Abstract
The AINTEGUMENTA-like (AIL) family plays a central role in regulating the growth and development of organs in many plants. However, little is known about the characteristics and functions of the AIL family in Chinese cabbage (Brassica rapa L. ssp. pekinensis). In this study, a genome-wide analysis was performed to identify the members of the AIL family in Chinese cabbage. We identified three ANT genes and six ANT-like genes of Chinese cabbage, most of which were differentially expressed in different organs or tissues. Furthermore, compared with the wild-type line, the size of different organs in the 35S-BrANT-1 line was significantly increased by promoting cell proliferation. Meanwhile, over-expression of BrANT-1 also increases the stomatal number and delays the leaf senescence. Transcriptome analyses revealed that a set of cell proliferation and stoma development genes were up-regulated, while the senescence-associated genes were down-regulated, suggesting these genes may be involved in BrANT-1 regulated processes for controlling organ size, stomatal density and leaf senescence. In summary, this study offers important insights into the characteristics and functions of the ANT genes in Chinese cabbage, and provides a promising strategy to improve yield or head size in Chinese cabbage breeding programs.
Similar content being viewed by others
Introduction
The size of the leafy head is an important economical trait of Chinese cabbage (Brassica rapa L. ssp. pekinensis). However, little is known about how the size of the leafy head is controlled. Therefore, understanding the molecular mechanism of the leafy head development has always been a key goal of genetic improvement in Chinese cabbage.
Plant organ size reflects both cell number and cell size which are regulated by a polygenic system through cell proliferation and cell expansion1,2. Previous reports have shown that many regulatory factors are involved in these two processes3, such as AINTEGUMENTA (ANT)4,5,6,7, ANT-like (AIL)8,9, auxin-regulated gene involved in organ size (ARGOS)10,11 and growth-regulating factors (GRFs)12,13. ANT-like (AIL) transcription factors are members of the APETALA2 (AP2) subfamily, belonging to the APETALA2/ETHYLENE RESPONSE FACTOR (AP2/ERF) super family14. The AP2 subfamily is divided into two groups, euAP2 group and ANT group. the euAP2 group is characterized by a microRNA binding site (miR172) in the post-domain region, and the ANT group is characterized by lineage-specific motifs or amino acid insertions in the two AP2 domains. The ANT group is further subdivided into basal ANT lineage and euANT lineage, according to the length of the pre-domain region sequences and other specific motifs. For example, the euANT lineage has a long pre-domain region (127–307 aa) and three specific motifs in the pre-domain region (the euANT2, 3, and 4 motifs: WLGFSLS, PKLEDFLG, and TFGQR), while the basal ANT lineage has a short pre-domain region (44–81 aa)15. A wealth of studies have shown that the AIL gene family members are involved in the ovule primordium initiation, female gametophyte formation, as well as organ growth and polarity4,5,7,16. For example, the transgenic plants over-expressing the ANT genes result in the production of organs with a large size7,17,18, while the loss-of-function mutations of the ANT gene have a smaller organ size4,5,7. Additionally, many studies revealed that ANT regulates organ size by changing the total cell numbers7. Furthermore, cells ectopically expressing ANT in fully differentiated organs exhibit neoplastic activity by producing calli and adventitious roots and shoots7. These studies strongly suggest that ANT most likely maintains ongoing cell proliferation coordinately with cell growth19. However, to our knowledge, the AIL family from Chinese cabbage has not been characterized. Therefore, identifying and analyzing the AIL family in Chinese cabbage is of great interest.
In this study, we investigated the BrAIL family members in Chinese cabbage through genome-wide bioinformatics analysis, including the identification and characterization of the AIL family members, gene structural analysis, phylogeny and motif analysis. The expression patterns of these genes were characterized in detail in response to the naphthaleneacetic acid (NAA) treatment in different tissues by quantitative real-time PCR (qRT-PCR). Our results show BrANT-1 had the highest sequence similarity to AtANT (AT4G37750), and hence was chosen for further functional analysis. Over-expression of BrANT-1 enhanced organ size by increasing cell number and stomatal density and delaying leaf senescence. Furthermore, by analyzing the potential pathways where BrANT-1 participates, it is possible to understand how the gene regulates Chinese cabbage yield and/or head size.
Results
Identification of AIL family members in Chinese cabbage
A total of nine BrAIL family members were identified in the Chinese cabbage genome, according to the taxonomy of the AP2 superfamily (Supplementary Tables S1 and S2). These included three ANT genes and six AIL genes (Table 1). The sequences of each member were downloaded from the Brassica database (http://brassicadb.org/brad/)20 and the two AP2 domains were confirmed according to the SMART database (http://smart.embl-heidelberg.de/). All nine BrAIL family members were named by the A. thaliana orthologs based on the sequence similarity of the protein sequences. The detailed description of sequence similarity between Arabidopsis thaliana and Brassica rapa is shown in Supplementary Table S3. When more than two Chinese cabbage genes were mapped to one homologous gene in A. thaliana, one additional number was added to the end of the gene name21. For example, three ANT genes, Bra017852, Bra011782 and Bra010610, were homologs of AtANT (AT4G37750). Accordingly, they were named BrANT-1, BrANT-2 and BrANT-3, respectively. The BrAIL family members were randomly mapped to different chromosomes of B. rapa (chromosome number: 01, 02, 03, 06, 08 and 10). The chromosome 02 and 03 contained three and two BrAIL genes, respectively, while the chromosome 01, 06, 08 and 10 contained only one BrAIL gene. Subsequent sequence analysis showed that the coding sequences of these nine BrAIL genes had a length of 1323 to 1725 bp, which encode a peptide of 440 to 574 aa. The encoded proteins had a predicted molecular weight varying from 49.6 to 64.1 kDa and a theoretical isoelectric point (pI) varying from 6.09 to 7.81.
Phylogenetic relationships and gene structure of the AIL protein in Arabidopsis, rice and Chinese cabbage
In order to understand the classification of the BrAIL genes in Chinese cabbage, the AIL genes from two other model plants were selected for comparative analyses, including a model C3 monocotyledon plant (rice) and a model eudicots plant (Arabidopsis). A phylogenetic tree was constructed based on the full-length protein sequences of Chinese cabbage, rice and Arabidopsis, by using the bootstrap–neighbor-joining method. The AIL genes of the other two species were obtained from previous reports on Arabidopsis22 and rice23. According to the phylogenetic analysis, the 21 members were divided into seven groups (Class A-Class G) (Fig. 1A). Low bootstrap values were obtained because the difference of the AP2 domain sequences among these three species was small, indicating that BrAILs have high similarity to AtAILs. On the contrary, BrAIL proteins were only remotely related to the OsAIL proteins. To further explore the evolutionary relationships of the coding sequences, the structural analyses of intron/exon of the three species were performed by an online tool GSDS (http://gsds.cbi.pku.edu.cn/). It was found that most of the AIL family members in the three species had at least three introns, except OsAP2/EREBP#058, while the number of introns in the BrAIL genes ranged from six to eight (Fig. 1B). Furthermore, most of the genes (7 out of 9) had six introns, except BrAIL1 and BrAIL5, which contained seven and eight introns, respectively.
Phylogenetic relationships and intron/exon structure analysis among three species (Chinese cabbage, Rice and Arabidopsis). (A) The phylogenetic tree was constructed by MEGA5 using the Neighbor-Joining method with 1000 bootstrap replicates of the AIL proteins among Chinese cabbage, rice and Arabidopsis, (B) Intron and exon structures of the AIL family members in these three species.
Syntenic analysis of the AIL family members in Chinese cabbage
To understand the evolution mechanism of the BrAIL genes in Chinese cabbage, the syntenic genes were analyzed between A. thaliana and B. rapa with the BRAD program (http://brassicadb.org/brad/searchSynteny.php)24. The results showed that nine BrAIL genes were from five blocks of four translocation Proto-Calepineae Karyotype (tPCK) chromosomes of the ancestor, respectively. All genes were uniformly distributed on three subgenomes, namely, less fractionized (LF), more fractionized 1 (MF1), and more fractionized 2 (MF2). Additionally, two sets of triplicated genes were found in the Chinese cabbage genomes: BrANT-1, BrANT-2 and BrANT-3 in the U block; BrAIL6-1, BrAIL6-2 and BrAIL6-3 in the R block (Table 2). The other genes (BrAIL1, BrAIL5 and BrAIL7) were singletons. Interestingly, the tandemly duplicated gene was not found in the BrAIL family.
Multiple sequence alignment and motif analysis
The nine BrAIL proteins were used for the multiple sequence alignment. From previous studies, the euANT proteins in Arabidopsis thaliana or Oryza sativa are divided into five regions, including the pro-domain region, the R1 domain, the linker region, the R2 domain, and the post-domain region. Accordingly, all BrANT proteins were divided into five regions (Supplementary Fig. S1). Interestingly, all BrAIL proteins belonged to the euANT subgroup according to the conserved motifs, including a 10-aa insertion in the R1 domain (the euANT1 motif: NS[C/F][K/R][K/R]EG[Q/H][S/A]R) and three insertions in the pre-domain region (the euANT2, 3, and 4 motifs: [W/L]L[G/T]FSLS, PK[L/M]E[D/N]F[L/F]G, and [T/S]FGQR). Subsequent amino acid sequence analysis was carried out on the two AP2 domains of the nine AIL family members. The E-value of the forward AP2 domain (R1) matched to the full sequences varied from 2.56e-26 to 1.54e-11, and the E-value of the subsequent AP2 domain (R2) varied from 2.35e-34 to 3.85e-30. The lengths of these two AP2 domains (AP2-R1 and AP2-R2) were constant. The length of most AP2-R1 domains (70 aa) was longer than that of AP2-R2 (65 aa), except BrANT-3. The detailed description of the two AP2 domains is shown in Supplementary Table S4. Additionally, the length of the linker region was conservative (30 aa). The motif of the BrAIL members was analyzed using an online tool MEME (meme.nbcr.net/meme/intro.html), and ten motifs were identified, including motif 1, 2, 3, 4 and 10 in nine BrAIL proteins, motif 5 and 6 in eight BrAIL proteins, motif 8 in seven BrAIL proteins, and motif 7 and 9 in three BrAIL proteins, respectively (Supplementary Fig. S2).
Expression patterns of the BrAIL family members in various organs
Accumulating experiments have shown that the AIL genes were expressed in multiple tissues and involved in regulating the development process of different tissues, such as root25, shoot7, floral organ26,27,28, leaf 29 and seed18. To explore the potential roles of the BrAIL genes in regulating the growth and development of Chinese cabbage, the expression patterns of the BrAIL genes was investigated in root (R), dwarf stem (DS), old leaf (OL), young leaf (YL), blooming flower (BFL) of Chinese cabbage by qRT-PCR. Eight genes were detected in different tissues, except BrAIL1, which was undetectable in all tissues (Fig. 2). For example, four BrAIL family members, including BrANT-2, BrANT-3, BrAIL6-1 and BrAIL6-2, showed higher expression levels in R than other tissues; three BrAIL genes (BrANT-1, BrAIL6-3 and BrAIL7) were mainly expressed in DS; BrAIL5 was mainly expressed in YL. Moreover, none of the genes were detected in old leaves.
Cluster analysis of the expression pattern of the AIL family members in different tissues of Chinese cabbage. Five different tissues (50 days after sowing) were surveyed by real-time PCR, including root (R), booming flower (BFL), dwarf stem (DS), young leaf (YL) and old leaf (OL). The expression level of different tissues was calculated by the 2−∆Ct method using the BrActin gene as the reference gene.
Expression profiles of the BrAIL family members under NAA treatments
Previous studies have shown that auxin plays an important role in plant growth and developmental processes30. In addition, the auxin signal may regulate organ growth by modulating ANT expression10. To understand the expression responses of the BrAIL members to auxin, we assessed the transcript levels of the BrAIL genes upon NAA treatments by qRT-PCR. As shown in Fig. 3, the transcriptional levels of most genes were induced by the NAA treatment, except BrAIL1. The expression level of BrAIL5 was only up-regulated at 3 h. Additionally, six genes were increased at 1 h and 3 h compared with the untreated control, including BrANT-1, BrANT-3, BrAIL6-1, BrAIL6-2, BrAIL6-3 and BrAIL7. However, the expression level of BrANT-2 showed a trend of decrease at 1 h, followed by an increase at 3 h.
Expression profile of the BrAIL family members under naphthaleneacetic acid (NAA) treatments. Plants at the four-leafed stage (21 day after sowing) were treated with 100 μM naphthaleneacetic acid or distilled water (DW) for 0, 1, and 3 h before the leaves at the same position were harvested. Relative expression level was calculated using the 2−ΔCt method. BrActin was used as the reference gene. The expression relative to CK was compared at 0 h.
Over-expression of the BrANT-1 gene caused a variety of phenotypic changes in Arabidopsis
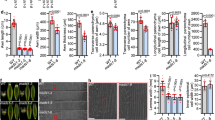
Compared with other BrAIL family members, BrANT-1 had the highest sequence similarity to Arabidopsis thaliana ANT (AT4G37750), while BrANT-1 was different from other AtAIL genes (Supplementary Table S5). Moreover, BrANT-1 was expressed in different tissues, suggesting that BrANT-1 might play an important role in growth and development of Chinese cabbage. To explore the potential function of BrANT-1, Arabidopsis plants were transformed with BrANT-1 under the control of the CaMV 35 S promoter. A series of pleiotropic effects were significantly distinguishable from the wild-type (WT) at 7 days after sowing, with the 35S-BrANT-1 transgenic seedling exhibiting a longer root (Fig. 4A). Additionally, the hypocotyl length of the transgenic seedling also showed a >24% increase compared with that of the WT (Fig. 4C). At 15 days after acclimating to the nutrient soil conditions, the seedling size of the transgenic plants was bigger than that of the WT (Fig. 4E). At 40 days after sowing, the leaf dimension of the transgenic plants was significantly increased compared with that of the WT (Fig. 5). Rachis length, flower dimension, seed size and silique were also enlarged (Fig. 6), while the seed number per silique of the 35S-BrANT-1 transgenic line was slightly more than that in the WT (Fig. 6D). Furthermore, SEM was used to compare the epidermal cells in the adaxial of mature leaves of 35S-BrANT-1 and WT. The results showed the numbers of the adaxial epidermal cells of the 35S-BrANT-1 transgenic line had a >50% increase while the cell size of the 35S-BrANT-1 transgenic line was modestly decreased (<6%) compared with the WT (Fig. 5E). However, the slightly smaller cell size could not account for the big leaf size. Therefore, we conclude that the enlarged leaf area was caused by the increase of the cell number (Fig. 5B). All phenotypic analysis data were shown in Supplementary Table S6.
Phenotype analyses induced by overexpression of BrANT-1 in Arabidopsis. (A) Root length of the WT and 35S-BrANT-1 transgenic plants at 7 day after sowing. Bar = 1 cm. (B,D) Root length and hypocotyl length. (C) Hypocotyl length of the WT and 35S-BrANT-1 transgenic plants at 3 day after germination under darkness condition. Bar = 1 cm. (E) The sizes of individual seedlings of the WT and 35S-BrANT-1 transgenic plants at 15 day after sowing in the nutrient soil. Bar = 1 cm. Each sample was repeated 20 times, and the error bars represent standard deviations.
Morphology and histological analysis of mature leaves of 40 day after sowing. (A) The 12 leaves of the WT and 35S-BrANT-1 transgenic Arabidopsis at 40 day after sowing. Bar = 1 cm. (B) Scanning electron micrographs of the largest rosette leaves of the WT and transgenic 35S-BrANT-1 transgenic Arabidopsis. Bar = 100 μm. (C) The petiole length, leaf length and leaf width of the largest rosette leaves of the WT and transgenic 35S-BrANT-1 transgenic Arabidopsis. Each sample was repeated 20 times. (D) The number of stomata and epidermal cell per unit area (mm2) in the adaxial surface of fully expanded largest leaves of the WT and 35S-BrANT-1 lines. (E) The cell size (μm²) of the epidermal cells in the adaxial surface of fully expanded largest leaves of the WT and 35S-BrANT-1 lines. Each sample was repeated six times, and the error bars represent standard deviations. *Indicates that the expression level is significantly different from that of the control (*p < 0.05, **p < 0.01).
Phenotype analyses of the 35S-BrANT-1 plants induced by the overexpression of BrANT-1 in Arabidopsis (40 day after sowing). (A) The height of the WT and 35S-BrANT-1 transgenic Arabidopsis. bar = 1 cm. (B) Plant height. Each sample was repeated 20 times. (C,E,F) A 35S-BrANT-1 transgenic plant produces large seed, flower and silique (C, bar = 100 μm; E, bar = 1 mm; F, bar = 1 cm). (D) Seed number (per silique). Each sample was repeated 20 times, and the error bars represent standard deviations. *Indicates that the expression level is significantly different from that of the control (*p < 0.05, **p < 0.01); the difference in seed number per silique is not statistically significant.
Additionally, at 40 days after sowing, the transgenic lines had a delayed leaf senescence phenotype, compared with the WT, in that the fifth rosette leaf (numbered from the bottom) became senescent, while the second rosette leaf was still green (Fig. 5A).
In addition, other interesting phenotypes were observed. For example, the stomatal density of the 35S-BrANT-1 line was significantly increased in the mature leaves compared with that of the WT (Fig. 5B, detailed in Supplementary Fig. S3). Consistently, the stomatal conductance (Gs) and transpiration rate (Tr) of the 40-day 35S-BrANT-1 plants were significantly increased compared with those of the WT, while the intracellular CO2 concentration (Ci) and net photosynthetic rate (Pn) between the WT and 35S-BrANT-1 plants were only slightly changed (Fig. 7).
The stomatal number has a positive effect on A. thaliana leaf stomatal conductance (Gs) and transpiration rate (Tr) under the light-saturated condition. (A) Net photosynthetic rate (Pn) and stomatal conductance (Gs); (B) Intracellular CO2 concentration (Ci); (C) Transpiration rate (Tr); (D) Stomatal conductance (Gs). Photosynthetic capacity analysis of mature leaves of the WT and 35S-BrANT-1 transgenic plants (40 day after germination) measured using a GFS-3000 gas exchange system under the light-saturated condition (PPFD = 500 μmol m−2 s−1), and CO2 concentration was set to 400 ppm. Each sample was repeated 20 times, and the error bars represent standard deviations. *Indicates that the expression level is significantly different from that of the control (*p < 0.05).
Transcriptome sequencing of the 35S-BrANT-1 and WT plants
To investigate the genome-wide effects of BrANT-1 on transcription, transcriptome sequencing was carried out for the WT and 35S-BrANT-1 in three independent biological replicates. A total of 533 differentially expressed genes (DEGs) were identified between the WT and 35S-BrANT-1 plants, including 94 up-regulated genes and 439 down-regulated genes (Supplementary Table S7). Additionally, ten DEGs were randomly selected for further verification by qRT-PCR; all genes exhibited the same expression tendency as shown in the original data (Fig. 8). According to the Arabidopsis genome sequence and the DEGs could be assigned to different families, such as MADS-box, TCP, extension, expansin, early auxin-responsive genes (small auxin-up RNA, SAUR), VQ, STOMAGEN, SAGs and transcription factors. The DEGs that were related to plant growth and development were selected and characterized (Table 3). Further analysis indicated many genes were up-regulated in 35S-BrANT-1, which were mainly involved in cell proliferation (MADS-box protein, AT1G59920; TCP21, AT5G08330) and stoma development (STOMAGEN, AT4G12970). On the other hand, the genes inhibiting plant growth (SAUR33, AT3G61900; VQ22, AT3G22160; VQ3, AT1G21326 and ANAC036, AT2G17040) and promoting leaf senescence (AtNAC2, AT5G39610; SUAR36, AT2G45210; SAG13, AT2G29350) were down-regulated in the 35S-BrANT-1 line.
qRT-PCR validation of the gene expression from RNA-Seq analyses. Ten DEGs (JAV1, AT3G22160; SAG13, AT2G29350; VQ10, AT1G78410; ZAT8, AT3G46080; SAUR36, AT2G45210; LRX5, AT4G18670; EXPB1, AT2G20750; STOMAGEN, AT4G12970; TCP21, AT5G08330; POE1;9, AT5G15780) were randomly selected for further verification by qRT-PCR. Relative expression levels were calculated using AtActin (At2g37620) as the reference gene by the method 2−ΔCt. Data presented are mean values of three biological replicates, and the error bars represent standard deviations.
Discussion
AIL genes duplication in Chinese cabbage
The genome of the mesopolyploid crop species Brassica rapa has undergone whole genome triplication (WGT) since divergence from Arabidopsis thaliana, resulting in a significant increase of the numbers of the duplicated genes31. However, only nine AIL family members were identified in Chinese cabbage, including three BrANTs and six BrAILs, which were only 1.5-fold more than those in Arabidopsis, suggesting many genes were lost during genome duplication32. Similar results were also found in other gene families, such as BrGRF and BrVQ genes, which were about 1.89- and 1.9-fold more than those in Arabidopsis, respectively13,33. Additionally, the expansion of the BrAIL family members mainly depended on the segmental duplication, because no tandem duplicated pairs were found. Many studies on genome duplications have shown that the genes involved in transcription, protein binding, response to biotic stimuli and signal transduction path are preferentially retained by segmental duplication34,35,36. Duplication events within a genome can result in paralogs, and these genes may have different expression patterns following duplication indicative of subfunctionalization37. For example, two duplicated genes, BrVQ22-1 and BrVQ22-2, are differentially expressed in different tissues33. In this study, the triplicated genes of BrAIL6-1/-2/-3 and BrANT-1/2/3 also exhibited a different expression pattern in different organs and in response to the auxin treatment. Additionally, similar cases have also been reported in the AP2/ERF family in Chinese cabbage38,39.
BrAIL family members were expressed in various tissues
By quantitative real time PCR, we have shown that most of the BrAIL family members were differentially expressed in both vegetative and reproductive tissues (Fig. 2). Similar to the AtAIL genes, the BrAIL genes had higher expression in young tissues (seedling and roots) and lower expression or absent in mature leaves40. Multiple BrAIL genes were expressed in different tissues, suggesting that the BrAIL genes play different roles in different organs, which has been confirmed in other plants. For example, ANT and AIL6 are involved in the regulation of flower or seed development in Arabidopsis or Medicago truncatula9,17,18 and the VviANT1 gene plays an important role in the regulation of berry size28. Additionally, we observed the BrAIL genes were lowly expressed in blooming flowers. A previous study also found that the AIL genes are mainly expressed in the advanced stage B (B2), flowers from inflorescences at stage G (G), and the early stage H (H1) in grapevine, while the expression of the AIL genes is low in blooming flowers28. Besides, AtANT, AtAIL5, AtAIL6, and AtAIL7 also exhibit the same expression tendency during the flower development40.
Transgenic lines over-expressing BrANT-1 had enlarged organs by increasing the cell number
Generally, the AIL genes play an important role in regulating organ growth through increasing cell number7 or cell size29. In the present study, the cell number in the leaf was increased in the 35S-BrANT-1 transgenic lines, implying that BrANT-1 positively regulates cell proliferation. This result was consistent with the function of the AtANT gene in the leaf or floral organ7. However, it is different from the roles of PnANTL1 and PnANTL2 genes, which increase leaf length through increasing cell size in tobacco29.
Recently, Liu et al.41 found that the OsMADS1 gene can positively regulate cell proliferation in rice. Therefore, the most up-regulated MADS-box gene (AT1G59920) among the DEGs may play critical roles in increasing the cell number in the 35S-BrANT-1 line. Additionally, previous studies have shown that the TCP gene can be divided into two classes (I and II). The class I genes like TCP20 function as positive regulators of cell growth42, while the class II genes like the Antirrhinum genes CINCINNATA (CIN) function as negative regulators of cell growth43. Here, AtTCP21 (AT5G08330), a member of the class I genes44 was found to be up-regulated, suggesting that it may partially promote cell proliferation in the 35S-BrANT-1 line. Apart from the up-regulated genes, some down-regulated genes, such as SAUR36 (AT2G45210)45, VQ22 (AT3G22160)46 and ANAC036 (AT2G17040)47, might also play an important role in increasing organ size in the 35S-BrANT-1 line, as indicated by previous studies, which have shown that some SAUR, VQ and NAC genes are involved in the negative control of plant growth46,47,48,49,50. Interestingly, some genes, such as LRX5 (AT4G18670)51 and expansinB1 (AT2G20750)52, positively controlling cell size were also identified from the up-regulated DEGs, as indicated in our histological results which showed a slight reduce in cell size between the 35S-BrANT-1 and WT plants. The result was consistent with our previous studies on the BrARGOS gene, which regulates the ANT gene10 and may also promote the transcription of AtEXP10 in transgenic Arabidopsis53. This is probably because meristematic competence is disrupted locally, cell division gradually ceases and differentiation begins with the expansion of the postmitotic cells19. In addition, cell proliferation is also coupled with a limited amount of cell expansion during the proliferation phase54. Therefore, we speculate that the extensin and expansin proteins promote cell expansion following a significant increase of cell proliferation. The modest reduction in cell size may compensate the increase in cell number19.
BrANT-1 might regulate leaf senescence
Leaf senescence constitutes the final phase of leaf development and is a highly complex but genetically programmed process involving the expression of many senescence-associated genes (SAGs)55,56. Our study shows that, compared with the wild type, leaf senescence in the 35S-BrANT-1 transgenic line was delayed, which is consistent with that in Arabidopsis over-expressing the AtANT gene57. Additionally, a number of SAGs such as AtNAC2 (AT5G39610)58, AtSAUR36(AT2G45210)45, and AtSAG13(AT2G29350)59 were prominently down-regulated in the 35S-BrANT-1 line. AtSAUR36 (or SAG201), a member of the early auxin-responsive gene family, was remarkably up-regulated during leaf senescence. In Arabidopsis, a saur36 knockout line shows a delayed leaf senescence phenotype, but the transgenic plant overexpressing SAUR36 displays an opposite phenotype45. In addition, SAG13 as an early senescence marker is strongly induced before visible yellowing60. Accordingly, we speculate that BrANT-1, similar to the AtANT gene, is one of the negative factors that prevent premature senescence.
BrANT-1 positively increased the stomatal number in Arabidopsis
Stomata is an important part of the epidermal tissues of leaves in plants, which controls gas exchange by paired subsidiary cells, and participates in the global carbon cycle61. Stomatal development in leaf is positively regulated by signaling factor STOMAGEN (At4g12970) through interacting with cell-surface receptor TOO MANY MOUTHS (TMM)62. In this study, it was found that the number of stoma significantly increased in the 35S-BrANT-1 line compared with the WT. Besides, the expression level of STOMAGEN (At4g12970) was significantly up-regulated in the transgenic line. The overexpression of STOMAGEN was found on many agminate stomata in mature leaves of Arabidopsis62. However, the homologous gene of AtANT regulating the stomatal development has not been reported. Additionally, previous studies have shown that the stomatal density in Arabidopsis influences the leaf photosynthetic capacity through regulating gas diffusion63. In this study, we assessed the leaf photosynthetic capacity of the BrANT-1-overexpressing transgenic and wild type lines. The result indicated that, with the increase of the stomatal number, the stomatal conductance (Gs) and transpiration rate (Tr) of mature leaves were significantly increased in the 35S-BrANT-1 transgenic plants. However, the net photosynthetic rate (Pn) was only slightly increased in the 35S-BrANT-1 plants, probably due to the low intracellular CO2 concentration (Ci). Severe stomatal patchiness results in the underestimation of Pn due to the lack of uniform Ci in the leaf 64. Taken together, these results indicated that BrANT-1 might be involved in regulation of the development of stomata by enhancing the expression level of STOMAGEN (At4g12970).
In summary, we identified three BrANT and six BrAIL proteins in Chinese cabbage, which can be classified into four subgroups. Phylogenetic analysis showed that the AIL genes of Chinese cabbage had high sequence similarity with those of Arabidopsis. Furthermore, multiple sequence alignment suggests that the nine genes belonged to the euANT subgroup according to the conserved motifs in Arabidopsis. Finally, ectopic expression of BrANT-1 in Arabidopsis controlled the organ size by regulating the cell number. BrANT-1 regulated the stomatal density and leaf senescence by increasing the expression of STOMAGEN and reducing the expression of SAGs. Taken together, these results not only enhance the understanding of the role of the AIL genes in controlling the organ size and other tissue growth, but also provide a promising tactic for Chinese cabbage molecular breeding program.
Methods
Identification and analysis of the AIL genes in Chinese cabbage
The gene and amino acid sequences of the AIL family members were confirmed according to the genome of the B. rapa line Chiifu (http://brassicadb.org). The difference of the AIL genes was analyzed using DNAMAN 6.0.40 (Lynnon Biosoft, USA). Gene Structure Display Server (GSDS) (http://gsds.cbi.pku.edu.cn/) was used to perform the intron/exon structure analysis. Phylogenetic trees were constructed based on the amino acid sequences using the neighbor-joining method by MEGA 5.0 software65. The physicochemical properties of the AIL proteins were calculated by using the ProtParam tool (http://web.expasy.org/protparam/). MEME (meme.nbcr.net/meme/intro.html) was used to analyze the conserved motifs of the AIL proteins in Chinese cabbage66. The AP2 domain sequences of the AIL proteins from Chinese cabbage were identified with SMART (http://smart.embl-heidelberg.de/).
Plant materials, growth conditions and plant phytohormone treatments
The wild-type and 35S-BrANT-1 transgenic Arabidopsis plants were pre-treated for 3 days at 4 °C under dark condition before transferring to pots with nutrient media (Sheng Xiang Agricultural Science and Technology Co, China; peat: vermiculite: pearlite = 1: 1: 1). The plants were cultivated in a growth room with a continuous artificial light period of 16 h and a dark period of 8 h, and a constant temperature between 19–23 °C. The Chinese cabbage line “Guangdongzao” was used for all stress experiments, and the growth condition was the same as that for Arabidopsis.
Chinese cabbage young seedlings at the four-leafed stage (21 days after sowing) were used for the phytohormone treatments, during which, the plant leaves were treated with 100 μM naphthaleneacetic acid or distilled water (DW), respectively. Plant leaves were harvested after 0, 1 and 3 h of the phytohormone treatment. For gene expression analysis of BrANTs in different tissues, the plants were pre-treated for 15 days at 4 °C under dark condition to accelerate the transition from the vegetative phase to the reproductive phase. Plant organs were harvested after the plants bloomed (50 days after sowing, including the pre-treated time), including root (R), dwarf stem (DS), old leaf (OL), young leaf (YL), and blooming flower (BFL). RNA preparation was performed for each organ in three biological replicates.
Plant transformation
The complete coding region (1671 bp) of Chinese cabbage ANT cDNA was amplified by using the following primers: BrANT-1-forward: 5′-GTTCTAGAATGAAGTCCTTTTGTGATAATGATG-3′ and BrANT-1-reverse: 5′-TAGTCGACTCAAGAATCAGCCCACGCAGCGAAA-3′. The italic sequences (TCTAGA and GTCGAC) were marked as the restriction sites for XbalI and SalI, respectively. The CAMV 35 S promoter was used instead of the intrinsic promoter. The product of polymerase chain reaction (PCR) was cloned into the binary vector pCAMBIA2300-35SOCS using the XbalI and SalI restriction sites. The constructed vector was used to transform Arabidopsis plants using the planta Agrobacterium-mediated method67 and the transgenic seedlings were selected by 1/2 MS agar plates containing 50 mg/mL kanamycin sulfate. Homozygous transgenic plants were used for further analyses.
RNA isolation and cDNA synthesis
The total RNA of each sample (in three biological replicates) was isolated using Trizol reagent (Invitrogen, Carlsbad CA, USA). The quality and concentration of the total RNA were measured separately using a 2100 Bioanalyzer RNA Nanochip (Agilent, Santa Clara, CA, USA) and a NanoDrop ND-2000 Spectrophotometer (Nano-Drop, Wilmington, DE, USA). The synthesis of cDNA was carried out using a PrimeScript™ RT reagent kit with a gDNA Eraser (Takara, Dalian, China).
Sequencing and data processing
The matured leaves of the WT and 35S-BrANT-1 transgenic Arabidopsis plants (40 days after sowing) were used for RNA-seq. Each line was biologically repeated three times. The sequencing of six cDNA libraries was performed at Beijing Genomics Institute (BGI, Shenzhen, China) using an Illumina HiSeqTM 2000 sequencing platform (Illumina Inc., San Diego, CA, USA). Clean reads were obtained from the raw reads that were clipped by abandoning the adaptor sequences and low-quality reads (reads with >10% ambiguous “N” bases or reads in which >50% of the bases had a Quality-score ≤ 5). The clean reads were mapped to the reference genome using a rapid short-read mapping program, namely SOAP aligner/soap268. More than two mismatches were abandoned in the sequence alignment. The quality of sequencing was controlled by quality assessment of reads, statistics of alignment, sequencing saturation analysis and randomness assessments.
Identification and functional annotation of differentially expressed genes
The reads per kb per Million reads (RPKM) method69 was used to calculate the expression level of each unigene. Therefore, the RPKM values can be directly used for comparing the difference of gene expression among the WT and 35S-BrANT-1 lines. The DEGs from these two lines (six cDNA libraries) were enriched for further analysis according to the standard with false discovery rate (FDR) ≤0.01, and the absolute value of log2 ratio ≥ 1.
Quantitative RT-PCR (qRT-PCR) analysis
To confirm the quality of the sequencing data and distinguish the expression level, the Chinese cabbage AIL genes were subjected to qRT-PCR analysis. qRT-PCR was performed under the following conditions: 94 °C for 2 min, followed by 45 cycles of reaction (94 °C for 20 s, followed by 60 °C for 34 s). The actin gene was used as a constitutive expression control. qRT-PCR was performed on an IQ5 Real-Time PCR System (BIO-RAD, Hercules, CA, USA). The specific primers for qRT-PCR (Supplementary Table S8) were designed using Primer Premier 5.0 (Premier Biosoft International, Palo Alto, CA) and synthesized by Shanghai Sangon Biological Engineering Technology & Services Company (Shanghai, China.). Three replicates were performed for each sample.
Scanning electron microscopy analysis
To observe the morphology of leaf blades, leaf sections were cut from the fully expanded eighth leaf of the WT and 35S-BrANT-1 plants (40 days after sowing), embedded in 100 mM sodium phosphate buffer (pH 7.2) containing 2% glutaraldehyde for 2 h at 4 °C. Then the samples were rinsed for 1 h by the same buffer and dehydrated in a graded ethanol series for 1 h at each gradation. The dehydrated samples were dried in a critical-point dryer with liquid CO2 as the transitional fluid and examined using a scanning electron microscope (SEM; JEOL JSM-7600 F, Japan) after coated with gold. The size and number of the epidermal cells on the adaxial side were determined in the middle region of a half leaf near the midvein. At least six leaves from each of the WT and transgenic plants were selected for cell number counting in a fixed area on the SEM images.
Data availability
The RNA-seq raw data are deposited in the Sequence Read Archive (SRA) under the number “SPR136061”. Phenotype datasets are available in this article and its supplementary files.
References
Sugimoto-Shirasu, K. & Roberts, K. “Big it up”: endoreduplication and cell-size control in plants. Curr. Opin. Plant Biol. 6, 544–553 (2003).
Horiguchi, G., Ferjani, A., Fujikura, U. & Tsukaya, H. Coordination of cell proliferation and cell expansion in the control of leaf size in Arabidopsis thaliana. J. Plant Res. 119, 37–42 (2006).
Powell, A. E. & Lenhard, M. Control of organ size in plants. Curr. Biol. 22, R360–R367 (2012).
Klucher, K. M., Chow, H., Reiser, L. & Fischer, R. L. The AINTEGUMENTA gene of Arabidopsis required for ovule and female gametophyte development is related to the floral homeotic gene APETALA2. Plant Cell 8, 137–153 (1996).
Elliott, R. C. et al. AINTEGUMENTA, an APETALA2-like gene of Arabidopsis with pleiotropic roles in ovule development and floral organ growth. Plant Cell 8, 155–168 (1996).
Krizek, B. A. Ectopic expression of AINTEGUMENTA in Arabidopsis plants results in increased growth of floral organs. Dev. Genet. 25, 224–236 (1999).
Mizukami, Y. & Fischer, R. L. Plant organ size control: AINTEGUMENTA regulates growth and cell numbers during organogenesis. Proc. Natl. Acad Sci. USA 97, 942–947 (2000).
Krizek, B. A. AINTEGUMENTA and AINTEGUMENTA-LIKE6 act redundantly to regulate Arabidopsis floral growth and patterning. Plant Physiol. 150, 1916–1929 (2009).
Krizek, B. A. AINTEGUMENTA and Aintegumenta-Like6 regulate auxin-mediated flower development in Arabidopsis. BMC Res. Notes 4, 176 (2011).
Hu, Y., Xie, Q. & Chua, N. H. The Arabidopsis auxin-inducible gene ARGOS controls lateral organ size. Plant Cell 15, 1951–1961 (2003).
Wang, B., Sang, Y. L., Song, J., Gao, X. Q. & Zhang, X. S. Expression of a rice OsARGOS gene in Arabidopsis promotes cell division and expansion and increases organ size. J. Genet. Genomics 36, 31–40 (2009).
Kim, J. H., Choi, D. & Kende, H. The AtGRF family of putative transcription factors is involved in leaf and cotyledon growth in Arabidopsis. Plant J. 36, 94–104 (2003).
Wang, F. D. et al. Genome-wide identification and analysis of the growth-regulating factor family in Chinese cabbage (Brassica rapa L. ssp. pekinensis). BMC Genomics 31, 1002–1011 (2014).
Shigyo, M. & Hasebe, M. M. Molecular evolution of the AP2 subfamily. Gene 366, 256–265 (2006).
Kim, S., Soltis, P. S., Wall, K. & Soltis, D. E. Phylogeny and domain evolution in the APETALA2-like gene family. Mol. Biol. Evol. 23, 107–120 (2006).
Nole-Wilson, S. & Krizek, B. A. AINTEGUMENTA contributes to organ polarity and regulates growth of lateral organs in combination with YABBY genes. Plant Physiol. 141, 977–987 (2006).
Dash, M. & Malladi, A. The AINTEGUMENTA genes, MdANT1 and MdANT2, are associated with the regulation of cell production during fruit growth in apple (Malus × domestica Borkh.). BMC Plant Biol. 12, 98 (2012).
Confalonieri, M. et al. Seed-Specific Expression of AINTEGUMENTA in Medicago truncatula Led to the Production of Larger Seeds and Improved Seed Germination. Plant Mol. Biol. Rep. 32, 957–970 (2014).
Mizukami, Y. A matter of size: developmental control of organ size in plants. Curr. Opin. Plant Biol. 4, 533–539 (2001).
Wang, X. et al. The Brassica rapa Genome Sequencing Project Consortium: The genome of the mesopolyploid crop species Brassica rapa. Nat. Genet. 43, 1035–1039 (2011).
Mohanta, T. K., Arora, P. K., Mohanta, N., Parida, P. & Bae, H. Identification of new members of the MAPK gene family in plants shows diverse conserved domains and novel activation loop variants. BMC Genomics 16, 58 (2015).
Horstman, A., Willemsen, V., Boutilier, K. & Heidstra, R. AINTEGUMENTA-LIKE proteins: hubs in a plethora of networks. Trends Plant Sci. 19, 146–57 (2014).
Rashid, M., Guangyuan, H., Guangxiao, Y., Hussain, J. & Xu, Y. AP2/ERF Transcription Factor inRice: Genome-Wide Canvas and Syntenic Relationships between Monocots and Eudicots. Evol. Bioinform. Online 8, 321–355 (2012).
Cheng, F. et al. BRAD: the genetics and genomics database for Brassica plants. BMC Plant Biol. 11, 136 (2011).
Rigal, A. et al. The AINTEGUMENTA LIKE1 Homeotic Transcription Factor PtAIL1 Controls the Formation of Adventitious Root Primordia in Poplar. Plant Physiol. 160, 1996–2006 (2012).
Nole-wilson, S., Azhakanandam, S. & Franks, R. G. Polar auxin transport together with aintegumenta and revoluta coordinate early Arabidopsis gynoecium development. Dev. Biol. 346, 181–195 (2010).
Chen, B. et al. Cloning and Expression Level Analysis of Two BnaANT Candidate Genes in Brassica napus. Agr. Sci. China 9, 488–496 (2010).
Chialva, C. et al. Expression of grapevine AINTEGUMENTA-like genes is associated with variation in ovary and berry size. Plant Mol. Biol. 91, 67–80 (2016).
Kuluev, B. R., Knyazev, A. V., Iljassowa, A. A. & Chemeris, A. V. Ectopic Expression of the PnANTL1 and PnANTL2 Black Poplar Genes in Transgenic Tobacco Plants. Russ. J. Genet. 48, 993–1000 (2012).
Santner, A., Calderon-Villalobos, L. & Estelle, M. Plant hormones are versatile chemical regulators of plant growth. Nat. Chem. Biol. 5, 301–307 (2009).
Saha, G. et al. Genome-wide identification and characterization of MADS-box family genes related to organ development and stress resistance in Brassica rapa. BMC Genomics 16, 178 (2015).
Mun, J. H. et al. Genome-wide comparative analysis of the Brassica rapa gene space reveals genome shrinkage and differential loss of duplicated genes after whole genome triplication. Genome Biol. 10, R111 (2009).
Zhang, G. Y. et al. Genome-Wide Identification and Analysis of the VQ Motif-Containing Protein Family in Chinese Cabbage (Brassica rapa L. ssp. Pekinensis). Int. J. Mol. Sci. 16, 28683–28704 (2015).
Seoighe, C. & Gehring, C. Genome duplication led to highly selective expansion of the Arabidopsis thaliana proteome. Trends Genet. 20, 461–464 (2004).
Blanc, G. & Wolfe, K. H. Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes. Plant Cell. 16, 1667–1678 (2004).
Maere, S. et al. Modeling gene and genome duplications in eukaryotes. Proc. Natl. Acad Sci. USA 102, 5454–5459 (2005).
Duarte, J. M. et al. Expression pattern shifts following duplication indicative of subfunctionalization and neofunctionalization in regulatory genes of Arabidopsis. Mol. Biol. Evol. 23, 469–478 (2006).
Song, X. M., Li, Y. & Hou, X. L. Genome-wide analysis of the AP2/ERF transcription factor superfamily in Chinese cabbage (Brassica rapa ssp. pekinensis). BMC Genomics 289, 77 (2014).
Liu, Z. N. et al. Genome-Wide Identification, Phylogeny, Evolution and Expression Patterns of AP2/ERF Genes and Cytokinin Response Factors in Brassica rapa ssp. pekinensis. PLoS One 8, e83444 (2013).
Nole-Wilson, S., Tranby, T. & Krizek, B. A. AINTEGUMENTA-like (AIL) genes are expressed in young tissues and may specify meristematic or division-competent states. Plant Mol. Biol. 57, 613–628 (2005).
Liu, Q. et al. G-protein βγ subunits determine grain size through interaction with MADS-domain transcription factors in rice. Nat. Commun. 2018, 852 (2018).
Li, C., Potuschak, T., Colon-Carmona, A., Gutierrez, R. A. & Doerner, P. Arabidopsis TCP20 links regulation of growth and cell division control pathways. Proc. Natl. Acad. Sci. USA 102, 12978–12983 (2005).
Crawford, B. C., Nath, U., Carpenter, R. & Coen, E. CINCINNATA controls both cell differentiation and growth in petal lobes and leaves of Antirrhinum. Plant Physiol 135, 244–253 (2004).
Aguilar-Martínez, J. A. & Neelima, S. Analysis of the role of Arabidopsis class I TCP genes AtTCP7, AtTCP8, AtTCP22, and AtTCP23 in leaf development. Front. Plant. Sci. 4, 406 (2013).
Hou, K., Wu, W. & Gan, S. S. SAUR36, a SMALL AUXIN UP RNA gene, is involved in the promotion of leaf senescence in Arabidopsis. Plant Physiol. 161, 1002–1009 (2013).
Cheng, Y. et al. Structural and functional analysis of VQ motif-containing proteins in Arabidopsis as interacting proteins of WRKY transcription factors. Plant Physiol. 159, 810–825 (2012).
Kato, H., Motomura, T., Komeda, Y., Saito, T. & Kato, A. Overexpression of the NAC transcription factor family gene ANAC036 results in a dwarf phenotype in Arabidopsis thaliana. J. Plant Physiol. 167, 571 (2010).
Kant, S., Bi, Y. M., Zhu, T. & Rothstein, S. J. SAUR39, a Small auxin-up RNA gene, acts as a negative regulator of auxin synthesis and transport in rice. Plant Signaling & Behavior. 151, 691–701 (2009).
Xu, Y. X. et al. The small auxin-up RNA OsSAUR45 affects auxin synthesis and transport in rice. Plant Mol. Biol. 94, 1–11 (2017).
Gargul, J. M., Mibus, H. & Serek, M. Manipulation of MKS1 gene expression affects Kalanchoe blossfeldiana and Petunia hybrida phenotypes. Plant Biotechnol. J. 13, 51–61 (2015).
Draeger, C. et al. Arabidopsis leucine-rich repeat extensin (LRX) proteins modify cell wall composition and influence plant growth. BMC Plant Biol. 15, 155 (2015).
Li, X., Zhao, J., Tan, Z., Zeng, R. & Liao, H. GmEXPB2, a cell wall β-Expansin gene, affects soybean 2 nodulation through modifying root architecture and promoting 3 nodule formation and development. Plant Physiol. 169, 2640–2653 (2015).
Wang, B., Zhou, X., Xu, F. & Gao, J. Ectopic expression of a Chinese cabbage BrARGOS gene in Arabidopsis increases organ size. Transgenic Res. 19, 461–472 (2010).
Anastasiou, E. & Lenhard, M. Control of plant organ size. Springer Berlin Heidelberg. 10, 25–45 (2008).
Michele, R. D. et al. Transcriptome analysis of Medicago truncatula leaf senescence: similarities and differences in metabolic and transcriptional regulations as compared with Arabidopsis, nodule senescence and nitric oxide signalling. New Phytol. 181, 563–575 (2009).
Guo, Y. F. & Gan, S. S. Convergence and divergence in gene expression profiles induced by leaf senescence and 27 senescence-promoting hormonal, pathological and environmental stress treatments. Plant Cell and Environ. 35, 644–655 (2012).
Feng, G., Xu, Q., Wang, Z. & Zhuoma, Q. AINTEGUMENTA negatively regulates age-dependent leaf senescence downstream of AUXIN RESPONSE FACTOR 2 in Arabidopsis thaliana. Plant Biotechnol. J. 33, 71–76 (2016).
Balazadeh, S. et al. A gene regulatory network controlled by the NAC transcription factor ANAC092/AtNAC2/ORE1 during salt-promoted senescence. Plant J. 62, 250–264 (2010).
Weaver, L. M., Gan, S., Quirino, B. & Amasino, R. M. A comparison of the expression patterns of several senescence associated genes in response to stress and hormone treatment. Plant Mol. Biol. 37, 455–469 (1998).
Schippers, J. H. M. et al. The Arabidopsis onset of leaf death5 mutation of quinolinate synthase affects nicotinamide adenine dinucleotide biosynthesis and causes early ageing. Plant Cell 20, 2909–2925 (2008).
Hetherington, A. M. & Woodward, F. I. The role of stomata in sensing and driving environmental change. Nature 424, 901–908 (2003).
Sugano, S. S. et al. Stomagen positively regulates stomatal density in Arabidopsis. Nature 463, 241–244 (2010).
Tanaka, Y., Sugano, S. S., Shimada, T. & Hara-Nishimura, I. Enhancement of leaf photosynthetic capacity through increased stomatal density in Arabidopsis. New Phytol. 198, 757–764 (2013).
Laisk, A. Calculation of Leaf Photosynthetic Parameters Considering the Statistical Distribution of Stomatal Apertures. J. Exp. Bot. 34, 1–5 (1983).
Tamura, K. et al. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Bailey, T. L., Williams, N., Misleh, C. & Li, W. W. MEME: Discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res. 34, W369–W373 (2006).
Bechtold, N., Ellis, J. & Pelletier, G. In planta Agrobacterium mediated gene transfer by infiltration of adult Arabidopsis thaliana plants. Methods Mol. Biol. 82, 259–266 (1993).
Li, R. Q. et al. SOAP2: An improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Mortazavi, A., Williams, B. A., Mccue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Acknowledgements
This study was supported by the National Natural Science Foundation of China [31471884]; the Modern Agricultural Industrial Technology System Funding of Shandong Province, China [SDAIT-05-04]; the National Key R&D Program of China [Pilot Project on Seven Main Crop Breeding]; the Natural Science Foundation of Shandong Province, China [ZR2016CB16]; the Young Talents Training Program of Shandong Academy of Agricultural Science, China [NKYSCS-01]; Taishan Scholar Program of Vegetable Genomics, China [2016–2020]; Agricultural scientific and technological innovation project of Shandong Academy of Agricultural Sciences, China [CXGC2016A02]; The National Key Research and Development Program of China [2017YFD0101801]; The Young Talents Training Program of Shandong Academy of Agricultural Science, China (NKYSCS-03).
Author information
Authors and Affiliations
Contributions
Q.D., F.D.W. and J.W.G. conceived and designed the experiments. Q.D. and F.D.W. performed the experiments. Q.D., J.J.L. and F.D.W. analyzed the data. B.C., Y.H.Z., H.Y.L., N.W.Q., L.F.L. and X.H.L. contributed reagents/materials/analysis tools. Q.D., F.D.W. and J.W.G. wrote the main manuscript text. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ding, Q., Cui, B., Li, J. et al. Ectopic expression of a Brassica rapa AINTEGUMENTA gene (BrANT-1) increases organ size and stomatal density in Arabidopsis. Sci Rep 8, 10528 (2018). https://doi.org/10.1038/s41598-018-28606-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-28606-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.