Abstract
Utilizing the publicly available neuroimaging database enabled by Alzheimer’s disease Neuroimaging Initiative (ADNI; http://adni.loni.usc.edu/), we have compared the performance of automated classification algorithms that differentiate AD vs. normal subjects using Positron Emission Tomography (PET) with fluorodeoxyglucose (FDG). General linear model, scaled subprofile modeling and support vector machines were examined. Among the tested classification methods, support vector machine with Iterative Single Data Algorithm produced the best performance, i.e., sensitivity (0.84) × specificity (0.95), by 10-fold cross-validation. We have applied the same classification algorithm to four different datasets from ADNI, Health Science Centre (Winnipeg, Canada), Dong-A University Hospital (Busan, S. Korea) and Asan Medical Centre (Seoul, S. Korea). Our data analyses confirmed that the support vector machine with Iterative Single Data Algorithm showed the best performance in prediction of future development of AD from the prodromal stage (mild cognitive impairment), and that it was also sensitive to other types of dementia such as Parkinson’s Disease Dementia and Dementia with Lewy Bodies, and that perfusion imaging using single photon emission computed tomography may achieve a similar accuracy to that of FDG-PET.
Similar content being viewed by others
Introduction
Alzheimer’s disease (AD) is the most prevalent form of age-related dementia. Currently, and based on clinical and methodological consensus, fluorodeoxyglucose (FDG) positron emission tomography (PET) is the most widely used brain imaging modality that complements clinical diagnosis of AD1. AD patients exhibit declines in FDG uptake within the parietotemporal, frontal and posterior cingulate cortices when compared to normal age-matched controls. There exists, however, significant individual variation in patient PET signal expression which limits the use of a definitive threshold for FDG-PET in AD diagnosis. Therefore, clinical reading of FDG-PET images is usually based on subjective impressions of the relative hypo-metabolism in key anatomical brain regions. Consequently, no quantifiable biomarker-related numeric value is produced and, as a result, it is often concluded that reports are equivocal. In addition, the sensitivity of subjective readings at an early disease stage remains challenging, while relatively high accuracy can be achieved for more advanced stages1.
Alternative diagnostic approaches include the use of a PET radiotracer that binds to AD pathology-specific proteins such as beta-amyloid (Aβ) and phosphorylated tau. Since their invention, Pittsburgh Compound-B (PiB)2 and other second-generation Aβ radioligands have been heavily researched within the field of molecular imaging, particularly over the past decade. Meta-analysis on longitudinal PiB studies estimates 94.7% sensitivity but 57.2% specificity3 which implies high false positive rates of Aβ-PET. This confounding factor is inevitable since it has been estimated that approximately 20–30% of cognitively normal subjects exhibit abnormally high levels of high Aβ protein4. For this reason, the purpose of the recent on-going IDEAS study (Imaging Dementia-Evidence For Amyloid Scanning; http://www.ideas-study.org), in which the benefit of Aβ-PET will be evaluated for Medicare coverage, had to be limited to assessing Aβ status rather than AD status. A number of groups are currently investigating tau-imaging agents with promising results in some instances, but more rigorous testing needs to be done in upcoming years for these radioligands to be proven to be of any notable clinical significance that can be translated into clinical utility in a healthcare setting5.
In a combined effort involving multiple research centers in North America, the AD Neuroimaging initiative (ADNI) has collected the largest available multi-modal longitudinal brain imaging database for patients with AD, MCI and normal control subjects (http://www.adni-info.org). Utilizing the ADNI database, a number of studies constructed quantitative biomarkers that may aid AD diagnosis6,7,8,9,10,11,12. One of the limitations of ADNI-based studies are the lack of other types of dementia. For example, it is unknown if the above-mentioned AD-related biomarkers generated from ADNI’s FDG-PET database are only unique to AD or it is commonly expressed in patients with dementia with Lewy body (DLB), the most frequently misdiagnosed diseases as AD.
In this study, we have constructed machine-based imaging biomarkers using ADNI’s FDG-PET dataset. We used general linear model (GLM), subprofile modeling (SSM)13, and support vector machine (SVM). GLM is the most fundamental method for constructing a prediction model that classifies data. SSM is a form of principal component analysis (PCA), and it has proven to be useful for differential diagnosis and prognosis in other neurodegenerative disorders14,15,16. SVM is one of the most widely used machine learning algorithms that are used to solve binary classification problems with a large dataset17. We have compared the performance of these machine-based classification methods between AD and age-matched normal subjects, evaluated if they predict AD conversion from the prodromal state, examined if they are sensitive to other types of dementia such as DLB and Parkinson’s disease dementia (PDD).
Results
Performance Comparisons of different classification machines
From the ADNI database, we identified 94 patients with AD and 111 age-matched control subjects who stayed cognitively healthy for >3 years after the FDG-PET scans (Table 1). We have investigated the classification performance of three different approaches; voxel-wise GLM (Fig. 1A), SSM/PCA and SVM. Two different approaches of SSM/PCA were utilized13; 1) selecting the single principal component (PC) that best discriminates the groups (SSM/PCA1); and 2) linear combination of PCs that significantly improved the model by stepwise regression (SSM/PCA2). In SSM/PCA1, the first PC (PC1) discriminated the two groups most significantly (Fig. 1B). In SSM/PCA2, 7 PCs were linearly combined (PC1-5, PC7 and PC10) with the biggest coefficient being PC1 (Fig. 1C). In SVM analysis, two different solvers were used: (1) Iterative Single Data Algorithm (ISDA18; Fig. 1D) and (2) sequential minimal optimization (SMO19; Fig. 1E).
Visualization of classification machines. (A) Beta coefficient of voxel-wise linear regression (left). Subject scores are defined as the dot-product between the Beta coefficient map and the subject’s proportionally scaled FDG-PET images. The subject scores are z-scored to the mean and standard deviation of normal subjects (middle). Receiver-operating-characteristic (ROC) curve analysis is performed to estimate the area-under-the-curve (AUC; 0.922). The optimal subject score threshold for group classification is determined (z = 0.987) at the biggest sensitivity (0.862) × specificity (0.856). (B) In SSM/PCA1, the topography was similar to the GLM’s beta-coefficient map (left). The subject scores of the first PC (middle) produced the greatest separation of the groups (z-threshold = 0.771, AUC = 0.852, sensitivity = 0.787, specificity = 0.802; right). (C) The combined PCs of SSM/PCA2 was topographically similar to both GLM’s beta coefficient map and the first PC (left). The PCs were weighted by stepwise regression coefficients that significantly improved the model (p < 0.05). The resulting combined PC was 4.204 × PC1 + 0.865 × PC2 + 1.126 × PC3 – 1.825 × PC4 + 1.470 × PC5 + 1.099 × PC7 + 1.118 × PC10. The group separation by subject scores (middle) were relatively better than SSM/PCA1 of the first PC (z-threshold = 1.101, AUC = 0.911, sensitivity = 0.819, specificity = 0.865; right). (D) The hyperplane constructed by SVM-ISDA was topographically distinct from GLM or SSM/PCA (left). While complete group separation was achieved (AUC = 1; right), 3 of 111 NL subjects were labeled as AD (middle). It should be noted, in SVM’s case, that no score threshold is determined by ROC curve, but the sign of dot-product between hyperplane and subjects FDG-PET image (proportionally scaled) determines the labelling. (E) The hyperplane constructed by SVM-SMO (left) was topographically similar to the SVM-ISDA’s. Complete group separation was achieved (middle/right). For visualization purposes, all images are z-scored to the mean and standard deviation of all the voxel values of the whole-brain.
Qualitatively, the produced patterns consistently show a well-characterized AD-like distribution of hypometabolism20 present within the precuneus, medial frontal lobes, the temporal lobes, and the cingulum (posterior and anterior). Relative hypermetabolism was observed in the somatosensory-motor areas, basal ganglia, thalamus and cerebellum. Our findings echo those observed in the previously identified AD-related metabolic patterns21. Quantitatively, the whole-brain metabolic patterns from GLM and SSM/PCA were topographically similar to each other (p < 0.01, corrected for autocorrelation22 and multiple comparisons), but the similarities were not significant when compared to the SVM-produced patterns (p > 0.1 corrected for autocorrelation22 and multiple comparisons; Table 2).
As expected, all five approaches reliably discriminated patients and controls (sensitivity > 0.78, specificity > 0.80). When compared with 10-fold cross-validation (Fig. 2), the subject scores of the first three methods were almost identical to the original scores shown in Fig. 1 in AD (GLM: r = 0.998, r < 0.001; SSM/PCA1: r = 0.986, p < 0.001; SSM/PCA2: r = 0.990, p < 0.001) as well as in NL (GLM: r = 0.998, r < 0.001; SSM/PCA1: r = 0.992, p < 0.001; SSM/PCA2: r = 0.994, p < 0.001). While strongly correlated, lesser agreement was observed between original subjects scores and the ones from 10-fold cross-validation in SVM methods in both AD (SVM-ISDA: r = 0.860, p < 0.001; SVM-SMO: r = 0.875, p < 0.001) and in NL (SVM-ISDA: r = 0.648, p < 0.001; SVM-SMO: r = 0.739, p < 0.001), suggesting some overfitting in SVM. Nevertheless, the area-under-the-curve (AUC) for receiver-operating-characteristic (ROC) plot was the greatest by SVM-ISDA by 10-fold cross-validation (Fig. 2F; Table 3). Utilizing the structural MRI in the spatial normalization during the pre-processing did not improve the classification performance23 (Supplementary Table S1).
Relationship with other risk factors
In normal control subjects, age was significantly correlated with subject scores determined by GLM (r = 0.350, p < 0.001, Pearson’s correlation; Fig. 3A), SSM/PCA1 (r = 0.370, p < 0.001) and SSM/PCA2 (r = 0.367, p < 0.001). Since SVM outcome is dichotomous, student t-test was performed for age between subjects who were classified as AD vs. non-AD within normal control (SVM classified AD vs. NL in 100% accuracy in the training set thus the AD-designation determined as a testing set in 10-fold cross-validation was used). Age was not significantly associated with SVM’s designation of AD (p > 0.1, student t-test). The subject scores determined by GLM, SSM/PCA1 and SSM/PCA2 were not correlated with any of the other variables (MMSE, Aβ, tau and phosphorylated tau; p > 0.12, Pearson’s correlation) and they were not significantly different between sexes (p > 0.09, student t-test) or the presence of APOE ɛ4 genotype (p > 0.15, student t-test; Fig. 4A-left). These variables were not associated with SVM’s AD designation, either (p > 0.11, student t-test or chi-square; Fig. 4B-left).
Effects of age. (A) In NL, age was significantly correlated with subject scores determined by GLM method (r = 0.350, p < 0.001). In AD, no correlation was observed (r = 0.113, p = 0.279). Similar level of correlations were also observed only in NL when SSM/PCA1 (r = 0.370, p < 0.001) and SSM/PCA2 (r = 0.367, p < 0.001) were used. (B) Likewise, similar level of correlations between subject scores and age were observed only in stable MCI (GLM: r = 0.387, p < 0.001; SSM/PCA1: r = 0.329, p < 0.001; SSM/PCA2: r = 0.401, p < 0.001) but not in prodromal AD (r < 0.28). (C) In stable MCI, patients who were designated as “AD” (n = 45 out of 141) by SVM-ISDA were significantly older than patients who were designated as “non-AD” (*t(184) = 2.762, p = 0.006). Similar trend was observed when SVM-SMO was used (t(184) = 1.767, p = 0.079). This age difference associated with “AD”-designation was not observed in prodromal AD patients in both SVM methods (p > 0.48).
Effects of APOE mutation. (A) When GLM was used, no significant effects of APOE genotype in subject scores were observed in any groups (p > 0.08, student t-test), which was replicated when SSM/PCA was used (p > 0.06). (B) The proportion of subjects classified as “AD” vs. “non-AD” by SVM-ISDA was significantly different between APOE ɛ4 carriers vs. non-carriers only in stable MCI patients (χ2 = 7.202, p = 0.007, Pearson Chi-Square). In other words, if a patient with stable MCI has ɛ4, it is more likely that the patient will be classified as “AD”. From the 118 stable MCI non-ɛ4 carriers, 21 patients (17.8%) were classified as AD, in contrast to 24 of 68 stable MCI ɛ4 carriers (35.3%) were classified as AD. Similar results were produced when SVM-SMO was used in stable MCI patients (χ2 = 10.690, p = 0.001). In contrast, 8–10% normal control subjects received AD-designation regardless of APOE status. In AD, 86–88% individuals received AD-designation regardless of APOE status. In prodromal AD, 61–72% individuals received AD-designation regardless of APOE status.
The same analysis was repeated in AD, and no significant association was found between FDG-PET-based (subject scores and AD-designation) and other variables (age, MMSE, Aβ, tau and phosphorylated tau, sex, APOE genotype; p > 0.19). No significant effect of anti-AD medication status was observed in subject scores determined by GLM, SSM/PCA1 and SSM/PCA2 (p > 0.83, student t-test) or AD-designation by SVM-ISDA and SVM-SMO (p > 0.86, chi-square).
Prediction of AD conversion at MCI stage
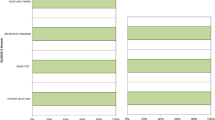
When the same machines were applied to predict future development of AD from 241 MCI patients (55 of whom developed AD and 186 of whom did not for the next 3 years; Table 1 - ADNI), a moderate sensitivity (0.563–0.673) and specificity (0.720–0.796) was achieved across all five different approaches (Fig. 5-left), which was comparable to prior studies7,24.
Prediction of future development of AD. Five different classification machines were applied to predict future development of AD from MCI patients who have been followed for >3 years. In the ADNI cohort, consistent group classification performance was achieved across 5 different approaches (left). When the perfusion SPECT data was used, GLM, SSM/PCA1 and SSM/PCA2 classified both prodromal AD and stable MCI patients as AD. Desirable performance (sensitivity × specificity > 0.5) was achieved in both SVM methods (right). It should be noted that the AD diagnosis was based on clinical examination, thus misdiagnosis cannot be ruled out, and may have contributed to the low sensitivity. It should be also noted that the limited follow-up period may have hindered the specificity of the method, i.e., if a longer follow-up was available, some of the stable MCI patients who have been classified as AD may have actually developed AD.
As in NL, age was correlated with subject scores (or AD-designation by SVM) in stable MCI patients (p < 0.001; Pearson’s correlation for GLM and SSM/PCA and student t-test for SVM; Fig. 3B,C), while Aβ, tau or phosphorylated tau concentrations were not associated with subject scores (p > 0.2, Pearson’s correlation for GLM and SSM/PCA) and AD-designation (p > 0.11, student t-test). APOE genotype was not associated with subject scores determined by GLM or SSM/PCA (p > 0.16, student t-test), but significantly more proportion of stable MCI patients had APOE ɛ4 allele when they were designates as AD than non-AD by both SVM methods (p < 0.007, chi-square; Fig. 4B).
In prodromal AD, age was very weakly correlated with GLM score (r = 0.271, p = 0.046; Fig. 3B) but not with SSM/PCA’s (p > 0.07). These subject scores were correlated with Aβ (r = −0.720, p = 0.019), tau (r = −0.595, p = 0.070) and phosphorylated tau (r = −0.749, p = 0.013) concentrations. Here, it should be noted that only 10 out of 55 prodromal AD patients had CSF measurements. As in AD, no significant difference was observed between the AD-designated patients and non-AD-designated patients in age, MMSE, Aβ, tau, phosphorylated tau (p > 0.075, student t-test), or APOE genotype (p > 0.3, chi-square), by either SVM methods.
Interestingly, when applied to the perfusion SPECT images from 38 patients with MCI (8 of whom later developed AD; 3-year follow-up; data from Dong-A University, Busan, S. Korea; Table 4), all patients (both stable MCI and prodromal AD) were classified as AD in GLM and SSM/PCA (Fig. 5-right). When SVM-ISDA was used, a higher level of sensitivity (0.875) was achieved but specificity was slightly poorer (0.600). When SVM-SMO was used, a similar level of sensitivity (0.625) and higher specificity (0.900) to the ADNI data was achieved. This result suggests a possibility that perfusion imaging may be as useful as FDG-PET for complementing AD diagnosis when analyzed by an advanced machine learning approach25.
Clinical Applications
In order to demonstrate the applicability of our proposed classification methods in a real clinical setting, a retrospective imaging study was conducted. In our picture archiving and communication system (PACS) at Health Science Centre (Winnipeg, Manitoba), we have identified 113 patients who referred to brain FDG-PET between 2010 and 2012 (Table 5). All patients were referred from our local memory clinic due to subjective memory complaints or MCI. Among these patients, 37 patients were subsequently diagnosed with AD, 19 patients with DLB or PDD, 4 patients with frontotemporal dementia (FTD; 1 patient excluded due to image quality issue), and 5 patients with vascular dementia (VaD). 21 patients received other diagnoses during the follow-up period. 18 patients did not develop any forms of dementia for >6 months, and 6 patients did not return for follow-up (thus excluded from our analysis). All five different approaches (GLM, SSM/PCA1, SSM/PCA2, SVM-ISDA and SVM-SMO) accurately classified the majority of patients who were later diagnosed with AD as AD (28–32 of 37; Fig. 6). The majority of patients who did not develop any form of dementia for >6 months were accurately classified as non-AD (13–16 of 22 stable MCI patients). Interestingly, most patients (16–17 of 19) who later diagnosed with DLB or PDD were also classified as AD, suggesting non-specificity of our method in different types of dementia, especially in alphasynucleinopathy-related dementia (PDD and DLB). The number of patients with FTD (n = 3) and VaD (n = 5) were very small, thus no formal conclusion can be drawn from this data. Nevertheless, more frequent “AD-designation” was observed in patients who were later diagnosed with VaD (3–4 of 5).
Patient classification performance in a real clinical setting. Four out of five approaches (GLM, SSM/PCA1, SVM-ISDA and SVM-SMO) achieved desirable performance (sensitivity × specificity >0.5) in classifying prodromal AD vs. stable MCI. Interestingly, >84% of DLB/PDD patients were also classified as “AD”, while low sample size limits definite conclusion on the cases of FTD and VaD.
AD-related metabolic pattern in DLB and PDD
In order to investigate the comorbid expression of an AD-like brain metabolic pattern in Parkinson’s disease (PD), we revisited the FDG-PET images from Asan Medical Centre (Seoul, S. Korea; Table 6)26. When SVM was used, none of the healthy control subjects (n = 18) were classified as AD, and only 2 of 17 non-demented PD (PDND) patients were classified as AD (Fig. 7). This rate was slightly increased in PD patients with MCI (7–8 of 26), while the majority of PDD (17 of 19) and DLB (14–15 of 19) patients were classified as AD (Fig. 4). The other methods (GLM, SSM/PCA1 and SSM/PCA2) were generally more sensitive but false positive identification from normal control subjects (1–6 of 18) was also more frequent (Fig. 7). Together with the above findings (Fig. 6), this result suggests that both PDD and DLB patients exhibit AD-like brain metabolic patterns while cognitively healthy PD patients do not (when examined by SVM).
AD-like glucose metabolic pattern expression in alphasynucleinopathy. As can be expected from Fig. 3, the majority of PDD (17 of 19) and DLB (14–15 of 19) patients were classified as AD, when analyzed by the SVM method. However the majority of non-demented PD (PDND; 15 of 17)) or PD patients with MCI (PD-MCI; 18–19 of 26) were not classified as AD. None of the healthy control subjects were classified as AD when SVM was used. When other methods (GLM and SSM/PCA) were used, the rate of AD designation was significantly higher in PD-MCI, PDND and NL, potentially suggesting false positive identification.
Discussion
When FDG-PET was used, the best classification performance (sensitivity × specificity) was achieved by the SVM-ISDA in all three cases (AD vs. NL from ADNI by 10-fold cross-validation; stable MCI vs. prodromal AD from ADNI; stable MCI vs. prodromal AD from Winnipeg) among the five different classification methods (Table 7). Thus, we recommend the use of SVM-ISDA in FDG-PET readings to complement AD diagnosis.
Of note, the GLM and SSM/PCA produced topographically similar metabolic patterns. This is not unexpected when a large training set was used (Eidelberg, personal communication). The region weights, in both GLM and SSM/PCA, are indicators of overall metabolic topography that differentiates AD from NL. In other words, the subject scores are computed as a dot-product between patients’ normalized FDG-PET (or perfusion SPECT) images and the metabolic “template” images (Fig. 1A–C). Thus, it quantitates to what degree subjects’ images are “look-a-likes” of those template images. And, this gives one a rationale to use it as a quantifiable biomarker to correlate with disease progression, symptom severity or treatment responses27. Indeed, the subject scores determined by GLM and SSM/PCA were significantly correlated with age in normal subjects and stable MCI, consistent with old age being one of the most prominent risk factor for AD.
In SVM, higher weights are assigned to the hyperplanes (Fig. 1D,E) if the value of the voxel (feature) is useful for group separation. In other words, the “importance” of each feature (i.e., voxel) is weighted in the hyperplane. And, the subject score itself is computed in the same manner as in GLM and SSM/PCA, i.e., dot-product with the subjects’ normalized images and the hyperplane. The difference is that only the sign of the subject score is used for group classification. Thus, the absolute size of the subject scores does not convey the same level of information (e.g., age or disease status) as the GLM and SSM/PCA does, but it only bears information about the dichotomic class designation. In this regard, it is not surprising that SVM-produced subject scores were not associated with age while GLM and SSM/PCA-produced subjects scores were. This fundamental difference resulted in a spatially less smooth representation of AD-related metabolic pattern, and dissimilar topographical profile from GLM and SSM/PCA (Table 2). Nevertheless, qualitatively, all five approaches were able to pinpoint relative hypometabolism in key brain regions commonly effected by AD: the occipito-parietal, frontal and temporal lobes and the cingulum28.
The robust performance of SVM-ISDA classification of prodromal AD vs. stable MCI patients from three different datasets (Figs 5 and 6), including the data archived from our community-based memory clinic demonstrates its immediate translational value. Interestingly, most patients with other forms of dementia (i.e., clinically determined during the follow-up period) also show positive AD designation. The DLB/PDD patients, especially, showed even higher proportion of positive results than AD patients. This was not surprising since DLB/PDD and AD often present with very similar clinical symptoms and neuropathology. Amyloid deposits and tau protein accumulation are commonly observed in patients with DLB, and it has been well established that these AD-related pathologies are also linked to the cognitive deficits that occur in DLB patients29. The similar neurodegenerative etiology of the two is also often reflected in FDG-PET scans30. The hypometabolic pattern observed in DLB is highly congruent with and can be easily superimposed onto that observed in AD patients with the only region exhibiting variation between the two being the occipital lobe31. The clinical overlap makes the two diseases fairly difficult to distinguish and accurate differential diagnosis between AD and DLB remains a popular arena of neurodegenerative disease research, and it is warranted that an additional step be taken for differential diagnosis, e.g., Parkinson’s disease-related metabolic pattern expression may be examined in these patients26. Other PET/SPECT radiotracers that target the dopaminergic32 system or tau33 have shown promising results for differentiating DLB from AD.
More interestingly, PDND and PD-MCI patients were rarely classified as “AD” by SVM-ISDA (Fig. 4). Taken together with the high rate of “AD-classification” of prodromal PDD/DLB patients (from HSC data), this result demonstrates a very important translational potential not only for predicting AD but also predicting PDD from non-demented PD patients. According to the dual syndrome hypothesis, PDD/DLB is more involved with posterior and temporal brain regions associated with visuospatial and attention deficits while non-progressing PD-MCI is more involved with fronto-subcortical circuits associated with executive function34,35,36,37. It is of interest that those visuospatial and attention deficits respond relatively well to the cholinergic treatment commonly used in AD36, prompting common pathology between AD and PDD/DLB. Nevertheless, there are autopsy-confirmed PDD patients without distinct AD-like pathology38. We have recently reported a brain metabolic pattern that was associated with cognitive decline in PDD/DLB, which was characterized by a hypometabolic caudate nucleus39. PDD/DLB patients also show further decreased caudate FP-CIT uptake (dopamine transporter availability) compared to the PDND patients26. Interactions between those two domains, AD-like vs. PDD/DLB-specific (e.g., involving the caudate nucleus) remain to be elucidated.
It is yet unknown why GLM and SSM/PCA designated more PDND and PD-MCI patients as AD (52–100%) while SVM didn’t (11–31%). One potential explanation is that the GLM and SSM/PCA-derived AD-related glucose metabolic patterns share some topographical property of “normal” aging-related metabolic pattern40,41 potentially due to the parallel processes between aging and AD (and other neurodegenerative disorders such as PD)42. Thus, it is possible that severer AD patients also showed higher aging-related changes, and their scans would also have contributed more to pattern topography. Of note, age was weakly but significantly correlated with GLM and SSM/PCA-derived pattern expression in normal control. On the other hand, SVM does not take the severity of the disease into consideration but instead focuses on the boundary between the two groups. The individuals who were used as support vectors are the ones who are adjacent to the marginal space. In other words, the hyperplane is formed based on the AD patients whose scans are better normal individual “look-a-likes”, as well as the normal individuals whose scans are better AD patient “look-a-likes”. Perhaps, the normal aging-related metabolic changes are similarly expressed in these two subpopulations, thus cancelling out when the hyperplane was defined. It must be noted that the SVM-ISDA is only designed for group separation and is therefore not designed to convey any information about disease status. For the purposes of monitoring the disease progression and/or treatment responses, GLM and SSM/PCA may be more suitable choices. Nevertheless, it is of interests that only the SVM’s AD designation was associated with APOE genotype in stable MCI, which is in agreement with the APOE ɛ4 status being one of the most significant risk factor for AD43, e.g., only 31 of 111 normal controls (28%) were ɛ4 carrier in contrast to 46 of 74 AD patients (61%).
When perfusion SPECT (from Dong-A University Hospital) was analyzed with SVM-ISDA, more prodromal AD patients were designated as AD (87%) than when FDG-PET (from ADNI) was analyzed (67%). This is in line with the recent observation that vascular changes precede metabolic changes44. However, it should be noted that there were also more false positives in perfusion SPECT, i.e., 40% of stable MCI were designated as AD in perfusion SPECT while only 24% were designated as AD in FDG-PET. Without longer follow-up duration and larger sample size, it is difficult to draw the conclusion that one method (perfusion SPECT) was superior to others (FDG-PET). It should be also noted that when SVM-SMO was used for analysis, perfusion SPECT’s sensitivity and specificity were similar to that of FDG-PET from the ADNI’s MCI patient database.
Materials and Methods
Data used in the preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). ADNI was launched in 2003 as a public-private partnership, led by Principal Investigator Michael W. Weiner, MD.
Subjects
A total of 763 subjects from the ADNI database who had available PET and MRI scans were included in this study (Supplementary Fig. S1). ADNI subjects came from one of three groups: AD, normal controls (NL), and MCI. MCI patients were further divided into stable MCI (who did not develop AD during the 3 years of follow-up period) and prodromal AD (who developed AD within 3 years). MCI patients who were not followed for more than 3 years were excluded. Demographic details regarding the ADNI cohort are given in Table 1.
To evaluate the utility of the proposed methods’ early diagnostic capacity using different imaging modalities, data from a total of 38 patients with MCI who underwent perfusion SPECT (TC-99m HMPAO) at Dong-A University Hospital in Busan, South Korea, then were subsequently followed for clinical diagnosis (>3 years) were included (Table 4). Patients were followed periodically over 1–2 years to monitor progression to dementia status. Among these patients, 8 patients were later diagnosed with AD in 19.0 ± 6.6 months after SPECT scan, and 30 patients were dementia-free for >3 years. Limited data has been published elsewhere45.
To demonstrate the feasibility of the proposed methods in a real clinical setting, a retrospective chart review study on a total of 113 patients was conducted. Subjects were patients of the Crescentwood Memory Clinic in Winnipeg, Manitoba, who were referred to the PET Centre at Health Science Centre (HSC) in Winnipeg, Manitoba, Canada, for scanning between 2010 and 2012 and subsequently diagnosed by a memory disorder specialist (PS) (Supplementary Fig. S2). HSC subjects were comprised of those who received a final diagnosis of MCI (n = 22), AD (n = 37), FTD (n = 4), DLB/PDD (n = 19) or VaD (n = 5) after having received a PET scan. Patients who were not followed for more than 6 months (n = 6) and who received other diagnoses (n = 21) were excluded. HSC cohort subject demographics are outlined in Table 5.
In order to test the performance of the proposed methods in other non-memory-oriented neurodegenerative disorders, data from a total of 97 subjects (18 NL, 17 PDND, 26 PD-MCI, 18 PDD, and 18 DLB) from a movement disorder clinic at Asan Medical Center in Seoul, South Korea were included (Table 6). All patients were scanned with FDG-PET, and their diagnoses were already well characterized at the time of scan. Limited data has been published elsewhere26,39.
All clinical and brain imaging data retrieval was completed in compliance with the Personal Health Information Act (PHIA), information regarding which can be found at http://www.gov.mb.ca/health/phia/. Ethics approval for this study was granted by the Biomedical Research Ethics Committee at the University of Manitoba, as well as institutional review boards at the Dong-A University Hospital and Asan Medical Center. Informed consent was obtained for all subjects who were enrolled in the study, either from the patient or from an individual designated as having power of attorney over the patient and able to make decisions regarding the patient’s health. All methods were performed in accordance with the relevant guidelines and regulations.
Image Acquisition
ADNI FDG-PET images were retrieved from the ADNI Laboratory of Neuroimaging (LONI) database in a format under which images had already been preprocessed through coregistration, averaging, and standardization. The detailed procedure can be found at http://adni.loni.usc.edu/methods/pet-analysis/pre-processing/.
DongA University’s SPECT images were acquired as described previously45. Patients were injected with 925 MBq of Tc-99m-HMPAO at resting state. Brain SPECT images were obtained from patients using a dual-head gamma camera (Multi-SPECT II, ICON, Siemens, USA) equipped with fan-beamcollimator.
HSC’s FDG-PET images were acquired using Siemens Biograph 16 HiRez PET/CT (Siemens Medical Solutions, Knoxville, TN) scanner. Patients were fasted for at least 6 hours before scanning. Patients were injected i.v. with 185 MBq of FDG and a 15-minute static image was acquired starting 40 minutes post-injection. A head CT scan was acquired for attenuation correction purposes.
Asan Medical Center’s FDG-PET images were acquired as described previously26,39. All subjects fasted for at least 6 h before scanning. A 5-min transmission scan using a 68Ge rotating pin source and a 15-min emission scan were acquired on the ECAT HR + scanner (Siemens Medical Systems, Hoffman Estate, IL, USA) at the Asan Medical Center, 40 min after intravenous injection of 370 MBq of FDG.
Image Preprocessing
All FDG-PET and perfusion SPECT image preprocessing was carried out using the standard procedure implemented in the statistical parametric mapping 12 (SPM) software (www.fil.ion.ucl.ac.uk/spm/) without the structural MRI (“old spatial normalization”). Images were normalized by warping to the MNI (Montreal Neurological Institute) standard space using a PET template available in SPM and then subsequently smoothed using an 8 mm Gaussian filter.
GLM-based classifier
Following image pre-processing, all images were proportionally scaled to the mean of all voxels within the whole-brain mask that is generated by relative thresholding at 25% of maximum as previously described13, which resulted in 189,629 voxels. Linear regression (y = β × x + c; where y is the observed image, β is the slope coefficient, x is the dummy variables for the group, and c is the constant) was performed for each voxel with the dummy variables that classifies the subject groups (AD vs. NL). The β map was saved (Fig. 1A), which was used to calculate the subject scores by dot-product with the proportionally scaled brain images. Thus, the subject score represents how similar the brain image is in terms of visual appearance to the β map.
SSM/PCA-based classifier
SSM/PCA was performed as described elsewhere13 using a customized MATLAB script. FDG-PET images of 111 NL and 94 AD subjects were log-transformed and double-centered. Singular value decomposition was performed to identify the top 10 PCs. The subject scores were calculated by dot-product between the scaled subprofile of each FDG-PET and PC. The PC that best discriminated the two groups was selected as the AD-related metabolic pattern for SSM/PCA1. For SSM/PCA2, stepwise regression was performed to select relevant PCs, then those PCs were linearly added with weights that were determined by the regression.
SVM-based classifier
The same proportionally scaled images were used as in GLM. SVM analysis was done using the fitcsvm function with linear kernelling implemented in the MATLAB Statistical Toolbox – Machine Learning. Assuming a 5% false classification in the training set (i.e., patients with other types of dementia who were misdiagnosed of AD; normal subjects who may develop AD beyond the 3-year follow-up period), the outlier fraction was set at 5%. Other parameters remained at default for both ISDA (no assumption in the initial estimates, alpha; misclassification cost = [0 1; 1 0]; tolerance for gradient difference = 0; Feasibility gap tolerance = 0; maximal number of numerical optimization iterations = 1,000,000; kernel offset parameter = 0.1; kernel scale = 1) and SMO (no assumption in the initial estimates, alpha; misclassification cost = [0 1; 1 0]; tolerance for gradient difference = 0.001; Feasibility gap tolerance = 0; maximal number of numerical optimization iterations = 1,000,000; kernel offset parameter = 0; kernel scale = 1).
Performance Evaluation and Statistical Analysis
The k-fold cross-validation (k = 10) was performed for all five methods (GLM, SSM/PCA1, SSM/PCA2, SVM-ISDA and SVM-SMO). For GLM and SSM/PCA, a receiver-operating curve (ROC) was examined to determine the optimal subject score threshold to produce the highest sensitivity and specificity in group discrimination. For SVM, the sign of the subject score was used to label each brain image as AD or non-AD. The performance characteristics of each classifier in group differentiation were evaluated by examining sensitivity × specificity.
The Student’s t-test was used to examine the group differences (AD vs. NL; stable MCI vs. prodromal MCI; across different neurodegenerative disorders) in age, MMSE, follow-up period, CSF markers (aβ, tau and phosphorylated tau), subject scores determined by GLM, SSM/PCA1 and SSM/PCA2. The subject scores (GLM and SSM/PCA) were compared by Student’s t-test between APOE ɛ4 carriers vs. non-carriers and male vs. female. These variables are correlated with Pearson’s correlation within each group. The differences is sex, APOE genotype and AD-designation (i.e., labels determined by SVM-ISDA and SVM-SMO) between groups are evaluated by chi-square test. All statistical analysis was performed using MATLAB R2014a and IBM SPSS Statistics version 24.
Data Availability Statement
The five models (Fig. 1) produced in the current study can be accessed via http://www.kolabneuro.com/. The raw data are not publicly available with the exception of ADNI data as they contain patient medical data which can only be accessed under the Personal Health Information Act (PHIA), information regarding which can be found at http://www.gov.mb.ca/health/phia/.
References
McConathy, J. & Sheline, Y. I. Imaging Biomarkers Associated With Cognitive Decline: A Review. Biol Psychiatry, https://doi.org/10.1016/j.biopsych.2014.08.024 (2014).
Klunk, W. E. et al. Imaging brain amyloid in Alzheimer’s disease with Pittsburgh Compound-B. Ann Neurol 55, 306–319, https://doi.org/10.1002/ana.20009 (2004).
Ma, Y. et al. Predictive accuracy of amyloid imaging for progression from mild cognitive impairment to Alzheimer disease with different lengths of follow-up: a systematic review. Medicine 93, e150, https://doi.org/10.1097/MD.0000000000000150 (2014).
Pontecorvo, M. J. & Mintun, M. A. PET amyloid imaging as a tool for early diagnosis and identifying patients at risk for progression to Alzheimer’s disease. Alzheimer’s research & therapy 3, 11, https://doi.org/10.1186/alzrt70 (2011).
Jagust, W. Time for tau. Brain 137, 1570–1571, https://doi.org/10.1093/brain/awu093 (2014).
Partovi, S. et al. Diagnostic performance of an automated analysis software for the diagnosis of Alzheimer’s dementia with (18)F FDG PET. Am J Nucl Med Mol Imaging 7, 12–23 (2017).
Shaffer, J. L. et al. Predicting cognitive decline in subjects at risk for Alzheimer disease by using combined cerebrospinal fluid, MR imaging, and PET biomarkers. Radiology 266, 583–591, https://doi.org/10.1148/radiol.12120010 (2013).
Presotto, L. et al. Validation of (18)F-FDG-PET Single-Subject Optimized SPM Procedure with Different PET Scanners. Neuroinformatics 15, 151–163, https://doi.org/10.1007/s12021-016-9322-9 (2017).
Chen, X. et al. Potential Clinical Value of Multiparametric PET in the Prediction of Alzheimer’s Disease Progression. PLoS One 11, e0154406, https://doi.org/10.1371/journal.pone.0154406 (2016).
Yamane, T. et al. Visual-statistical interpretation of (18)F-FDG-PET images for characteristic Alzheimer patterns in a multicenter study: inter-rater concordance and relationship to automated quantitative evaluation. AJNR. American journal of neuroradiology 35, 244–249, https://doi.org/10.3174/ajnr.A3665 (2014).
Dukart, J. et al. Meta-analysis based SVM classification enables accurate detection of Alzheimer’s disease across different clinical centers using FDG-PET and MRI. Psychiatry Res 212, 230–236, https://doi.org/10.1016/j.pscychresns.2012.04.007 (2013).
Zhang, D. & Shen, D., Alzheimer’s Disease Neuroimaging, I. Predicting future clinical changes of MCI patients using longitudinal and multimodal biomarkers. PLoS One 7, e33182, https://doi.org/10.1371/journal.pone.0033182 (2012).
Spetsieris, P. et al. Identification of Disease-related Spatial Covariance Patterns using Neuroimaging Data. J Vis Exp, https://doi.org/10.3791/50319 (2013).
Tang, C. C. et al. Differential diagnosis of parkinsonism: a metabolic imaging study using pattern analysis. Lancet Neurol 9, 149–158 (2010).
Holtbernd, F. et al. Abnormal metabolic network activity in REM sleep behavior disorder. Neurology. https://doi.org/10.1212/wnl.0000000000000130 (2014).
Niethammer, M. et al. A disease-specific metabolic brain network associated with corticobasal degeneration. Brain 137, 3036–3046, https://doi.org/10.1093/brain/awu256 (2014).
Cristianini, N. & Shawe-Taylor, J. xiii, 189 p. (Cambridge University Press, Cambridge; New York, 2000).
Kecman, V., Huang, T. & Vogt, M. In Support vector machines: theory and applications (ed Lipo Wang) 255–274 (Springer, 2005).
Fan, R. E., Chen, P. H. & Lin, C. J. Working set selection using second order information for training support vector machines. J Mach Learn Res 6, 1889–1918 (2005).
Mosconi, L. Brain glucose metabolism in the early and specific diagnosis of Alzheimer’s disease - FDG-PET studies in MCI and AD. European Journal of Nuclear Medicine and Molecular Imaging 32, 486–510, https://doi.org/10.1007/s00259-005-1762-7 (2005).
Mattis, P. J. et al. Distinct brain networks underlie cognitive dysfunction in Parkinson and Alzheimer diseases. Neurology 87, 1925–1933, https://doi.org/10.1212/WNL.0000000000003285 (2016).
Ko, J. H., Spetsieris, P., Ma, Y., Dhawan, V. & Eidelberg, D. Quantifying significance of topographical similarities of disease-related brain metabolic patterns. PLoS One 9, e88119, https://doi.org/10.1371/journal.pone.0088119 (2014).
Peng, S. et al. Characterization of disease-related covariance topographies with SSMPCA toolbox: Effects of spatial normalization and PET scanners. Hum Brain Mapp 35, 1801–1814, https://doi.org/10.1002/hbm.22295 (2014).
Cabral, C., Morgado, P. M., Campos Costa, D. & Silveira, M. & Alzheimers Disease Neuroimaging, I. Predicting conversion from MCI to AD with FDG-PET brain images at different prodromal stages. Comput Biol Med 58, 101–109, https://doi.org/10.1016/j.compbiomed.2015.01.003 (2015).
Collij, L. E. et al. Application of Machine Learning to Arterial Spin Labeling in Mild Cognitive Impairment and Alzheimer Disease. Radiology 281, 865–875, https://doi.org/10.1148/radiol.2016152703 (2016).
Ko, J. H., Lee, C. S. & Eidelberg, D. Metabolic network expression in parkinsonism: Clinical and dopaminergic correlations. J Cereb Blood Flow Metab 37, 683–693, https://doi.org/10.1177/0271678X16637880 (2017).
Eidelberg, D. Metabolic brain networks in neurodegenerative disorders: a functional imaging approach. Trends Neurosci 32, 548–557 (2009).
Burns, C. M. et al. Higher serum glucose levels are associated with cerebral hypometabolism in Alzheimer regions. Neurology 80, 1557–1564 (2013).
Ishii, K. PET Approaches for Diagnosis of Dementia. American Journal of Neuroradiology 35, 2030 (2014).
Kono, A. K. et al. Fully automatic differential diagnosis system for dementia with Lewy bodies and Alzheimer’s disease using FDG-PET and 3D-SSP. European Journal of Nuclear Medicine and Molecular Imaging 34, 1490–1497, https://doi.org/10.1007/s00259-007-0380-y (2007).
Chiba, Y. et al. Early differential diagnosis between Alzheimer’s disease and dementia with Lewy bodies: Comparison between F-18-FDG PET and I-123-IMP SPECT. Psychiatry Research-Neuroimaging 249, 105–112, https://doi.org/10.1016/j.pscychresns.2015.12.007 (2016).
Johnson, K. A. et al. Combined dopamine transporter and FDG PET IN DLB, AD, and PD. Neurobiology of Aging 25, S86–S87, https://doi.org/10.1016/s0197-4580(04)80293-8 (2004).
Brosch, J. R., Farlow, M. R., Risacher, S. L. & Apostolova, L. G. Tau Imaging in Alzheimer’s Disease Diagnosis and Clinical Trials. Neurotherapeutics 14, 62–68, https://doi.org/10.1007/s13311-016-0490-y (2017).
Rinne, J. O. et al. Cognitive impairment and the brain dopaminergic system in Parkinson disease: [18F]fluorodopa positron emission tomographic study. Arch Neurol 57, 470–475 (2000).
Brück, A. et al. Positron emission tomography shows that impaired frontal lobe functioning in Parkinson’s disease is related to dopaminergic hypofunction in the caudate nucleus. Neurosci Lett 311, 81–84, doi:S0304-3940(01)02124-3 (2001).
Kehagia, A. A., Barker, R. A. & Robbins, T. W. Cognitive impairment in Parkinson’s disease: the dual syndrome hypothesis. Neurodegener Dis 11, 79–92, https://doi.org/10.1159/000341998 (2013).
Nobili, F. et al. Cognitive-nigrostriatal relationships in de novo, drug-naive Parkinson’s disease patients: a [I-123]FP-CIT SPECT study. Mov Disord 25, 35–43, https://doi.org/10.1002/mds.22899 (2010).
Burack, M. A. et al. In vivo amyloid imaging in autopsy-confirmed Parkinson disease with dementia. Neurology 74, 77–84 (2010).
Ko, J. H. et al. Distinct brain metabolic patterns separately associated with cognition, motor function, and aging in Parkinson’s disease dementia. Neurobiol Aging 60, 81–91, https://doi.org/10.1016/j.neurobiolaging.2017.08.020 (2017).
Spetsieris, P. G. et al. Metabolic resting-state brain networks in health and disease. Proceedings of the National Academy of Sciences of the United States of America 112, 2563–2568, https://doi.org/10.1073/pnas.1411011112 (2015).
Moeller, J. R. et al. The Metabolic Topography of Normal Aging. Journal of Cerebral Blood Flow & Metabolism 16, 385–398, https://doi.org/10.1097/00004647-199605000-00005 (1996).
Udeochu, J. C., Shea, J. M. & Villeda, S. A. Microglia communication: Parallels between aging and Alzheimer’s disease. Clin Exp Neuroimmunol 7, 114–125, https://doi.org/10.1111/cen3.12307 (2016).
Bu, G. Apolipoprotein E and its receptors in Alzheimer’s disease: pathways, pathogenesis and therapy. Nat Rev Neurosci 10, 333–344, https://doi.org/10.1038/nrn2620 (2009).
Iturria-Medina, Y. et al. Early role of vascular dysregulation on late-onset Alzheimer’s disease based on multifactorial data-driven analysis. Nature Communications 7, 11934, https://doi.org/10.1038/ncomms11934 (2016).
Park, K. W. et al. Regional cerebral blood flow differences in patients with mild cognitive impairment between those who did and did not develop Alzheimer’s disease. Psychiatry Res 203, 201–206, https://doi.org/10.1016/j.pscychresns.2011.12.007 (2012).
Acknowledgements
Data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at: http://adni.loni.usc.edu/wp-content/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf. Financial support for this study was generously provided by The Dr. Paul H.T. Thorlakson Foundation, Brain Canada, Natural Sciences and Engineering Research Council of Canada, University of Manitoba and Alzheimer Society of Manitoba. We thank Erica Sheffield (Crescentwood Memory Clinic) for HSC data compiling. Data collection and sharing for part of this project was funded by the Alzheimer’s Disease Neuroimaging Initiative (ADNI) (National Institutes of Health Grant U01 AG024904) and DOD ADNI (Department of Defense award number W81XWH-12-2-0012). ADNI is funded by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, and through generous contributions from the following: AbbVie, Alzheimer’s Association; Alzheimer’s Drug Discovery Foundation; Araclon Biotech; BioClinica, Inc.; Biogen; Bristol-Myers Squibb Company; CereSpir, Inc.; Cogstate; Eisai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F. Hoffmann-La Roche Ltd and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd.; Janssen Alzheimer Immunotherapy Research & Development, LLC.; Johnson & Johnson Pharmaceutical Research & Development LLC.; Lumosity; Lundbeck; Merck & Co., Inc.; Meso Scale Diagnostics, LLC.; NeuroRx Research; Neurotrack Technologies; Novartis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; Takeda Pharmaceutical Company; and Transition Therapeutics. The Canadian Institutes of Health Research is providing funds to support ADNI clinical sites in Canada. Private sector contributions are facilitated by the Foundation for the National Institutes of Health (www.fnih.org). The grantee organization is the Northern California Institute for Research and Education, and the study is coordinated by the Alzheimer’s Therapeutic Research Institute at the University of Southern California. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of Southern California.
Author information
Authors and Affiliations
Contributions
A.K. collected, organized and analyzed the data and drafted the manuscript. P.S., A.L.G., D.L. and B.B. contributed to the conception of the research idea and collected the data. M.A. assisted the data analysis. H.Y., D.K., S.K. and C.L. collected and shared the data. J.K. conceived the research idea, analyzed the data and revised the manuscript.All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Katako, A., Shelton, P., Goertzen, A.L. et al. Machine learning identified an Alzheimer’s disease-related FDG-PET pattern which is also expressed in Lewy body dementia and Parkinson’s disease dementia. Sci Rep 8, 13236 (2018). https://doi.org/10.1038/s41598-018-31653-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-31653-6
This article is cited by
-
Radionuclide Imaging of the Neuroanatomical and Neurochemical Substrate of Cognitive Decline in Parkinson’s Disease
Nuclear Medicine and Molecular Imaging (2024)
-
Diagnostic performance of molecular imaging methods in predicting the progression from mild cognitive impairment to dementia: an updated systematic review
European Journal of Nuclear Medicine and Molecular Imaging (2024)
-
Functional brain networks in the evaluation of patients with neurodegenerative disorders
Nature Reviews Neurology (2023)
-
Machine Learning for Dementia Prediction: A Systematic Review and Future Research Directions
Journal of Medical Systems (2023)
-
Identification and validation of Alzheimer’s disease-related metabolic brain pattern in biomarker confirmed Alzheimer’s dementia patients
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.