Abstract
Mutations in ASXL1 and SETBP1 genes have been frequently detected and often coexist in myelodysplastic syndrome (MDS) and acute myeloid leukaemia (AML). We previously showed that coexpression of mutant ASXL1 and SETBP1 in hematopoietic progenitor cells induced downregulation of TGFβ pathway genes and promoted the development of MDS/AML in a mouse model of bone marrow transplantation. However, whether the repression of TGFβ pathway in fact contributes to leukaemogenesis remains unclear. Moreover, mechanisms for the repression of TGFβ pathway genes in ASXL1/SETBP1-mutated MDS/AML cells have not been fully understood. In this study, we showed that expression of a constitutively active TGFβ type I receptor (ALK5-TD) inhibited leukaemic proliferation of MDS/AML cells expressing mutant ASXL1/SETBP1. We also found aberrantly reduced acetylation of several lysine residues on histone H3 and H4 around the promoter regions of multiple TGFβ pathway genes. The histone deacetylase (HDAC) inhibitor vorinostat reversed histone acetylation at these promoter regions, and induced transcriptional derepression of the TGFβ pathway genes. Furthermore, vorinostat showed robust growth-inhibitory effect in cells expressing mutant ASXL1, whereas it showed only a marginal effect in normal bone marrow cells. These data indicate that HDAC inhibitors will be promising therapeutic drugs for MDS and AML with ASXL1 and SETBP1 mutations.
Similar content being viewed by others
Introduction
Mutations in ASXL1 and SETBP1 genes have been frequently detected and often coexist in a variety of myeloid neoplasms, including myelodysplastic syndrome (MDS) and acute myeloid leukaemia (AML)1,2,3. ASXL1 gene is located on chromosome 20q11 and encodes additional sex combs like 1 (ASXL1), which contains a highly conserved ASX homology (ASXH) domain at the N-terminal region and a plant homeodomain (PHD) finger at the C-terminal region4,5. ASXL1 interacts with multiple epigenetic regulators, such as EZH2 and BAP1, thereby regulates epigenetic marks and transcription of several target genes, including Hox genes6,7. Most ASXL1 mutations exist in exon 12 of the gene, generating C-terminally truncated mutations. The mutant ASXL1 gains novel functions to form a hyper active complex with BAP1 and to interact with BRD48,9,10. SETBP1 gene is located on chromosome 18q21.1 and encodes SET binding protein 1 (SETBP1), which contains a SKI homologous region and a SET-binding region11. SETBP1 binds an oncoprotein SET and the resulting heterodimer inhibits a phosphatase PP2A that acts as a tumour suppressor in many cancer cells12,13. Mutations of SETBP1 in the SKI homologous region inhibits its ubiquitination and degradation, resulting in increased expression of SETBP114.
Leukaemic transformation of MDS has had the most impact on the mortality of MDS patients1,2,15. A key mechanism of leukaemic transformation of MDS into AML is dysregulation of TGFβ pathway16,17. We previously reported that forced expression of a C-terminally truncated ASXL1 mutant in hematopoietic progenitor cells induced MDS-like diseases, and SETBP1 mutations drove leukaemic transformation in ASXL1-mutated MDS in mouse models18,19. We also showed global downregulation of TGFβ pathway genes, including Tgfbr1, Tgfbr2, Tgfbr3, Smad1, Smad3, Smad4 in cells expressing both ASXL1 and SETBP1 mutations19. However, whether the repression of TGFβ pathway in fact contributes to leukaemogenesis induced by ASXL1/SETBP1 mutations remains unclear. Furthermore, mechanisms for the repression of TGFβ pathway genes in ASXL1/SETBP1-mutated MDS/AML cells have not been fully understood.
In this study, we showed that activation of TGFβ pathway indeed inhibits leukaemogenesis induced by ASXL1 and SETBP1 mutations. The repression of TGFβ pathway genes are associated with histone deacetylation at their promoter regions, which can be reversed by treatment with the histone deacetylase (HDAC) inhibitor vorinostat.
Results
Activation of TGFβ pathway inhibits leukaemogenesis induced by ASXL1 and SETBP1 mutations
We first assessed the role of TGFβ pathway in leukaemogenesis using murine bone marrow cells transformed by a C-terminally truncated form of ASXL1 mutant [ASXL1-MT cells: cells expressing ASXL1 mutation (ASXL1-MT)]18 or those transformed by combined expression of SETBP1-D868N and ASXL1-MT (cSAM cells: cells with combined expression of SETBP1 and ASXL1 Mutations)19. SETBP1-D868N is an oncogenic mutation of SETBP1, and ASXL1-MT is a leukaemia-associated ASXL1 mutant [ASXL1 (1900–1922del; E635RfsX15)]. In a previous study, we showed that TGFβ pathway genes were specifically downregulated in cSAM cells but not in ASXL1-MT cells19. Consistent with this observation, TGFβ inhibited the growth of normal bone marrow c-Kit+ cells and ASXL1-MT cells in a dose-dependent manner, whereas it showed little effect on the growth of cSAM cells (Fig. 1a). Thus, cSAM cells that express low level of TGFβ pathway genes are poorly responsive to TGFβ-induced growth suppression. To assess the effect of forced activation of TGFβ pathway in cSAM cells, we next transduced vector or ALK5-TD (a constitutively active form of Tgfb receptor 1) into cSAM cells, and cultured the cells in vitro or directly transplanted them into recipient mice (Fig. 1b). ALK5-TD-transduced cSAM cells grew slower than vector-transduced cells in vitro (Fig. 1c). Furthermore, we observed impaired engraftment and significantly delayed leukaemia development in mice receiving ALK5-TD-transduced cSAM cells relative to those receiving control cells (Fig. 1d–f). These data indicate that repression of TGFβ pathway contributes to leukaemic proliferation of cSAM cells both in vitro and in vivo.
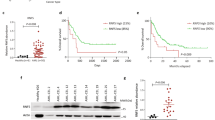
ALK5-TD inhibits leukaemic proliferation of cSAM cells. (a) CellTiter-Gro Luminescent Cell Viability Assay using bone marrow c-Kit+ cells, ASXL1-MT cells, and cSAM cells, treated with TGFβ at indicated concentrations for 72 h in triplicate. Data are normalized to vehicle control (0 μM group), and are shown as mean ± s.d. (b) Experimental scheme used in (c–f). cSAM cells were transduced with vector or ALK5-TD, and were cultured in the presence of 1 ng/ml IL-3, or were injected (5 × 105 cells/mouse) into recipient mice. (c) The growth of 5 × 104/ml cSAM cells transduced with vector or ALK5-TD was estimated using CellTiter-Glo Luminescent Cell Viability Assay from day 1 to day 3. Data are shown as mean ± s.e.m. from triplicated experiments. (d) Wright-Giemsa staining of spleen cells collected from moribund mice transplanted with cSAM cells. Scale bar: 50 μm. (e) Percentages of leukaemic (GFP+) cells in peripheral blood at 16 days post transplantation in mice transplanted with vector- or ALK5-TD-transduced cSAM cells. N = 4 per group. (f) Survival curves of the mice transplanted with vector- or ALK5-TD-transduced cSAM cells. N = 9 per group.
Vorinostat promotes transcription of TGFβ pathway genes in cSAM cells
Histone acetylation is associated with active transcription, and it was recently shown that SETBP1 represses Runx1 expression through histone deacetylation20. We therefore examined levels of histone acetylation around the promoter regions of TGFβ pathway genes in c-Kit+ cSAM cells and normal c-Kit+ bone marrow progenitor cells. Consistent with our previous report19, multiple TGFβ pathway genes (Tgfbr1, Tgfbr2, Tgfbr3, Smad1, Smad3, and Smad4) were downregulated, whereas Hox genes (Hoxa9 and Hoxa10) were upregulated in cSAM cells compared with control cells (Fig. 2a,b). Chromatin immunoprecipitation coupled with quantitative PCR (ChIP-qPCR) revealed the decreased acetylation of several histone lysine residues, especially of histone H3 lysine 14 (H3K14) and histone H4 lysine 5 (H4K5), at the vicinity of transcription starting sites (TSSs) of multiple TGFβ pathway genes in cSAM cells. Treatment of cSAM cells with vorinostat resulted in enhanced acetylation of H3K14 and H4K5, particularly at the promotor regions of TGFβ pathway genes (Fig. 2c–e, Supplemental Fig. 1). We also observed decreased binding of a histone acetyltransferase p300 at the same promoter regions, which was reversed by vorinostat (Fig. 2f). Vorinostat treatment also induced upregulation of Tgfbr1, Tgfbr2, Tgfbr3, Smad3 and Smad4 in cSAM cells (Fig. 3a). In contrast, expression of Hoxa9 and Hoxa10 were not upregulated, but were rather downregulated by vorinostat in cSAM cells (Fig. 3b). We then assessed responsiveness of vehicle- or vorinostat-treated cSAM cells to TGFβ. As expected, TGFβ-induced phosphorylation of Smad2, a key intracellular mediator of TGFβ signaling, was substantially increased in vorinostat-treated cSAM cells (Fig. 3c). These results indicate that repression of TGFβ pathway genes are mainly caused by histone deacetylation, which can be reversed by the HDAC inhibitor vorinostat.
Vorinostat increases acetylation of lysine residues on histone H3 and H4 in cSAM cells. (a,b) Expression of TGFβ pathway genes was decreased (a), while expression of Hoxa9 and Hoxa10 was increased (b) in cSAM cells compared to normal bone marrow c-Kit+ cells. Data are shown as mean ± s.e.m. from duplicate experiments. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s t-test. (c) Western blotting for histone acetylation in 32D cells and cSAM cells. Vorinostat increased both H3K14ac and H4K5ac in cSAM cells. Full-length blots are shown in Supplemental Fig. 2. (d–f) Genomic DNA fragments from control [normal bone marrow (BM) c-kit + cells] and cSAM cells cultured with 1 µM vorinostat or vehicle control (DMSO) were immunoprecipitated with anti-H3K14ac (d), anti-H4K5ac (e) and anti-p300 (f) antibodies. Enrichments of H3K14ac and H4K5ac at transcription starting sites (TSS) of Tgfbr1, Tgrbr2, Tgfbr3, Smad1, Smad3, and Smad4 were measured by qPCR. Data are shown as mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s t-test.
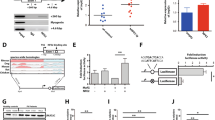
Vorinostat upregulated TGFβ pathway genes and activates TGFβ pathway in cSAM cells. (a,b) cSAM cells were cultured with 1 or 4 µM vorinostat or vehicle control (DMSO) for 6 hours, and expression of TGFβ pathway genes (a) and Hox genes (b) was analyzed by qPCR. Results were normalized to GAPDH, with the relative mRNA level in vehicle-treated cells set at 1. Data are shown as mean ± s.e.m. of duplicate wells. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s t-test. (c) cSAM cells were treated with vorinostat (1 μM) for 24 hours, following TGFβ (100 ng/ml) stimulation for 45 minutes. TGFβ-induced phosphorylation of Smad2 was substantially increased in vorinostat-treated cSAM cells.
Vorinostat inhibits leukaemogenesis induced by ASXL1 and SETBP1 mutations
Next, we examined the effect of vorinostat on the growth of cSAM cells. Addition of low-dose vorinostat (0.5 μM) was sufficient to inhibit proliferation of cSAM cells in culture, whereas even high dose vorinostat (1 μM) only showed a marginal effect on the growth of a mouse myeloid cell line, 32D (Fig. 4a). We then performed colony replating assay using cSAM cells in the presence or absence of vorinostat. Again, addition of vorinostat (0.1 μM) reduced colony forming capability of cSAM cells (Fig. 4b). To characterize the decreased cell growth of cSAM cells induced by vorinostat, we analyzed cell cycle status, differentiation, and apoptosis in vehicle- or vorinostat-treated cSAM cells. Vorinostat treatment consistently decreased the proportion of S phase cells (Fig. 4c) and the frequency of immature cells (c-Kit+Gr-1− cells) in culture (Fig. 4d). In contrast, vorinostat did not induce apoptosis of cSAM cells (Fig. 4e). Thus, vorinostat inhibits the growth of cSAM cells mainly by inducing cell cycle arrest and differentiation.
Vorinostat inhibits leukaemic proliferation of cSAM cells. (a) cSAM cells and 32D cells were cultured in the presence of 1 ng/ml IL-3 together with 0.5 or 1 μM vorinostat or vehicle control (DMSO). The growth of the cells was estimated using CellTiter-Glo Luminescent Cell Viability Assay. All data with error bars indicate the mean ± s.e.m. from triplicated experiments. (b) c-kit+ cSAM cells were serially replated in methylcellulose supplemented with SCF (100 ng/ml), IL-3 (10 ng/ml), together with vorinostat (0.1 μM) or vehicle control (DMSO). Shown are colony counts per 3 × 102 replated cells (mean ± s.e.m) from duplicate plates at 1st, 2nd and 3rd rounds. *P < 0.05, **P < 0.01, Student’s t-test. (c–e) Cell-cycle status, differentiation and apoptosis were assessed after 6 hours culture with vorinostat (1 μM). (c) FACS profiles of vehicle- or vorinostat-treated cSAM cells are shown. The numbers indicate the percentages of cells in the G0/G1, S and G2/M phases. (d) Shown are FACS profiles of Gr-1 and c-Kit expression, and Wright-Giemsa staining of cytospin preparations of vehicle- or vorinostat-treated cSAM cells. The numbers indicate the percentages of cells in each gate. Scale bars; 50 μm. (e) Shown are FACS profiles of Annexin V and DAPI expression of vehicle- or vorinostat-treated cSAM cells. The numbers indicate the percentages of Annexin V+ cells.
To assess the effect of vorinostat on the development of cSAM cell-driven MDS/AML in vivo, we performed two independent assays (Fig. 5a). First, we pretreated cSAM cells with 1 μM vorinostat or vehicle control for 6 hours and transplanted these cells into recipient mice. As shown in Fig. 5b, pretreatment of cSAM cells with vorinostat significantly delayed the development of leukaemia in vivo. Second, we directly transplanted cSAM cells into recipient mice, and the mice were treated with vehicle or vorinostat from day 2 to day 9. This in vivo treatment with vorinostat also prolonged survival of the recipient mice transplanted with cSAM cells (Fig. 5c). Thus, vorinostat has the robust in vivo effect to inhibit leukaemogenesis induced by ASXL1 and SETBP1 mutations.
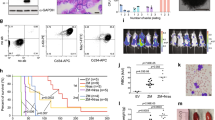
Vorinostat inhibits the development of cSAM cell-driven MDS/AML in vivo. (a) Experimental scheme used in (b,c). cSAM cells were pretreated with 1 mM vorinostat or vehicle control (DMSO) for 6 hours before transplantation, or directly injected into recipient mice following oral administration of vorinostat (50 mg/kg) to the recipient mice. (b) Overall survival of mice transplanted with vehicle- or vorinostat-pretreated cSAM cells. N = 7 per group. (c) Overall survival of mice that were orally treated with vehicle or vorinostat after transplantation. N = 11 (vehicle) or 12 (vorinostat). P value was calculated using a log-rank test.
Vorinostat inhibits the growth of other MDS/AML cells expressing mutant ASXL1
Finally, we assessed the effect of vorinostat on other murine MDS/AML cells expressing mutant ASXL1; ASXL1-MT cells and cRAM cells. cRAM (combined expression of RUNX1 and ASXL1 Mutations) cells were generated by transducing a C-terminally truncated RUNX1 mutant (RUNX1-S291fsX30021) into bone marrow progenitors derived from the recently established conditional knock-in mice expressing ASXL1-MT22. We cultured these MDS/AML cells and bone marrow c-Kit+ cells in the presence/absence of vorinostat. Addition of vorinostat in culture resulted in increased acetylation of H4K5 in all these cells (Fig. 6a) and effectively inhibited the growth of cRAM cells and ASXL1-MT cells (Fig. 6b). In contrast, bone marrow c-Kit+ cells grew normally in the presence of vorinostat (Fig. 6b). Vorinostat also induced upregulation of Tgfbr1, Tgfbr2, Tgfbr3, Smad1, Smad3, and Smad4 in both cRAM and ASXL1-MT cells (Fig. 6c). These data suggest that vorinostat upregulates TGFβ pathway genes and inhibits cell growth in a wide range of MDS/AML cells.
Vorinostat inhibits the growth of other MDS/AML cells expressing ASXL1-MT. (a) Western blotting for histone acetylation in bone marrow c-Kit+ cells, cRAM cells and ASXL1-MT cells. Full-length blots are shown in Fig. S3. Vorinostat increased H4K5ac in all these cells. (b) Bone marrow c-Kit+ cells, cRAM cells and ASXL1-MT cells were cultured in the presence of 1 or 10 ng/ml IL-3 together with 0.5 or 1 μM vorinostat or vehicle control (DMSO). The growth of the cells was estimated using Cell Counting Kit-8. All data with error bars indicate the mean ± s.e.m. from triplicated experiments. (c) cRAM cells and ASXL1-MT cells were cultured with 1 µM vorinostat or vehicle control (DMSO) for 6 hours, and expression of TGFβ pathway genes was analyzed by qPCR. Results were normalized to GAPDH, with the relative mRNA level in vehicle-treated cells set at 1. Data are shown as mean ± s.e.m. of duplicate wells. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s t-test.
Discussion
Clinical data revealed frequent coexistence of ASXL1 and SETBP1 mutations in myeloid neoplasms, and we previously demonstrated their cooperation to promote leukaemic transformation in mouse MDS/AML models19. We extended the research, and showed that activation of TGFβ pathway inhibits leukaemogenesis induced by both mutations. Aberrant TGFβ signaling has been implicated in a variety of haematopoietic neoplasms. Loss of Smad3 promotes the development of T-cell lineage acute lymphoblastic leukaemia in concert with loss of p27(kip1)23. Several leukaemia-associated oncogenes, EVI1, RUNX1-EVI1 and RUNX1-ETO, interact with Smad3 to repress TGFβ signaling in AML and chronic myeloid leukaemia23,24,25. Mutations in Smad4 gene that disrupt its transcriptional activity has been detected in AML26. Loss of TGFBRI and TGFBRII expression was shown to trigger the development of myeloid or lymphocytic leukaemia27. Here we identified epigenetic repression of multiple TGFβ pathway genes as another mechanism to promote leukaemogenesis.
Recently, Vishwakarma et al. reported that SETBP1 recruits a nucleosome remodeling deacetylase (NuRD) complex containing HDAC1 to Runx1 promoters, thereby represses its expression. They also showed that HDAC inhibitors restore Runx1 expression and inhibit leukaemogenesis driven by SETBP1 overexpression20. These findings, together with our data, indicate that SETBP1 plays a key role in inducing aberrant histone acetylation around promoter regions of several tumor suppressor genes. However, vorinostat treatment also induced upregulation of TGFβ pathway genes in MDS/AML cells that express only the ASXL1 mutant (ASXL1-MT cells and cRAM cells). In addition, our previous Liquid Chromatography-Mass Spectrometry analysis revealed physical interaction between mutant ASXL1 and HDAC110. These observations suggest the possible involvement of ASXL1 mutations in the regulation of histone acetylation. How the complexes containing ASXL1, SETBP1 and HDAC1 select their target genes warrants further investigation.
Interestingly, vorinostat treatment resulted in significant downregulation of Hoxa9 and Hoxa10 in cSAM cells. Given that both ASXL1 and SETBP1 mutations were shown to upregulate Hox gene expression19,28, vorinostat may directly modulate activities of ASXL1 and/or SETBP1 mutants through histone acetylation-independent functions. Such effects of HDAC inhibitors have been repeatedly shown for a variety of non-histone proteins29,30,31. The potential role of acetylation to regulate activities and stabilities of ASXL1 and SETBP1 proteins will merit future studies.
Importantly, neither ALK5-TG overexpression nor vorinostat treatment completely inhibited the development of MDS/AML induced by cSAM cells. We also found that vorinostat did not induce apoptosis, whereas it effectively induced cell cycle arrest and differentiation in cSAM cells. The lack of apoptosis-inducing effect of vorinostat on cSAM cells may account for its limited therapeutic efficacy. Combined therapy with vorinostat and inhibitors of anti-apoptosis proteins, such as venetoclax32, could show synergistic effects against MDS/AML cells.
In summary, we showed decreased histone acetylation around promoters of TGFβ pathway genes in MDS/AML cells expressing mutant ASXL1 and SETBP1. The HDAC inhibitor vorinostat increased levels of histone acetylation, reversed transcription of the TGFβ pathway genes, and inhibited leukaemic proliferation of ASXL1/SETBP1-mutated cells. Vorinostat also inhibited the growth of other MDS/AML cells expressing ASXL1-MT. These findings suggest that HDAC inhibitors will be promising therapeutic drugs to treat myeloid neoplasms with ASXL1 and/or SETBP1 mutations.
Materials and Methods
Mice
C57BL/6 (Ly5.1) mice (Sankyo Labo Service Corporation, Tokyo, Japan) and C57BL/6 (Ly5.2) mice (Charles River Laboratories Japan, Yokohama, Japan) were used for bone marrow transplantation assays. All animal studies were approved by the Animal Care Committee of the Institute of Medical Science at the University of Tokyo (approval number: PA13-19, PA16-31, PA17-75), and were conducted in accordance with the Regulation on Animal Experimentation at University of Tokyo based on International Guiding Principles for Biomedical Research Involving Animals.
Plasmids
The plasmid pcDNA3-HASL-ALK5(TD) was provided by Dr. Miyazono K. (Tokyo university, Japan), and we cloned it into the vector pMYs-IRES-Puromicin-resistant gene (pMYs-IP).
Retrovirus and transduction
As described previously21, retroviruses were generated by transient transfection of Plat-E packaging cells with the calcium-phosphate coprecipitation method.
Establishment of murine MDS/AML cells
To generate cSAM cells, we transduced ASXL1-MT and SETBP-D868N into mouse bone marrow progenitor cells, and transplanted these cells into sublethally irradiated recipient mice. Leukaemic cells were isolated from bone marrow of the moribund mice, and their leukaemogenic activity was confirmed by serial transplantations19. ASXL-MT cells were established through serial passages in mice using mouse bone marrow progenitor cells transduced with ASXL1-MT alone18. To generate cRAM cells, we transduced RUNX1-S291fsX30021 into mouse bone marrow progenitor cells derived from the recently established conditional knock-in mice expressing ASXL1-MT22. The cells expressing RUNX1 and ASXL1 mutations were transplanted into sublethally irradiated recipient mice to establish mouse MDS/AML cells.
Cell culture
cSAM cells, ASXL1-MT cells, cRAM cells and 32D cells were cultured in RPMI-1640 medium supplemented with 10% fetal bovine serum and 1 ng/ml interleukin-3 (IL-3) (R&D Systems, catalog #403-ML). Mouse bone marrow c-Kit+ cells were cultured in RPMI-1640 medium supplemented with 10% fetal bovine serum and 10 ng/ml interleukin-3 (IL-3) (R&D Systems, catalog #403-ML) . In some experiments, we added the indicate concentrations of TGFβ (R&D Systems, catalog #70-MB) in the culture. The growth of viable cells in culture was measured using the CellTiter-Gro Luminescent Cell Viability Assay (Promega, Madison, WI, USA) or the Cell Counting Kit-8 (Dojindo, Kumamoto, Japan).
Colony-forming assay
c-Kit+ cells were isolated from mouse bone marrow cells and cSAM cells using MACS (magnetic-activated cell sorting), and were cultured in Methocult 3231 (StemCell Technologies, Vancouver, BC, Canada) supplemented with 20 ng/ml SCF and 10 ng/ml mouse IL-3. A total of 300 cells were plated in duplicates for each round of plating. Colony counting and replating were performed every 5 days.
Transplantation assay
cSAM cells were infected for 72 hours with the retroviruses harbouring pMYs-IP and pMYs-IP-ALK5-TD, using 6-well dishes coated with RetroNectin (Takara Bio, Otsu, Japan). The infected cells were selected using 1 μg/ml puromycin, and 5 × 105 infected cells were transplanted intravenously into non-radiated recipient mice. For experiments using vorinostat, cSAM cells were cultured in RPMI-1640 medium supplemented with 1 ng/ml IL-3 and 1 μM vorinostat or DMSO for 6 hours. These precultured cells (2 × 105/mouse) were transplanted into recipient mice. In some experiments, cSAM cells were directly transplanted into recipient mice (2 × 105/mouse), and vehicle control or vorinostat (50 mg/kg/day) were orally administered to these mice from day 2 to day 9. Overall survival of transplanted mice was calculated using the Kaplan-Meier method.
Western blot analysis
For analyzing histone modifications, cells were cultured with 1 µM vorinostat or DMSO for 6 hours, and were lysed in IP buffer (150 mM NaCl, 50 mM Tris pH 7.5, 1 mM EDTA, 1% Triton, 2 mM sodium orthovanadate, 2 mM PMSF, 50 mM sodium fluoride)18. Pelleted nuclei were resuspended with 0.2 M sulfuric acid, and histones were precipitated from the supernatant with trichloroacetic acid (TCA). In other experiments, cells were cultured with 1 µM vorinostat or DMSO for 24 hours, or with 100 ng/ml TGFβ (R&D Systems, catalog #70-MB) for 45 minutes, and were lysed directly in 1x Laemmli Sample Buffer. Whole-cell lysates were subjected to sodium dodecyl sulfate–polyacrylamide gel electrophoresis and transferred to a polyvinylidene fluoride membrane (Bio-Rad). The blot was incubated with the anti-histone antibodies, anti-acetylated histone antibodies, anti-Smad2/3 (#3102; Cell Signaling Technology, Beverly, MA), anti-phospho-Smad2 (Ser465/Ser467, clone E8F3R, #18338, Cell Signaling Technology, Beverly, MA) and anti-Tubulin (clone B-5-1-2; Santa Cruz Biotechnology Santa Cruz, CA). Signals were detected with SuperSignalWest Pico (Pierce, Thermo Fisher Scientific) and visualized on films or with imagequant LAS 4000 (Fujifilm Life Science, Roche Diagnostics). All antibodies for histones (H3, H4, H3K14ac, H4K5ac, H4K8ac, H4K16ac) were kind gifts from Dr. Kimura H. (Tokyo Institute of Technology, Japan)33.
Flow cytometry
Cells were stained by fluoro-conjugated antibodies [anti-mouse CD117 (BioLegend, catalog #105808, clone 2B8, 1:400), and anti-mouse Gr-1 (BioLegend, catalog #108411, clone RB6-8C5, 1:400)] for 30 min at 4 °C. After staining, cells were washed with cold PBS several times, and were resuspended with PBS containing 2% FBS. Cells were analyzed on a FACS Verse (BD Biosciences, San Jose, CA, USA). Cell cycle analysis (Cycletesttm Plus DNA Reagent Kit, BD Biosciences) and apoptosis analysis (Annexin V-APC kit; BD Biosciences) were performed according to the manufacturer’s recommendations. The data were analyzed using FlowJo software (Treestar, Inc., San Carlos, CA).
Quantitative PCR
As described previously33, total RNA was extracted from BM cells using the RNeasy Mini kit (QIAGEN), and reverse-transcribed using the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA) with the deoxyribonuclease I (Invitrogen - Thermo Fisher Scientific - MA, USA). Quantitative PCR (qPCR) was performed using SYBR Premix EX Taq (Takara Bio) and Rotor-Gene Q (Qiagen, Venlo, The Netherlands). Sequences of the primers used for qPCR in this study, from 5′ to 3′ are as follows:
Tgfbr1 Fw: TCTGCATTGCACTTATGCTGA, Tgfbr1 Rv: AAAGGGCGATCTAGTGATGGA, Tgfbr2 Fw: CCGCTGCATATCGTCCTGTG, Tgfbr2 Rv: AGTGGATGGATGGTCCTATTACA, Tgfbr3 Fw: GGTGTGAACTGTCACCGATCA, Tgfbr3 Rv: GTTTAGGATGTGAACCTCCCTTG, Smad1 Fw: GCTTCGTGAAGGGTTGGGG, Smad1 Rv: CGGATGAAATAGGATTGTGGGG, Smad3 Fw: CACGCAGAACGTGAACACC, Smad3 Rv: GGCAGTAGATAACGTGAGGGA, Smad4 Fw: ACACCAACAAGTAACGATGCC, Smad4 Rv: GCAAAGGTTTCACTTTCCCCA.
Chromatin Immunoprecipitation assay (ChIP)
cSAM cells were cultured with 1 µM vorinostat or vehicle control (DMSO) for 6 hours. Genomic DNA fragments from these c-kit+ cSAM cells and normal c-kit+ bone marrow progenitor cells were immunoprecipitated using anti-acetylated-histone (same antibodies used for western blotting) and anti-p300 antibodies (Santa Cruz 584) as described previously34. Quantitative PCR was performed with a Rotor-Gene Q (Qiagen) using SYBR Premix EX Taq (Takara). Sequences of the primers used for ChIP-qPCR in this study, from 5′ to 3′ are as follows:
Tgfbr1 Fw: CAGTTACAAAGGGCCGGAGC, Tgfbr1 Rv: CAACACGATGAGGAGCTGCG, Tgfbr2 Fw: TGTAGAGTCCAGGCAAGGCT, Tgfbr2 Rv: GACTCACTCATCAACTTTACCCG, Tgfbr3 Fw: CCCTTTCGCTGATTGCTGTG, Tgfbr3 Rv: CTTCTCCCTCTCCACCTCTCT, Smad1 Fw: GGGACTAGAGTCAGAGGAAGG, Smad1 Rv: CGAGCCTGGATTGATCTGGT, Smad3 Fw: CAGAGGAGGAGGAGGAGGAG, Smad3 Rv: GTGGCAGTAGAAAGTTTGGGTT, Smad4 Fw: CTCGCTCGCTGCTCAAACTC, Smad4 Rv: GGTAATTTCAGGGTTTGGCCC.
Morphological analysis
Cytospin preparations were stained with Giemsa. Images were obtained with a BX51 microscope and a DP12 camera (Olympus).
Statistics
Statistical significance was determined by the indicated tests for independent variables using GraphPad Prism 7 (GraphPad Software Inc., La Jolla, CA). Statistical analyses for evaluating differences between two groups were performed by the unpaired and two-tailed Student’s t test. The survival distributions were compared by the log-rank test. No specific statistical methods were used to predetermine the sample size.
Data Availability
All data generated or analyzed during this study are included in this published article. The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Papaemmanuil, E. et al. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood 122, 3616–3627; quiz 3699, https://doi.org/10.1182/blood-2013-08-518886 (2013).
Cazzola, M., Della Porta, M. G. & Malcovati, L. The genetic basis of myelodysplasia and its clinical relevance. Blood 122, 4021–4034, https://doi.org/10.1182/blood-2013-09-381665 (2013).
Goyama, S. & Kitamura, T. Epigenetics in normal and malignant hematopoiesis: An overview and update 2017. Cancer science 108, 553–562, https://doi.org/10.1111/cas.13168 (2017).
Fisher, C. L., Berger, J., Randazzo, F. & Brock, H. W. A human homolog of Additional sex combs, ADDITIONAL SEX COMBS-LIKE 1, maps to chromosome 20q11. Gene 306, 115–126 (2003).
Fisher, C. L., Randazzo, F., Humphries, R. K. & Brock, H. W. Characterization of Asxl1, a murine homolog of Additional sex combs, and analysis of the Asx-like gene family. Gene 369, 109–118, https://doi.org/10.1016/j.gene.2005.10.033 (2006).
Fisher, C. L. et al. Additional sex combs-like 1 belongs to the enhancer of trithorax and polycomb group and genetically interacts with Cbx2 in mice. Developmental biology 337, 9–15, https://doi.org/10.1016/j.ydbio.2009.10.004 (2010).
Abdel-Wahab, O. et al. ASXL1 mutations promote myeloid transformation through loss of PRC2-mediated gene repression. Cancer Cell 22, 180–193, https://doi.org/10.1016/j.ccr.2012.06.032 (2012).
Balasubramani, A. et al. Cancer-associated ASXL1 mutations may act as gain-of-function mutations of the ASXL1-BAP1 complex. Nature communications 6, 7307, https://doi.org/10.1038/ncomms8307 (2015).
Yang, H. et al. Gain-of-function of ASXL1 truncating protein in the pathogenesis of myeloid malignancies. Blood, https://doi.org/10.1182/blood-2017-06-789669 (2017).
Asada, S. et al. Mutant ASXL1 cooperates with BAP1 to promote myeloid leukaemogenesis. Nature Communications 9, https://doi.org/10.1038/s41467-018-05085-9 (2018).
Makishima, H. et al. Somatic SETBP1 mutations in myeloid malignancies. Nature genetics 45, 942–946, https://doi.org/10.1038/ng.2696 (2013).
Cristobal, I. et al. PP2A impaired activity is a common event in acute myeloid leukemia and its activation by forskolin has a potent anti-leukemic effect. Leukemia 25, 606–614, https://doi.org/10.1038/leu.2010.294 (2011).
Trimarchi, T., Ntziachristos, P. & Aifantis, I. A new player SETs in myeloid malignancy. Nature genetics 45, 846–847, https://doi.org/10.1038/ng.2709 (2013).
Piazza, R. et al. Recurrent SETBP1 mutations in atypical chronic myeloid leukemia. Nature genetics 45, 18–24, https://doi.org/10.1038/ng.2495 (2013).
Raza, A. & Galili, N. The genetic basis of phenotypic heterogeneity in myelodysplastic syndromes. Nature reviews. Cancer 12, 849–859, https://doi.org/10.1038/nrc3321 (2012).
Lin, H. K., Bergmann, S. & Pandolfi, P. P. Deregulated TGF-beta signaling in leukemogenesis. Oncogene 24, 5693–5700, https://doi.org/10.1038/sj.onc.1208923 (2005).
Dong, M. & Blobe, G. C. Role of transforming growth factor-beta in hematologic malignancies. Blood 107, 4589–4596, https://doi.org/10.1182/blood-2005-10-4169 (2006).
Inoue, D. et al. Myelodysplastic syndromes are induced by histone methylation-altering ASXL1 mutations. The Journal of clinical investigation 123, 4627–4640, https://doi.org/10.1172/jci70739 (2013).
Inoue, D. et al. SETBP1 mutations drive leukemic transformation in ASXL1-mutated MDS. Leukemia 29, 847–857, https://doi.org/10.1038/leu.2014.301 (2015).
Vishwakarma, B. A. et al. Runx1 repression by histone deacetylation is critical for Setbp1-induced mouse myeloid leukemia development. Leukemia 30, 200–208, https://doi.org/10.1038/leu.2015.200 (2016).
Watanabe-Okochi, N. et al. AML1 mutations induced MDS and MDS/AML in a mouse BMT model. Blood 111, 4297–4308, https://doi.org/10.1182/blood-2007-01-068346 (2008).
Nagase, R. et al. Expression of mutant Asxl1 perturbs hematopoiesis and promotes susceptibility to leukemic transformation. The Journal of experimental medicine 215, 1729–1747, https://doi.org/10.1084/jem.20171151 (2018).
Wolfraim, L. A. et al. Loss of Smad3 in acute T-cell lymphoblastic leukemia. The New England journal of medicine 351, 552–559, https://doi.org/10.1056/NEJMoa031197 (2004).
Kurokawa, M. et al. The oncoprotein Evi-1 represses TGF-beta signalling by inhibiting Smad3. Nature 394, 92–96, https://doi.org/10.1038/27945 (1998).
Jakubowiak, A. et al. Inhibition of the transforming growth factor beta 1 signaling pathway by the AML1/ETO leukemia-associated fusion protein. The Journal of biological chemistry 275, 40282–40287, https://doi.org/10.1074/jbc.C000485200 (2000).
Imai, Y. et al. Mutations of the Smad4 gene in acute myelogeneous leukemia and their functional implications in leukemogenesis. Oncogene 20, 88–96, https://doi.org/10.1038/sj.onc.1204057 (2001).
Le Bousse-Kerdiles, M. C. et al. Differential expression of transforming growth factor-beta, basic fibroblast growth factor, and their receptors in CD34+ hematopoietic progenitor cells from patients with myelofibrosis and myeloid metaplasia. Blood 88, 4534–4546 (1996).
Oakley, K. et al. Setbp1 promotes the self-renewal of murine myeloid progenitors via activation of Hoxa9 and Hoxa10. Blood 119, 6099–6108, https://doi.org/10.1182/blood-2011-10-388710 (2012).
He, L. Z. et al. Histone deacetylase inhibitors induce remission in transgenic models of therapy-resistant acute promyelocytic leukemia. The Journal of clinical investigation 108, 1321–1330, https://doi.org/10.1172/jci11537 (2001).
Ramsey, J. M. et al. Entinostat prevents leukemia maintenance in a collaborating oncogene-dependent model of cytogenetically normal acute myeloid leukemia. Stem cells (Dayton, Ohio) 31, 1434–1445, https://doi.org/10.1002/stem.1398 (2013).
Bots, M. et al. Differentiation therapy for the treatment of t(8;21) acute myeloid leukemia using histone deacetylase inhibitors. Blood 123, 1341–1352, https://doi.org/10.1182/blood-2013-03-488114 (2014).
Mihalyova, J. et al. Venetoclax: A new wave in hematooncology. Experimental Hematology 61, 10–25, https://doi.org/10.1016/j.exphem.2018.02.002 (2018).
Kimura, H., Hayashi-Takanaka, Y., Goto, Y., Takizawa, N. & Nozaki, N. The organization of histone H3 modifications as revealed by a panel of specific monoclonal antibodies. Cell structure and function 33, 61–73 (2008).
Enomoto, Y. et al. Characterization of leukocyte mono-immunoglobulin-like receptor 7 (LMIR7)/CLM-3 as an activating receptor: its similarities to and differences from LMIR4/CLM-5. The Journal of biological chemistry 285, 35274–35283, https://doi.org/10.1074/jbc.M110.137166 (2010).
Acknowledgements
We thank Shiori Shikata for expert technical assistance. We thank Dr. Miyazono K. (Tokyo university, Japan) for plasmids. We thank Dr. Kimura H. (Tokyo Institute of Technology, Japan) for antibodies. We also thank the Flow Cytometry Core, the Mouse Core at The Institute of Medical Science, The University of Tokyo for their help. This work was supported by a grant from The Tokyo Biochemical Research Foundation (TK), Grant-in-Aid for Scientific Research (B) (15H04855, TK), and Grant-in-Aid for Challenging Exploratory Research (16K15499, SG).
Author information
Authors and Affiliations
Contributions
Study design: M.S., D.I., T.K., S.G. Conducting experiments: M.S., D.I., R.N., N.S., A.T., T.Y. Data analysis: M.S., D.I., T.K., S.G. Manuscript preparation: M.S., T.K., S.G.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Saika, M., Inoue, D., Nagase, R. et al. ASXL1 and SETBP1 mutations promote leukaemogenesis by repressing TGFβ pathway genes through histone deacetylation. Sci Rep 8, 15873 (2018). https://doi.org/10.1038/s41598-018-33881-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-33881-2
Keywords
This article is cited by
-
Modeling and therapeutic targeting of t(8;21) AML with/without TP53 deficiency
International Journal of Hematology (2024)
-
SETBP1 is dispensable for normal and malignant hematopoiesis
Leukemia (2023)
-
Clinical features and next-generation sequencing landscape of essential thrombocythemia, prefibrotic primary myelofibrosis, and overt fibrotic primary myelofibrosis: a Chinese monocentric retrospective study
Journal of Cancer Research and Clinical Oncology (2023)
-
Prospective Identification of Prognostic Hot-Spot Mutant Gene Signatures for Leukemia: A Computational Study Based on Integrative Analysis of TCGA and cBioPortal Data
Molecular Biotechnology (2023)
-
Proposals for Clinical Trials in Chronic Myelomonocytic Leukemia
Current Treatment Options in Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.