Abstract
Biological Ecosystem Networks (BENs) are webs of biological species (nodes) establishing trophic relationships (links). Experimental confirmation of all possible links is difficult and generates a huge volume of information. Consequently, computational prediction becomes an important goal. Artificial Neural Networks (ANNs) are Machine Learning (ML) algorithms that may be used to predict BENs, using as input Shannon entropy information measures (Shk) of known ecosystems to train them. However, it is difficult to select a priori which ANN topology will have a higher accuracy. Interestingly, Auto Machine Learning (AutoML) methods focus on the automatic selection of the more efficient ML algorithms for specific problems. In this work, a preliminary study of a new approach to AutoML selection of ANNs is proposed for the prediction of BENs. We call it the Net-Net AutoML approach, because it uses for the first time Shk values of both networks involving BENs (networks to be predicted) and ANN topologies (networks to be tested). Twelve types of classifiers have been tested for the Net-Net model including linear, Bayesian, trees-based methods, multilayer perceptrons and deep neuronal networks. The best Net-Net AutoML model for 338,050 outputs of 10 ANN topologies for links of 69 BENs was obtained with a deep fully connected neuronal network, characterized by a test accuracy of 0.866 and a test AUROC of 0.935. This work paves the way for the application of Net-Net AutoML to other systems or ML algorithms.
Similar content being viewed by others
Introduction
Many important molecular, living, economical, and other complex systems may be described as complex networks of i parts or nodes interconnected by links, edges, bonds, ties, or relationships1,2,3,4,5,6,7. The volume of information about all these collections of nodes and links is so large that it is impossible for a single person to remember and rationalize all possible connections in known networks. Consequently, it is even more difficult to assign/predict correct connections in new cases. This problem can be solved using Machine Learning (ML) models. In this area, ML models used as input variables are able to quantify structural information of the system. The process has been applied to multiple levels, ranging from the prediction of drug-target networks in molecules to the construction of complex biological networks8,9,10,11.
Specifically, a molecular or living complex system can be explained using numerical parameters that quantify information about the structure of the system. In information theory, Shannon entropy quantifies the information contained in a message, usually in bits. The concept was introduced by Claude E. Shannon in his 1948 paper “A Mathematical Theory of Communication”12. With the pass of time, the Shannon entropy information measures (Shk) of different types and other related information measures have become commonly used indices in quantifying information of the system under study in ML modelling13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29.
In any case, developing ML models using as input Shk values involves, as in other ML problems, the application of data pre-processing variable selection and other techniques. Next, it is necessary to a priori select one or more ML algorithms and train/validate them to seek the final ML model. Consequently, non-experts in ML may encounter difficulties to accomplish this goal. Specifically, in the case of complex molecular and living systems, a non-expert may find it difficult to decide a priori which ML algorithms should be selected to develop the model. In this context, Automated Machine Learning (AutoML) may have an important role in automatically selecting ML algorithms during the development of practical ML applications by non-experts30,31.
This work proposes for the first time the use of Shk values to quantify both the structure of the complex biological system to be predicted and the structure of the ML algorithm to be selected for this task. To this end, a preliminary proof-of-concept experiment is carried out, focusing on a specific class of complex biological systems, and a specific type of ML algorithms. Biological Ecosystem Networks (BENs) have been selected to play the role of a complex biological system.
In addition, Artificial Neural Networks (ANN) have been selected to represent the ML algorithms. The current study uses the entropy values Shk(Ai) and Shk(Bj) as inputs for different pairs of species in the BENs, of the systems under study. The Shk(ANNj) values calculated are also used as inputs for different ANN topologies. In fact, Ecosystems represent one of the most important examples of complex systems. They are a clear example of network-like structures with known procedures to calculate the Shk values32,33,34. In this sense, our group reported different ML models that evaluate the structure of parasite-host webs to predict the interactions between species in different networks35,36,37. In one of our previous works, special emphasis has been placed on the use of Shk information measures to codify structural information in this type of ML studies38. On the other hand, ANNs have been selected because of their more apparent network-like structure, and because they are a useful tool to solve this kind of ML problems. In fact, ANNs are powerful bio-inspired algorithms able to learn/infer large datasets39,40,41,42. ANNs are also able to learn topology patterns in large datasets of bio-systems and other complex networks43. This work proposes the calculation of Shk information indices in both sets of networks: BENs and ANNs. That is why, it has been called a Net-Net AutoML approach. Last, an AutoML linear model is sought using these indices as input. This Net-Net AutoML model could be employed to screen different ANN topologies in order to pre-select the one expected to correctly predict BEN structures before training it.
Results
This work introduces for the first time a new type of algorithm to find the best ANNs that predict BENs. The main steps of the methodology are described in Fig. 1. This is the first report of a Net-Net AutoML model for ANN screening, with the subsequent saving of time and computational resources in the prediction of Complex Networks. The BEN node pairs and the ANN classifiers that were trained for the prediction of BEN node connectivity were turned into Shk descriptors that encoded information for the BEN nodes and the entire ANN topology. Shk were calculated for each node with the MI-NODES software44. In the case of ANN classifiers, the average of all the values of Shk for all the neurons in the ANN was used as input. For the MI-NODES descriptors, the Markov chains theory was applied and, therefore, they were calculated for each k values ranging between 0 and 5 (k = node distance of interaction) as Shk45. These descriptors were linearly combined to find a model (AutoML) that was able to predict how a specific ANN topology would evaluate BEN node connectivity. Thus, AutoML could be used to screen which is the best ANN classifier topology for BEN node connectivity prediction. The AutoML methodology used for the prediction of BEN connectivity includes the following steps with their respective results.
First, Shk values were calculated for a large number of nodes in 69 BENs using the MI-NODES software. We created a dataset of biological systems (bsi dataset) using 33,805 pairs of nodes selected randomly from the 69 BENs. If we consider the adjacency matrix (A) as the mathematical representation of all pairs of Ai vs. Bj nodes in the BEN, the output variable of this dataset are the elements Aij of this matrix. These values quantify the structure (connectivity) of the BEN with values Aij = 1 for the pairs of nodes that are connected (interacting biological species) and Aij = 0 otherwise (non-interacting biological species).
Next, the bsi dataset was expanded with node differences as input variables ∆Shkij = Shki − Shkj for each pair, where Shki is the Shk for the first node and Shkj the Shk for the second node (k = 0–5). As a result, there are bsiNvar = 6·3 = 18 input features for the Aij output for each pair of BEN nodes. The variables Sh0 quantify information for an isolated node, Sh1 refers to the nodes with direct link, Sh2 to nodes that have other nodes between them, and so on.
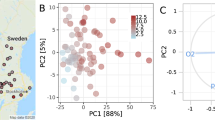
Figure 2 illustrates the distribution of three Shk parameters (k = 2, 3, 5) for both BENs and ANN classifiers to predict them. This dataset was used to train 10 different ANNs. Next, the ANN screening model testing dataset (mt dataset) was made up. The output variable of the mt dataset represents the values of correct or incurred prediction of BEN connectivity Aij by a specific ANN classifier topology, P(ANNAij) = 1 when the ANN topology correctly predicts the observed BEN nodes connectivity Aij (Aij = 1 or 0 in the original bsi dataset). On the contrary, P(ANNAij) = 0 when a specific ANN topology fails to correctly classify the observed Aij of 1 or 0 from the original bsi dataset. The mt dataset contains the predictions of 338,050 node pairs from 69 BENs using 10 different trained ANN classifiers (different topologies). The input variables of the mt dataset are the original variables for each pair of nodes and the values of information indices ANNShk (average value of Sh of all ANN neurons): mtNvar = 6*(Shki + Shkj + ∆Shkij + ANNShk) = 24. The last step consists of the dataset analysis to find the best linear AutoML model for ANN classifier screening (see previous Fig. 1).
Discussion
There are at least two major problems if ANNs are used to predict node connectivity in complex networks. First, the information in complex systems should be turned into numerical input parameters to the future ANN classifiers for node connectivity. Secondly, many ANN classifiers with different topologies should be trained in order to find the best ANN topology that can learn the complex system structure patterns. The first problem can be solved by quantifying the structural information of the complex system (Brain, Ecological, Social, etc.) with Shk information measures46. The classical solution for the second problem is the training of different ANNs to find the best topology. This step involves the use of High Performance Computing (HPC) services if the aim is to test a high number of ANNs for many complex systems.
The current study proposes a new methodology to evaluate how a new ANN classifier could predict the BEN node connectivity, without the need for ANN training. Thus, two types of information descriptors were used: node descriptors for BEN complex networks and the average of ANN neuron descriptors. If ANNs are networks with nodes (neurons) and links (weights), the same mathematical processing as in the case of the BEN complex network could be applied. Therefore, it is possible to quantify topological (connectivity) information of both the BEN complex networks under study and a set of ANNs trained using Shk descriptors. Thus, each node of the complex networks encoded information into Shk descriptors and each ANN classifier was characterized as an entire network by the average of Shk.
The new AutoML methodology proposed a screening of ANN classifier topologies for BEN node connectivity prediction. The current work applied the AutoML methodology to the Ecological systems. Consequently, the AutoML output P(ANNAij) predicted the propensity of a specific ANN topology to predict the biological interaction Aij between the species Ai and Bj from the ecological web (Aij = 1 or not Aij = 0). The best linear AutoML with maximum values of Ac, Sp, and Sn (training and external validation series) is described by Eq. 1. The best linear AutoML was made up of only 5 features: two Shk (k = 0, 3) for each node Ai and Bj, and a Shk (k = 3) for the ANN classifier.
In each BEN, the connections (Aij = 1) indicated the existence of a biological interaction between the organisms of i biological species with the organisms of j species. Table 1 shows the ANN topologies trained to predict BEN connectivity. Thus, the Net-Net AutoML model was able to predict whether a new ANN topology could correctly predict the connectivity between a pair of BEN nodes, prior to training. We introduced variability in the ANN topologies using ANNs without hidden layers (no. 8, 9, 10), with only one hidden layer (no. 1, 2, 7, 5) and with two hidden layers (no. 3, 4, 6). Future work should include different ANN topologies such as skip layer connections, drop out neurons, deep ANNs. These will enable a wider search in the space of possible networks.
The LDA model showed significant goodness-of-fit, also illustrated by Accuracy (Ac), Sensitivity (Sn), and Specificity (Sp) classification values, both in training and external validation series (see Table 2). The proof-of-concept AutoML model fit very well 338,050 outcomes predicted with 10 (previously trained) ANNs. These results were obtained after training the 10 ANNs to learn to discriminate between biological interactions (predation, parasitism, mutualism, etc.) which were connected (Aij = 1) or not (Aij = 0) in BENs of many ecosystems. The mission of the AutoML did not consist of the prediction of BEN connectivity, and, therefore, Sn referred to the number of times that the AutoML was able to evaluate whether a given ANN topology could correctly predict BEN nodes connectivity. The same analogy applied to Sp and Ac. Using Net-Net AutoML methodology, one could decide which ANN will receive more computing resources for training and which one can be used to predict different links (Aij = 1 or 0). The parameter Sh3i quantified the information related to the position of i organism and their neighbours (k = 3) placing a topological distance d ≤ 3 in the BEN. ANNSh3 is also similar but quantifies information for the neurons in a specific ANN topology and not for the organisms in the biological network. Figure 2 illustrates the distribution of the Shk values for all the BENs and ANN topologies studied herein.
The LDA model is a base line classifier that was compared to 11 complex classifiers obtained with 9 ML methods, such as Bayesian Nets, Naïve Bayes Nets, Logistic Regression, Decision Table, Multilayer Perceptron (MLP), Random Forest, Bagging, AdaBoost, and Deep Fully Connected (FC) Networks. All 18 descriptors were used as inputs. The test accuracy (ACC) and AUROC values are presented in Table 3.
It should be observed that the Bayesian methods, Decision Table and Logistic Regression provided accuracies lower than the LDA model. With the MLP, by introducing hidden layers in Artificial Neural Networks, the accuracies and AUROC were improved, with values over 0.8, better than the LDA classifier. More complex models such as ensemble classifiers based on MLP with only one hidden layer (Bagging MLP and AdaBoostM1 MLP) could produce slightly better results. By introducing more hidden layers with MLP 2 H and Deep FC Nets, the test accuracy was increased up to 0.866 (test AUROC = 0.935). Random Forest was not the best model, but it was able to provide a test accuracy of 0.832 (AUROC = 0.914). The ensemble classifier based on simple REP trees, such as Bagging REP, had a performance similar to the MLP 1 H (only one hidden layer).
Therefore, Deep Nets provided the best results, starting with MLP 2H with only 18 neurons (=number of input features) in the first hidden layer, and 9 neurons in the second hidden layer (ACC = 0.827, AUROC = 0.902), leading to the more complex Deep FC Nets with 200, 400 and 200 neurons in the hidden layers 1, 2 and 3 (ACC = 0.866, AUROC = 0.935). An accuracy increase of 4% was obtained with more complex topology of the neural network from 18–9 to 200–400–200 neurons. The DL model was obtained using 10 different network topologies, from one to three hidden layers, with different optimization algorithms, dropout rates, and other hyperparameters. The best DL model had the hidden layer topology n-n*2-n, with activation functions = tanh, n = 200, dropout rate = 0.5, optimizer algorithm = Adam, initialization of weights = glorot_normal, batch size = 4096, epochs = 500, training AUROC = 0.963, training ACC = 0.897, test AUROC = 0.935 and test ACC = 0.866.
The current method used different applications such as MI-NODES for descriptors, STATISTICA, Weka, and Python/Keras for ML classifiers. If the user does not test deep learning classifiers for the final Net-Net model, there is no need for programming. In Weka it is possible to test a deeper MLP. Therefore, scientists without advanced knowledge of programming are able to implement this methodology for specific BENs. An optimal implementation of the method should be performed using a unique python code for all the Net-Net methodology steps. This is the next step for the future version of this method.
Conclusions
The current study confirms that Markov chains are useful to calculate Shk information indices in order to quantify the connectivity patterns of both BENs and ANNs. The new Net-Net AutoML methodology demonstrated how to develop a linear AutoML model, able to select which ANN topology would correctly predict the connectivity of BEN nodes before training it. The best AutoML model demonstrated an accuracy over 86% in test subsets. In conclusion, Net-Net AutoMLs with Shk information indices could be used to screen ANN topologies that can predict the links in biological networks. This may lead to an optimization of computing resources with the prioritization of the training of the best ANN topologies.
Methods
Biological ecosystem networks dataset
A number of 69 Ecosystems or Food Webs were used. The network files in.net format were assembled by our group in a previous work47. The datasets were downloaded from the Interaction Web Database (IWDB): http://www.nceas.ucsb.edu/interactionweb/index.html.
Computational model
Markov-Shannon Entropy Centralities from MI-NODES tool
In the present work, the classical Markov matrix (1Π) was constructed for each network (BEN complex networks and ANNs). In the case of BENs, the adjacency/connectivity matrix were downloaded from public resources as A (n by n matrix, where n is the number of nodes/vertices). Next, the Markov matrix Π was calculated. It contains the vertices probability (pij) based on A. The probability matrix was raised to the power k, resulting (1Π)k, and it was multiplied by the vector of the initial probabilities (0pj). The resulting vectors kP contained the absolute probabilities to reach the nodes moving throughout a walk of length k from node j (kpj) for each k (Eqs 2 and 3). The entropy of graph Shk(G) could be calculated based on the entropy of each node Shkj:
Net-Net AutoML models
Due to the dimension of the dataset and the complexity of the models, for the current calculations two systems were used:
-
BioCAI cluster of RNASA-IMEDIR group (UDC) with 200 CPU cores;
-
A desktop computer with processor i7 (3.60 GHz × 4 physical cores), 16 G RAM, and a GPU NVIDIA Titan Xp (Pascal architecture, dedicated memory of 12 G G5X, 3840 CUDA cores, boost clock 1,582 MHz).
The GPU was particularly useful for the Deep Learning calculations with Keras. All the calculations could be carried out with a desktop computer over a larger period a time, especially due to the Deep Learning calculations.
Once the Shk values of both the ANNs and BENs have been obtained, a Linear Discriminant Analysis (LDA) implemented in the STATISTICA software48 can be run. Let P(ANNAij) be the output of a screening model used to predict the ability of a given ANN topology to correctly classify the BEN connectivity Aij between two nodes i and j (Aij = 1 or 0). Eq. 4 describes the general formula for the LDA model using the following coefficients: aki as coefficients of the Sh for the i node (Shki), bkj as coefficients of the Sh for the j node (Shkj), ckij as coefficients of the differences between the Sh of the nodes i and j (∆Shkij = Shki − Shkj), ANNdk as coefficient of the average Sh for a specific ANN topology, and e0 as the free term coefficient. The k index indicates that this Shk value codifies information for all nodes placed at least at topological distance k from the reference node.
Different statistical parameters can be used to evaluate the statistical significance and validate the goodness-of-fit of LDA equation: n = number of cases, χ2 = Chi-square, p = error level, as well as the Accuracy (Ac), Specificity (Sp), and Sensitivity (Sn) of both training and external validation series49.
Several complex classifiers were tested (see Table 3):
-
Bayesian Nets = weka.classifiers.bayes.BayesNet -D -Q weka.classifiers.bayes.net.search.local.K2–P 1 -S BAYES -E weka.classifiers.bayes.net.estimate.SimpleEstimator –A 0.
-
Naive Bayes Nets = weka.classifiers.bayes.NaiveBayes
-
Logistic Regression = weka.classifiers.functions.Logistic -R 1.0E-8 -M -1 -num-decimal-places 4
-
Decision Table = weka.classifiers.rules.DecisionTable -X 1 -S “weka.attributeSelection.BestFirst -D 1 -N 5”
-
MLP 1H (Multilayer Perceptron 1 hidden layer) = weka.classifiers.functions.MultilayerPerceptron -L 0.3 -M 0.2 -N 1000 -V 0 -S 0 -E 20 -H a
-
MLP 2H (Multilayer Perceptron 2 hidden layers) = weka.classifiers.functions.MultilayerPerceptron -L 0.3 -M 0.2 -N 5000 -V 0 -S 0 -E 20 -H “18, 9” -batch-size 500
-
Random Forest = weka.classifiers.trees.RandomForest -P 100 -I 500 -num-slots 1 -K 0 -M 1.0 -V 0.001 -S 1
-
Bagging REP = weka.classifiers.meta.Bagging -P 100 -S 1 -num-slots 1 -I 10 -W weka.classifiers.trees.REPTree –M 2 -V 0.001 -N 3 -S 1 -L -1 -I 0.0
-
Bagging MLP = weka.classifiers.meta.Bagging -P 100 -S 1 -num-slots 1 -I 10 -W weka.classifiers.functions.MultilayerPerceptron -batch-size 4000 –L 0.3 -M 0.2 -N 500 -V 0 -S 0 -E 20 -H a
-
AdaBoostM1 MLP = weka.classifiers.meta.AdaBoostM1 -P 100 -S 1 -I 10 -W weka.classifiers.functions.MultilayerPerceptron -batch-size 4000 –L 0.3 -M 0.2 -N 500 -V 0 -S 0 -E 20 -H a
-
Deep FC Nets (Deep Learning Fully Connected Networks) = n-n*2-n′ hidden layer topology (n = 200).
Deep Learning FC Nets were programmed in Python with Keras, and the other classifiers were obtained with the Weka tool. For the DL models, different hyperparameter values were tested:
-
n = Number of neurons in a hidden layer: 10, 18, 50, 100, 200, 500.
-
Network topologies: ‘n’,‘n-n’, ‘n-n-n’, ‘n-n*2’, ‘n-n*2-n’, ‘n*2’, ‘n*2-n’, ‘n-n:2’, ‘n:2’, ‘n-n:2-n:4’ (this notation does not include the input layer with 18 neurons = no. of features and the output layer with a neuron for the class).
-
Neuron dropout rate: 0.0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9.
-
Optimizers: ‘RMSprop’, ‘Adagrad’, ‘Adadelta’, ‘Adam’, ‘Adamax’, ‘Nadam’.
-
Weight initialization for hidden layer neurons: ‘uniform’, ‘lecun_uniform’, ‘normal’, ‘glorot_normal’, ‘glorot_uniform’, ‘he_normal’, ‘he_uniform’.
-
Batch size for training = 1024, 2048, 4096.
-
Training epochs = 20, 50, 100, 200, 300, 400, 500.
-
Training cross validation: 3-folds (default value in Keras).
The Net-Net AutoML algorithm shown in Fig. 1 could be described as follows:
-
(1)
For each BEN:
-
(1.1)
Get the connectivity matrix.
-
(1.2)
Add weights for the BEN connections (if present).
-
(1.3)
For each node A:
-
(1.3.1)
Calculate node Shannon Entropies with MI-NODES: Shk(Ai).
-
(1.3.2)
Create pairs of entropies for all the other nodes B: Shk(Ai) − Shk(Bj).
-
(1.3.1)
-
(1.1)
-
(2)
Find different ANN classifiers to predict BEN node Aj − Bj connectivity:
-
(2.1)
For each ANNj classifier:
-
(2.1)
-
(2.1.1)
Calculate network Shannon Entropy: Shk(ANNj).
-
(3)
Merge BEN node descriptors with ANN descriptors into Net-Net dataset: Shk(Ai), Shk(Bj), Shk(ANNj).
-
(4)
Split Net-Net dataset into training and test subsets.
-
(5)
Find the best Net-Net classifier to evaluate whether a specific ANN can predict the BEN connectivity:
-
(5.1)
For each ML method (Bayesian, Trees, Artificial Neural Networks, etc.)
-
(5.1.1)
For each set of model parameters (ex: topology, activation function, etc.)
-
(5.1.1.1)
Use a Net-Net subset to train the classifier
-
(5.1.1.2)
Evaluate the model with test subset calculating accuracy (ACC) and AUROC.
-
(6)
Choose the best Net-Net classifier with the best ACC and AUROC.
Steps (5) and (6) used Weka and Python/Keras scripts. In the future version of the method, different classifiers will be tested for the BEN connectivity prediction (not only ANNs). This involves the adaptation of MI-NODES application. The main advantage of the Net-Net methodology is that it can build a Net-Net classifier able to screen ANN classifiers which predict BEN node connectivities.
Data Availability
All data generated or analyzed during this study were included in this article (along with its Supplementary Information files) and they are publicly available at Figshare repository with https://doi.org/10.6084/m9.figshare.6238424.
References
Sandhu, K. S. et al. Large-scale functional organization of long-range chromatin interaction networks. Cell Rep 2, 1207–1219, https://doi.org/10.1016/j.celrep.2012.09.022 (2012).
Gaspar, M. E. & Csermely, P. Rigidity and flexibility of biological networks. Brief Funct Genomics 11, 443–456, https://doi.org/10.1093/bfgp/els023 (2012).
Csermely, P., Korcsmaros, T., Kiss, H. J., London, G. & Nussinov, R. Structure and dynamics of molecular networks: A novel paradigm of drug discovery: A comprehensive review. Pharmacol. Ther. 138, 333–408, https://doi.org/10.1016/j.pharmthera.2013.01.016 (2013).
Vidal, M., Cusick, M. E. & Barabasi, A. L. Interactome networks and human disease. Cell 144, 986–998, https://doi.org/10.1016/j.cell.2011.02.016 (2011).
Barabasi, A. L., Gulbahce, N. & Loscalzo, J. Network medicine: a network-based approach to human disease. Nat Rev Genet 12, 56–68, https://doi.org/10.1038/nrg2918 (2011).
Barabasi, A. L. & Oltvai, Z. N. Network biology: understanding the cell’s functional organization. Nat Rev Genet 5, 101–113, https://doi.org/10.1038/nrg1272 (2004).
Strogatz, S. H. Exploring complex networks. Nature 410, 268–276, https://doi.org/10.1038/35065725 (2001).
Riera-Fernandez, P. et al. From QSAR models of Drugs to Complex Networks: State-of-Art Review and Introduction of New Markov-Spectral Moments Indices. Curr Top Med Chem 12, 927–960, https://doi.org/10.2174/156802612800166819 (2012).
Gonzalez-Diaz, H. QSAR and Complex Networks in Pharmaceutical Design, Microbiology, Parasitology, Toxicology, Cancer and Neurosciences. Current Pharmaceutical Design 16, 2598–U2524, https://doi.org/10.2174/138161210792389261 (2010).
González-Díaz, H., Prado-Prado, F., Pérez-Montoto, L. G., Duardo-Sánchez, A. & López-Díaz, A. QSAR Models for Proteins of Parasitic Organisms, Plants and Human Guests: Theory, Applications, Legal Protection, Taxes, and Regulatory Issues. Curr Proteomics 6, 214–227, https://doi.org/10.2174/157016409789973789 (2009).
Prado-Prado, F. J., Ubeira, F. M., Borges, F. & Gonzalez-Diaz, H. Unified QSAR & Network-Based Computational Chemistry Approach to Antimicrobials. II. Multiple Distance and Triadic Census Analysis of Antiparasitic Drugs Complex Networks. J. Comput. Chem. 31, 164–173, https://doi.org/10.1002/jcc.21292 (2010).
Shannon, C. E. A Mathematical Theory of Communication. The Bell System Technical Journal 27, 379–423, https://doi.org/10.1002/j.1538-7305.1948.tb01338.x (1948).
Dehmer, M. & Emmert-Streib, F. Analysis of Complex Networks. From Biology to Linguistics. (WILEY-VCH Verlag GmbH & Co. KGaA, 2009).
Dehmer, M., Grabner, M. & Varmuza, K. Information indices with high discriminative power for graphs. PLoS ONE 7, e31214, https://doi.org/10.1371/journal.pone.0031214 (2012).
Dehmer, M., Varmuza, K., Borgert, S. & Emmert-Streib, F. On entropy-based molecular descriptors: statistical analysis of real and synthetic chemical structures. Journal of chemical information and modeling 49, 1655–1663 (2009).
Estrada, E. & Avnir, D. Continuous symmetry numbers and entropy. J Am Chem Soc 125, 4368–4375, https://doi.org/10.1021/ja020619w (2003).
Graham, D. J., Grzetic, S., May, D. & Zumpf, J. Information properties of naturally-occurring proteins: Fourier analysis and complexity phase plots. The protein journal 31, 550–563, https://doi.org/10.1007/s10930-012-9432-7 (2012).
Graham, D. J. & Greminger, J. L. On the information expressed in enzyme structure: more lessons from ribonuclease A. Mol. Divers. 15, 769–779, https://doi.org/10.1007/s11030-011-9307-4 (2011).
Graham, D. J. & Greminger, J. L. On the information expressed in enzyme primary structure: lessons from Ribonuclease A. Mol. Divers. 14, 673–686, https://doi.org/10.1007/s11030-009-9211-3 (2010).
Graham, D. J. & Kim, M. Information and classical thermodynamic transformations. The journal of physical chemistry 112, 10585–10593, https://doi.org/10.1021/jp7119526 (2008).
Graham, D. J., Malarkey, C. & Sevchuk, W. Experimental investigation of information processing under irreversible Brownian conditions: work/time analysis of paper chromatograms. The journal of physical chemistry 112, 10594–10602, https://doi.org/10.1021/jp711953r (2008).
Graham, D. J. Information Content in Organic Molecules: Brownian Processing at Low Levels. Journal of chemical information and modeling 47, 376–389 (2007).
Graham, D. J. Information content in organic molecules: aggregation states and solvent effects. Journal of chemical information and modeling 45, 1223–1236, https://doi.org/10.1021/ci050101m (2005).
Graham, D. J. & Schulmerich, M. V. Information Content in Organic Molecules: Reaction Pathway Analysis via Brownian Processing. J Chem Inf Comput Sci 44 (2004).
Graham, D. J., Malarkey, C. & Schulmerich, M. V. Information Content in Organic Molecules: Quantification and Statistical Structure via Brownian Processing. J. Chem. Inf. Comput. Sci. 44 (2004).
Graham, D. J. Information and organic molecules: structure considerations via integer statistics. J. Chem. Inf. Comput. Sci. 42, 215–221 (2002).
Graham, D. J. & Schacht, D. V. Base information content in organic formulas. J. Chem. Inf. Comput. Sci. 40, 942–946 (2000).
Barigye, S. J. et al. Shannon’s, Mutual, Conditional and Joint Entropy Information Indices. Generalization of Global Indices Defined from Local Vertex Invariants. Curr Comput Aided Drug Des (2013).
Aguiar-Pulido, V. et al. Naïve Bayes QSDR classification based on spiral-graph Shannon entropies for protein biomarkers in human colon cancer. Mol Biosyst, https://doi.org/10.1039/c2mb25039j (2012).
Kotthoff, L., Thornton, C., Hoos, H. H., Hutter, F. & Leyton-Brown, K. Auto-WEKA 2.0: Automatic model selection and hyperparameter optimization in WEKA. Journal of Machine Learning Research 18, 1–5 (2017).
Feurer, M., Klein, A., Eggensperger, K., Springenberg, J. & Blum, M. Efficient and Robust Automated Machine Learning. Advances in Neural Information Processing Systems 28, 2962–2970 (2015).
Borenstein, E. & Feldman, M. W. Topological signatures of species interactions in metabolic networks. J Comput Biol 16, 191–200, https://doi.org/10.1089/cmb.2008.06TT (2009).
Ulanowicz, R. E. Quantitative methods for ecological network analysis. Comput Biol Chem 28, 321–339, https://doi.org/10.1016/j.compbiolchem.2004.09.001 (2004).
Olff, H. et al. Parallel ecological networks in ecosystems. Philos Trans R Soc Lond B Biol Sci 364, 1755–1779, https://doi.org/10.1098/rstb.2008.0222 (2009).
Gonzalez-Diaz, H., Riera-Fernandez, P., Pazos, A. & Munteanu, C. R. The Rucker-Markov invariants of complex Bio-Systems: applications in Parasitology and Neuroinformatics. Biosystems 111, 199–207, https://doi.org/10.1016/j.biosystems.2013.02.006 (2013).
Gonzalez-Diaz, H. & Riera-Fernandez, P. New Markov-Autocorrelation Indices for Re-evaluation of Links in Chemical and Biological Complex Networks used in Metabolomics, Parasitology, Neurosciences, and Epidemiology. J. Chem. Inf. Model. 52, 3331–3340, https://doi.org/10.1021/ci300321f (2012).
Riera-Fernandez, I. et al. From QSAR models of Drugs to Complex Networks: State-of-Art Review and Introduction of New Markov-Spectral Moments Indices. Curr. Top. Med. Chem. (2012).
Riera-Fernandez, P. et al. New Markov-Shannon Entropy models to assess connectivity quality in complex networks: From molecular to cellular pathway, Parasite-Host, Neural, Industry, and Legal-Social networks. Journal of Theoretical Biology 293, 174–188, https://doi.org/10.1016/j.jtbi.2011.10.016 (2012).
Gonzalez-Diaz, H. et al. ANN-QSAR model for selection of anticancer leads from structurally heterogeneous series of compounds. European Journal of Medicinal Chemistry 42, 580–585, https://doi.org/10.1016/j.ejmech.2006.11.016 (2007).
Jalali-Heravi, M. & Fatemi, M. H. Prediction of thermal conductivity detection response factors using an artificial neural network. J. Chromatogr. A 897, 227–235 (2000).
Prado-Prado, F. J., Garcia-Mera, X. & Gonzalez-Diaz, H. Multi-target spectral moment QSAR versus ANN for antiparasitic drugs against different parasite species. Bioorganic & Medicinal Chemistry 18, 2225–2231, https://doi.org/10.1016/j.bmc.2010.01.068 (2010).
Tenorio-Borroto, E. et al. ANN multiplexing model of drugs effect on macrophages; theoretical and flow cytometry study on the cytotoxicity of the anti-microbial drug G1 in spleen. Bioorganic & Medicinal Chemistry 20, 6181–6194, https://doi.org/10.1016/j.bmc.2012.07.020 (2012).
Gonzalez-Diaz, H. et al. MIANN models in medicinal, physical and organic chemistry. Curr Top Med Chem 13, 619–641 (2013).
Duardo-Sanchez, A. et al. Modeling complex metabolic reactions, ecological systems, and financial and legal networks with MIANN models based on Markov-Wiener node descriptors. Journal of chemical information and modeling 54, 16–29, https://doi.org/10.1021/ci400280n (2014).
Duardo-Sanchez, A., Gonzalez-Diaz, H. & Pazos, A. MI-NODES Multiscale Models of Metabolic Reactions, Brain Connectome, Ecological, Epidemic, World Trade, and Legal-Social Networks. Curr. Bioinf. 10, 692–713, https://doi.org/10.2174/1574893610666151008013413 (2015).
Shannon, C. E., Weaver, W., Blahut, R. E. & Hajek, B. The mathematical theory of communication. Vol. 117 (University of Illinois press Urbana, 1949).
Riera-Fernández, P. et al. Definition of Markov-Harary Invariants and Review of Classic Topological Indices and Databases in Biology, Parasitology, Technology, and Social-Legal Networks. Current Bioinformatics 6, 94–121 (2011).
STATISTICA (data analysis software system), version 6. 0, www.statsoft.com.Statsoft, Inc. v. 6.0 (2002).
Hill, T. & Lewicki, P. STATISTICS Methods and Applications. A Comprehensive Reference for Science, Industry and Data Mining. Vol. 1 (StatSoft, 2006).
Acknowledgements
The authors acknowledge Basque Government (Eusko Jaurlaritza) grant (IT1045-16) - 2016–2021 for consolidated research groups. This work was supported by the “Collaborative Project in Genomic Data Integration (CICLOGEN)” PI17/01826 funded by the Carlos III Health Institute, as part of the Spanish National plan for Scientific and Technical Research and Innovation 2013–2016 and the European Regional Development Funds (FEDER). This project was also supported by the General Directorate of Culture, Education and University Management of Xunta de Galicia ED431D 2017/16 and “Drug Discovery Galician Network” Ref. ED431G/01 and the “Galician Network for Colorectal Cancer Research” (Ref. ED431D 2017/23), and finally by the Spanish Ministry of Economy and Competitiveness for its support through the funding of the unique installation BIOCAI (UNLC08-1E-002, UNLC13-13-3503) and the European Regional Development Funds (FEDER) by the European Union. CR Munteanu acknowledges the support of NVIDIA Corporation with the donation of the Titan Xp GPU used for this research.
Author information
Authors and Affiliations
Contributions
E.B. performed the data analysis and participated in the writing of the paper. C.R.M. coded MI-NODES scripts for the calculation of Shk values and participated in the writing of the paper. M.C.M. supervised the work of E.B. on his internship and participated in the writing of the paper. A.P. contributed to the discussion and participated in the writing of the paper. H.G.D. proposed the Net-Net AutoML algorithm idea, performed the data analysis, and participated in the writing of the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Barreiro, E., Munteanu, C.R., Cruz-Monteagudo, M. et al. Net-Net Auto Machine Learning (AutoML) Prediction of Complex Ecosystems. Sci Rep 8, 12340 (2018). https://doi.org/10.1038/s41598-018-30637-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-30637-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.