Abstract
Firefly luciferases produce yellow-green light under physiological and alkaline conditions, however at acidic pH, higher temperatures or in the presence of heavy metals the color changes to red, a property called pH-sensitivity. Despite many decades of studies, the proton and metal binding sites responsible for pH-sensitivity remain enigmatic. Previously we suggested that the salt bridge E311/R337 keeps a closed conformation of the luciferin phenolate binding site. Here we further investigated the effect of this salt bridge and mutations of the neighbor residues H310 and E/N354, on metal and pH-sensitivity of firefly luciferases emitting distinct bioluminescence colors (Cratomorphus distinctus: 548 nm; Macrolampis sp2: 569 nm). The substitutions of H310 and E/N354 modulate metal sensitivity, whereas the carboxylate of E311 may work as the catalytic base essential for green bioluminescence and pH-sensitivity. Modeling studies showed that H310, E311 and E354 side-chains coordinate Zinc, constituting the metal binding site and the pH-sensor. Electrostatic potential and pKa calculations suggest that the external couple H310/E354 is affected by pH, whereas E311/R337 make a stabilized internal pair which retains excited oxyluciferin ejected proton near its phenolate group into a high energy state, promoting yellow-green bioluminescence. Protonation or metal binding weaken these electrostatic gates and their ability to retain the excited oxyluciferin released proton near its phenolate, promoting red light emission.
Similar content being viewed by others
Introduction
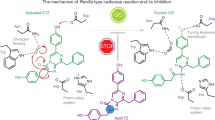
Luciferases are the enzymes which elicit the beautiful yellow-green flashes of fireflies during summer nights around the word. They catalyze an ATP-dependent oxidation of a benzothiazolic luciferin1,2. Whereas the normal bioluminescence color produced by firefly luciferases under physiological and alkaline conditions is usually yellow-green, at lower pH, in the presence of heavy metals such as Zn2+, Ni2+ and Hg2+ or at high temperatures, the color changes to bright orange-red, a property that has been called pH-sensitivity or pH-dependency3,4. Although many studies during the last decades have attempted to elucidate the mechanisms of bioluminescence color determination in beetle luciferases, the specific structural targets and mechanism of pH-sensitivity have remained elusive. Because firefly luciferases and their genes are widely used as bioanalytical reagents, reporter genes and biosensors5,6, and more recently are emerging as novel intracellular ratiometric biosensors for pH and heavy metals ions7,8, there is a lot of interest to better understand the origin of pH-sensitivity.
The color of bioluminescence depends on both the chemical structure of the emitter and on the surrounding luciferase active site microenvironment. The emitter, excited oxyluciferin, has potentially 6 forms including tautomers and anionic species9,10,11. Most recent experimental and theoretical studies indicate that the keto-phenolate form is the most likely emitter11,12, but the enol-phenolate and enolate-phenolate forms were also proposed as potential emitters13,14,15. Three general mechanisms have been quoted to explain bioluminescence color determination by the luciferase active site7: (I) non-specific solvent and polarizability effects16,17; (II) specific acid-base and electrostatic interactions of active site residues with excited oxyluciferin9,18 and (III) the conformation of the active site affecting the rigidity of the microenvironment and the rotation of thiazinic rings of excited oxyluciferin19. Among the specific interactions, the presence of bases near the thiazinic side of the luciferin binding site assisting the tautomerization between a keto and enol forms has been originally claimed to explain green to red bioluminescence color change in firefly luciferases9,13. More recently, interactions influencing the resonance forms of excited oxyluciferin20, specific acid-base and finally electrostatic effects around oxyluciferin 6′ phenol group are being considered to determine bioluminescence colors18,21,22.
More than 30 beetle luciferases have been already cloned, sequenced and investigated in the past 20 years23,24,25,26,27,28,29,30,31,32,33,34,35, most of them from fireflies. The three-dimensional structures have been solved for the North-American firefly luciferase Photinus pyralis (Ppy) in the absence of substrates36, in the presence of bromophorm37 and with the C-terminal trapped in a closed conformation38, for the Japanese L. cruciata firefly luciferase complexed with the luciferyl-adenylate analogue DLSA in a closed conformation and with oxyluciferin and AMP in an open conformation39, and finally, for Lampyris turkanensis luciferase40. The luciferin binding-site is well conserved among different beetle luciferases and consists mainly of the following residues and segments (P. pyralis numeration): R218, the motif 244HHGF247, the loop 314SGGAPLS320, and the β-hairpin motif 340YGLTETTS34741,42,43.
In firefly luciferases, several single-point mutations in the luciferin binding site and in other regions located far from the active site resulted in red mutants41,42,43,44,45,46, whereas a few of the respective mutations in the pH-insensitive click beetle and railroadworm luciferases affected the bioluminescence color47,48,49,50, indicating that the active sites of pH-sensitive firefly luciferases is somehow less rigid than that of pH-insensitive luciferases50. Mutation of the luciferin binding site residues, including histidines H244 and H245, which supposedly could assist oxyluciferin tautomerization in the thiazolyl part of the active site, resulted in red mutants in firefly luciferases41,42,43. However, H245 is invariant and H244 is conserved in beetle luciferases displaying distinct bioluminescence colors, therefore none of them could play the role of basic residues assisting tautomerization of excited oxyluciferin13. Furthermore, studies with the 5,5-dimethylluciferin-adenylate indicated that there is no need of C5 proton abstraction and tautomerization of oxyluciferin for pH-sensitive spectral changes20. Interactions of main-chain amide bonds with oxyluciferin phenolate44 and the active site conformation, were also proposed to be important for bioluminescence colors.
Noteworthy, a group of mutations which affected spectra in both pH-sensitive and pH-insensitive luciferases was found to be clustered in the loops 223–23551,52 and 351–36033,53. The motif 227Y(F/V)GN(T)229 in the loop 223–235 was shown to be critical for bioluminescence colors and pH-sensitivity. In the loop 351–360, mutation of E354, which is conserved in the great majority of firefly luciferases, affected thermostability54 whereas the natural substitution E354N was shown to be responsible for the red-shifted, broader and more pH-sensitive bioluminescence spectrum in Macrolampis sp2 firefly luciferase33. These loops, participate in a network of polar interactions with E311 and R33720,53 (Fig. 1), in the same bromophorm binding cavity described by Franks et al.37, leading to a first suggestion that this network could be the pH-sensor of firefly luciferases52. In this amphiphilic cavity, E311 and R337 are located internally close to oxyluciferin phenolate, and the residues H310 and E354 are located more externally. We recently provided evidences that the salt bridge between E311 and R337 is essential to stabilize a closed hydrophobic green emitting conformation in firefly luciferases55. However, the specific function of these residues and the identity of the specific targets responsible for pH- and metal-sensitivity remain unsolved.
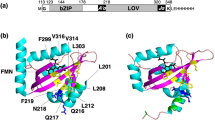
(Upper panel) Beetle luciferases primary structures multialignment, showing the residues H310, E311, R337 and E/N354 and interacting residues: (gray shadow) conserved residues of the pH-sensor in firefly luciferases; (yellow shadow) variable residues and (red shadow) unique substitutions affecting pH and metal sensitivity; (Lower panel) three-dimensional homology model of Cratomorphus distinctus firefly luciferase (83% identity with P. pyralis luciferase) showing E311/R337 and H310/E354 salt bridges and the network of polar interactions with the loop between residues 223–235 according to Viviani et al.52 which lead to the identification of the metal binding site.
Considering that H310, E311 and E354 display both prototropic and metal chelating side-chains, we decided to investigate whether they are involved in metal and pH-sensitivity. The mutation of H310 and N354 by other residues with metal chelating groups in Macrolampis firefly luciferase was recently shown to affect the metal-sensitivity in firefly luciferases8, indicating that this region is indeed critical for metal-sensitivity in firefly luciferases. Here we compared the effect of natural and artificial substitutions of these residues on metal- and pH-sensitivities of three Brazilian firefly luciferases displaying different bioluminescence colors (Amydetes vivianii: 539 nm; Cratomorphus distinctus: 548 nm and Macrolampis sp2: 569 nm; Fig. 2), modelled this region with Zinc ion, and calculated the associated electrostatic potential and pKas, providing convincing evidences that this region indeed constitutes the metal binding site and main pH-sensing moiety of firefly luciferases.
(Upper panel) Fireflies which emit different bioluminescence colors: (A) Amydetes vivianiifirefly, (B) Cratomorphus distinctus larva and (C) Macrolampis sp2 firefly, and (D) pH effect on in vitro bioluminescence elicited by Macrolampis sp2 firefly luciferase. (Lower panel) Bioluminescence spectra of 4 firefly luciferases displaying distinct bioluminescence colors at different pHs: (I) Amydetes vivianii; (II) Cratomorphus distinctus; (III) Photinus pyralis and (IV) Macrolampis sp2; (a) 0.10 M Tris-HCl buffer pH 8.0; (b) 0.10 M Phosphate buffer pH 8.0; (c) phosphate buffer pH 7.0; (d) phosphate buffer pH 6.0.
Results
Firefly luciferases bioluminescence spectra display different pH-sensitivities
When comparing the bioluminescence spectra of four firefly luciferases displaying different colors and pH-sensitivities (Fig. 2: Amydetes vivianii, 538 nm; Cratomorphus distinctus, 548 nm; Photinus pyralis, 558 nm, and Macrolampis, 564 nm), the bioluminescence spectrum of Amydetes vivianii luciferase was the most blue-shifted with the lowest amount of red light at pH 8.0, and is the less sensitive to pH, whereas that of Macrolampis sp2 luciferase has the broadest one with largest amount of red light even at pH 8, with Photinus pyralis and Cratomorphus disctinctus luciferases displaying intermediate values (Fig. 2). According to the solvent effect on the fluorescence spectra of luciferin and derivatives16,17, the results indicate that the active site of Amydetes luciferase is the most hydrophobic for both green and red emitters at pH 8 and 6. Recent quantum yield measurements by Ando et al.56 showed that the pH affects only the quantum yield of green emission, whereas the quantum yield of red emission is almost unaffected. Thus, we can assume that in other firefly luciferases too, only the amount of green emitter changes at different pHs. The results reinforce our previous proposal that in firefly luciferases the emission spectra are determined mainly by the ratio of green and red emitters52, although both green and red emitters may also experience slightly different environments in each luciferase, influencing the overall bioluminescence spectrum.
Firefly luciferases display different metal sensitivities
Considering that the structural targets of sensitivity to pH, metal ions and temperature must be the same, and that firefly luciferases display different degrees of pH-sensitivity, we compared the effect of Zn2+, as a representative metal íon, on the bioluminescence spectra of the above firefly luciferases and their mutants. Similarly to the pH sensitivity, quantum yield measurements showed that the binding of these metal ions to P. pyralis luciferase induce decrease of the quantum yields of the green emission component, resulting in the remaining red emission spectra57.
C. distinctus luciferase, similarly to Photinus pyralis luciferase, showed a larger red shift than Macrolampis sp2 luciferase in the presence of Zn2+ (Fig. 3). The luciferase of Amydetes firefly was much less to this metal (results not shown). Therefore, among these luciferases, that of C. distinctus was the most sensitive to these metal ions.
(Upper panel) Effect of ZnSO4 at the concentration of 2 mM (gray lines) in the bioluminescence spectra of distinct firefly luciferases and their mutants (black lines): (A) Cratomorphus distinctus; (B) Macrolampis sp2; (C) Macrolampis sp2 N354E; (D) Cratomorphus distinctus E354N; (Lower panel) Effect of pH on the bioluminescence spectra of Macrolampis firefly luciferase mutants: (A) wild-type; (B) Mac-H310C; (C) Mac-N354C and (D) Mac-N354H.
Effect of E354 and H310 mutations on bioluminescence spectra and pH-sensitivity
Considering H310 and E354 are located close to E311 and R337, and that natural substitutions and mutations of E354 were shown to affect bioluminescence spectra and pH-sensitivity, we investigated the effect of mutations by residues which lack or replace the acid-base side chains of E354 and H310 in firefly luciferases pH-sensitivity. All mutants of H310 and E354 (H310A, H310C, N354C, N354H, N354E), independently of being substituted by acid-base or non-acid-base residues, still displayed pH-sensitivity (Fig. 3; Table 1). The wild-type C. distintus luciferase, was more sensitive to pH than Macrolampis luciferase, which naturally displays the substitution E354N. The most pH-sensitive mutants of Macrolampis firefly luciferase were N354C > N354E > H310C/N354C > N354H > WT.
Whereas the mutant H310A has little effect on the bioluminescence spectrum of Macrolampis sp2 luciferase at pH 8.033, the mutation H310A in Cratomorphus luciferase, resulted in a considerable broadening and red-shifting of the spectrum (results not shown). The absence of effect of H310A in Macrolampis sp2 luciferase, which naturally has the substitution E354N, and the broadening effect of H310A in Cratomorphus luciferase, which has E354, supports the existence of a critical stabilizing interaction between H310 and E354 in Cratomorphus luciferase (and perhaps P. pyralis luciferase) which may stabilize a closed green-emitting conformation, differently from Macrolampis sp2 firefly luciferase which lacks the interaction and produces a broader and red-shifted emission. Furthermore, the mutation of H310R in Macrolampis sp2 luciferase, which inserts a permanent positive charge in this region, was previously shown to result in a very broad and red-shifted spectrum20, indicating that a positive charge at this position change the distribution of emission toward the red.
Altogether, these results indicate that although mutations the positions 310 and 354 influence spectral distribution, they are not determinants of pH-sensitivity.
H310 and N354 are metal-sensitive sites
Considering that substitutions of E354 affect the bioluminescence spectrum and pH-sensitivity of firefly luciferases, we decided to investigate whether this position is also involved with metal sensitivity. The mutant Mac-N354E in Macrolampis firefly luciferase displayed a larger red-shift in presence of Zn2+, similar to that observed for the wild-type C. distinctus luciferase that naturally displays the substitution N354E (Fig. 3; Table 1). The reverse mutant, E354N in C. distinctus luciferase, had opposite effect, undergoing a smaller red shift, similar to that observed for the wild-type Macrolampis sp2 that displays N354 at this position. These results indicate that the position 354 is indeed important for metal sensitivity.
Since in Cratomorphus distinctus and Photinus pyralis luciferases E354 is at hydrogen bonding distance to H310, potentially making a salt bridge, and histidines have strong affinity to divalent metals, we also investigated whether the mutation of H310 affects metal sensitivity. The mutant H310A8, in which the metal chelating imidazole side-chain is removed, indeed was much less sensitive to metal ions, indicating that H310 is also important to bind metal ions such as Zn2+. These results prompted us to investigate the effect of other substitutions of H310 and N354 by residues with metal chelating side-chains such as His, Cys or Glu on the metal-sensitivity of Macrolampis firefly luciferase, resulting in a recently published paper in which we produced mutants with different sensitivities to Zn2+, Hg2+ Cd2+ and Ni2+ which could be used to ratiometrically estimate these metal concentrations8.
Altogether, the above results clearly indicate that H310 and especially E354 are important sites for metal-sensitivity. However, the fact that the mutation of these residues by residues with non-chelating side-chains does not completely abolish metal-sensitivity, indicate that these residues are not the only targets for metal binding, and that there must be at least another metal binding group in the neighborhood.
Structure of the metal binding site
Thus, in order to see whether the divalent heavy metal ions can bind to H310, E354 and surrounding region, modeling studies of beetle luciferases docked with Zn2+ were done (Fig. 4). Indeed, Zn+2 complexed with H310 N and carbonyl groups, with the side-chains of N354 in Macrolampis firefly luciferase or E354 in Cratomorphus firefly luciferase, and furthermore with E311 carboxylate, supporting the experimental results, showing that H310, E311 and E/N354 constitute the metal binding site.
Structure of the metal binding site in firefly luciferases: (A) zoom showing Cratmorphus distinctus firefly luciferase luciferin binding site (dark blue) and metal binding site (colored side-chains); (B) zoom showing Cratomorphus distinctus firefly luciferase complexed with Zinc; (C) Zoom showing Macrolampis sp2 firefly luciferase complexed with Zinc. The dark blue area in the three figures represent the oxyluciferin phenolate binding pocket; yellow represent His310, green Glu 311 and pink Glu(Asn) 354.
Role of the salt bridge between E311 and R337
Previously, we showed that the salt bridge between E311 and R337 is essential to stabilize the closed hydrophobic conformation in firefly luciferases, and that mutations which removed the charge or decreased the size of the side-chains, resulted in red light emission in firefly luciferases54. In order to investigate whether the salt bridge between E311 and R337 has just a conformational effect on bioluminescence color, or whether these residues display additional catalytic effects on bioluminescence color, we inverted the identity of these residues by constructing the double mutant E311R/R337E. Such double mutant is expected to keep the salt bridge, and therefore a closed conformation. If the color of bioluminescence was related exclusively to the maintenance of the closed conformation, we would expect that the double mutant would also produce yellow-green light at pH 8.0, such as the wild-type enzyme. However, the double mutant produced weak red bioluminescence (601 nm; Table 1). These results clearly indicate that green emission does not depend exclusively on a closed conformation kept by the salt bridge, but rather on additional specific interactions that such residues may play, especially E311.
E311 is the main proton binding site
The above structural and site-directed mutagenesis results indicate that E311 display a key role in bioluminescence color. The mutants E311A and E311Q, in which the negative charge was removed, besides being the most red-shifted ones, are totally pH-insensitive and completely lost the bioluminescence activity at pH 6.0. The mutant E311R completely lost the activity, indicating that charge reversal is incompatible at this position. Only the mutants E311D and R337K, in which the side chains were shortened, but the charge preserved, still displayed a considerable degree of pH-sensitivity: at pH 6.0 E311D spectrum shifts about 17 nm to the red and R337K about 10 nm (Fig. 5). These results indicate that E311 negative charge is essential for green-yellow bioluminescence and also for pH-sensitivity in firefly luciferases. Furthermore, in contrast to R337 mutants, which independently of charge removal or reversal still produce weak red bioluminescence activity, the charge reversal of E311 (E311R), or removal (E311Q and E311A) under acidic pH, completely abolished the bioluminescent activity. Altogether, these results provide compelling evidences that E311 is the primary base responsible for green bioluminescence, and for proton and metal binding responsible for pH- and metal-sensitivityies, playing also a catalytic role in bioluminescence.
Bioluminescence spectra of mutants with 6′Amino-luciferin
Previously we showed that the 6′-amino-luciferin, which has the 6′ hydroxil group substituted by an amino group, is a pH-insensitive luciferin analog which is useful to probe the oxyluciferin phenolate binding site microenvironment polarity21. Therefore, in order to check whether the mutations of H310 and N354 cause substantial polarity changes, we analyzed the bioluminescence spectra of this analog with mutants of these residues. However, the bioluminescence spectra peaks for all the mutants were very close to each other (Table 1), indicating that the microenvironment polarity is quite similar and is not considerably affected by these mutations.
pKa analysis of the side-chains of pH-sensor
In order to better understand the involvement of the above residues and their mutants in pH-sensitivity, we calculated the pKa of the identified residues in the wild-type luciferases and mutants, and correlated them with the electrostatic potential and bioluminescence spectra at different pHs and in the presence of metals (Fig. 6; Table 2 Supplemental Materials).
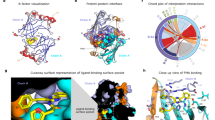
Electrostatic and van der Waals potential for the four pH-sensor residues microenvironment considering the solvent effects: (A) Energy distributions for all positions; (B) linear model showing the relationship between spectrum wavelength emission and electrostatic potential. Microenvironments of pH-sensor residues; (C) Structural relationships between pH-sensor microenvironment and active site (surface; blue) of four luciferases. Microenvironment for position 310 is represented in yellow, position 311 in green, position 337 in cyan and position 354 in magenta (numbering is based on Cratomorphus distinctus luciferase); (D–G) Representations for pH-sensor residues of Amydetes vivianii, Cratomorphus distinctus, Macrolampis sp2 and Phrixotrix vivianii luciferases, respectively. Residues in each microenvironment are represented by sticks in the same color of microenvironment. Putative position for metal binding is indicated by red sphere. Arrow indicates position of phenolate involved in deprotonation. Volume of active site is 600 A3 66.
Among the wild-type residues, H310 was the one which displayed the largest shift of pKa value in relation to the reported value in water (~7.0), displaying a much more basic value (7.85 for Macrolampis luciferase and 8.95 for Cratomorphus luciferase). The pKa values for E311 and R337 were 4.53 and 13, respectively, close to the expected values in water.
As expected, the mutation of these residues affected the pKa of nearby side-chains. Only R337 pKa (13.0) was insensitive to mutations of nearby residues. Among the side-chains that had their pKa values affected by nearby mutations, again the imidazolium side-chain of H310 was the most sensitive site, changing several pH units upon mutations of E311 and R337, whereas the pKa of E311 carboxylate had its pKa only slightly changed upon the mutations of H310 and N354, but much less upon mutation of R337. The largest effects on the pKa of H310 were observed upon the mutations E311A and E311Q which removed the negative charge and decreased the value more than 3 pH units, below 5.4, whereas the mutation E311D, which preserved the negative charge, but with shorter side chain, also considerably lowered the pKa of H310. Altogether, these results are fully consistent with the need of the negative charge of E311 carboxylate to counterbalance the positive charge of R337 guanidinium group. Upon the removal of the negative charge of the E311 carboxylate by the mutations E311A or E311Q, the H310 imidazolium group becomes much more acidic, apparently to loose the extra positive charge in the neighborhood of the unshielded R337 guanidinium ion. Mutation of R337K, which preserved the positive charge but shortened the side chain, also had considerable lowering effect of H310 pKa. In agreement with the above results, charge reversal upon the mutation R337E resulted in a considerable increase of pKa of H310, which is consistent with the need of an additional neighbor positive charge in H310 to counterbalance the extra negative charge of E337. Altogether the results clearly indicate that all these side-chains are close enough to influence each other, especially E311 and R337 strongly influencing H310.
Electrostatic potential calculation
To verify the effects of microenvironment in pH sensor residues, we calculated the electrostatic and van der Waals potential for each one of four putative residues considering the solvent effects. It can be observed that the positions 310 and 354 have similar contributions for pH-sensor site, whereas positions 311 and 337 display opposite potentials (Fig. 6-psloA). Furthermore, taking the total potential of four residues and the difference of these potentials at pH 6 and 8, unveil a clear relationship between increasing potential difference and a larger spectral shift, reinforcing the role of these residues on pH sensitivity.
To find the main potential components involved in the spectral change, the electrostatic potential for each candidate residue was combined in all possible pairs for the three wild-type pH-sensitive proteins of this study (Amydetes vivianii, Cratomorphus distinctus, Macrolampis sp2 firefly luciferases) and comparatively with the Phrixotrix hirtus railroadworm luciferase, which is an extreme model of pH-insensitive luciferase which naturally emits red light. At pH 8.0, of eleven evaluated regression models (Spectrum function as a function of energy), one model showed the best fitting: energy as the sum of electrostatic potential of the residue pair 310/354 (R2: 0.99) (Fig. 6_psloB). The green emitting luciferases of Amydetes and Cratomorphus display a stabilized electrostatic pair E311/R337, as indicated by favorable micro-environment (lowest potential value, about −900 kJ/mol). Here, the pair H310/E354 acts as strong acid-base pair, ultimately receiving the proton of phenolic hydroxyl group of oxyluciferin. In the more red-shifted Macrolampis luciferase, there is just one strong base (E312; equivalent to position 311 in Photinus pyralis luciferase) which makes an electrostatic pair with R338 (equivalent position to 337). Absence of the second base at position 355 (equivalent to position 354), increased the microenvironment potential, decreasing the strength to retain the proton ejected from excited oxyluciferin phenolate in the neighborhood. The red-emitting luciferase of Phrixotrix does not display the electrostatic pair (308/334 corresponding to 311/337) due the natural substitution of arginine by leucine (L334 corresponding to R337), and the entrance of this cavity (T307 and N351 corresponding to the couple H310/E354) do not have bases to assist residue E308 (equivalent to position 311) to trap oxyluciferin phenolate released proton. Not surprisingly, this luciferase displays the worst microenvironment potential (~−500 kJ/mol), with insensitivity to pH changes. Furthermore, although the mutation L334R slightly blue-shifts the spectrum of Phrixotrix luciferase, its microenvironment potential remained almost unaffected (−503 kJ/mol). Taken together, these results indicate that measurements of the microenvironment potential of pair 310/354 can provide a good estimate about pH sensitivity.
Discussion
The structural identity of the pH- and metal-sensing moiety, as well as the underlying mechanisms of pH-sensitivity and bioluminescence colors in firefly luciferases, have remained elusive for decades.
Three mechanisms to explain pH-sensitivity have been proposed: (1) presence of basic residues near oxyluciferin C59, promoting tautomerization, or phenolic hydroxyl group12 which could be protonated at lower pH; (2) conformational changes mediated by protonation of specific basic residues important to keep the active-site conformation, leading to entry of water molecules increasing the polarity of the oxyluciferin phenolate binding site52; (3) direct electrostatic interactions with oxyluciferin phenolate53,57. These hypotheses are not mutually exclusive.
The presence of bases, which has been originally suggested for the tautomerization hypothesis, are also required to accept the released proton from excited oxyluciferin phenol group. According to the hypothesis by Hirano et al.12, a base close to the phenolic hydroxyl group of oxyluciferin may serve to receive the proton, and the proximity of the conjugate acid to the excited oxyluciferin phenolate, may increase the degree of covalent bond character of the O···H bond between the phenolate group and the hydrogen of the conjugate acid. The compromise between the strength of the base and the polarity of the environment, may affect the degree of covalent character of the O···H bond and therefore modulating bioluminescence spectrum. Protonation of such bases at low pH may therefore affect bioluminescence spectra. However, any protic group, including water and main-chain amide groups, may serve as weak temporary bases to accept the proton ejected from the strongly acidic phenolic hydroxyl group of excited oxyluciferin (pKa ~ 1.0). The main-chain amide group of S/C/T314 (Photinus pyralis sequence number) indeed have already been shown to be important as putative assisting base near luciferin phenol group44.
pH-sensitivity was also explained on the basis of protonation of basic residues involved in keeping a closed active site conformation, promoting a conformational change to a more open and polar conformation, in the benzothiazolyl part of oxyluciferin binding site53. Nakatsu et al.39, showed that L. cruciata luciferase displays a close active site conformation with the critical luciferyl-adenylate intermediate analog, DLSA, and an open one with the products AMP and oxyluciferin. Later studies with 6′-aminoluciferin analogs supported the existence of two binding modes for oxyluciferin in firefly luciferases active site, an apolar (N) responsible for green light emission and another polar (P) responsible for red emission21,22. The active site in firefly luciferases may undergo conformational changes between these two extremes, and the light emitting step may take place somewhere between these two conformations. Whether such polar and apolar binding modes correspond to distinct conformations or sites remain to be elucidated.
Finally, electrostatic effects have been also claimed to play important roles in emission spectra modulation53,57,58,59. The presence of positive charges near oxyluciferin phenolate were suggested to blue shift the emission spectrum53,57. In this regard, the absence of an arginine at position 334 in the red emitting luciferase of Phrixotrix railroadworm luciferase was shown to be responsible for the very red-shifted spectrum of this enzyme, and inclusion of an arginine upon the mutation L334R (corresponding to R337 in P. pyralis luciferase) blue-shifted the emission55. In the pH-sensitive luciferases and their mutants, although the calculation of the overall microenvironment charge, considering the four residues H310, E311, R337 and E354, showed a trend for blue-shifted emission in electrostatically more negative environments and red-shifted emission for more positive environments (Table 2), the mutant R337E, in which there is a substantial increase of the overall negative charge character of the environment, also resulted in red light.
Here we show for the first time that the pH-sensing moiety and metal binding site responsible for pH-sensitivity involve the residues H310, E311, R337 and E(N)354. These four residues, with their acid-base side-chains are involved in two main electrostatic interactions that keep a closed and tight active site conformation favorable for green light emission: E311/R337 and H310/R354. The residues E311 and R337 make a more stable and internal salt bridge closer to oxyluciferin phenolate, whereas H310 and E354 are located in a more external and polar part of this cavity, making a second potential gate which is more susceptible to external pH and temperature changes.
The divalent metal ions such as Zn2+, Cu2+, Hg2+, Pb2+, Cd2+ clearly fit in this amphyphilic cavity, coordinating especially with E311 and E354 carboxylates, which are located close to each other (3.5 Å), and also with H310 imidazolium in most firefly luciferases (Fig. 4). The imidazole and carboxylate side chains of residues at positions 310 and 354 are responsible for modulating the sensitivity of Macrolampis and Cratomorphus firefly luciferases to metal ions such as nickel and zinc8. The interaction of the divalent metal ions with these glutamates in most firefly luciferases disrupt the salt bridges between E311/R337 and H310/E354, unshielding the positive charges of H310 and especially of R337, polarizing the environment (Fig. 7). Such excess of positive charges increases the polarity of the oxyluciferin phenolate binding pocket by creating a hydration shell around R337, resulting in red light emission. Noteworthy, the mutation H310R in Macrolampis luciferase which naturally lacks E354, partially simulates the effect of the metal, by inserting an additional positive charge in the neighborhood, and thus broadening and red-shifting the spectra.
Despite their importance for metal sensitivity, neither H310 nor E354 can be considered the only proton and metal binding sites, since H310 and E354 are not strictly conserved in other firefly luciferases which are also pH-sensitive, and their mutations by non-acid-base and residues do not abolish either pH-and metal-sensitivity. The residue H310 is substituted by threonine or valine in some firefly and other beetle luciferases. In Macrolampis firefly luciferase the lack of E354 negative charge upon the natural substitution E354N is responsible for the observed weaker metal sensitivity and for the naturally increased proportion of red light observed in its bioluminescence spectrum in relation to the close C. distinctus and P. pyralis firefly luciferases. Therefore, other groups could be important for metal binding, including the main chain amide bond of C/S/T314 which has been already shown to be located near oxyluciferin phenolate and to be important for bioluminescence color modulation44.
The internal salt bridge between E311 and R337 was previously shown to play a critical role in bioluminescence color, by keeping a closed hydrophobic conformation favorable for green light emission55. Although we already suggested that these residues could play additional specific functions besides the salt bridge, the effective contributions of the salt bridge in keeping a closed conformation, or specific interactions of E311 and R337 in bioluminescence color determination, remained to be elucidated. Here we showed that the exchange of these residues in the double mutant E311R/R337E, with the aim to keep the salt bridge and a closed conformation, resulted in a red emitting mutant, indicating that green light emission does not depends exclusively on a closed active site conformation, but also on more specific effects of E311 and R337.
The results shown in this manuscript bring compelling evidences that E311 carboxylate is the main base responsible for green light emission and pH-sensitivity, and the third metal binding site in firefly luciferases. The acid-base properties of E311 carboxylate, the residual pH-sensitivity upon its substitution by the conservative aspartate (E311D), the complete abolishment of pH-sensitivity upon substitution by other residues that lack negative charge (E311A/Q), provide compelling evidences that E311 carboxylate, most likely through a water molecule, acts as a main base in bioluminescence color determination. Although, the estimated pKa value of E311 carboxylate in this environment (~4.5) is apparently lower than the expected value for a critical base mediating the pH effect between 6.0–8.0, the close proximity to oxyluciferin phenol which acts as a strong acid upon chemiexcitation, may affect the pKa of E311 increasing the sensitivity of this group to external pH-changes.
Furthermore, the complete abolishment of bioluminescence activity upon charge reversal in the mutant E311R, and in the negative charge lacking mutants E311A and E311Q at pH 6.0, indicate the unforeseen possibility of a more integral role of E311 carboxylate in bioluminescence catalysis. All beetle luciferases have E311, and in most of them the negative charge of E311 carboxylate is counterbalanced by the positive charge of guanidinium group of residue R337 (only Phrixotrix luciferases do not). According to CTIL (charger transfer induced luminescence) mechanism, during the chemiexcitation step, the phenolic hydroxyl group of the dioxetanone intermediate must be necessarily deprotonated to give phenolate, in order to induce intramolecular charge transfer, promoting its decomposition into carbon dioxide and the excited singlet oxyluciferin which decays with emission of light8. The carboxylate of E311, assisted by a water molecule, may play the essential role of catalytic base for proton abstraction of the phenolic hydroxyl group of the dioxetanone intermediate, generating excited oxyluciferin phenolate during the chemiexcitation step. Because excited oxyluciferin phenol is a much stronger acid (pKa < 1.0) than E311 carboxylate (pKa ~ 4.5), it is likely that E311 carboxylate will work as a base retaining the proton. Under such circumstances, E311 carboxylate will be protonated and R337 guanidinium ion will in turn counter stabilize the negative charge of oxyluciferin phenolate. In contrast, in the electropositive microenvironment of the mutants E311R, or E311A/Q at pH 6.0, the phenol proton abstraction necessary for the generation of the critical phenolate ion during the dioxetanone intermediate activation could be halted, severely impacting bioluminescence activity.
At alkaline pH, the electrostatic couple E311/R337, reinforced by the external H310/E354 salt bridge, may work as a gate which keeps a closed conformation, as well as an electrostatic cushion that keeps the oxyluciferin ejected proton near its phenolate into a high energy state (Fig. 7). Acidic pH, the presence of heavy metals or higher temperatures weaken such electrostatic gates, decreasing the force to retain the oxyluciferin excited state released proton near its phenolate group. The few surrounding organized active site water molecules (Fig. 7) may escape as hydronium ions and be exchanged with water, polarizing and increasing the mobility of the microenvironment. Altogether these factors will decrease the energy of excited state resulting in red light emission. Similarly, in the absence of the external salt bridge between H310 and E354 in some beetle luciferase and mutants, the more internal salt bridge between E311 and R337 could be also weakened and the environment relaxed, increasing the proportion of red light. The lack of these salt bridges is observed in Macrolampis firefly luciferase which has E354 substituted by asparagine and produces a broader bioluminescence spectrum, and in a more extreme case, Phrixotrix hirtus railroadworm red emitting luciferase, which also displays asparagine at the corresponding position 351 and, and additionally lacks arginine at position 334 (L334 corresponding to R337 of P. pyralis luciferase), missing both salt bridges corresponding to E311/337 and H310/E354.
Material and Methods
Plasmids and beetle luciferases cDNAs
The cDNAs for Macrolampis sp2 luciferase was originally inserted in pPro vector (Invitrogen) and then was subcloned into NdeI site of pCold vector (Takara, Japan), the cDNA of Cratomorphus distincus luciferase in pBluescript vector32, the cDNA of Amydetes vivianii luciferase was cloned in pSport vector (Invitrogen)35.
Site-directed mutagenesis
The mutants Mac H310A, H310C, N354E, N354C, N354H, E311A, E311Q, E311D were previously constructed8 by site-directed mutagenesis using an Agilent site-mutagenesis kit (Catalog 200518). The following primers were designed for generating new mutations: (E311R) CT AAT T TG CAC AGA ATT GCT TCT GG; (R337E) CCA GGT ATA GAA CAA GGA TAT GGG C. To prepare the double mutant Mac-E311R/R337E we performed mutagenesis using the mutant E311R and the primers Mac-R337E. The plasmids containing the luciferase cDNAs were amplified using Pfu turbo polymerase and 2 complementary primers containing the desired mutation, using a thermal cycler (1 cycle 95 °C; 25 cycles 95 °C, 30 s; 55 °C, 1 min and 68 °C 7 min). After amplification, mutated plasmids containing staggered nicks were generated. The products were treated with Dpn I in order to digest non-mutated parental plasmids, and used directly to transform E. coli XL1-Blue cells.
Luciferase expression and purification
For luciferase expression, transformed E. coli BL21-DE3 cells were grown in 500–1000 mL of LB medium at 37 °C up to OD600 = 0.4, and then induced at 18 °C with 0.4 mM IPTG during 18 h. Cells were harvested by centrifugation at 2,500 g for 15 min and resuspended in extraction buffer consisting of 0.10 M sodium phosphate buffer, 1 mM EDTA, 1 mM DTT and 1% Triton X-100, 10% glycerol and protease inhibitor cocktail (Roche), pH 8.0, lysed by ultrasonication and centrifuged at 15,000 g for 15 min at 4 °C. The N-terminal histidine-tagged Macrolampis sp2 luciferase was further purified by agarose-Nickel affinity chromatography followed by dialysis and anion-exchange chromatography, according to established procedures54. The concentrations of purified luciferases were between 0.5 and 1 mg/mL, and the estimated purity, according to SDS-PAGE gels were about 90%.
Measurement of luciferase activity
Luciferase bioluminescence intensities were measured using a AB2200 (ATTO; Tokyo, Japan) luminometer. The assays were performed by mixing 5 μL of 40 mM ATP/80 mM MgSO4 with a solution consisting of 5 μL of luciferase and 85 μl of 0.5 mM luciferin in 0.10 M Tris-HCl pH 8.0 at 22 °C. All assays were measured in triplicate.
Bioluminescence spectra
Bioluminescence spectra reported here were recorded in ATTO Lumispectra spectroluminometer (Tokyo, Japan) with cooled CCD camera whereas the previously reported spectra used the same equipment or a Hitachi F4500 spectrofluorometer. The bioluminescence spectra reported here using the CCD-based spectroluminometer are red-shifted in relation to previously reported values measured with a spectrofluorometer. Such differences are likely to be caused by distortions associated to the longer scanning time and photosensitivity correction in the older spectrofluorometer in relation to the new equipment. For the in vitro bioluminescence recorded using the spectroluminometer, 5.0 μL of luciferases were mixed with 90 μL of 0.10 M Tris-HCl pH 8.0, 5 μL of 10 mM D-luciferin or 6′-aminoluciferin, and 5 μL of 40 mM ATP/80 mM MgSO4. The pH effect on bioluminescence spectra was analyzed in 0.10 M Phosphate buffer pH 8.0 and 6.0, and 0.10 M Tris-HCl pH 8.0. The 6′-aminoluciferin analog was synthetized by T. Hirano.
Homology Modelling
Homology-based models of Phrixotrix hirtus, Pyrearinus termitilluminans, Amydetes vivianii and Macrolampis sp2. luciferases were constructed using as template the three-dimensional structure of Luciola cruciata luciferase in the presence of DLSA (PDB code 2D1S) and of oxyluciferin and AMP (PDB code – 2D1R) as previously described55, Modeller v9.9 was used to align the sequences (using align2d function) and to construct 200 three dimensional models of each sequence Visualization and analyses of the best model of each luciferase were performed using PyMol version 1.4.160.
pKa estimation and system neutralization
Initially, side-chain conformations for pH-sensor residues were optimized using SCWALL method of Optimize function in YASARA61. To achieve theoretical pKa values for each residues of studied luciferase we used the routine “Experiment Neutralization” of software YASARA62. Initially, the hydrogen bonding network was optimized61. The method to predict pKa is based on a empirical function that uses hydrogen bonds, Coulomb potential and surface accessible area parameters weighted by experimental data. Electrostatic terms are calculated using Ewald summation63,64 using force field AMBER03. pKa values for Asp, Glu, His and Lys residues were predicted on the pH 6.0 and 8.0. During model construction, protonation states were assigned using these rule: Protonate Asp and Glu if the predicted pKa is higher than the pH; Protonate His if the predicted pKa is higher than the pH and His is not a hydrogen bond acceptor or deprotonate otherwise. Cys is deprotonated if the selected pH is higher than 8.7. Deprotonate Lys if the predicted pKa is lower than the pH. After that, simulation box was filled with water molecules, and ions are placed at the positions to neutralize highest and lowest electrostatic potential residues. A short MD simulation was ran for the solvent, and water molecules are subsequently deleted until the water density has reached 1.0 g/ml.
Potential Energy calculation
To calculate the potential energy of the selected residues using parameters from force field Amber03 and cutoff distance of 8.0 A. Energies were calculated for wild-type luciferases of Amydetes, Cratomorphus, Macrolampis and Phrixotrix. Statistical software R65,66 was used to create a script that processed energy data for each pH-sensor residue. To check the combination of residues that better explained the spectrum shift, the energies of four residues were combined in pairs and trios, creating eleven microenvironment putative potentials. Using linear model that relate emission wavelength of a given luciferase against each microenvironment potential, it was defined the best combination using the explained variance statistic (R2). The results are plotted using ggplot2 library67.
Concluding Remarks
The proton and metal binding site responsible for pH- and metal-sensitive bioluminescence color change in firefly luciferases involves mainly the side-chains of H310, E311 and E354 near the oxyluciferin phenolate binding site. These residues are involved in two critical salt bridges (E311/R337 and H310/E354) which keep oxyluciferin phenolate binding site electrostatically closed, promoting green light emission. The carboxylate of E311 is the main base responsible for green light emission and pH-sensitivity, playing also a critical role as a water mediated catalytic for bioluminescence activity base, by assisting proton abstraction of the dioxetanone intermediate phenol group during the chemiexcitation step. Under physiological and alkaline pH conditions, the stable salt bridge between the residues E311 and R337, aided by the more external salt bridge between E354 and H310 (in many firefly luciferases), keep a closed hydrophobic active site environment retaining the excited oxyluciferin released proton near its phenolate group into a high energy state, promoting green light emission. At lower pH or upon metal binding to H310, E311 and E354, the salt bridge between E311 carboxylate and R337 guanidinium is disrupted, reducing the ability of E311 to retain the excited oxyluciferin released proton near its phenolate group, polarizing the environment and thereof resulting in red light emission.
References
Wood, K. V. The chemical mechanism and evolutionary development of beetle bioluminescence. Photochem. Photobiol. Sci. 62, 662–673 (1995).
Viviani, V. R. The origin, diversity and structure function relationships of insect luciferases. Cell. Mol. Life Sci. 59, 1833–1850 (2002).
Seliger, H. H. & McElroy, W. D. The colors of firefly bioluminescence: Enzyme configuration and species specificity. Proc.Natl. Acad. Sci. USA 52, 75–81 (1964).
Viviani, V. R. & Bechara, E. J. H. Bioluminescence of Brazilian fireflies (Coleoptera: Lampyridae): spectral distribution and pH effect on luciferase-elicited colors. Comparison with elaterid and phengodid luciferases. Photochem. Photobiol. Sci. 62, 490–495 (1995).
Roda, A., Pasini, P., Mirasole, M., Michelini, E. & Guardigli, M. Biotechnological application of bioluminescence and chemiluminescence. Trends Biotechnol. 22, 295–303 (2004).
Viviani, V. R. & Ohmiya, Y. Beetle luciferases: Colorful lights on biological processes and diseases of referencing Photoproteins in Bioanalysis, (ed. Daunert, S. & Deo, S. K.), Wiley, New York, 49–60 (2006).
Gabriel, G. V. & Viviani, V. R. Novel application of pH-sensitive firefly luciferases as dual reporter genes for simultaneous ratiometric analysis of intracellular pH and gene expression/location. Photochemical & Photobiological Sciences. 13, 1661–1670 (2014).
Gabriel, G. V. & Viviani, V. R. Engineering the metal sensitive sites in Macrolampis sp2 firefly luciferase and use as a novel bioluminescent ratiometric biosensor for heavy metals. Analytical and Bioanalytical Chemistry. 408, 8881–8893 (2016).
White, E. H., Rapaport, E., Hopkins, T. A. & Seliger, H. H. Chemi- and bioluminescence of firefly luciferin. Jam. Chem. Soc. 91, 2178–2180 (1968).
Cheng, Y. Y. What exactly is the light-emitter of a firefly? J. Chem. Theory Comput. 11, 5360–5370 (2015).
Orlova, G., Goddard, J. D. & Brovko, L. Y. Theoretical study of the amazing firefly bioluminescence: the formation and structure of the light emitters. JACS. 125, 6962–6971 (2003).
Hirano, T. et al. Spectroscopic studies of the light-color modulation mechanism of firefly (beetle) bioluminescence. J. Am. Chem. Soc. 131, 2385–2396 (2009).
White, E. H. & Branchini, B. Modification of firefly luciferase with luciferin analog: a red light producing enzyme. J. Am. Chem. Soc. 97, 1243–1245 (1975).
Naumov, P., Ozawa, Y., Ohkubo, K. & Fukuzumi, S. Structure and spectroscopy of oxyluciferin, the light emitter of the firefly bioluminescence. J. Am. Chem. Soc. 131, 11590–11605 (2009).
Ghose, A. et al. Emission properties of oxyluciferin and its derivatives in water: Revealing the nature of the emissive species in firefly bioluminescence. Phys. Chem. 119, 2638–2649 (2015).
Ugarova, N. N. & Brovko, L. Y. Protein structure and bioluminescence spectra for firefly bioluminescence. Luminescence 17, 321–330 (2002).
DeLuca, M. Hydrophobic nature of the active site of firefly luciferase. Biochem. 8, 160–166 (1969).
Silva, L. P. & da Silva, E. J.C.G. Chemiexcitation Induced proton transfer: enolate oxyluciferin as the firefly bioluminophore. Phys. Chem. 119, 2140–2148 (2015).
McCapra, F., Gilfoyle, D. J., Young, D. W., Church, N. J. & Spencer, P. The chemical origin of color diferences in beetle bioluminescence of referencing. In Bioluminescence and Chemiluminescence: Fundamental and Applied Aspects (eds Campbell, A. K., Kricka, L. J. & Stanley, P. E.) pp 387–391, (Wiley & Sons, Chichester, U.K, 1994).
Branchini, B. R. et al. An alternative mechanism of bioluminescence color determination in firefly luciferase. Biochemistry 43, 7255–7262 (2004).
Viviani, V. R. et al. Bioluminescence of beetle luciferases with 6′-Amino-D-luciferin analogues reveals excited keto-oxyluciferin as the emitter and phenolate/luciferin binding site interactions modulate bioluminescence colors. Biochemistry 53, 5208–5220 (2014).
Kakiuchi, M. et al. Spectroscopic Properties of Amine-substituted Analogues of Firefly Luciferin and Oxyluciferin. Photochem Photobiol. 93(2), 486–494 (2016).
De Wet, J. R., Wood, K. V., Helinsky, D. R. & DeLuca, M. Cloning of firefly luciferase cDNA and expression of active luciferase in Escherichia coli. Proc. Natl. Acad. Sci. 82, 7870–7873 (1985).
Tatsumi, H., Masuda, T., Kajiyama, N. & Nakano, E. Luciferase cDNA from Japanese firefly Luciola cruciata: cloning, structure and expression in E. coli, J. Biolum. Chemilum. 3, 75–78 (1989).
Tatsumi, H., Kajiyama, N. & Nakano, E. Molecular cloning and expression in E. coli of a cDNA enconding luciferase of a firefly Luciola lateralis. Biochem. Biophys. Acta. 1131, 161–165 (1992).
Devine, J. H., Kutuzova, G. D., Green, V. A., Ugarova, N. N. & Baldwin, T. O. Luciferase from the East European firefly Luciola mingrelica: cloning and nucleotide sequence of cDNA, overexpression in E. coli and purification of the enzyme. Biochem. Biophys. Acta. 1173, 121–132 (1993).
Ohmiya, Y., Ohba, N., Toh, H. & Tsuji, F. I. Cloning, expression and sequence analysis of cDNA for the Japanese fireflies, Pyrocoelia miyako and Hotaria parvula. Photochem. Photobiol. Sci. 62, 309–313 (1995).
Sala-Newby, G. B., Thomson, C. M. & Campbell, A. K. Sequence and biochemical similarities between the luciferases of the glow-worm Lampyris noctiluca and the firefly, Photinus pyralis. Biochem. J. 313, 761–767 (1996).
Li, Y., Buck, L. M., Scaeffer, H. J. & Leach, F. R. Cloning and sequencing of a cDNA for the firefly luciferase from Photuris pennsilvanica. Biochem. Biophys. Acta. 1339, 39–52 (1997).
Viviani, V. R. et al. Cloning and molecular characterization of the cDNA for the Brazilian larval Click-beetle Pyrearinus termitilluminans luciferase. Photochem. Photobiol. Sci. 70, 254–260 (1999).
Viviani, V. R., Bechara, E. J. H. & Ohmyia, Y. Cloning, sequence analysis and expression of active Phrixothrix railroad-worms luciferases: relationship between bioluminescence spectra and primary structures. Biochem. 38, 8271–8279 (1999).
Viviani, V. R., Arnoldi, F. G., Brochetto-Braga, M. & Ohmiya, Y. Cloning and characterization of the cDNA for the Brazilian Cratomorphus distinctus larval firefly luciferase: similarities with European Lampyris noctiluca and Asiatic Pyrocoelia luciferases. Comp. Biochem. Physiol., Part B: Biochem. Mol. Biol. 139, 151–156 (2004).
Viviani, V. R., Oehlmeyer, T. L., Arnoldi, F. G. C. & Brochetto- Braga, M. R. A new firefly luciferase with bimodal spectrum: identification of structural determinants of spectral pH-sensitivity in firefly luciferases. Photochem. Photobiol. Sci. 81, 843–848 (2005).
Alipour, B. S. et al. Molecular cloning, sequence analysis, and expression of a cDNA encoding the luciferase from the glow-worm Lampyris turkestanicus. Biochem Biophys. Res. Commun. 325, 215–222 (2004).
Viviani, V. R., Amaral, D. T., Prado, R. A. & Arnoldi, F. G. C. A new blue-shifted luciferase from the Brazilian Amydetes fanestratus (Coleoptera: Lampyridae) firefly: molecular evolution and structural/functional properties. Photoch. Photobiol. Sci. 10, 1879–1886 (2011).
Conti, E., Franks, N. P. & Brick, P. Crystal structure of firefly luciferase throws light on a superfamily of adenylate-forming enzymes. Structure. 4, 287–298 (1996).
Franks, N. P., Jenkins, A., Conti, E., Lieb, W. R. & Brick, P. Strucutral basis for the inhibition of firefly luciferase by a general anesthetic. Biophys. J. 75, 2205–2211 (1998).
Branchini, B. R. et al. Bioluminescence is produced from a trapped firefly luciferase conformation predicted by the domain alternation mechanism. Journal of the American Chemical Society 133, 11088–11091 (2011).
Nakatsu, T. et al. Structural basis for the spectral difference in luciferase bioluminescence. Nature 13, 372–376 (2006).
Kheirabadi, M. et al. Crystal structure of native and a mutant of Lampyris turkestanicus luciferase implicate in bioluminescence color shift. Biochimica et Biophysica Acta - Proteins and Proteomics 1834, 2729–2735 (2013).
Branchini, B. R., Magyar, R. A., Murtiashaw, M. H., Anderson, S. M. & Zimmer, M. Site-directed mutagenesis of Histidine 245 in firefly luciferase: a proposed model of the active site. Biochemistry 37, 15311–15319 (1998).
Sandalova, T. P. & Ugarova, N. N. Model of the active site of firefly luciferase. Biochemistry (Moscow, Russ. Fed.) 64, 962–967 (1999).
Branchini, B. R., Southworth, T. L., Murtiashaw, M. H., Boije, H. & Fleet, S. E. A mutagenesis study of the luciferin binding site residues of firefly luciferase. Biochem. 42, 10429–10436 (2003).
Viviani, V. R., Amaral, D. T., Neves, D. R., Simões, A. & Arnoldi, F. G. C. The luciferin binding site residues C/T311 (S314) influence the bioluminescence color of beetle luciferase through main-chain interaction with oxyluciferin phenolate. Biochem. 52, 19–27 (2013).
Kajiyama, N. & Nakano, E. Isolation and characterization of mutants of firefly luciferase which produce different colors of light. Protein Eng. 4, 691–693 (1991).
Koksharov, M. I. & Ugarova, N. N. Thermostabilization of firefly luciferase by in vivo directed evolution. Protein Eng., Des. and Sel. 24, 835–844 (2011).
Viviani, V. R. & Ohmiya, Y. Bioluminescence color determinants of Phrixothrix railroadworm luciferases: chimeric luciferases, site-directed mutagenesis of Arg215 and guanidine effect. Photochem. Photobiol. Sci. 72, 267–271 (2000).
Viviani, V. R., Uchida, A., Viviani, A. & Ohmiya, Y. The influence of Ala243(Gly247), Arg 215 and Thr226(Asn230) on the bioluminescence spectra and pH-sensitivity of railroad worm, click beetle and firefly luciferases. Photochem. Photobiol. Sci. 76, 538–544 (2002).
Viviani, V. R., Arnoldi, F. G. C., Ogawa, F. T. & Brochetto-Braga, M. R. Few substitutions affect the bioluminescence spectra of Phrixotrix (Coleoptera: Phengodidae) luciferases: a site-directed mutagenesis survey. Luminescence 22, 362–369 (2007).
Viviani, V. R. et al. Active-site properties of Phrixotrix railroad worm green and red bioluminescence-eliciting luciferases. The Journal of Biochemistry 140, 467–474 (2006).
Viviani, V. R., Silva Neto, A. J., Arnoldi, F. G. C., Barbosa, J. A. R. G. & Ohmiya, Y. The influence of the loop between residues 223-235 in beetle luciferases bioluminescence spectra: a solvent gate for the active site of pH-sensitive luciferases. Photochem. Photobiol. Sci. 84, 138–144 (2007).
Viviani, V. R. et al. The structural origin and biological function of pH-sensitivity in firefly luciferases. Photochem. Photobiol. Sci. 7, 159–169 (2008).
Moradi, A., Hosseinkhani, S., Naderi-Manesh, H., Sadeghizadeh, M. & Alipour, B. A. Effect of charge distribution in a flexible loop on the bioluminescence color of fireflies luciferase. Biochem. 48, 575–582 (2009).
White, P. J., Squirrell, D. J., Arnaud, P., Lowe, C. R. & Murray, J. A. H. Improved thermostability of the North American firefly luciferase: saturation mutagenesis at position 354. Biochem. J. 319, 343–350 (1996).
Viviani, V. R. et al. Glu311 and Arg337 stabilize a closed conformation and provide a critical catalytic base and countercation for green bioluminescence in beetle luciferases. Biochemistry 55, 1–8 (2016).
Ando, Y. et al. Firefly bioluminescence quantum yield and colour change by pH-sensitive green emission. Nature Photonics 2, 44–47 (2008).
Wang, Y., Akiyama, H., Terakado, K. & Nakhatsu, T. Impact of site-directed mutant luciferase on quantitative green and orange/red emission intensities in firefly bioluminescence. Scientific Reports 3, 2490–2495 (2013).
Min, C. G. et al. A time-dependent density functional theory investigation on the origin of red chemiluminescence. Chemphyschem. 11(1), 251–9 (2010).
Sali, A. & Blundell, T. L. Comparative protein modeling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
DeLano, W. L. The Pymol Molecular Graphics System, DeLano Scientific, San Carlos, CA (2002).
Krieger, E. et al. Improving physical realism, stereochemistry, and side-chain accuracy in homology modeling: Four approaches that performed well. CASP8. Proteins 77(9), 114–122 (2009).
Krieger, E., Dunbrack, R. L. Jr, Hooft, R. W. & Krieger, B. Assignment of protonation states in proteins and ligands: combining pKa prediction with hydrogen bonding network optimization. Methods Mol Biol. 819, 405–421 (2012).
Krieger, E., Nielsen, J. E., Spronk, C. A. & Vriend, G. Fast empirical pKa prediction by Ewald summation. J.Mol.Graph.Model. 25, 481–486 (2006).
YASARA View - molecular graphics for all devices - from smartphones to workstations Krieger, E., Vriend, G. Bioinformatics 30, 2981–2982 (2014).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria, https://www.R-project.org/ (2017).
Oliveira, S. H. P. KVFinder: steered identification of protein cavities as a PyMOL plugin. BMC Bioinformatics, 15(1), 197 (2014).
H. Wickham. Springer-Verlag New York, ggplot2: Elegant Graphics for Data Analysis (2009).
Acknowledgements
To FAPESP (2013/09594-0; 2010/05426-8), CNPq (401867/2016-1) and CAPES for financial support.
Author information
Authors and Affiliations
Contributions
Vadim R. Viviani idealized the research, prepared mutants H310A, H310R, E311A, E311R, E311R/R337E and Crt-E354E, recorded the bioluminescence spectra of these mutants, prepared Figures 1 and 2, and wrote the manuscript. Gabriele V. Gabriel prepared and characterized mutants H310C, N354C, N354H and measured the effect of metals on the bioluminescence spectra of mutants and prepared Figure 3. Vanessa R. Bevilaqua obtained and analyzed the bioluminescence spectra for the mutants and wild-type luciferases with amino-analogs, analyzed bioluminescence spectra and prepared Figure 5. A. F. Simões prepared and characterized mutants E311D, R337K and R337E. Takashi Hirano synthesized amino-analogs, prepared mechanistic Figure 7 and participated on the discussion of paper. Paulo S. Lopes de Oliveira constructed homology-based models of beetle luciferases complexed with Zinc, calculated the potentials and pKa of the side-chains of the pH-sensor residues, prepared Figures 4 and 6, participating on the mechanism proposal.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Viviani, V.R., Gabriel, G.V.M., Bevilaqua, V.R. et al. The proton and metal binding sites responsible for the pH-dependent green-red bioluminescence color tuning in firefly luciferases. Sci Rep 8, 17594 (2018). https://doi.org/10.1038/s41598-018-33252-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-33252-x
Keywords
This article is cited by
-
Role of Histidine 310 in Amydetes vivianii firefly luciferase pH and metal sensitivities and improvement of its color tuning properties
Photochemical & Photobiological Sciences (2024)
-
The orange light emitting luciferase from the rare Euryopa clarindae adult railroadworm (Coleoptera:Phengodidae): structural/functional and evolutionary relationship with green and red emitting luciferases
Photochemical & Photobiological Sciences (2024)
-
Effect of pH on the secondary structure and thermostability of beetle luciferases: structural origin of pH-insensitivity
Photochemical & Photobiological Sciences (2023)
-
Cloning and molecular properties of a novel luciferase from the Brazilian Bicellonycha lividipennis (Lampyridae: Photurinae) firefly: comparison with other firefly luciferases
Photochemical & Photobiological Sciences (2022)
-
Synthesis of bioluminescent gold nanoparticle–luciferase hybrid systems for technological applications
Photochemical & Photobiological Sciences (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.