Abstract
Pw1/Peg3 is an imprinted gene expressed from the paternally inherited allele. Several imprinted genes, including Pw1/Peg3, have been shown to regulate overall body size and play a role in adult stem cells. Pw1/Peg3 is expressed in muscle stem cells (satellite cells) as well as a progenitor subset of muscle interstitial cells (PICs) in adult skeletal muscle. We therefore examined the impact of loss-of-function of Pw1/Peg3 during skeletal muscle growth and in muscle stem cell behavior. We found that constitutive loss of Pw1/Peg3 function leads to a reduced muscle mass and myofiber number. In newborn mice, the reduction in fiber number is increased in homozygous mutants as compared to the deletion of only the paternal Pw1/Peg3 allele, indicating that the maternal allele is developmentally functional. Constitutive and a satellite cell-specific deletion of Pw1/Peg3, revealed impaired muscle regeneration and a reduced capacity of satellite cells for self-renewal. RNA sequencing analyses revealed a deregulation of genes that control mitochondrial function. Consistent with these observations, Pw1/Peg3 mutant satellite cells displayed increased mitochondrial activity coupled with accelerated proliferation and differentiation. Our data show that Pw1/Peg3 regulates muscle fiber number determination during fetal development in a gene-dosage manner and regulates satellite cell metabolism in the adult.
Similar content being viewed by others
Introduction
Genomic imprinting is a mammalian-specific form of gene regulation in which one allele is repressed depending upon parental origin1. Although about 100–200 parentally imprinted genes have been identified to date, it remains unclear how parental imprinting contributes to gene function and how this form of epigenetic regulation was evolutionarily selected1,2. In addition, during development, loss or ‘relaxation’ of imprinting in specific tissue and cell types leads to bi-allelic expression of imprinted genes3,4,5,6. This absence of imprinting regulates specific biological processes such as the generation and maintenance of the postnatal neural stem cell pool4,7. Furthermore, the regulation of imprinting is proposed to maintain gene dosage in central nervous system (CNS) stem cells during development and adult life8.
Pw1/Peg3 was isolated from a screen designed to identify genes that regulate skeletal muscle lineage commitment9, as well as being discovered an imprinted gene expressed primarily from the paternal allele10. During embryogenesis, Pw1/Peg3 is expressed at high levels upon gastrulation and down-regulated during fetal and postnatal development9. In addition to its expression during development, we found that Pw1/Peg3 is expressed in adult stem cells in all tissues examined thus far including skeletal muscle, skin, blood and CNS11. In adult skeletal muscle, Pw1/Peg3 is expressed in satellite cells, which give rise to new muscle fibers during regeneration, as well as in a subpopulation of interstitial progenitor cells (PICs) that consist of several non-muscle progenitor lineages12,13.
Several Pw1/Peg3 mutant mouse lines have been generated, including a recent line generated by our laboratory. While some differences in phenotypes have been described, all the mice share a defect in postnatal growth14,15,16,17,18. It has previously been shown that loss of Pw1/Peg3 function results in reduced postnatal growth with a decrease in lean mass and a concomitant increase in body fat17. This work highlights a central role for Pw1/Peg3 in regulating body metabolic pathways, consistent with the emerging role of imprinted genes as key players in mammalian metabolism19. Previous reports demonstrate that PW1 regulates two key cell stress pathways via interactions with the TNF receptor-associated factor2 (TRAF2) and p53-mediated cell death. By direct interaction with Siah1 (Seven in absentia homolog 1) and BAX (Bcl2-associate X) proteins, PW1 participates in cell death and growth arrest20,21,22. In addition, Pw1/Peg3 has been described as a tumor suppressor in glioma cell lines and human ovarian cancer23,24. Moreover, we note that PW1 contains 12 Krüppel-like DNA binding zinc fingers9,10 and chromosomal immunoprecipitation assays reveal that a large number of its potential gene targets are involved in mitochondrial function, suggesting a link between Pw1/Peg3 function and cell metabolism25. To support this hypothesis other studies have shown that Pw1/Peg3 regulates genes involved in lipid metabolism and plays a central role in catabolic processes15,26,27. Together, these studies suggest that Pw1/Peg3 controls not only whole body metabolic pathways but also the metabolic state of the cell.
Here, we investigated the role of Pw1/Peg3 specifically in skeletal muscle including postnatal growth and adult muscle progenitor function. We used a mutant floxed allele for Pw1/Peg3 (referred to henceforth as Pw1), that recombines exons 8 and 9 removing >90% of the coding domain. This mouse line was used to generate both a constitutive Pw1 loss-of-function mouse18 and to delete Pw1 function specifically in muscle satellite cells.
We report here that Pw1 mutant mice exhibit a decrease in myofiber number as compared to wildtype and this difference is established at birth. Interestingly, we observed that the Pw1 maternal inherited allele is expressed at very low levels, and its loss alone has no detectable phenotype. However, deletion of both Pw1 alleles in homozygotes has a more profound effect on myofiber number when compared to the deletion of only the paternal allele, revealing a functional contribution for maternally-inherited Pw1 when the paternal allele is deleted. In addition to a role in fiber number determination, we found that Pw1 deletion leads to a decline in satellite cell number and disrupts the balance between self-renewal and differentiation following injury. Transcriptome analyses comparing mutant and wildtype satellite cells reveals a down-regulation of gene expression involved in cell death and mitochondrial organization. Consistent with this, we observe that mutant satellite cells display an increase in mitochondrial activity and exit the quiescent state more rapidly than wildtype cells. Our study shows that Pw1 gene dosage regulates skeletal muscle growth and loss of Pw1 function abrogates satellite cell renewal and proper mitochondrial function. These findings provide further insights into the importance of imprinted genes in muscle development and homeostasis, and represent another example of selective biallelic expression of an imprinted gene in an adult stem cell niche.
Results
Pw1 gene-dosage regulates skeletal muscle mass and fiber number
Skeletal muscle represents ~50% of total body mass, therefore we investigated whether the decrease in lean mass and overall body size of Pw1 mutant mice was due to changes in muscle tissue. Hind limb skeletal muscles from all 4 genotypes (wild type, Pw1+/+; heterozygotes with the paternally inherited allele deleted, Pw1+/p−; heterozygotes with the maternally inherited allele deleted, Pw1m−/+; homozygotes for the mutant allele, Pw1m−/p−) of 3 month old male mice were examined. Pw1+/p− and Pw1m−/p− muscles displayed an overall reduction in weight and cross-sectional area as compared to wildtype (Pw1+/+), whereas no differences were detected between Pw1m−/+ and wildtype mice (Fig. 1A,B). In particular, the larger limb muscles, such as the Tibialis Anterior, Quadriceps and Gastrocnemius, were significantly reduced in size and mass in Pw1m−/p− mice as compared to the same muscle from Pw1+/p− mice, revealing a contribution of the Pw1 maternal allele to muscle growth in the absence of the canonically expressed paternally inherited copy (Fig. 1A,B). Since the overall body size and weight of Pw1 mutant mice are decreased18, we normalized muscle mass to total body weight and observed that, with the exception of the Quadriceps, muscle mass was reduced proportionally to body mass. We next measured myofiber cross-sectional area (CSA) and fiber number in the Tibialis Anterior (TA) muscle. Pw1+/p− and Pw1m−/p− fiber size distributions were unaffected as compared to wildtype and Pw1m−/+ (Fig. 1A,C). In contrast, the total number of TA fibers was significantly lower in the Pw1+/p− and Pw1m−/p− mice, when compared to the wildtype (Fig. 1D). The decrease in myofiber number was not accompanied by any change in relative distribution of fiber types (Type2A, 2B and 1 fibers) (Fig. S1A). Moreover, the reduction in fiber number was more pronounced in mice lacking both the paternal and maternal alleles (Pw1m−/p−) than when only the paternal allele was deleted (Pw1+/p−) (Fig. 1D), further supporting a role for the maternal allele in this process. Taken together, these results show that both Pw1 alleles can participate in the establishment of muscle fiber number whereas fiber type and size are unaffected.
Pw1 is required for the establishment of muscle size and myofiber number. (A) Representative photomicrographs at lower (upper panel; scale bars = 200 μm) and higher magnification (lower panel; scale bars = 50 μm) of TA muscles cross-sections from 3 month Pw1+/+, Pw1+/p−, Pw1m−/+ and Pw1m−/p− mice stained with hematoxylin and eosin. (B) Histograms showing the muscle weight (left panel) and muscle/body weight ratio (right panel) of Tibialis Anterior (TA), Extensor Digitorum Longus (EDL), Quadriceps (QDC), Gastrocnemius (GAS), Soleus (SOL) muscles from 3 month Pw1+/+, Pw1+/p−, Pw1m−/+ and Pw1m−/p− mice (n = 10 for each genotype). Pw1+/p− and Pw1m−/p− muscles are smaller than Pw1+/+ and Pw1m−/+ muscles. Pw1m−/p− TA, QDC and GAS muscles are smaller than Pw1+/p− TA, QDC and GAS muscles. (C) Fiber size distribution in 3 month Pw1+/+ (dash), Pw1+/p− (red), Pw1m−/+ (grey) and Pw1m−/p− (green) TA. Values represent the mean number ± s.e.m. per 100 fibers (n = 5 for each genotype). Statistical analyses were performed using Student’s t-test. Pw1 depletion did not result in fibers area differences. (D) Histograms representing the fiber number in 3 month Pw1+/+, Pw1+/p−, Pw1m−/+ and Pw1m−/p− TA muscles sectioned through the midbelly region (n = 7 for each genotype). The number of fibers is decreased in Pw1+/p− and Pw1m−/p− TA muscles as compared to Pw1+/+ and Pw1m−/+ TA muscles. Pw1m−/p− TA muscles have reduced number of muscle fibers as compared to Pw1+/p−. (E) Left panel-Histograms showing total body weight of postnatal day 0 (P0) Pw1+/+, Pw1+/p−, Pw1m−/+, and Pw1m−/p− mice (n = 14 for each genotype). Pw1+/p− and Pw1m−/p− are smaller as compared to Pw1+/+ and Pw1m−/+ mice. Right panel-Histograms representing the fibers number in P0 Pw1+/+, Pw1+/p−, Pw1m−/+, and Pw1m−/p− mice TA muscles (n = 5 for each genotype). Pw1+/p− and Pw1m−/p− TA have less fiber as compared to Pw1+/+. Pw1m−/p− TA displays less number of fibers as compared to Pw1+/p− TA. Values are expressed as mean ± s.e.m. Statistical analyses were performed using one-way ANOVA and Tukey’s post-test for multiple comparison *P < 0.05, **P < 0.01 and ***P < 0.001 (n ≥ 4); n.s., not significant. In all the graphs Pw1+/+ (+/+), Pw1+/p− (+/p−), Pw1m−/+ (m−/+), and Pw1m−/p− (m−/p−).
The mechanisms underlying myofiber number determination are not fully elucidated, however several studies suggest that skeletal muscle fiber number is determined during embryonic/fetal development28,29. Comparison of the body weight of newborn (P0) mice in all four genotypes indicated that Pw1+/p− and Pw1m−/p− mice display a slightly reduced weight compared to wildtype (Fig. 1E-left panel). We quantified total myofiber number in newborn TAs from all four genotypes. The fiber number was significantly lower in the TA from Pw1+/p− and Pw1m−/p− mice as compared to Pw1m−/+ and Pw1+/+mice (Fig. 1E-right panel). Additionally, newborn Pw1m−/p− mice displayed a higher reduction on the fiber number as compared to Pw1+/p−. Taken together, these results show that Pw1 plays a role during fetal development in the determination of muscle fiber number and while loss of the maternal allele alone has no observable phenotype, the loss of both alleles results in a stronger phenotype demonstrating a contribution of the maternal allele to the establishment of myofiber number.
Pw1 is transcribed from the maternal allele in skeletal muscle
Pw1 is considered to be expressed from the paternal allele, however the observation that the homozygous mutant (Pw1m−/p−) has a stronger fiber number phenotype as compared to the loss of only the paternal allele (Pw1+/p−), suggests that the maternally inherited allele has a required function. We showed previously that neonatal muscle expresses higher levels of Pw1 as compared to the adult9,20, however expression levels are elevated in the adult in response to injury with a peak of expression occurring five days after muscle damage (Fig. 2A).
Paternal and maternal Pw1 alleles are transcribed in skeletal muscle. (A) Expression levels of Pw1 wildtype allele normalized to Hprt1 gene expression from real time PCR at three, five, and seven days after CTX injury in adult Pw1+/+ TA muscle (n = 3 for each genotype). Pw1 has a peak of expression five days after muscle injury. (B) Expression levels of Pw1 wildtype allele from real time PCR normalized to Hprt1 gene expression in uninjured and five days after injury adult Pw1+/+, Pw1+/p−, Pw1m−/+, and Pw1m−/p− TA (n = 3 for each genotype). Maternal Pw1 transcript is detected in injured Pw1+/p− muscles. (C) Pw1 allele specific expression in adult TA muscle before and five days after CTX injury from reciprocal hybrid offspring of C57BL6/J and CAST/EiJ mice (n = 3 for each genotype). Bi-allelic Pw1 expression was observed in adult injured and uninjured muscle tissue. Values are expressed as mean ± s.e.m. In all the graphs statistical analyses were performed using Student’s t-test *P < 0.05, **P < 0.01 and ***P < 0.001. In all the graphs Pw1+/+ (+/+), Pw1+/p− (+/p−), Pw1m−/+ (m−/+), and Pw1m−/p− (m−/p−).
In order to assess the absence or presence of maternal Pw1 transcripts in skeletal muscle, we used real time PCR to detect the Pw1 wildtype allele in adult uninjured and injured skeletal muscle from 3 month Pw1+/+, Pw1+/p−, Pw1m−/+, and Pw1m−/p− TA. Using primers specific for the Pw1 wildtype allele, we observed maternal Pw1 transcripts in adult uninjured as well as 5 days following cardiotoxin (CTX) injury in Pw1+/p− muscle (Fig. 2B). We next stained muscle tissue sections from Pw1+/+, Pw1+/p−, Pw1m−/+ and Pw1m−/p− newborn muscle for PW1 expression but did not observe detectable levels of PW1 protein in either Pw1+/p− or Pw1m−/p− muscles samples as compared to Pw1m−/+ and Pw1+/+ muscle section (Fig. S1B). These results suggest that the levels of PW1 protein are either below detection limits or that the maternal transcript is not translated. Taken together, these data show an activation of the Pw1 paternal allele upon injury and, in the absence of an intact paternal allele, the maternal copy is also activated.
To determine whether the maternal expression observed was also evident in the presence of an intact paternally inherited Pw1 allele, we assessed allele-specific Pw1 expression in muscle tissue before and after injury in hybrid offspring from reciprocal crosses of Mus musculus domesticus (C57BL6/J) and Mus musculus castaneus (CAST/EiJ) strains. As a negative control (background), we measured the expression of CAST/EiJ Pw1 allele transcript from C57BL6/J X C57BL6/J crosses and vice versa in adult muscle tissue before and after injury (Fig. S2A–C). A comparable significant increase above background was quantified between pre- and post-injury states (Fig. S2B,C). These data show that in wild type animals, Pw1 transcription derives predominantly from the paternally inherited allele but that there is a low level of transcription from the maternal allele (5–15% depending on genetic background) (Fig. 2C). Importantly however, activation of the maternal allele was not observed after injury in the presence of an intact paternally inherited Pw1. Hence, maternal allelic activation upon injury is only evident in Pw1 mutants.
In adult skeletal muscle, Pw1 expression is restricted to two progenitor populations: satellite cells and PICs (PW1+ interstitial cells)12,30. Therefore, we asked whether the Pw1 maternal transcript distribution is cell-type specific. Using real time PCR, we analyzed Pw1 wildtype allele expression in satellite cells and the fibro adipogenic progenitor (FAPs), which represent a subpopulation of PICs, isolated from Pw1+/+ and Pw1+/p− adult muscle and observed that both stem cell populations display maternal Pw1 transcript expression (Fig. S3A). Taken together, these results reveal that the maternal Pw1 transcript is constitutively expressed at low levels in skeletal muscle throughout postnatal life, at least, and while overall levels increase in response to injury, the maternal transcript remains low and may not be translated.
Pw1 mutant satellite cells display impaired self-renewal
Satellite cells are required for proper regeneration31,32. As satellite cells express Pw1, we tested muscle regeneration in wildtype and Pw1m−/p− mice. The TA muscles were injured using cardiotoxin (CTX) and examined two weeks later. We observed no overt differences between mutant and wildtype muscles, nor did we observe significant levels of fibrosis or fat infiltration, which are features of muscle regenerative defects (Fig. S4A). In addition, myofiber size (cross-sectional area, CSA) post-regeneration, was unaffected by the loss of Pw1 function (Fig. S4B).
We next investigated the number of satellite cells based upon the expression of PAX7 in wildtype and mutant muscles before and after injury. These analyses revealed a ~10% higher number of PAX7+ cells in mutant muscle as compared to the wildtype prior to injury (Fig. 3A). We did not detect any co-staining of PAX7 and KI67 nor do we observe any staining for MYOD in Pw1 mutant satellite cells in uninjured muscles (data not shown) revealing that there is no overt chronic cell cycle activation in the absence of injury that could account for this increased number of satellite cells in Pw1 mutant muscles, however we cannot rule out an accumulation of satellite cells due to a low level of cell cycling in the absence of Pw1 function. While we detected an overall increase in the number of PAX7+ cells as compared to the steady-state, we observed a ~30% decrease in satellite cell number in the mutant muscle as compared to wildtype two weeks following CTX injury (Fig. 3A). These results suggest that loss of Pw1 function disrupts the maintenance of the satellite cell pool following injury via a loss in self-renewal capacity. To test this, we quantified the absolute number and percentage of self-renewing satellite cells in CTX injured muscle 5 days after injury by flow cytometry (Fig. 3B). Satellite cells capable of replenishing the stem cell pool (quiescent satellite cells) after acute injury are defined based on the surface expression of α7-integrin and CD34 and the absence of TER119, CD45 and SCA1 expression (after viability dye exclusion)33. Consistent with a reduction in self-renewal capacity, there was a marked decrease in the satellite stem cell pool in Pw1m−/p− as compared to Pw1+/+ muscles (Fig. 3B).
Pw1 null satellite cells display altered self-renewal properties. (A) Quantification of PAX7 positive cells in Pw1+/+ and Pw1m−/p− 3 month TA muscle before and after two weeks of a single CTX injury (n = 4 for each genotype). NI = non injured; SI = single injured. Values represent the mean number of positive cells ± s.e.m. per 100 fibers. The number of PAX7 positive cells is higher in uninjured Pw1m−/p− muscle as compared to Pw1+/+. In contrast, the number of PAX7 positive cells in injured Pw1m−/p− muscle is significantly lower as compared to Pw1+/+ two weeks after injury. (B) Left Panel: Flow cytometric analyses of single cells from Pw1+/+ and Pw1m−/p− mice TA muscles five days after injury gated for α7-integrin+/CD34+/TER119−/CD45−/SCA1−. The gate used to identify CD34+/SCA− quiescent satellite cells is shown in blue. Right Panel: Histograms showing percentages (left) and absolute cell numbers (right) of quiescent satellite cells CD34+/SCA− fraction analyzed as shown in (left panel) (n = 3 for each genotype). Quiescent satellite cells CD34+/SCA− population is reduced in Pw1m−/p− injured muscle as compared to Pw1+/+. (C) Left Panel: Representative cross-sections of Pw1+/+ and Pw1m−/p− TA muscles one week after two consecutive CTX injury immunostained for PAX7 (red), MYOD (green), and LAMININ (orange). Nuclei were visualized by DAPI. White and yellow arrows denote PAX7+/MYOD− (self-renewing) and PAX7+/MYOD+ (proliferating) myogenic cells respectively. Scale bar = 50 μm. Right Panel: Histograms showing proportions of myogenic cells PAX7+/MYOD− (red), PAX7+/MYOD+ (green) and PAX7−/MYOD+ (yellow). The proportion of self-renewing cells is decreased in Pw1m−/p− compared Pw1+/+, whereas the proliferating and committed myoblasts show a concomitant increase (n = 3 for each genotype). (D) Histograms showing proportions of PAX7+/KI67− cells (quiescent satellite cells, red), PAX7+/KI67+ cells (proliferating satellite cells, green) and PAX7−/KI67+ cells (proliferating myoblast, yellow) from Pw1+/+ and Pw1m−/p− TA muscles one week after two consecutive CTX injury (n = 3 for each genotype). The proportion of quiescent satellite cells is decreased in Pw1m−/p− compared Pw1+/+. (E) Histograms showing proportions of myogenic cells MYOD+/KI67− (committed cells, red), MYOD+/KI67+ (proliferating and committed cells, green) and MYOD−/KI67+ (proliferating cells, yellow) from Pw1+/+ and Pw1m−/p− TA muscles one week after two consecutive CTX injury (n = 3 for each genotype). The proportion of proliferating and committed cells is increased in Pw1m−/p− compared Pw1+/+. Values are expressed as mean ± s.e.m. Statistical analyses were performed using Student’s t-test *P < 0.05, **P < 0.01 and ***P < 0.001. In all the graphs Pw1+/+ (+/+) and Pw1m−/p− (m−/p−).
Satellite cell self-renewal can be measured by tracking the populations of PAX7−/MYOD+, PAX7+/MYOD+ and PAX7+/MYOD− cells one week following injury. This corresponds to a stage during which regeneration is ongoing, consisting of committed (PAX7−/MYOD+) and expanding satellite cell derived myoblasts that will form new myofibers (PAX7+/MYOD+), as well as a smaller population of cells that restore the satellite cell population (PAX7+/MYOD−)34,35. Using this approach, we observed that all three populations of cells are present in wildtype and mutant muscles after injury, however the population of self-renewing satellite cells (PAX7+/MYOD−) was markedly decreased in mutant muscle, while the number of proliferating and differentiation-committed myoblasts were markedly increased (Fig. 3C). In addition, using a marker for cell proliferation (KI67), we noted a reduced percentage of non-cycling satellite cells (PAX7+ KI67−) during muscle regeneration in the mutant muscles and an increase in the percentage of committed/proliferating (KI67+ MYOD+) cells (Fig. 3D,E). Taken together, these data reveal that Pw1 regulates satellite cell number and self-renewal capacity.
Pw1 loss-of-function abrogates muscle regeneration after multiple injuries
The reduced number of satellite cells following a single muscle injury in Pw1 mutant mice suggests that satellite cell self-renewal is compromised even though muscle regeneration appeared to be normal (Fig. S4A). Since stem cell self-renewal is essential for tissue regeneration in response to multiple injuries36,37,38, we investigated the effect of Pw1 loss in adult muscle regeneration after two consecutive injuries with CTX (double injury). H&E staining of doubly injured TA muscles revealed a low level of fibrosis and fat infiltration in Pw1 mutant muscle as compared to wildtype (Fig. S5A,B) suggesting compromised regeneration. In addition, we measured fiber cross-sectional area (CSA) after double injury and we observed a decrease of the percentage of newly formed small fibers (<800 μm) in Pw1m−/p− as compared to Pw1+/+ muscles as well as an increase of the percentage of newly formed bigger fibers (>35000 μm) (Fig. S5C). Furthermore, we noted an increased number of centrally located nuclei per fiber in doubly injured Pw1 mutant muscle as compared to wildtype (Fig. S5D), however fiber number was maintained in both genotypes (Fig. S5E). Lastly, we observed a reduction of PAX7+ cells in mutant muscles after two injuries (Fig. S5F) consistent with results obtained following a single injury (Fig. 3A) indicating that Pw1 deletion leads to the exhaustion of the satellite cell pool with the consecutive impairment of muscle regeneration. The larger fiber CSA coupled with an increase in myonuclear content suggested that the reduced satellite cells pool observed after multiple injuries in Pw1 null muscle has an enhanced differentiation capacity disrupting the normal balance between satellite cell expansion and terminal differentiation.
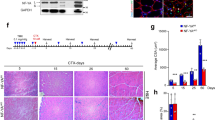
In order to confirm a Pw1 specific role in satellite cells, we used the Pax7-CreERT2 mice which carry a tamoxifen-inducible CRE recombinase-estrogen receptor fusion protein cassette driven by Pax731 crossed with the Pw1fl/fl mice18. The resultant Pax7-CRE:: Pw1fl/fl mice were used to delete Pw1 expression specifically in satellite cells following tamoxifen (TM) administration. Immunohistochemistry of 1 month old Pax7-CRE::Pw1fl/fl and Pax7-CRE TA revealed that PW1 expression was ablated in PAX7+ cells with very high efficiency one week following TM treatment (Fig. 4A,B). We next injured TA muscles of TM-treated Pax7-CRE and Pax7-CRE::Pw1fl/fl mice and analyzed muscle samples two weeks later (Fig. 4C). Interestingly, we observed that specific ablation of Pw1 expression in satellite cells led to a more severe defect in muscle regeneration including a marked increase in ectopic fat deposition and fibrosis even following a single injury (Fig. 4D). This stronger phenotype may reflect compensatory mechanisms that are established during development and postnatal life in the constitutive Pw1m−/p− mouse model as compared to the conditional Pax7-CRE::Pw1fl/fl targeted specifically to satellite cells. We note that as Pw1 is also expressed in the interstitial muscle cell population, these cells may participate in this compensation. Furthermore, as the Cre protein is expressed in the conditional allele in a genetic background in which one of the Pax7 alleles is recombined, there may be an additive effect that is not seen when the Pw1 allele is not recombined. Consistent with our previous results, we observed a decline in PAX7+ cells following injury in Pax7-CRE:: Pw1fl/fl (cKO) as compared to Pax7-CRE (CNT) (Fig. 4E). While the Pax7-CRE model leads to a loss of Pw1 expression in satellite cells, we noted that the PW1+ PAX7− interstitial cells (PICs) were also reduced in number (Fig. 4F), suggesting that loss of Pw1 function in satellite cells has an effect on neighboring niche populations.
Specific Pw1 deletion in PAX7 positive cells leads to impairment in satellite cell self-renewal. (A) Representative cross-section of 4 weeks-old Pax7-Cre and Pax7-Cre::Pw1fl/fl TA one week after TM injection stained for PAX7 (red), PW1 (green), LAMININ (orange) and DAPI (blue). White and yellow arrows in Pax7-Cre::Pw1fl/fl TA one week after TM injection indicate Pw1 deletion in satellite and interstitial PW1+ cells (PICs) respectively. Scale bar = 10 µm. (B) Percentage of PW1+ PAX7+ cells in 4 weeks-old Pax7-Cre and Pax7-Cre::Pw1fl/fl TA one week after TM injection stained as shown in (A) (n = 3 for each assay and genotype). All PAX7+ cell from Pax7-Cre::Pw1fl/fl TA do not express PW1 as compared to Pax7-Cre TA. (C) Schematic representation of experimental strategy: seven weeks old Pax7-Cre and Pax7-Cre::Pw1fl/fl mice were injected intraperitoneally with TM for three days. One day after the last TM injection, TA muscles were injured by a single CTX injection. Muscles were collected two weeks after injury. (D) Cross-sections images of TA muscles from Pax7-Cre and Pax7-Cre::Pw1fl/fl mice two weeks after CTX injury stained with hematoxylin and eosin (upper panels), Sirius Red (middle panels) and Oil-Red O (lower panels). Specific deletion of PW1 in satellite cells impairs skeletal muscle regeneration. (E,F) Quantification of satellite cells (PAX7+) and PICs (PW1+) per 100 fibers in TA from Pax7-Cre and Pax7-Cre::Pw1fl/fl mice two weeks after CTX injury (n = 3 for each assay and genotype). Regenerating myofiber in Pw1 depleted satellite cells have less number of satellite cells and PICs. Values represent the mean number of positive cells ± s.e.m. per 100 fibers. Statistical analyses were performed using Student’s t-test *P < 0.05, **P < 0.01 and ***P < 0.001.
Pw1 regulates mitochondrial function metabolically primes satellite cell activation
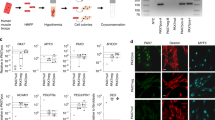
To elucidate how Pw1 affects satellite cell function, transcriptome analyses was performed on satellite cells from Pw1+/+ and Pw1m−/p− mice by RNA-sequencing (RNA-seq) (Table S1). Satellite cells were freshly isolated from whole hind-limb muscles of 3 months mice. Pw1m−/p− satellite cells displayed 211 downregulated genes (p < 0.05, log2 fold change <−0.5) and 208 upregulated genes (p < 0.05, log2 fold change >0.5) when compared to wildtype mice. Gene Ontology (GO) analyses revealed downregulation of the expression of genes related to mitochondrial organization and activity as well as cell death (Fig. 5A) (Figs 5B and S6A) (Table S2). Among the list of genes involved in mitochondrial organisation were known inhibitors of mitochondrial function, suggesting an increase in mitochodrial activity in satellite cells lacking Pw1 expression39,40,41. Consistent with this, we observed an increase in mitochondrial membrane potential in freshly isolated mutant satellite cells as compared to wildtype (Fig. 5C) as measured by Mitotracker staining42.
Pw1 deletion results in satellite cell activation. (A) Gene Ontology (GO) analysis of downregulated genes in RNA-seq analysis of Pw1m−/p− satellite cell. GO term represent the biological processes, dashed histograms indicates genes involved in mitochondrial activity and red histograms denote genes implicated in cell death. The x-axis shows the p-value (−log10). (B) GO clustering analysis for mitochondrial organization gene categories. Data were obtained from RNA-seq analysis of FACS sorted satellite cells from Pw1+/+ and Pw1m−/p− mice. The gene tree is shown on the left and gene coloring was based on normalized signals as shown. Each row represents one gene, with red representing high expression and blue representing low expression (n = 3 for each genotype). (C) Left panel: Representative FACS analysis of Mitotracker fluorescence in Pw1+/+ (black) and Pw1m−/p− (green) satellite cells. Right panel: Histogram represent Mitotracker mean fluorescence intensity (MFI) of Pw1m−/p− satellite cells relative to Pw1+/+ (n = 6 for each genotype). Loss of PW1 corresponds to an increase of mitochondrial activity in satellite cell. (D) Left panel: Representative images of FACS isolated Pw1+/+ and Pw1m−/p− satellite cells grown for 24 h in growth media (GM) and stained with DAPI. Right panel: Histogram showing quantitative evaluation of FACS isolated Pw1+/+ and Pw1m−/p− satellite cells area (in square micrometers), grown for 24 h in GM (n = 3 for each genotype). Pw1m−/p− satellite cells are larger than wildtype. (E) Histograms showing quantifications of the number of satellite cells per colonies (Left panel), percentages of satellite cell colonies bigger than 10 cells (Middle panel), and percentages of satellite cell colonies smaller than 4 cells (Right panel) from FACS purified satellite cells from adult (3 month) wildtype and Pw1m−/p− muscles grown for 5 days in GM. Pw1m−/p− satellite cells display increased proliferative capacity as compared to wildtype (n = 3 for each genotype). (F) Expression levels of PGC-1α from real time PCR normalized to Tbp gene expression from FACS isolated Pw1+/+ and Pw1m−/p− satellite cells satellite cells grown for 5 days in GM (n = 3 for each genotype). Pw1m−/p− satellite cells display increased expression level of PGC-1α gene as compared to wildtype. (G) Lactate production from FACS isolated Pw1+/+ and Pw1m−/p− satellite cells grown for 5 days in GM (n = 4 for each genotype). Pw1m−/p− satellite cells display decrease secretion of lactate as compared to wildtype. (H) Left panel: Representative images of FACS isolated Pw1+/+ and Pw1m−/p− satellite cells immunostained for MHC (red) and counterstained with DAPI (blue). Satellite cells were grown for 48 h in differentiation media (DM). Scale bar = 200 µm. Right panel: Fusion index (mean percentage of nuclei incorporated into MHC+ cells) of differentiating wildtype and Pw1m−/p− satellite cells stained as shown in left panel. Pw1m−/p− differentiated satellite cells show increased Myogenic differentiation (n = 4 for each genotype). (I) Expression levels of Pw1, Pax7, MyoD, Myogenin, Mck and Mhc1 from real time PCR normalized to Tbp gene expression in differentiated FACS isolated Pw1+/+ and Pw1m−/p− satellite cells. Differentiating Pw1m−/p− satellite cells display increased expression level of Myogenin and Mck gene as compared to wildtype (n = 3 for each genotype). Value are expressed as mean ± s.e.m. In all the graphs statistical analyses were performed using Student’s t-test *P < 0.05, **P < 0.01 and ***P < 0.001. In all the graphs Pw1+/+ (+/+) and Pw1m−/p− (m−/p−).
Recent studies have described a cell cycle phase that occurs prior to G1 entry referred to as GAlert defined by cellular metabolic activation43. Specifically, GAlert entry entails an increase in mitochondrial activity and an increase in satellite cell size with entry into this phase being regulated by mTORC1 activation and its downstream targets including phospho-S6 (pS6)43,44,45. While no significant differences in the size of freshly sorted Pw1+/+ and Pw1m−/p− satellite cells were observed (Fig. S6B), we detected a striking difference in size of Pw1 mutant and wildtype satellite cells one day after plating (Fig. 5D). The percentages of PS6+ PAX7+ cells in mutant and wildtype adult uninjured muscle are similar indicating that the cells are not fully in a GAlert phase as previously described, but rather are metabolically primed (Fig. S6C). Consistent with this, mutant satellite cells form more and larger colonies as compared to wildtype (Fig. 5E). It has been shown that satellite cell activation involves a switch from glycolytic to mitochondrial metabolism46 and that several mitochondrial genes such as PGC-1α regulate mitochondrial biogenesis and respiration to direct cells to oxidative metabolism47,48. We observed an increased in PGC-1α gene expression in Pw1m−/p− proliferating satellite cells as compared to Pw1+/+ (Fig. 5F). Furthermore, a reduction in lactate production in proliferating mutant satellite cells was observed, indicating an accelerated switch from glycolysis to mitochondrial metabolism in Pw1m−/p− satellite cells as compared to Pw1+/+ (Fig. 5G). In addition, Pw1m−/p− satellite cells form larger myotubes as compared to wildtype (Fig. 5H) and upregulate myogenic markers such as myogenin and Mck (Fig. 5I) consistent with a higher state of cell activation and a shift towards differentiation commitment. Taken together, these data suggest that Pw1 represses satellite cell activation and participates in maintaining a quiescent state.
Discussion
We generated a novel conditional mutant mouse model for Pw118 to analyze the role of Pw1 during skeletal muscle growth and in adult muscle stem cell function. Previous studies, including our initial description of a Pw1 mutant allele, have shown that constitutive loss of Pw1 function results in a postnatal growth defect14,17,18,49,50. In this study, we show that the loss of Pw1 causes a reduction in muscle mass accompanied by a ~30% decrease in myofiber number, but not in the myofiber cross-sectional area. Previous studies have shown that skeletal muscle fiber number is determined during embryogenesis in two waves of myogenesis (ED 11–14 and ED 14–16)51,52 corresponding to a stage when Pw1 also shows a peak of expression9. Other imprinted genes have been implicated in modulating muscle mass and myofiber number. This includes the polar overdominance muscle hypertrophy phenotype of the Callipyge sheep and related phenotypes associate with the altered dosage of imprinted genes at the Dlk1-Dio3 cluster in mice53,54. Loss of the maternally expressed genes Grb10 and H19 results in an overgrowth phenotype associated with an increase in myofiber number, whereas deletion of the paternally expressed genes Mest and Dlk1 leads to a decrease in myofiber number55,56,57,58. In these studies and the present analysis, the precise cellular basis for myofiber number differences has not been elucidated, and likely reflects complex interaction between the developing muscle connective tissue and forming myofibers. As Pw1 is expressed in both the muscle interstitium and myogenic cells, further analyses targeting loss-of-function in the interstitium will be required. Odd skip related 1 (Osr1) has been shown recently to be expressed in the developing muscle connective tissue, as well as the emerging fibroadipogenic population that constitutes a large portion of the PICs population, and loss of Osr1 function results in disorganized and poorly formed myofibers59. Thus, it is possible that the decrease in myofiber number observed in Pw1 mutant muscle results from a loss of Pw1 function in the muscle interstitium. Our data point to a key contributing factor in the postnatal growth defects observed in the Pw1 mutant mice; namely, while newborn mutant and wildtype mice do not show pronounced body size differences, the observation that mutant mice possess a decreased myofiber number poses a limit on final achievable body size.
Genomic imprinting is a form of epigenetic regulation that is limited to 100–200 mammalian genes1 and the selective advantage of parental imprinting remains obscure. Emerging evidence suggests that genomic imprinting is a mechanism of gene dosage control that regulates body growth and stem cell function4,7,60. In the present study, we found that the reduction in muscle fiber number is more pronounced in mice lacking both paternal and maternal Pw1 alleles as compared to muscle where only the paternal allele is deleted, suggesting that the maternal allele contributes to muscle growth in contexts where the paternal allele is absent. Recent studies have reported that Pw1 undergoes a relaxation of imprinting leading to the expression of the canonically repressed maternal allele in newborn and adult brain18,61. We confirmed a low level of Pw1 transcription from the maternal allele by assessing its imprinting status in the adult muscle of hybrid offspring from reciprocal crosses of Mus musculus domesticus (C57BL6/J) and Mus musculus castaneus (CAST/EiJ) strains in addition to verifying that either the wildtype maternal or the recombined maternal (truncated) allele are expressed in our heterozygous mutant mice. We note however that in mice carrying the mutant paternal allele and the wildtype maternal allele, we do not observe detectable levels of PW1 protein expression. Therefore, while our results demonstrate that Pw1 gene dosage participates in the establishment of muscle fiber number, it is not clear whether this is a result of overall protein levels or an interaction between the two Pw1 alleles via an undetermined mechanism. Regardless, our results, combined with the results of others55,56,57,58 reveal an interesting pattern in which maternally and paternally imprinted genes exert opposite effects upon muscle mass through the control of myofiber number.
Growing evidence points to a role for imprinted genes in adult stem cells11,62,63,64. Pw1 is part of a group of imprinted genes, referred to as the “imprinted gene network” (IGN). Members of the IGN are expressed at high levels during embryonic development, and whereas the overall expression levels decline postnatally, they remain highly expressed in adult stem cells63. In vivo and in vitro deletion of different members of the IGN reveal key roles in adult stem cell maintenance and self-renewal4,7,56,65,66. We have shown that Pw1 is specifically expressed in a wide range of adult stem cells11. Furthermore, it has been recently reported that Pw1 regulates adult mesoangioblast competence and that PW1 expressing cells correspond to competent and self-renewing cells11,67,68. In skeletal muscle, Pw1 expression is confined to two progenitor cell populations: satellite cells and interstitial cells (PICs)12,13. We observe here that Pw1 loss of function leads to a decline in muscle regenerative capacity coupled with fat deposition and the exhaustion of the satellite cell pool. Furthermore, our data indicate that Pw1 has a distinct role during fetal muscle development in the determination of muscle fiber number versus its role in adult satellite cells. To properly address this issue will require specific conditional alleles to the various cells types in developing skeletal muscle since the connective (interstitial) cells are known to play a role in fiber number determination and also express Pw1 during development and in the adult59. We note that recent results regarding the role of Osr1 during muscle development and in the adult reveal similarly distinct developmental roles and we further note that Osr1 and Pw1 expression largely overlap in the interstitial muscle cell population12,13,30,59,69.
Our findings raises the question regarding how Pw1 regulates adult stem cell function, and in particular, stem cell competence and self-renewal. Satellite cells are a quiescent cell population in adult skeletal muscle that is activated in response to muscle injury70. Replenishment of the satellite cell population by self-renewal is pivotal for skeletal muscle homeostasis and defects in this process compromise muscle regeneration71,72,73. Accumulating evidence points to a key role for the satellite cell niche in satellite cell fate determination73,74,75. We report here that Pw1 mutant satellite cells display an impaired self-renewal and specific mouse crosses that ablate Pw1 expression exclusively in satellite cells have a profound impact on muscle regeneration with fat and fibrotic tissue deposition. The intramuscular fat infiltration observed in Pw1 null regenerating muscle may result from a disruption of the muscle stem cell niche as well as satellite cell transdifferentiation along the adipocyte program76. These studies reveal that Pw1 participates in the regulation of satellite cell activation and in turn, controls self-renewal capacity. RNA-seq analyses of purified satellite cells from wildtype and Pw1 null mutant (Pw1m−/p−) mice revealed that multiple genes involved in mitochondrial organization and cell death are downregulated in the absence of Pw1. Our findings are consistent with a previous report showing that PW1 interacts with the mitochondrial cell death pathway22,77,78. These results are also consistent with previous ChIP-sequencing analyses in the adult brain that demonstrated that PW1 binds the promoters of multiple genes involved in mitochondrial function79. The link between Pw1 and mitochondrial function is further supported by the in vivo phenotypes reported here, revealing an increase in mitochondrial activity in Pw1 mutant satellite cells. Recent studies have uncovered a novel satellite cell cycle state in adult resting muscle characterized by elevated mitochondrial activity referred to as “GAlert”43,45. This state occurs in satellite cells that are distal to the site of injury and has been proposed to ‘prime’ satellite cells to enter the cell cycle. It should be noted that Pw1 mutant satellite cells exhibit some, but not all, of the characteristics of GAlert. It is likely that there are multiple steps involved in the transition of satellite cells from a quiescent state towards cell cycle activation in response to injury and that loss of Pw1 in the absence of injury metabolically primes satellite cells but is insufficient to enter a GAlert state identical to what has been previous described. We note however that mitochondrial function and metabolic activation are critical to cell cycle activation and mitochondrial activity plays a key role during stem cell fate regulation including governing a switch from glycolytic to oxidative (mitochondrial) metabolism80. In the case of satellite cells, mitochondrial activity is associated with their capacity to differentiate rather than self-renew81. Our previous results showing that Pw1 is expressed in all stem cells coupled with results presented in this study demonstrating that Pw1 regulates satellite cell self-renewal coupled with an activation of mitochondrial function provide a crucial link towards understanding a more global role for Pw1 in stem cell regulation.
Several studies have demonstrated that loss of Pw1 function disrupts overall body metabolism including an increase in body fat and a reduction in lean mass17. To date, constitutive mutants have been used primarily to study the roles of parentally imprinted genes. While these studies have proven invaluable, complex traits ranging from behavior to body growth and body composition have been ascribed to be under the control of many parentally imprinted genes expressed quite widely, including Pw162. Hence, the mutant phenotypes reported to date may not be due to the action of a single parentally imprinted gene in any one cell type or tissue, but rather could result from complex cell and tissue interactions. Comparison of constitutive and conditionally targeted mutants provides valuable insights in this regard. However, further work will be needed in order to understand whether the defects observed in the Pw1 constitutive mutant are due or not to a specific stem cell population. Nonetheless, using a conditional allele to target disruption of Pw1 function in a single tissue progenitor cell type we have shown that stem cell function is disrupted demonstrating a clear role for Pw1 in postnatal stem cell function in vivo further illustrating the importance of imprinted genes in the regulation of the postnatal stem cell niche.
Material And Methods
Mice
Mouse models used were Pw1 floxed (Pw1fl/fl) mice and constitutive Pw1 knock-out (Pw1m−/p−)18. Mice were maintained on a C57BL6 background. All four genotypes, Pw1+/+(wildtype), Pw1m+/p− (paternal deletion), Pw1m−/p+ (maternal deletion) and Pw1m−/p− (homozygous deletion), were analyzed. Tg: Pax7CreERT2 mice82; Tg: Pax7CreERT2 mice were crossed with Pw1 floxed (Pw1fl/fl) mice to obtain specific deletion of Pw1 in PAX7+ cells. Approval for the animal (mouse) work performed in this study was obtained through review by the French Ministry of Education (Agreement#A751320).
RNA extraction and real time PCR
Total RNA was extracted using RNeasy Micro Kit (Qiagen) and RNeasy Mini Kit (Qiagen) according to manufacturer guidelines. RNA was treated with RNase-free DNase I (Qiagen) to remove genomic DNA. RNA was reverse-transcribed using SuperScript III First-Strand Synthesis System (Thermo Fisher). Cycling conditions and primers were used as previously described18.
Real time PCR primers
Gabarapl1 forward CATCGTGGAGAAGGCTCCTA, Gabarapl1 reverse ATACAGCTGGCCCATGGTAG
Ppif forward TGGCTCTCAGTTCTTTATCTGC, Ppif reverse ACATCCATGCCCTCTTTGAC
Akt3 forward GGATCACAGATGCAGCTACC, Akt3 reverse GTAGAAAGGCAACCTTCCACAC
Atp2a1 forward ACACAGACCCTGTCCCTGAC, Atp2a1 reverse TGCAGTGGAGTCTTGTCCTG
Vps13c forward CACAAGCATTGAAGATAGAAGCAAAA, Vps13c reverse AGTGATGGCACAATGTCTTGTTG
Mgarp forward AAAGAACAAACAAAGGCGGAGTTG, Mgarp reverse CACACTTGCTCGGCTTCTGC
Hspa1l forward AGAGTTGTGTGCAGACCTGT, Hspa1l reverse CCGGGTTGGTTGTCAGAGTA
Tamoxifen treatment
7 weeks-old Pax7CreERT2::Pw1fl/fl mice were injected intraperitoneally daily for 3 days with tamoxifen (TM) (150 μl, 20 mg/ml; Sigma Aldrich) diluted in sunflower seed oil/5% ethanol.
Regeneration assays
Skeletal muscle regeneration was induced by intramuscular CTX injection (0.06 mg/ml, Sigma) and muscles were analyzed 14 days after injury. To analyze satellite cell self-renewal, muscle regeneration was induced by focal freeze crush, a second injury performed with a 15 days interval, and muscles were collected 7 days following the second injury. To analyze tissue regeneration, multiple injuries experiments were performed using CTX and muscle were collected 14 days following the second injury. All techniques used are described in detail35.
Histological and cells analyses
Muscles were weighed and frozen in liquid nitrogen-cooled isopentane as previously described49. Transverse cryosections (10 mm) were stained with haematoxylin and eosin. Collagen deposition was detected by Sirius Red staining83, fat tissue was stained by Oil Red O and hematoxylin84.
Transverse TA cryosection muscles were stained with LAMININ (Sigma) antibody to assess muscle fiber cross-sectional area (CSA) and number of muscle fiber. MHC isoforms were measured from cryosections obtained from the mid-belly of TA stained with MHC2b/BFF3, MHC1/BAD5 and MHC2a/SC71 antibodies (Hybridoma bank) as previously described35. Images were captured on a Zeiss AxioImagerZ1 microscope, and morphometric analysis was performed using MetaMorph7.5 (Molecular Devices). Entire muscle sections were analyzed from 3 to 5 animals per group. Sections and cultured cells were stained with antibodies for PW1, PAX7 (Developmental Studies HybridomaBank), KI67 (BD Biosciences and Abcam), MYOD (BD Biosciences and Santa Cruz), MF20 (Developmental Studies Hybridoma Bank), CASPASE3 (BD Biosciences) and PS6 (Cell signaling), species specific secondary antibodies coupled to AlexaFluor 488 (Molecular Probes), Cy3 or Cy5 (Jackson Immunoresearch), and nuclei were counterstained with DAPI (Sigma). For quantitative analysis, positive cells in at least 700 fibers from randomly chosen fields were counted from at least three animals per group.
FACS analysis and primary culture
Hind-limb muscles were processed to obtain single cells as previously described11,12. In order to isolate satellite cells, cells were incubated with rat anti-mouse CD45-PE-Cy7 (eBiosciences), rat anti-mouse TER119-APC (Becton Dickinson), rat anti-mouse CD34-brilliant violet (Becton Dickinson), rant anti-mouse α7-integrin-A700, rat anti-mouse SCA1-FITC (eBiosciences). Satellite cells were isolated by α7-integrin+/CD34+/TER119−/CD45−/SCA1−. Primary antibodies were used at a concentration of 10 ng.ml-1. DAPI and 7AAD were used to collect live intact cells. To stain mitochondria we used 25 nM MitoTracker Red CMXRos (M7512). Flow cytometry was performed on a FACS Aria (Becton Dickinson) and for flow cyotmetry data analysis we used Flowjo software.
Cells were grown in high-glucose Dulbecco’s modified eagle medium (DMEM, Gibco) supplemented with 2.5 ng.ml-1 bFGF (Invitrogen), 20% heat-inactivated FBS (Invitrogen), 10% heat-inactivated horse serum (Gibco), 1% (v/v) penicillin-streptomycin (Gibco), 1% (v/v) LGlutamine (Gibco) and 1% (v/v) Na-pyruvate (Gibco). Medium was changed every 2 days. For clonal analysis, purified cell populations were grown on gelatin-coated dishes at low density for 5 days. For myogenic differentiation, three thousand satellite cells were seeded in 48-well plates for 1 week and transferred to differentiation medium (DM) for 2 days: DMEM containing 5% (v/v) horse serum and 1% (v/v) penicillinstreptomycin. To inhibit glycolysis proliferating satellite cells were treated for 24 h with 2-deoxiglucose (5 mM). To analyze the area of satellite cells, freshly sorted satellite cells were grown for 24 h and fixed with PFA 4%. Satellite cells area was measured using the ImageJ software. Proliferative capacity was quantified by counting at least 100–150 colonies, from at least three independent experiments. Fusion indexes were quantified by counting the number of nuclei in MF20+ cells per total number of nuclei12,20.
Lactate assay
Three thousand purified satellite cells were seeded in 48-well plates. Lactate concentration was tested on cell culture medium 5 days after seeding. JM-K607–100 Lactate Assay Kit from cliniscience was used for the analysis.
Allele-specific determination assays
To estimate the proportional allelic expression of Pw1, mouse hybrids of CAST/EiJ and C57BL/6 strains were used. RNA was isolated and cDNA generated from the hybrid 7–9 week old male mouse muscles, Allelic expression was estimated by pyrosequencing. Briefly, the cDNA was amplified by PCR in a reaction mixture (Bioline reagents, 21060) consisting of 3 µl of 10X reaction buffer, 0.6 µl of primer mix (10 µM forward and reverse primers), 0.9 µl of 50 mM MgCl2, 1.2 µl of dNTPs 2.5 mM, 0.125 µl of 5U/µl Taq polymerase, 23.175 µl of water and 1 µl of cDNA (diluted 20X). Reaction parameters were; 94 °C for 2 minutes, 40 cycles of 94 °C for 30 seconds, 60 °C for 30 seconds and 72 °C for 30 seconds, followed by 72 °C for 5 minutes. Amplification primers were F(biotinilated): AAGGCTCTGGTTGACAGTCGTG and R: TTCTCCTTGGTCTCACGGGC. The biotin conjugated forward strand of the amplicon was separated from the reverse strand and cleaned: The PCR mixture, 30 µl, was mixed with 1 µl of streptavidine sepharose beads (GE, 17-5113-01), 39 µl binding buffer (QIAGEN, 979006) and 10 µl water. The mixture was incubated on a shaker for 5 minutes and processed on a pyrosequencing vacuum station (QIAGEN, 9001529). The biotinilated DNA strands were captured (via their binding with streptavidin), washed in 70% ethanol, 0.2 M NaOH and 10 mM Tris-acetate (pH 7.6) for about 30 seconds each, the vacuum removed and the DNA deposited in a plate containing annealing buffer (QIAGEN, 979009) and a sequencing primer (AATGAAAGACTCCCCAC). The plate was incubated at 80 °C for three minutes before analysis on the pyrosequencer. Pyrosequencing was conducted using the PyroMark gold reagents (QIAGEN, 972812).
RNA-Sequencing
Satellite cells were isolated by FACS from hindlimb muscle from 3 month Pw1+/+ and Pw1m−/p− mice. We purified RNA from muscle stem cell population using RNAqueous-Micro total RNA isolation Kit (life-technologies) with a gDNA degradation step.
Directional libraries were prepared using Truseq Stranded mRNA sample preparation kit following the manufacturer’s instructions (Illumina). Libraries were checked for concentration and quality on DNA chips with the Bioanalyser Agilent (Illumina).
The libraries were quantified by fluorimetric measurements with the Qubit® dsDNA HS Assay Kit (ThermoFisher). 51-bp Single Read sequences were generated on the Hiseq2500 sequencer according to manufacturer’s instructions (Illumina). The multiplexing level was 2 samples per lane.
Reads were cleaned of adapter sequences and low-quality sequences using an in-house program (https://github.com/baj12/clean_ngs). Only sequences at least 25 nucleotides in length were considered for further analysis. Tophat version 1.4.1.185, with default parameters, was used for alignment on the reference genome (GRCm38 from Ensembl database version 74). Genes were counted using HTSeq-count version 0.6.186 (parameters: -t exon -i gene_id -m intersection-nonempty -s yes).The package used for the statistical analysis is DESeq2. For this analysis, a BH p-value adjustment was performed and the level of controlled false positive rate was set to 0.05. REVIGO software was used for the GO analysis.
References
Ferguson-Smith, A. C. Genomic imprinting: the emergence of an epigenetic paradigm. Nature reviews. Genetics, https://doi.org/10.1038/nrg3032 (2011).
Barlow, D. P. & Bartolomei, M. S. Genomic imprinting in mammals. Cold Spring Harbor perspectives in biology, https://doi.org/10.1101/cshperspect.a018382 (2014).
Charalambous, M. et al. An enhancer element at the Igf2/H19 locus drives gene expression in both imprinted and non-imprinted tissues. Developmental biology, https://doi.org/10.1016/j.ydbio.2004.04.022 (2004).
Ferron, S. R. et al. Postnatal loss of Dlk1 imprinting in stem cells and niche astrocytes regulates neurogenesis. Nature, https://doi.org/10.1038/nature10229 (2011).
Hagege, H. et al. The 3′ portion of the mouse H19 Imprinting-Control Region is required for proper tissue-specific expression of the Igf2 gene. Cytogenetic and genome research, https://doi.org/10.1159/000090837 (2006).
DeChiara, T. M., Robertson, E. J. & Efstratiadis, A. Parental imprinting of the mouse insulin-like growth factor II gene. Cell (1991).
Ferron, S.R. et al. Differential genomic imprinting regulates paracrine and autocrine roles of IGF2 in mouse adult neurogenesis. Nature communications, https://doi.org/10.1038/ncomms9265 (2015).
Perez, J. D., Rubinstein, N. D. & Dulac, C. New Perspectives on Genomic Imprinting, an Essential and Multifaceted Mode of Epigenetic Control in the Developing and Adult Brain. Annu Rev Neurosci, https://doi.org/10.1146/annurev-neuro-061010-113708 (2016).
Relaix, F. et al. Pw1, a novel zinc finger gene implicated in the myogenic and neuronal lineages. Developmental biology, https://doi.org/10.1006/dbio.1996.0172 (1996).
Kuroiwa, Y. et al. Peg3 imprinted gene on proximal chromosome 7 encodes for a zinc finger protein. Nat Genet, https://doi.org/10.1038/ng0296-186 (1996).
Besson, V. et al. PW1 gene/paternally expressed gene 3 (PW1/Peg3) identifies multiple adult stem and progenitor cell populations. Proceedings of the National Academy of Sciences of the United States of America, https://doi.org/10.1073/pnas.1103873108 (2011).
Mitchell, K. J. et al. Identification and characterization of a non-satellite cell muscle resident progenitor during postnatal development. Nature cell biology. https://doi.org/10.1038/ncb2025 (2010).
Pannerec, A., Formicola, L., Besson, V., Marazzi, G. & Sassoon, D. A. Defining skeletal muscle resident progenitors and their cell fate potentials. Development, https://doi.org/10.1242/dev.089326 (2013).
Li, L. et al. Regulation of maternal behavior and offspring growth by paternally expressed Peg3. Science, (1999).
Kim, J. et al. Peg3 mutational effects on reproduction and placenta-specific gene families. PloS one, https://doi.org/10.1371/journal.pone.0083359 (2013).
Perera, B. P., Teruyama, R. & Kim, J. Yy1 gene dosage effect and bi-allelic expression of Peg3. PloS one, https://doi.org/10.1371/journal.pone.0119493, (2015).
Curley, J. P. et al. Increased body fat in mice with a targeted mutation of the paternally expressed imprinted gene Peg3. FASEB journal: official publication of the Federation of American Societies for Experimental Biology, https://doi.org/10.1096/fj.04-3216fje (2005).
Denizot, A. L. et al. A Novel Mutant Allele of Pw1/Peg3 Does Not Affect Maternal Behavior or Nursing Behavior. PLoS genetics, https://doi.org/10.1371/journal.pgen.1006053 (2016).
Peters, J. The role of genomic imprinting in biology and disease: an expanding view. Nature reviews. Genetics, https://doi.org/10.1038/nrg3766 (2014).
Schwarzkopf, M., Coletti, D., Sassoon, D. & Marazzi, G. Muscle cachexia is regulated by a p53-PW1/Peg3-dependent pathway. Genes & development, https://doi.org/10.1101/gad.412606 (2006).
Relaix, F., Wei, X. J., Wu, X. & Sassoon, D.A. Peg3/Pw1 is an imprinted gene involved in the TNF-NFkappaB signal transduction pathway. Nat Genet, https://doi.org/10.1038/ng0398-287 (1998).
Relaix, F. et al. Pw1/Peg3 is a potential cell death mediator and cooperates with Siah1a in p53-mediated apoptosis. Proceedings of the National Academy of Sciences of the United States of America, https://doi.org/10.1073/pnas.040378897 (2000).
Kohda, T. et al. Tumour suppressor activity of human imprinted gene PEG3 in a glioma cell line. Genes to cells: devoted to molecular & cellular mechanisms (2001).
Feng, W. et al. Imprinted tumor suppressor genes ARHI and PEG3 are the most frequently down-regulated in human ovarian cancers by loss of heterozygosity and promoter methylation. Cancer, https://doi.org/10.1002/cncr.23323 (2008).
Thiaville, M. M. et al. DNA-binding motif and target genes of the imprinted transcription factor PEG3. Gene, https://doi.org/10.1016/j.gene.2012.10.005, (2013).
Buraschi, S. et al. Decorin causes autophagy in endothelial cells via Peg3. Proceedings of the National Academy of Sciences of the United States of America, https://doi.org/10.1073/pnas.1305732110 (2013).
Neill, T., Sharpe, C., Owens, R. T. & Iozzo, R. V. Decorin-evoked paternally expressed gene 3 (PEG3) is an upstream regulator of the transcription factor EB (TFEB) in endothelial cell autophagy. The Journal of biological chemistry, https://doi.org/10.1074/jbc.M116.769950 (2017).
Goldspink, G. & Ward, P. S. Changes in rodent muscle fibre types during post-natal growth, undernutrition and exercise, J Physiol (1979).
Rowe, R. W. & Goldspink, G. Muscle fibre growth in five different muscles in both sexes of mice. II. Dystrophic mice. J Anat (1969).
Pannerec, A., Marazzi, G. & Sassoon, D. Stem cells in the hood: the skeletal muscle niche. Trends in molecular medicine, https://doi.org/10.1016/j.molmed.2012.07.004 (2012).
Lepper, C., Partridge, T. A. & Fan, C. M. An absolute requirement for Pax7-positive satellite cells in acute injury-induced skeletal muscle regeneration. Development, https://doi.org/10.1242/dev.067595 (2011).
von Maltzahn, J., Jones, A. E., Parks, R. J. & Rudnicki, M. A. Pax7 is critical for the normal function of satellite cells in adult skeletal muscle. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1307680110 (2013).
Ieronimakis, N. et al. Absence of CD34 on murine skeletal muscle satellite cells marks a reversible state of activation during acute injury. PLoS One, https://doi.org/10.1371/journal.pone.0010920 (2010).
Zammit, P. S. et al. Pax7 and myogenic progression in skeletal muscle satellite cells. Journal of cell science, https://doi.org/10.1242/jcs.02908 (2006).
Didier, N., Hourde, C., Amthor, H., Marazzi, G. & Sassoon, D. Loss of a single allele for Ku80 leads to progenitor dysfunction and accelerated aging in skeletal muscle. EMBO Mol Med, https://doi.org/10.1002/emmm.201101075 (2012).
Becker, A. J., Mc, C. E. & Till, J. E. Cytological demonstration of the clonal nature of spleen colonies derived from transplanted mouse marrow cells. Nature (1963).
Worton, R. G., McCulloch, E. A. & Till, J. E. Physical separation of hemopoietic stem cells differing in their capacity for self-renewal. J Exp Med 1969).
Ploemacher, R. E. & Brons, R. H. Separation of CFU-S from primitive cells responsible for reconstitution of the bone marrow hemopoietic stem cell compartment following irradiation: evidence for a pre-CFU-S cell. Exp Hematol (1989).
Boyer-Guittaut, M. et al. The role of GABARAPL1/GEC1 in autophagic flux and mitochondrial quality control in MDA-MB-436 breast cancer cells. Autophagy, https://doi.org/10.4161/auto.28390 (2014).
Chami, M. et al. Role of SERCA1 truncated isoform in the proapoptotic calcium transfer from ER to mitochondria during ER stress. Molecular cell, https://doi.org/10.1016/j.molcel.2008.11.014 (2008).
Baines, C.P. et al. Loss of cyclophilin D reveals a critical role for mitochondrial permeability transition in cell death. Nature, https://doi.org/10.1038/nature03434 (2005).
Cottet-Rousselle, C., Ronot, X., Leverve, X. & Mayol, J. F. Cytometric assessment of mitochondria using fluorescent probes. Cytometry A, https://doi.org/10.1002/cyto.a.21061 (2011).
Rodgers, J. T. et al. mTORC1 controls the adaptive transition of quiescent stem cells from G0 to G(Alert). Nature, https://doi.org/10.1038/nature13255 (2014).
Ryall, J. G. Metabolic reprogramming as a novel regulator of skeletal muscle development and regeneration. FEBS J, https://doi.org/10.1111/febs.12189 (2013).
Rodgers, J. T., Schroeder, M. D., Ma, C. & Rando, T. A. HGFA Is an Injury-Regulated Systemic Factor that Induces the Transition of Stem Cells into GAlert. Cell reports, https://doi.org/10.1016/j.celrep.2017.03.066 (2017).
Sin, J. et al. Mitophagy is required for mitochondrial biogenesis and myogenic differentiation of C2C12 myoblasts. Autophagy, https://doi.org/10.1080/15548627.2015.1115172 (2016).
Puigserver, P. & Spiegelman, B. M. Peroxisome proliferator-activated receptor-gamma coactivator 1 alpha (PGC-1 alpha): transcriptional coactivator and metabolic regulator. Endocr Rev, https://doi.org/10.1210/er.2002-0012 (2003).
Scarpulla, R. C. Metabolic control of mitochondrial biogenesis through the PGC-1 family regulatory network. Biochimica et biophysica acta, https://doi.org/10.1016/j.bbamcr.2010.09.019 (2011).
Nicolas, N., Marazzi, G., Kelley, K. & Sassoon, D. Embryonic deregulation of muscle stress signaling pathways leads to altered postnatal stem cell behavior and a failure in postnatal muscle growth. Developmental biology, https://doi.org/10.1016/j.ydbio.2005.02.022 (2005).
Kim, J. et al. Imprinting control region (ICR) of the Peg3 domain. Human molecular genetics, https://doi.org/10.1093/hmg/dds092 (2012).
Kelly, A. M. & Zacks, S. I. The histogenesis of rat intercostal muscle. J Cell Biol (1969).
Bentzinger, C. F., Wang, Y. X. & Rudnicki, M.A. Building muscle: molecular regulation of myogenesis. Cold Spring Harbor perspectives in biology, https://doi.org/10.1101/cshperspect.a008342 (2012).
Cockett, N. E. et al. Polar overdominance at the ovine callipyge locus. Science (1996).
Georgiades, P., Watkins, M., Surani, M. A. & Ferguson-Smith, A.C. Parental origin-specific developmental defects in mice with uniparental disomy for chromosome 12. Development (2000).
Holt, L. J. et al. Grb10 regulates the development of fiber number in skeletal muscle. FASEB journal: official publication of the Federation of American Societies for Experimental Biology, https://doi.org/10.1096/fj.11-199349 (2012).
Martinet, C. et al. H19 controls reactivation of the imprinted gene network during muscle regeneration. Development, https://doi.org/10.1242/dev.131771 (2016).
Hiramuki, Y., Sato, T., Furuta, Y., Surani, M.A. o Sehara-Fujisawa, A. Mest but Not MiR-335 Affects Skeletal Muscle Growth and Regeneration. PloS one, https://doi.org/10.1371/journal.pone.0130436 (2015).
Waddell, J. N. et al. Dlk1 is necessary for proper skeletal muscle development and regeneration. PLoS One, https://doi.org/10.1371/journal.pone.0015055 (2010).
Vallecillo García, P. et al. Odd skipped-related 1 identifies a population of embryonic fibro-adipogenic progenitors regulating myogenesis during limb development. Nature Communications, in press (2017).
da Rocha, S. T. et al. Gene dosage effects of the imprinted delta-like homologue 1 (dlk1/pref1) in development: implications for the evolution of imprinting. PLoS Genet, https://doi.org/10.1371/journal.pgen.1000392 (2009).
Perera, B.P. & Kim, J. Alternative promoters of Peg3 with maternal specificity. Scientific reports, https://doi.org/10.1038/srep24438 (2016).
Plasschaert, R. N. & Bartolomei, M. S. Genomic imprinting in development, growth, behavior and stem cells. Development. https://doi.org/10.1242/dev.101428 (2014).
Berg, J. S. et al. Imprinted genes that regulate early mammalian growth are coexpressed in somatic stem cells. PloS one, https://doi.org/10.1371/journal.pone.0026410 (2011).
Zacharek, S. J. et al. Lung stem cell self-renewal relies on BMI1-dependent control of expression at imprinted loci. Cell stem cell, https://doi.org/10.1016/j.stem.2011.07.007 (2011).
Venkatraman, A. et al. Maternal imprinting at the H19-Igf2 locus maintains adult haematopoietic stem cell quiescence. Nature, https://doi.org/10.1038/nature12303 (2013).
Matsumoto, A. et al. p57 is required for quiescence and maintenance of adult hematopoietic stem cells. Cell stem cell, https://doi.org/10.1016/j.stem.2011.06.014 (2011).
Besson, V. et al. Expression Analysis of the Stem Cell Marker Pw1/Peg3 Reveals a CD34 Negative Progenitor Population in the Hair Follicle. Stem Cells, https://doi.org/10.1002/stem.2540 (2017).
Bonfanti, C. et al. PW1/Peg3 expression regulates key properties that determine mesoangioblast stem cell competence. Nat Commun, https://doi.org/10.1038/ncomms7364 (2015).
Stumm, J. et al. Odd skipped-related 1 (Osr1) identifies muscle-interstitial fibro-adipogenic progenitors (FAPs) activated by acute injury. Stem Cell Research, https://doi.org/10.1016/j.scr.2018.08.010 (2018).
Charge, S. B. & Rudnicki, M. A. Cellular and molecular regulation of muscle regeneration. Physiological reviews, https://doi.org/10.1152/physrev.00019.2003 (2004).
Kuang, S. & Rudnicki, M. A. The emerging biology of satellite cells and their therapeutic potential. Trends in molecular medicine, https://doi.org/10.1016/j.molmed.2007.12.004 (2008).
Kuang, S., Gillespie, M. A. & Rudnicki, M. A. Niche regulation of muscle satellite cell self-renewal and differentiation. Cell stem cell, https://doi.org/10.1016/j.stem.2007.12.012 (2008).
Liu, W. et al. Hypoxia promotes satellite cell self-renewal and enhances the efficiency of myoblast transplantation. Development, https://doi.org/10.1242/dev.079665 (2012).
Urciuolo, A. et al. Collagen VI regulates satellite cell self-renewal and muscle regeneration. Nature communications, https://doi.org/10.1038/ncomms2964 (2013).
Yin, H., Price, F. & Rudnicki, M. A. Satellite cells and the muscle stem cell niche. Physiological reviews, https://doi.org/10.1152/physrev.00043.2011 (2013).
Sciorati, C., Clementi, E., Manfredi, A. A. & Rovere-Querini, P. Fat deposition and accumulation in the damaged and inflamed skeletal muscle: cellular and molecular players. Cellular and molecular life sciences: CMLS, https://doi.org/10.1007/s00018-015-1857-7 (2015).
Deng, Y. & Wu, X. Peg3/Pw1 promotes p53-mediated apoptosis by inducing Bax translocation from cytosol to mitochondria. Proceedings of the National Academy of Sciences of the United States of America, https://doi.org/10.1073/pnas.97.22.12050 (2000).
Theka, I., Sottile, F., Aulicino, F., Garcia, A. C. & Cosma, M.P. Reduced expression of Paternally Expressed Gene-3 enhances somatic cell reprogramming through mitochondrial activity perturbation. Sci Rep, https://doi.org/10.1038/s41598-017-10016-7 (2017).
Lee, S., Ye, A. & Kim, J. DNA-Binding Motif of the Imprinted Transcription Factor PEG3. PloS one, https://doi.org/10.1371/journal.pone.0145531 (2015).
Wanet, A., Arnould, T., Najimi, M. & Renard, P. Connecting Mitochondria, Metabolism, and Stem Cell Fate. Stem Cells Dev, https://doi.org/10.1089/scd.2015.0117 (2015).
Theret, M. et al. AMPKalpha1-LDH pathway regulates muscle stem cell self-renewal by controlling metabolic homeostasis. The EMBO journal, https://doi.org/10.15252/embj.201695273 (2017).
Lepper, C., Conway, S. J. & Fan, C. M. Adult satellite cells and embryonic muscle progenitors have distinct genetic requirements. Nature, https://doi.org/10.1038/nature08209 (2009).
Lopez-De Leon, A. & Rojkind, M. A simple micromethod for collagen and total protein determination in formalin-fixed paraffin-embedded sections. J Histochem Cytochem (1985).
Koopman, R., Schaart, G. & Hesselink, M. K. Optimisation of oil red O staining permits combination with immunofluorescence and automated quantification of lipids. Histochem Cell Biol (2001).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics, https://doi.org/10.1093/bioinformatics/btp120 (2009).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics, https://doi.org/10.1093/bioinformatics/btu638 (2015).
Acknowledgements
We thank C. Blanc and B. Hoareau (Flow Cytometry Core CyPS. Pierre et Marie Curie University, Paris, France) for FACS assistance. We thank animal facility platform of Pierre et Marie Curie University for their help handling mouse lines. We thank C. Proux, R. Legendre and H. Varet (Institut Pasteur, Transcriptome and EpiGenome, BioMics, Center for Innovation and Technological Research, F-75015, Paris, France) for library preparation and high-throughput sequencing. RMC and BTA were supported by INGENIUM grant (FP7 MC-ITN INGENIUM project no. 290123). DO and MV were supported by funding from REVIVE. Lastly, we thank the Brack laboratory (UCSF) for their comments and critiques of this work prior to submission.
Author information
Authors and Affiliations
Contributions
All authors designed experiments, analyzed and interpreted data. R.M.C., D.O., M.V., A.M. and B.T.A. performed experiments. R.M.C., G.M. and D.A.S. prepared the manuscript with the assistance from A.F.S.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Correra, R.M., Ollitrault, D., Valente, M. et al. The imprinted gene Pw1/Peg3 regulates skeletal muscle growth, satellite cell metabolic state, and self-renewal. Sci Rep 8, 14649 (2018). https://doi.org/10.1038/s41598-018-32941-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-32941-x
Keywords
This article is cited by
-
Effects of paternal exposure to cigarette smoke on sperm DNA methylation and long-term metabolic syndrome in offspring
Epigenetics & Chromatin (2022)
-
Hypoxia promotes a perinatal-like progenitor state in the adult murine epicardium
Scientific Reports (2022)
-
MiR-200a-3p Aggravates DOX-Induced Cardiotoxicity by Targeting PEG3 Through SIRT1/NF-κB Signal Pathway
Cardiovascular Toxicology (2021)
-
Mouse CD146+ muscle interstitial progenitor cells differ from satellite cells and present myogenic potential
Stem Cell Research & Therapy (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.