biol Chem 280, 30909–30915, https://doi.org/10.1074/jbc.M500287200 (2005).
Kobayashi, T. & Solaro, R. J. Increased Ca2+ affinity of cardiac thin filaments reconstituted with cardiomyopathy-related mutant cardiac troponin I. J Biol Chem 281, 13471–13477, https://doi.org/10.1074/jbc.M509561200 (2006).
Kohler, J. et al. Familial hypertrophic cardiomyopathy mutations in troponin I (K183D, G203S, K206Q) enhance filament sliding. Physiol Genomics 14, 117–128, https://doi.org/10.1152/physiolgenomics.00101.2002 (2003).
Kimura, A. et al. Mutations in the cardiac troponin I gene associated with hypertrophic cardiomyopathy. Nature genetics 16, 379–382, https://doi.org/10.1038/ng0897-379 (1997).
Luthra, M. & Balasubramanian, D. Nonenzymatic glycation alters protein structure and stability. A study of two eye lens crystallins. J Biol Chem 268, 18119–18127 (1993).
Metskas, L. A. & Rhoades, E. Conformation and dynamics of the troponin I C-terminal domain: Combining single-molecule and computational approaches for a disordered protein region. J Am Chem Soc 137, 11962–11969, https://doi.org/10.1021/jacs.5b04471 (2015).
Cheng, Y. & Regnier, M. Cardiac troponin structure-function and the influence of hypertrophic cardiomyopathy associated mutations on modulation of contractility. Archives of Biochemistry and Biophysics 601, 11–21, https://doi.org/10.1016/j.abb.2016.02.004 (2016).
Van Eyk, J. E. & Hodges, R. S. The biological importance of each amino acid residue of the troponin I inhibitory sequence 104-115 in the interaction with troponin C and tropomyosin-actin. Journal of Biological Chemistry 263, 1726–1732 (1988).
Gilda, J. E. et al. The functional significance of the last 5 residues of the C-terminus of cardiac troponin I. Archives of Biochemistry and Biophysics 601, 88–96, https://doi.org/10.1016/j.abb.2016.02.023 (2016).
Galinska, A. et al. The C terminus of cardiac troponin I stabilizes the Ca2+-activated state of tropomyosin on actin filaments. Circ Res 106, 705–711, https://doi.org/10.1161/CIRCRESAHA.109.210047 (2010).
Na, I., Kong, M. J., Straight, S., Pinto, J. R. & Uversky, V. N. Troponins, intrinsic disorder, and cardiomyopathy. Biological chemistry 397, 731–751, https://doi.org/10.1515/hsz-2015-0303 (2016).
Hoffman, R. M. & Sykes, B. D. Isoform-specific variation in the intrinsic disorder of troponin I. Proteins 73, 338–350, https://doi.org/10.1002/prot.22063 (2008).
Iakoucheva, L. M. et al. The importance of intrinsic disorder for protein phosphorylation. Nucleic Acids Research 32, 1037–1049, https://doi.org/10.1093/nar/gkh253 (2004).
Pejaver, V. et al. The structural and functional signatures of proteins that undergo multiple events of post-translational modification. Protein Science: A Publication of the Protein Society 23, 1077–1093, https://doi.org/10.1002/pro.2494 (2014).
Li, M. X. et al. Phosphorylation and mutation of human cardiac troponin I deferentially destabilize the interaction of the functional regions of troponin I with troponin C. Biochemistry 42, 14460–14468, https://doi.org/10.1021/bi035408y (2003).
Brownlee, M. Biochemistry and molecular cell biology of diabetic complications. Nature 414, 813–820 (2001).
Hung, Y. C., Yang, H. T. & Yin, M. C. Asiatic acid and maslinic acid protected heart via anti-glycative and anti-coagulatory activities in diabetic mice. Food Funct 6, 2967–2974, https://doi.org/10.1039/c5fo00549c (2015).
Liu, Z., Miers, W. R., Wei, L. & Barrett, E. J. The ubiquitin-proteasome proteolytic pathway in heart vs skeletal muscle: effects of acute diabetes. Biochem Biophys Res Commun 276, 1255–1260, https://doi.org/10.1006/bbrc.2000.3609 (2000).
Mellor, K. M., Reichelt, M. E. & Delbridge, L. M. D. Autophagy anomalies in the diabetic myocardium. Autophagy 7, 1263–1267, https://doi.org/10.4161/auto.7.10.17148 (2011).
Gundry, R. L. et al. Preparation of proteins and peptides for mass spectrometry analysis in a bottom-up proteomics workflow. Curr Protoc Mol Biol Chapter 10, Unit10 25, https://doi.org/10.1002/0471142727.mb1025s88 (2009).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J. V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat Protoc 1, 2856–2860, https://doi.org/10.1038/nprot.2006.468 (2006).
Xu, H. & Freitas, M. A. A mass accuracy sensitive probability based scoring algorithm for database searching of tandem mass spectrometry data. BMC Bioinformatics 8, 133, https://doi.org/10.1186/1471-2105-8-133 (2007).
Acknowledgements
Technical expertise from the Mass Spectrometry and Proteomics Facility, Bio21, University of Melbourne is acknowledged.
Author information
Authors and Affiliations
Contributions
K.M.M., L.M.D. and J.V.J. designed the study. J.V.J. and B.M. led the experiments and analyzed the data. M.A.B. and J.E.V.E. contributed technical expertise. K.M.M. and J.V.J. compiled the data and drafted the manuscript. L.M.D., K.M.M., J.V.J. and J.E.V.E. interpreted the data and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
JVJ, JEVE, KMM and LMD are co-inventors on a provisional patent filed by the University of Melbourne and Auckland Uniservices Limited (App #: 2018903745). Some of the claims in this application are supported by the present work. All other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Janssens, J.V., Ma, B., Brimble, M.A. et al. Cardiac troponins may be irreversibly modified by glycation: novel potential mechanisms of cardiac performance modulation. Sci Rep 8, 16084 (2018). https://doi.org/10.1038/s41598-018-33886-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-33886-x
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.