Abstract
In this study it was theoretically shown that discovered by us recently (Brovarets’ et al., Frontiers in Chemistry, 2018, 6:8; doi: 10.3389/fchem.2018.00008) high-energetical, significantly non-planar (symmetry C1), short-lived wobbled conformers of the classical Watson-Crick А·Т(WC), reverse Watson-Crick А·Т(rWC), Hoogsteen А·Т(Н) and reverse Hoogsteen А·Т(rН) DNA base pairs are the intermediates of their pairwise А∙Т(WC)/А∙Т(rWC) ↔ А∙Т(H)/А∙Т(rH) conformational transformations. These transitions do not require for their realization the energy-consumable anisotropic rotation of the amino group of A around the exocyclic C6-N6 bond. They are controlled by the non-planar transition states with quasi-orthogonal geometry (symmetry C1) joined by the single intermolecular (Т)N3H···N6(А) H-bond (~4 kcal∙mol−1). The Gibbs free energies of activation for these non-dissociative, dipole-active conformational transitions consist 7.33 and 7.81 kcal∙mol−1, accordingly. Quantum-mechanical (QM) calculations in combination with Bader’s quantum theory of “Atoms in Molecules” (QTAIM) have been performed at the MP2/aug-cc-pVDZ//B3LYP/6-311++G(d,p) level of QM theory in the continuum with ε = 4 under normal conditions.
Similar content being viewed by others
Introduction
Spontaneous transition of the DNA base pairs from the Watson-Crick (WC) to Hoogsteen (H) configuration and vice versa is one of the functionally-important physico-chemical properties of DNA1,2,3,4,5,6,7,8,9. It was shown by NMR methods1,2,3,4,5 that Watson-Crick ↔ Hoogsteen breathing in DNA duplex containing A∙T rich region occurs via the switching of the Watson-Crick DNA base pair (bp) from the anti- to syn-conformation with the probability ~10−2 and represents one of the pathways for the reaction of formaldehyde with DNA10. Thorough calculations by the method of molecular dynamics indicate that А·Т(WC) ↔ А·Т(Н) transitions of actually bps and anti ↔ syn transitions of the A around the glycosidic bond are closely correlated processes, for which Gibbs free energy of activation is 10–11 kcal∙mol−1 under normal conditions8.
Based on analysis of the microstructural nature of these transitions, it is quite logical to connect it with the analogical properties of the isolated DNA bps11,12,13. Comprehensive analysis of the current literature data showed that the nature of these biologically-important processes has not been investigated at all. Currently in the literature there is only one single theoretical work devoted to the study of the anti ↔ syn non-dissociative transitions in irregular pairs of nucleotide bases that do not have an exocyclic amino group in its composition14.
Recently, we have theoretically revealed novel high-energetic, significantly non-planar (symmetry C1), short-lived wobbled (w) conformers – А·Т(wWC), А·Т(wrWC), А·Т(wН) and А·Т(wrН) for each of the four classical А·Т(WC) DNA bps – Watson-Crick А·Т(WC), reverse Watson-Crick А·Т(rWC), Hoogsteen А·Т(Н) and reverse Hoogsteen А·Т(rН)11. It is known from the literature data, that these bps joined by different H-bonds are formed due to the rotation of the DNA base relative to the other on 180° around: the (A)N1–N3(T) axis for the reverse Watson-Crick А·Т(rWC) or in other terms Donohue DNA bp15,16,17,18,19,20,21,22,23; the (A)C9′-N9 axis for the Hoogsteen A·T(H) bp1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30 and the (A)N7–N3(T) axis for the reverse Hoogsteen A·T(rH) or in other terms Haschemeyer–Sobell bp31,32,33,34.
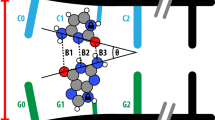
It was found that revealed А·Т(wWC), А·Т(wrWC), А·Т(wН) and А·Т(wrН) conformers have essentially non-planar structure joined by the two anti-parallel N6H/N6H′···O4/O2 and N3H···N6 H-bonds (the N6H′ chemical bond has trans-orientation relative to the N1C6 chemical bond of A). These specific intermolecular contacts involve pyramidalized A amino group, acting simultaneously as an acceptor and a donor of the H-bonding. The transition states (TSs) – TSА·Т(WC)↔А·Т(wWC), TSА·Т(rWC)↔А·Т(wrWC), TSА·Т(Н)↔А·Т(wН) and TSА·Т(rН)↔А·Т(wrН) – of the dipole-active conformational transformations of the basic, plane-symmetric state of the classical А·Т DNA bps into the high-energetic, essentially non-planar wobbled bps and vice versa possess wobble structures (symmetry C1) and are joined by the N6H/N6H′···O4/O2 and N3H···N6 H-bonds. The А·Т(wWC), А·Т(wrWC), А·Т(wН) and А·Т(wrН) conformers was found to be dynamically stable structures with short lifetime τ = (1.4–3.9) ps. It was assumed that these conformational transitions are directly related to the thermally-driven fluctuational behavior of DNA – pre-melting and breathing6,7.
In this work it was established for the first time that just-mentioned novel conformers А·Т(wWC), А·Т(wrWC), А·Т(wН) and А·Т(wrН) control the А·Т(wWC)/А·Т(wrWC) ↔ А·Т(wН)/А·Т(wrН) conformational transitions. Moreover, in view of the recently discovered conformational transitions for the classical A·T DNA bps - А·Т(WC) ↔ А·Т(wWC), А·Т(rWC) ↔ А·Т(wrWC), А·Т(Н) ↔ А·Т(wН) and А·Т(rН) ↔ А·Т(wrН)11, they are also intermediates of the biologically-important А·Т(WC)/А·Т(rWC) ↔ А·Т(Н)/А·Т(rН) conformational transitions.
Energetically favorable mechanism of the conformational pairwise transformation of the intermediates А∙Т(wWC) ↔ А∙Т(wH) and А∙Т(wrWC) ↔ А∙Т(wrH), and together with them conformational transition of the А∙Т DNA bps – А∙Т(WC)/А∙Т(rWC) ↔ А∙Т(H)/А∙Т(rH), does not require for their realization the rotation of the amino group of A around the exocyclic C6N6 bond35.
In this case conformational transformations are controlled by the soft, non-planar TSs, stabilized by the participation of the single intermolecular (Т)N3H···N6(А) H-bond between the imino group of T and pyramidilized amino group of A. The Gibbs free energies of activation for these non-dissociative, dipole-active conformational transitions consist 7.33 and 7.81 kcal∙mol−1, accordingly.
Two other mechanisms – the А∙Т(wWC) ↔ А∙Т(wH) and А∙Т(wrWC) ↔ А∙Т(wrH) – are realized via the anisotropic rotation of the amino group of A (together with T interacting with A through two intermolecular antiparallel (A)N6H/N6H′···O4/O2(T) and (T)N3H···N6(A) H-bonds) around the exocyclic C6N6 bond. In TSs of these conformational transitions the pyramidality of the amino group of A significantly increases: this causes increase of the energy of the N3H···N6 H-bond and decrease of the energy of the intermolecular N6H/N6H′···O4/O2 H-bond. The transitions states of these reactions – TScysА·Т(wWC)↔А·Т(wН), TStransА·Т(wWC)↔А·Т(wН) and TScysА·Т(wrWC)↔А·Т(wrН), TStransА·Т(wrWC)↔А·Т(wrН) – have close energy in corresponding conformational transformations (14.9 and 15.0 kcal∙mol−1, accordingly). Thus, these TSs of the mutual conformational transformation of the wobble intermediates – А∙Т(wWC) ↔ А∙Т(wH) and А∙Т(wrWC) ↔ А∙Т(wrH) of the classical А∙Т DNA bps – А∙Т(WC)/А∙Т(rWC) ↔ А∙Т(H)/А∙Т(rH) – determine their conformational transformations.
Computational Methods
We have calculated geometries of the basic and high-energetic conformers and transition states (TSs) of their mutual conformational transformations together with their harmonic vibrational frequencies at the B3LYP/6–311++G(d,p) level of theory36,37,38,39,40, using Gaussian’09 package41, in the continuum with ε = 4, which is typical for the processes in real biological complexes and taking into account the structural and functional characteristics of the bases in the duplex DNA and at the same time satisfactorily reflecting the environment in the essentially hydrophobic base-pair recognition pocket of the high-fidelity DNA-polymerase42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66. Considered level of theory has been successfully applied for the calculations of the similar tasks and systems47,48,49,50,51,52,53,54,55. A scaling factor of 0.966855,56,57,58,59,60,61 has been used in order to correct the harmonic frequencies of all bps and TSs of the transitions between them. The local minima or TSs, localized by Synchronous Transit-guided Quasi-Newton method62, have been appointed to the complexes on the potential energy landscape containing any or one imaginary frequency in their vibrational spectra, accordingly. We used TS theory in order to estimate the activation barriers of the conformational transformations63. Electronic energy calculations have been performed at the single point at the MP2/aug-cc-pVDZ level of theory67,68.
The Gibbs free energy G for all structures has been received at the MP2/6-311++G(2df,pd) level of theory by the formula:
where Eel – electronic energy, while Ecorr – thermal correction.
The electronic energies of interaction ∆Eint have been obtained at the MP2/6-311++G(2df,pd) level of theory as a difference between the BSSE-corrected69,70,71,72 electronic energy of the bp and electronic energies of the isolated bases.
Bader’s quantum theory of Atoms in Molecules (QTAIM)73,74,75,76,77,78 has been applied for the analysis of the electron density distribution by AIMAll program package79, using wave functions calculated at the B3LYP/6-311++G(d,p) level of theory. We considered the presence of the (3, −1) bond critical point (BCP), a bond path between the donor and acceptor of the intermolecular contact and positive value of the Laplacian at this BCP (Δρ > 0) as criteria for the existence of the H-bond or attractive van der Waals contact formation73,74,75,76,77,78,79,80,81,82,83,84.
The energies of the attractive van der Waals contacts85,86 in TSs of the conformational transitions have been estimated by the Espinosa-Molins-Lecomte (EML) formula87,88:
where V(r) – value of a local potential energy at the (3, −1) BCP.
The energies of the conventional AH···B H-bonds have been calculated by the Iogansen’s formula89:
where Δν – frequency shift of the stretching mode of the H-bonded AH group involved in the AH···B H-bond relatively the unbound group. We applied the partial deuteration in order to avoid the effect of vibrational resonances90,91.
In this study the numeration for the DNA bases is generally accepted92.
In this study we have provided investigations at the basic, but sufficient level of the isolated H-bonded pairs of nucleotide bases, that adequately simulates the processes in real biological systems93,94,95, in particular in the base-pair recognition pocket of the high-fidelity DNA-polymerase42,43,44,45,46. At this, we have relied on the experience received in the previous works11,96,97,98 on the related topic and systems, in which the negligibly small impact of the stacking and sugar-phosphate backbone on the tautomerisation processes has been shown.
Results and Their Discussion
In our previous paper11 we have succeed to establish in the classical А∙Т DNA bps with Cs symmetry – Watson-Crick (WC), reverse Watson-Crick А·Т(rWC), Hoogsteen А·Т(Н) and reverse Hoogsteen А·Т(rН) DNA bps – novel high-energetic, dynamically-stable, mirror-symmetrical А∙Т(wWC)R,L, А∙Т(wH)R,L, А∙Т(wrWC)R,L and А∙Т(wrH)R,L conformational states. Their distinguished feature independently of the pair, in which they are realized, is significantly non-planar structure (С1 symmetry), caused by the pyramidal structure of the ≥C6N6H2 amino fragment of the A DNA base, in which the amino group acts simultaneously as a donor and an acceptor of the specific intermolecular interaction with T through the two (Т)N3H···N6(A) and (A)N6H/N6H′···O4/O2(T) H-bonds. Each of the four А∙Т Watson-Crick DNA bps transfers into the aforementioned conformer via two mirror-symmetric pathways through the TSА∙Т(WC)↔А∙Т(wWC)R,L, TSА∙Т(rWC)↔А∙Т(wrWC)R,L, TSА∙Т(H)↔А∙Т(wrH)R,L and TSА∙Т(rH)↔А∙Т(wrH)R,L (C1 symmetry). At this, the structures, which names differ from each other only by the subscripts R and L, are mirror-symmetrical, that is enantiomers. It is well known that enantiomers have identical scalar physico-chemical characteristics and differ only by the direction of the dipole moment.
Let analyze the biological significance of these non-usual conformers of the classical А∙Т DNA bps.
In this context it was fixed important result – these conformers are responsible for the two different WC/rWC ↔ H/rH mechanisms of the non-dissociative conformational transformation of the А∙Т DNA bps (Fig. 1, Tables 1–3).
Geometrical structures of the stationary points on the reaction pathways of the discovered conformational transitions of the four biologically important А·Т DNA bps. Electronic energies of the interaction ΔEint (MP2/6-311++G(2df,pd)//B3LYP/6-311++G(d,p) level of theory, in kcal∙mol−1), relative Gibbs free energies ∆G and electronic energies ∆E (in kcal∙mol−1), imaginary frequencies νi at the TSs of the conformational transitions (MP2/aug-cc-pVDZ//B3LYP/6-311++G(d,p) level of theory in the continuum with ε = 4 at T = 298.15 К) are presented below complexes in brackets. Dotted lines indicate AH···B H-bonds and attractive A···B van der Waals contacts – their lengths are presented in angstroms (for their more detailed physico-chemical characteristics see Table 2); carbon atoms are in light-blue, nitrogen – in dark-blue, hydrogen – in grey and oxygen – in red. Exclusively enantiomers of one type are presented.
First of these conformational transformations, which are the most energetically favorable mechanisms, are controlled by the soft TSА∙Т(wWC)R,L↔А∙Т(wH)R,L and TSА∙Т(wrWC)R,L↔А∙Т(wrH)R,L (C1 symmetry) with low values of imaginary frequency (7.7 i and 16.1 i cm−1, accordingly). Both of them are joined by the one-single intermolecular (T)N3H···N6(A) H-bond (~4 kcal∙mol−1) between the imino group of T and pyramidilized amino group of A. In this case, conformational transformations of the А∙Т DNA bps are realized by the following non-dissociative scenario (each of them – by the mirror-symmetric pathways): А∙Т(WC) (0.00) ↔ TSА∙Т(WC)↔А∙Т(wWC)R,L (7.13) ↔ А∙Т(wWC)R,L (5.36)11 ↔ TSА∙Т(wWC)R,L↔А∙Т(wH)R,L (7.33) ↔ А∙Т(wH)R,L (5.35) ↔ TSА∙Т(wH)R,L↔А∙Т(H) (7.24) ↔ А∙Т(H) (−0.44)11 and А∙Т(rWC) (0.00) ↔ TSА∙Т(rWC)↔А∙Т(wrWC)R,L (7.26) ↔ А∙Т(wrWC)R,L (5.97)11 ↔ TSА∙Т(wrWC)R,L↔А∙Т(wrH)R,L (7.81) ↔ А∙Т(wrH)R,L (5.79) ↔ TSА∙Т(wrH)R,L↔А∙Т(rH) (7.41) ↔ А∙Т(rH) (−0.03)11. Notably, obtained energetic barriers are in good coincidence with the molecular-dynamic data for the А∙Т(WC) ↔ А∙Т(H) transition (10-11 kcal∙mol−1 under normal conditions8).
Herewith, some R structures transform into the other R structures, the same concerns L-structures. Saying in other words, pathways of these dipole-active conformational transformations are mirror-symmetric. In fact, the TSА∙Т(wWC)R,L↔А∙Т(wH)R,L and TSА∙Т(wrWC)R,L↔А∙Т(wrH)R,L, which pairwise link the А∙Т(wWC)R,L and А∙Т(wH)R,L, А∙Т(wrWC)R,L and А∙Т(wrH)R,L conformers, are transition states of the WC/rWC ↔ H/rH conformational transformations of the classical А∙Т DNA bps.
High-energetic mechanism of the WC/rWC ↔ H/rH conformational transitions of the А∙Т DNA bps is connected with anisotropic rotation of the amino group of A around the exocylic С6-N6 bond35 and is controlled by the TScysА∙Т(wWC)R,L↔А∙Т(wH)L,R, TStransА∙Т(wWC)R,L↔А∙Т(wH)L,R and TScysА∙Т(wrWC)R,L↔А∙Т(wrH)L,R, TStransА∙Т(wrWC)R,L↔А∙Т(wrH)L,R, that have non-planar structure (С1 symmetry) and quite high values of the imaginary frequencies (~252 i cm−1). These TSs are joined by the two anti-parallel intermolecular (Т)N3H···N6(A) and (A)N6H/N6H′···O4/O2(T) H-bonds; notably, first of them is significantly stronger than the second one. The attractive O2···N7 and O4···N7 van der Waals contacts with weak energy (~0.18 kcal∙mol−1) also participate in the stabilization of the TScysА∙Т(wWC)R,L↔А∙Т(wH)L,R and TScysА∙Т(wrWC)R,L↔А∙Т(wrH)L,R, accordingly.
In this case, the R structures transform into the L structures and vice versa and WC/rWC ↔ H/rH conformational transitions of the classical А∙Т DNA bps occur in such a case (each of them through two energetically and topologically non-equivalent ways):
А∙Т(WC) (0.00) ↔ TSА∙Т(WC)↔А∙Т(wWC)R,L (7.13) ↔ А∙Т(wWC)R,L (5.36)11 ↔ TScysА∙Т(wWC)R,L↔А∙Т(wH)L,R (14.89) ↔ А∙Т(wH)L,R (5.35) ↔ TSА∙Т(wH)L,R↔А∙Т(H) (7.24) ↔ А∙Т(H) (−0.44)11;
А∙Т(WC) (0.00) ↔ TSА∙Т(WC)↔А∙Т(wWC)R,L (7.13) ↔ А∙Т(wWC)R,L (5.36)11 ↔ TStransА∙Т(wWC)R,L↔А∙Т(wH)L,R (14.88) ↔ А∙Т(wH)L,R (5.35) ↔ TSА∙Т(wH)L,R↔А∙Т(H) (7.24) ↔ А∙Т(H) (−0.44)11;
А∙Т(rWC) (0.00) ↔ TSА∙Т(rWC)↔А∙Т(wrWC)R,L (7.26) ↔ А∙Т(wrWC)R,L (5.97)11 ↔ TScysА∙Т(wrWC)R,L↔А∙Т(wrH)L,R (15.01) ↔ А∙Т(wrH)L,R (5.79) ↔ TSА∙Т(wrH)L,R↔А∙Т(rH) (7.41) ↔ А∙Т(rH) (−0.03)11 and
А∙Т(rWC) (0.00) ↔ TSА∙Т(rWC)↔А∙Т(wrWC)R,L (7.26) ↔ А∙Т(wrWC)R,L (5.97)11 ↔ TStransА∙Т(wrWC)R,L↔А∙Т(wrH)L,R (15.00) ↔ А∙Т(wrH)L,R (5.79) ↔ TSА∙Т(wrH)L,R↔А∙Т(rH) (7.41) ↔ А∙Т(rH) (−0.03)11 (relative Gibbs free energy is presented after each structure in brackets at the MP2/aug-cc-pVDZ//B3LYP/6-311++G(d,p) level of QM theory in the continuum with ε = 4 under normal conditions).
It should be noted that the orientation of the methyl group of the T DNA base does not alter in the course of all reactions of conformational transitions. At this, the heterocycles of the DNA bases, capable for the out-of-plane bending99,100,101, stay planar.
So, obtained by us results launch the conception of the “mechanics” of the non-dissociative WC/rWC ↔ H/rH conformational transformations of the classical А∙Т DNA bps.
Of course, in the composition of DNA these conformational transitions represent a self-consistent transformation of the bps, the anti ↔ syn transition of A around the glycosidic bond (ΔΔGTS = 3.4 kcal∙mol−1 at χTS = 121◦ for BI-conformer of the isolated 2′-deoxyadenosine102) and reorganization of stacking and hydratation8. Simple comparison of the energetics, determining these processes, clearly indicates that the first two of them plays a leading role. This fact gives hope that obtained in this paper data are closely related to the nature of the А∙Т(WC) ↔ А∙Т(H) thermal fluctuation process, which occurs in DNA1,2,3,4,5,6,7. This conclusion can be verified, applying the newest methods of ab initio dynamics for the short fragments of DNA.
Conclusions
By applying developed by us novel ideas according the high-energetic conformers of the classical А∙Т DNA bps11, we offered novel non-dissociative mechanisms of the А∙Т(WC) ↔ А∙Т(H) and А∙Т(rWC) ↔ А∙Т(rH) conformational transitions, that do not require for their realization energy-consuming anisotropic rotation of the amino group of the A DNA base around the C6-N6 exocyclic bond. Figuratively speaking, at the transformation of the A base from the anti- to syn-conformation leading to the formation of the Hoogsteen А∙Т(H) and reverse Hoogsteen А∙Т(rH) bps, it dynamically relies as on the support on the T DNA base through the pyramidilized amino group of A, interacting with it in the TS region by one single (Т)N3H···N6(А) H-bond.
In the light of the obtained by us results, it could be suggested that the А∙Т(WC) ↔ А∙Т(H) conformational transition in DNA duplex, which was registered experimentally1,2,3,4,5,6,7, most likely occurs by the non-dissociative mechanism: A, rotating from the anti- to syn-configuration, interacts with T via the intermolecular H-bonds along the entire process of the conformational transformation.
References
Nikolova, E. N. et al. Transient Hoogsteen base pairs in canonical duplex DNA. Nature 470, 498–504 (2011).
Alvey, H. S., Gottardo, F. L., Nikolova, E. N. & Al-Hashimi, H. M. Widespread transient Hoogsteen base pairs in canonical duplex DNA with variable energetics. Nat. Commun. 5, 4786–4793 (2014).
Zhou, H. et al. New insights into Hoogsteen base pairs in DNA duplexes from a structure-based survey. Nucleic Acids Res. 43, 3420–3433 (2015).
Zhou, H. Occurrence and function of Hoogsteen base pairs in nucleic acids. PhD Thesis: Duke University, Durham (2016).
Sathyamoorthy, B. et al. Insights into Watson–Crick/Hoogsteen breathing dynamics and damage repair from the solution structure and dynamic ensemble of DNA duplexes containing m1A. Nucleic Acids Res. 45, 5586–5601 (2017).
von Hippel, P., Johnson, N. P. & Marcus, A. H. Fifty years of DNA “breathing”: reflections on old and new approaches. Biopolymers 99, 923–953 (2013).
Frank-Kamenetskii, M. & Prakash, S. Fluctuations in the DNA double helix: a critical review. Phys. Life Rev. 11, 153–170 (2014).
Yang, C., Kim, E. & Pak, Y. Free energy landscape and transition pathways from Watson–Crick to Hoogsteen base pairing in free duplex DNA. Nucl. Acids Res. 43, 7769–7778 (2015).
Cubero, E., Abrescia, N. G., Subirana, J. A., Luque, F. J. & Orozco, M. Theoretical study of a new DNA structure: the antiparallel Hoogsteen duplex. J. Am. Chem. Soc. 125, 14603–14612 (2003).
Bohnuud, T. et al. Computational mapping reveals dramatic effect of Hoogsteen breathing on duplex DNA reactivity with formaldehyde. Nucl. Acids Res. 40, 7644–7652 (2012).
Brovarets’, O. O., Tsiupa, K. S. & Hovorun, D. M. Surprising conformers of the biologically important A·T DNA base pairs: QM/QTAIM proofs. Front. Chem. 6(8), https://doi.org/10.3389/fchem.2018.00008 (2018).
Lavery, R. Modeling nucleic acids: fine structure, flexibility, and conformational transitions. Adv. Comput. Biol. 1, 69–145 (1994).
Stofer, E., Chipot, C. & Lavery, R. Free energy calculations of Watson−Crick base pairing in aqueous solution. J. Am. Chem. Soc. 121, 9503–9508 (1999).
Brovarets’, O. O. & Hovorun, D. M. How do long improper purine-purine pairs of DNA bases adapt the enzymatically competent conformation? Structural mechanism and its quantum-mechanical grounds. Ukr. J. Phys. 60, 748–756 (2015).
Donohue, J. & Trueblood, K. N. Base pairing in DNA. J. Mol. Biol. 2, 363–371 (1960).
Tchurikov, N. A., Chernov, B. K., Golova, Y. B. & Nechipurenko, Y. D. Parallel DNA: Generation of a duplex between two Drosophila sequences in vitro. FEBS Lett. 257, 415–418 (1989).
Cubero, E., Luque, F. J. & Orozco, M. Theoretical studies of d(A:T)-based parallel-stranded DNA duplexes. J. Am. Chem. Soc. 123, 12018–12025 (2001).
Parvathy, V. R. et al. NMR structure of a parallel-stranded DNA duplex at atomic resolution. Nucleic Acids Res. 30, 1500–1511 (2002).
Poltev, V. I. et al. Analysis of the conformational features of Watson–Crick duplex fragments by molecular mechanics and quantum mechanics methods. Biophysics 61, 217–226 (2016).
Ye, M. Y. et al. Adaptively recognizing parallel-stranded duplex structure for fluorescent DNA polarity analysis. Anal. Chem. 89, 8604–8608 (2017).
Szabat, M. & Kierzek, R. Parallel‐stranded DNA and RNA duplexes: structural features and potential applications. FEBS J. 284, 3986–3998 (2017).
Brovarets’, O. O. Under what conditions does G·C Watson-Crick DNA base pair acquire all four configurations characteristic for A·T Watson-Crick DNA base pair? Ukr. Biochem. J. 85, 98–103 (2013).
Brovarets’, O. O. Structural and energetic properties of the four configurations of the A·T and G·C DNA base pairs. Ukr. Biochem. J. 85, 104–110 (2013).
Hoogsteen, K. The crystal and molecular structure of a hydrogenbonded complex between 1-methylthymine and 9-methyladenine. Acta Cryst. 16, 907–916 (1963).
Abrescia, N. G., Thompson, A., Huynh-Dinh, T. & Subirana, J. A. Crystal structure of an antiparallel DNA fragment with Hoogsteen base pairing. Proc. Natl. Acad. Sci. USA 99, 2806–2811 (2002).
Abrescia, N. G., Gonzalez, C., Gouyette, C. & Subirana, J. A. X-ray and NMR studies of the DNA oligomer d(ATATAT): Hoogsteen base pairing in duplex DNA. Biochemistry 43, 4092–4100 (2004).
Pous, J. et al. Stabilization by extra-helical thymines of a DNA duplex with Hoogsteen base pairs. J. Am. Chem. Soc. 130, 6755–6760 (2008).
Campos, L. et al. Overview of the structure of all AT oligonucleotides: organization in helices and packing interactions. Biophys. J. 91, 892–903 (2006).
Nikolova, E. N. et al. A historical account of Hoogsteen base-pairs in duplex DNA. Biopolymers 99, 955–968 (2014).
Acosta-Reyes, F. J., Alechaga, E., Subirana, J. A. & Campos, J. L. Structure of the DNA duplex d(ATTAAT)2 with Hoogsteen hydrogen bonds. PLOS ONE 10, e0120241 (2015).
Haschemeyer, A. E. V. & Sobell, H. M. The crystal structure of an intermolecular nucleoside complex: Adenosine and 5-bromouridine. Proc. Natl Acad. Sci. USA 50, 872–877 (1963).
Sühnel, J. Beyond nucleic acid base pairs: From triads to heptads. Biopolymers 61, 32–51 (2002).
Zagryadskaya, E. I., Doyon, F. R. & Steinberg, S. V. Importance of the reverse Hoogsteen base pair 54–58 for tRNA function. Nucleic Acids Res. 31, 3946–3953 (2003).
Liu, K., Miles, H. T., Frazier, J. & Sasisekharan, V. A novel DNA duplex. A parallel-stranded DNA helix with Hoogsteen base pairing. Biochemistry 32, 11802–11809 (1993).
Hovorun, D. M., Mishchuk, Y. R. & Kondratyuk, I. V. On a quantum-chemical nature of a stereochemical nonrigidity of canonical nucleotide bases. Biopol. Cell 12, 5–12 (1996).
Tirado-Rives, J. & Jorgensen, W. L. Performance of B3LYP Density Functional Methods for a large set of organic molecules. J. Chem. Theory Comput. 4, 297–306 (2008).
Parr, R. G. & Yang, W. Density-functional theory of atoms and molecules. Oxford: Oxford University Press (1989).
Lee, C., Yang, W. & Parr, R. G. Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys. Rev. B. 37, 785–789 (1988).
Hariharan, P. C. & Pople, J. A. The influence of polarization functions on molecular orbital hydrogenation energies. Theor. Chim. Acta 28, 213–222 (1973).
Krishnan, R., Binkley, J. S., Seeger, R. & Pople, J. A. Self‐consistent molecular orbital methods. XX. A basis set for correlated wave functions. J. Chem. Phys. 72, 650–654 (1980).
Frisch, M. J., et al. GAUSSIAN 09 (Revision B.01). Wallingford CT: Gaussian Inc (2010).
García-Moreno, B. E. et al. Experimental measurement of the effective dielectric in the hydrophobic core of a protein. Biophys. Chem. 64, 211–224 (1997).
Bayley, S. T. The dielectric properties of various solid crystalline proteins, amino acids and peptides. Trans. Faraday Soc. 47, 509–517 (1951).
Dewar, M. J. S. & Storch, D. M. Alternative view of enzyme reactions. Proc. Natl. Acad. Sci. USA 82, 2225–2229 (1985).
Mertz, E. L. & Krishtalik, L. I. Low dielectric response in enzyme active site. Proc. Natl. Acad. Sci. USA 97, 2081–2086 (2000).
Petrushka, J., Sowers, L. C. & Goodman, M. Comparison of nucleotide interactions in water, proteins, and vacuum: Model for DNA polymerase fidelity. Proc. Natl. Acad. Sci. USA 83, 1559–1562 (1986).
Matta, C. F. How dependent are molecular and atomic properties on the electronic structure method? Comparison of Hartree-Fock, DFT, and MP2 on a biologically relevant set of molecules. J. Comput. Chem. 31, 1297–1311 (2010).
Danilov, V. I., Anisimov, V. M., Kurita, N. & Hovorun, D. MP2 and DFT studies of the DNA rare base pairs: the molecular mechanism of the spontaneous substitution mutations conditioned by tautomerism of bases. Chem. Phys. Lett. 412, 285–293 (2005).
Rutledge, L. R. & Wetmore, S. D. A computational proposal for the experimentally observed discriminatory behavior of hypoxanthine, a weak universal nucleobase. Phys. Chem. Chem. Phys. 14, 2743–2753 (2012).
Brovarets’, O. O. & Hovorun, D. M. Quantum-chemical investigation of tautomerization ways of Watson-Crick DNA base pair guanine-cytosine. Ukr. Biochem. J. 82, 55–60 (2010).
Brovarets’, O. O. & Hovorun, D. M. Quantum-chemical investigation of the elementary molecular mechanisms of pyrimidine·purine transversions. Ukr. Biochem. J. 82, 57–67 (2010).
Brovarets’, O. O., Zhurakivsky, R. O. & Hovorun, D. M. Is there adequate ionization mechanism of the spontaneous transitions? Quantum-chemical investigation. Biopol. Cell 26, 398–405 (2010).
Brovarets’, O. O. & Hovorun, D. M. How the long G·G* Watson-Crick DNA base mispair comprising keto and enol tautomers of the guanine tautomerises? The results of the QM/QTAIM investigation. Phys. Chem. Chem. Phys. 6, 15886–15899 (2014).
Brovarets’, O. O., Zhurakivsky, R. O. & Hovorun, D. M. The physico-chemical “anatomy” of the tautomerisation through the DPT of the biologically important pairs of hypoxanthine with DNA bases: QM and QTAIM perspectives. J. Mol. Model. 19, 4119–4137 (2013).
Brovarets’, O. O. Microstructural mechanisms of the origin of the spontaneous point mutations. DrSci Thesis: Taras Shevchenko National University of Kyiv, Kyiv, Ukraine (2015).
Brovarets’, O. O. & Hovorun, D. M. The nature of the transition mismatches with Watson-Crick architecture: the G*·T or G·T* DNA base mispair or both? A QM/QTAIM perspective for the biological problem. J. Biomol. Struct. & Dynam. 33, 925–945 (2015).
Brovarets’, O. O., Zhurakivsky, R. O. & Hovorun, D. M. DPT tautomerisation of the wobble guanine·thymine DNA base mispair is not mutagenic: QM and QTAIM arguments. J. Biomol. Struct. & Dynam. 33, 674–689 (2015).
Brovarets’, O. O. & Hovorun, D. M. Stability of mutagenic tautomers of uracil and its halogen derivatives: the results of quantum-mechanical investigation. Biopol. Cell 26, 295–298 (2010).
Brovarets’, O. O. & Hovorun, D. M. Intramolecular tautomerization and the conformational variability of some classical mutagens – cytosine derivatives: Quantum-chemical study. Biopol. Cell 27, 221–230 (2011).
Palafox, M. A. Molecular structure differences between the antiviral nucleoside analogue 5-iodo-2′-deoxyuridine and the natural nucleoside 2′-deoxythymidine using MP2 and DFT methods: conformational analysis, crystal simulations, DNA pairs and possible behavior. J. Biomol. Struct. & Dynam. 32, 831–851 (2014).
El-Sayed, A. A., Tamara Molina, A., Alvarez-Ros, M. C. & Palafox, M. A. Conformational analysis of the anti-HIV Nikavir prodrug: comparisons with AZT and thymidine, and establishment of structure-activity relationships/tendencies in other 6′-derivatives. J. Biomol. Struct. & Dynam. 33, 723–748 (2015).
Peng, C., Ayala, P. Y., Schlegel, H. B. & Frisch, M. J. Using redundant internal coordinates to optimize equilibrium geometries and transition states. J. Comput. Chem. 17, 49–56 (1996).
Atkins, P. W. Physical chemistry. Oxford: Oxford University Press (1998).
Brovarets’, O. O. & Hovorun, D. M. Can tautomerisation of the A∙T Watson-Crick base pair via double proton transfer provoke point mutations during DNA replication? A comprehensive QM and QTAIM analysis. J. Biomol. Struct. & Dynam. 32, 127–154 (2014).
Brovarets’, O. O., Zhurakivsky, R. O. & Hovorun, D. M. The physico-chemical mechanism of the tautomerisation via the DPT of the long Hyp*·Hyp Watson-Crick base pair containing rare tautomer: a QM and QTAIM detailed look. Chem. Phys. Lett. 578, 126–132 (2013).
Brovarets’, O. O. & Hovorun, D. M. Does the G·G*syn DNA mismatch containing canonical and rare tautomers of the guanine tautomerise through the DPT? A QM/QTAIM microstructural study. Mol. Phys. 112, 3033–3046 (2014).
Frisch, M. J., Head-Gordon, M. & Pople, J. A. Semi-direct algorithms for the MP2 energy and gradient. Chem. Phys. Lett. 166, 281–289 (1990).
Kendall, R. A., Dunning, T. H. Jr & Harrison, R. J. Electron affinities of the first-row atoms revisited. Systematic basis sets and wave functions. J. Chem. Phys. 96, 6796–6806 (1992).
Boys, S. F. & Bernardi, F. The calculation of small molecular interactions by the differences of separate total energies. Some procedures with reduced errors. Mol. Phys. 19, 553–566 (1970).
Gutowski, M., Van Lenthe, J. H., Verbeek, J., Van Duijneveldt, F. B. & Chalasinski, G. The basis set superposition error in correlated electronic structure calculations. Chem. Phys. Lett. 124, 370–375 (1986).
Sordo, J. A., Chin, S. & Sordo, T. L. On the counterpoise correction for the basis set superposition error in large systems. Theor. Chim. Acta 74, 101–110 (1988).
Sordo, J. A. On the use of the Boys–Bernardi function counterpoise procedure to correct barrier heights for basis set superposition error. J. Mol. Struct. 537, 245–251 (2001).
Bader, R. F. W. Atoms in molecules: A quantum theory. Oxford: Oxford University Press (1990).
Matta, C. F. & Hernández-Trujillo, J. Bonding in polycyclic aromatic hydrocarbons in terms of the electron density and of electron delocalization. J. Phys. Chem. A 107, 7496–7504 (2003).
Matta, C. F., Castillo, N. & Boyd, R. J. Atomic contributions to bond dissociation energies in aliphatic hydrocarbons. J. Chem. Phys. 125, 204103 (2006).
Cukrowski, I. & Matta, C. F. Hydrogen–hydrogen bonding: A stabilizing interaction in strained chelating rings of metal complexes in aqueous phase. Chem. Phys. Lett. 499, 66–69 (2010).
Matta, C. F. Modeling biophysical and biological properties from the characteristics of the molecular electron density, electron localization and delocalization matrices, and the electrostatic potential. J. Comput. Chem. 35, 1165–1198 (2014).
Lecomte, C., Espinosa, E. & Matta, C. F. On atom–atom ‘short contact’ bonding interactions in crystals. IUCrJ 2, 161–163 (2015).
Keith, T. A. AIMAll (Version 10.07.01). Retrieved from aim.tkgristmill.com (2010).
Matta, C. F., Castillo, N. & Boyd, R. J. Extended weak bonding interactions in DNA: π-stacking (base-base), base-backbone, and backbone-backbone interactions. J. Phys. Chem. B 110, 563–578 (2006).
Brovarets’, O. O., Zhurakivsky, R. O. & Hovorun, D. M. Is the DPT tautomerisation of the long A·G Watson-Crick DNA base mispair a source of the adenine and guanine mutagenic tautomers? A QM and QTAIM response to the biologically important question. J. Comput. Chem. 35, 451–466 (2014).
Brovarets’, O. O. & Hovorun, D. M. Tautomeric transition between wobble А·С DNA base mispair and Watson-Crick-like A·C* mismatch: miscrostructural mechanism and biological significance. Phys. Chem. Chem. Phys. 17, 15103–15110 (2015).
Brovarets’, O. O. & Hovorun, D. M. Wobble ↔ Watson-Crick tautomeric transitions in the homo-purine DNA mismatches: a key to the intimate mechanisms of the spontaneous transversions. J. Biomol. Struct. & Dyn. 33, 2710–2715 (2015).
Brovarets’, O. O. & Hovorun, D. M. DPT tautomerisation of the G·Asyn and A*·G*syn DNA mismatches: A QM/QTAIM combined atomistic investigation. Phys. Chem. Chem. Phys. 16, 9074–9085 (2014).
Matta, C. F. & Boyd, R. J. The Quantum Theory of Atoms in Molecules: from solid state to DNA and drug design. Wiley-VCH Verlag GmbH & Co. KGaA (2007).
Brovarets’, O. O., Zhurakivsky, R. O. & Hovorun, D. M. Structural, energetic and tautomeric properties of the T·T*/T*·T DNA mismatch involving mutagenic tautomer of thymine: a QM and QTAIM insight. Chem. Phys. Lett. 2014(592), 247–255 (2014).
Espinosa, E., Molins, E. & Lecomte, C. Hydrogen bond strengths revealed by topological analyses of experimentally observed electron densities. Chem. Phys. Lett. 285, 170–173 (1998).
Mata, I., Alkorta, I., Espinosa, E. & Molins, E. Relationships between interaction energy, intermolecular distance and electron density properties in hydrogen bonded complexes under external electric fields. Chem. Phys. Lett. 507, 185–189 (2011).
Iogansen, A. V. Direct proportionality of the hydrogen bonding energy and the intensification of the stretching ν(XH) vibration in infrared spectra. Spectrochim. Acta Part A: Mol. Biomol. Spectrosc. 55, 1585–1612 (1999).
Brovarets’, O. O. & Hovorun, D. M. New structural hypostases of the A·T and G·C Watson-Crick DNA base pairs caused by their mutagenic tautomerisation in a wobble manner: a QM/QTAIM prediction. RSС Adv. 5, 99594–99605 (2015).
Brovarets’, O. O. & Hovorun, D. M. A novel conception for spontaneous transversions caused by homo-pyrimidine DNA mismatches: a QM/QTAIM highlight. Phys. Chem. Chem. Phys. 17, 21381–21388 (2015).
Saenger, W. Principles of nucleic acid structure. New York: Springer (1984).
Kimsey, I. J., Petzold, K., Sathyamoorthy, B., Stein, Z. W. & Al-Hashimi, H. M. Visualizing transient Watson-Crick-like mispairs in DNA and RNA duplexes. Nature 519, 315–320 (2015).
Szymanski, E. S., Kimsey, I. J. & Al-Hashimi, H. M. Direct NMR evidence that transient tautomeric and anionic states in dG∙dT form Watson–Crick-like base pairs. J. Am. Chem. Soc. 139, 4326–4329 (2017).
Maximoff, S. N., Kamerlin, S. C. L. & Florián, J. DNA Polymerase λ active site favors a mutagenic mispair between the enol form of deoxyguanosine triphosphate substrate and the keto form of thymidine template: a free energy perturbation study. J. Phys. Chem. B 121, 7813–7822 (2017).
Zoete, V. & Meuwly, M. Double proton transfer in the isolated and DNA-embedded guanine-cytosine base pair. J. Chem. Phys. 121, 4377–4388 (2004).
Negi, I., Kathuria, P., Sharma, P. & Wetmore, S. D. How do hydrophobic nucleobases differ from natural DNA nucleobases? Comparison of structural features and duplex properties from QM calculations and MD simulations. Phys. Chem. Chem. Phys. 19, 16365–16374 (2017).
Felske, L. R., Lenz, S. A. P. & Wetmore, S. D. Quantum chemical studies of the structure and stability of N-methylated DNA nucleobase dimers: insights into the mutagenic base pairing of damaged DNA. J. Phys. Chem. A 122, 410–419 (2018).
Govorun, D. M. et al. AM1 calculation of the nucleic acid bases structure and vibrational spectra. J. Mol. Struct. 267, 99–103 (1992).
Nikolaienko, T. Y., Bulavin, L. A. & Hovorun, D. M. How flexible are DNA constituents? The quantum-mechanical study. J. Biomol. Struct. Dynam. 29, 563–575 (2011).
Hovorun, D. M., Gorb, L. & Leszczynski, J. From the nonplanarity of the amino group to the structural nonrigidity of the molecule: a post-Hartree-Fock ab initio study of 2-aminoimidazole. Int. J. Quantum. Chem. 75, 245–253 (1999).
Zhurakivsky R. O. Conformational properties of elementary structural units of nucleic acids: the nonempirical quantum-mechanical study. PhD Thesis: National Taras Shevchenko University of Kyiv, Kyiv (2011).
Acknowledgements
The authors gratefully appreciate technical support and computational facilities of joint computer cluster of SSI “Institute for Single Crystals” of the National Academy of Sciences of Ukraine (NASU) and Institute for Scintillation Materials of the NASU incorporated into Ukrainian National Grid. DrSci Ol’ha Brovarets’ expresses sincere gratitude to organizing committee headed by Prof. Karl Kuchler (Medical University Vienna, Austria) for the kind invitation and financial support (ABC fellow) of the participation in the “ATP-Binding Cassette (ABC) Proteins: From Multidrug Resistance to Genetic Disease” (March 6–12, 2018, Innsbruck, Austria) and to Chemistry Biological Interface Division of the Royal Society of Chemistry (RSC, UK) for the RSC Travel Grant for the participation in the “3rd Green and Sustainable Chemistry Conference” (May 13–16, 2018, Hotel Intercontinental, Berlin, Germany). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
D.M.H. proposed the idea of the study, formulated the task and suggested possible pathways of the non-dissociative structural transitions of the Watson-Crick and reverse Watson-Crick А·Т DNA base pairs into the Hoogsteen and reverse Hoogsteen forms and partly wrote the text of the manuscript. O.O.B. localized the pathways of the investigated interconversions, prepared numerical data for Tables and graphical materials for Figures, performed QM/QTAIM calculations and partly wrote the text of the manuscript. K.S.T. performed QM/QTAIM calculations, participated in the preparation of the text of the manuscript. All authors were involved in the proofreading of the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Brovarets’, O.O., Tsiupa, K.S. & Hovorun, D.M. Non-dissociative structural transitions of the Watson-Crick and reverse Watson-Crick А·Т DNA base pairs into the Hoogsteen and reverse Hoogsteen forms. Sci Rep 8, 10371 (2018). https://doi.org/10.1038/s41598-018-28636-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-28636-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.