Abstract
Mutations in multiple RNA/DNA binding proteins cause Amyotrophic Lateral Sclerosis (ALS). Included among these are the three members of the FET family (FUS, EWSR1 and TAF15) and the structurally similar MATR3. Here, we characterized the interactomes of these four proteins, revealing that they largely have unique interactors, but share in common an association with U1 snRNP. The latter observation led us to analyze the interactome of the U1 snRNP machinery. Surprisingly, this analysis revealed the interactome contains ~220 components, and of these, >200 are shared with the RNA polymerase II (RNAP II) machinery. Among the shared components are multiple ALS and Spinal muscular Atrophy (SMA)-causative proteins and numerous discrete complexes, including the SMN complex, transcription factor complexes, and RNA processing complexes. Together, our data indicate that the RNAP II/U1 snRNP machinery functions in a wide variety of molecular pathways, and these pathways are candidates for playing roles in ALS/SMA pathogenesis.
Similar content being viewed by others
Introduction
The neurodegenerative disease Amyotrophic Lateral Sclerosis (ALS) has no known treatment, and elucidation of disease mechanisms is urgently needed. This problem has been especially daunting, as mutations in greater than 30 genes are ALS-causative, and these genes function in numerous cellular pathways1. These include mitophagy, autophagy, cytoskeletal dynamics, vesicle transport, DNA damage repair, RNA dysfunction, apoptosis, and protein aggregation2,3,4,5,6. The discovery that mutations in two RNA/DNA binding proteins, FUS and TARDBP, are ALS-causative first raised the possibility that dysfunction of RNA-related processes plays a role in the disease7,8,9,10,11. This hypothesis gained traction when additional ALS-causative RNA/DNA binding proteins (EWSR1, TAF15, HNRNPA1, HNRNPA2B1, MATR3 and TIA1) were identified12,13,14,15,16. At present, however, the roles of these proteins in ALS pathogenesis are not known.
FUS, EWSR1 and TAF15 constitute the FET family of structurally related proteins17,18. They share in common RNA binding motifs and low complexity domains. Similar to the FET family members, MATR3 also contains both types of domains19. Although ample evidence exists that all four of these ALS-causative proteins function in transcription and splicing, much less is known about how their functions are distinguished from one another in these processes. We recently found that the four ALS-causative proteins associate with the RNAP II machinery and that several other ALS-causative proteins, including HNRNPA120, HNRNPA2B120, TIA116 and VCP21, do as well (BC et al., submitted). Moreover, multiple proteins that are mutated in the childhood motor neuron disease cause Spinal Muscular Atrophy (SMA) associate with the RNAP II machinery, including SMN1, EXOSC822, HSPB123,24 and two components (ASCC1 and TRIP4)25,26 of the ASC-1 transcriptional co-activator (BC et al., submitted). To investigate the roles of ALS-causative proteins within the RNAP II machinery, we used CRISPR to knock out the 3 FET family members or MATR3 in HeLa cells and then characterized the RNAP machinery isolated from these cell lines. One of the notable conclusions from this study was that all four ALS-causative proteins are required for interaction of the SMA-causative ASC-1 complex with RNAP II (BC et al., submitted). The observation that two different components of the ASC-1 complex are mutated to cause SMA and that the ALS-causative proteins mediate the association of the ASC-1 complex with RNAP II provide excellent examples of the importance of identifying interaction partners of ALS/SMA-causative proteins, as these interaction partners themselves are candidates for causing the diseases. In addition, identification of their interaction partners will assist in identifying molecular pathways involved in the pathogenesis of motor neuron disease.
In the present study, we report the interactomes of FUS, EWSR1, TAF15 and MATR3, and show that all four of these proteins associate with U1 snRNP. Unexpectedly, comparison of the interactome of the U1 snRNP machinery with that of the RNAP II machinery shows that virtually the entire U1 snRNP machinery overlaps with the RNAP II machinery. Among the proteins present in the U1 snRNP/RNAP II machinery are multiple ALS/SMA-causative proteins. These data raise the possibility that the RNAP II/U1 snRNP machinery and the pathways in which it functions may underlie the pathogenesis caused by a host of motor neuron disease-causative proteins.
Results and Discussion
FUS, EWSR1, TAF15 and MATR3 associate with U1 snRNP
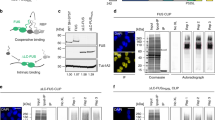
To characterize the interactomes of FUS, EWSR1, TAF15 and MATR3 (hereafter referred to as ALS proteins) we immunopurified (IP’d) these proteins from HeLa cell nuclear extracts. To identify the highly abundant interactors, we excised individual bands from a Coomassie-stained gel and carried out mass spectrometry. This analysis revealed that U1 snRNP components are enriched in the FUS, EWSR1 and TAF15 IPs. These components include all of the U1 snRNP-specific proteins (SNRNP70, SNRPA, SNRPC) as well as the snRNP core proteins (SNRP proteins) (Fig. 1a, lanes 1–3). U1 snRNP components were not observed in the MATR3 IP (Fig. 1a, lane 4). We next carried out reciprocal IP/westerns using an antibody against the SNRPC core component of U1 snRNP. As shown in Fig. 1b, the three FET family members and MATR3 co-IP’d with U1 snRNP, but not with the negative control nuclear protein EIF4A3. Although U1 components were not detected on the Coomassie gel in the MATR3 IP, this may be due to a buried epitope (see below for mass spectrometry data of the U1 snRNP machinery that support this conclusion).
FET proteins and MATR3 associate with U1 snRNP. (a) Immunoprecipitations (IPs) were carried out with antibodies to FET proteins or MATR3 followed by analysis on a Coomassie-stained gel. Molecular weight markers and protein identified by mass spectrometry are indicated. (b) IPs were carried out from nuclear extract using a negative control antibody (EIF4A3) or an antibody to the SNRPC subunit of the U1 snRNP followed by Westerns with the indicated antibodies. (c) IPs were carried out with the indicated antibodies from nuclear extract treated with a U1 snRNA AMO or a negative control AMO followed by Western using the SNRPC antibody. (d) Same as (c) except that total RNAs from the IPs were examined on a denaturing gel stained with ethidium bromide.
To determine whether the association between the ALS proteins and U1 snRNP was specific, we treated nuclear extracts with an anti-sense morpholino (AMO) that binds to the 5’ end of U1 snRNA and blocks splicing27. This U1 AMO also disrupted the association of FUS with U1 snRNP28. We obtained the same results in the present study (Fig. 1c, lanes 5–8). In addition, the U1 AMO disrupted the interactions between U1 snRNP and TAF15, EWSR1 and MATR3 (Fig. 1c, lanes 9–14). We further confirmed these associations by carrying out IPs and analyzing total RNA on an ethidium bromide stained gel. As shown in Fig. 1d, all of the ALS proteins co-IP’d with U1 snRNA, and the interaction was specific as it was disrupted by the U1 AMO. We conclude that FUS, TAF15, EWSR1 and MATR3 associate with U1 snRNP.
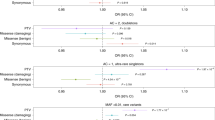
Top hit interactors in the FET proteins and MATR3 interactomes. The top ranked (by total peptide number) proteins in each interactome are shown. The rank, HGNC official symbol, calculated molecular weight, best-known function, total and unique peptide counts are shown. Functions in splicing (pink), transcription (txn, orange), DNA damage response (green), neuronal (blue) and other (black) are indicated. The symbols of ALS-causative proteins are in red. The stars indicate U1 snRNP components.
FUS, EWSR1, TAF15 and MATR3 interactomes
To gain further insight into the interactomes of the four ALS-causative proteins, we carried out shotgun mass spectrometry of each IP. These data revealed 156, 68, 132 and 91 interactors for FUS, EWSR1, TAF15 and MATR3, respectively (Table S1). We listed the well-known functions and/or functions potentially relevant to motor neuron disease in the table for all of the interactors (color coded in Table S1). The top 15 hits in each interactome are shown in Fig. 2. Consistent with the Coomassie gel, canonical U1 snRNP components are among the top hits in the FUS, EWSR1 and TAF15 interactomes (marked by stars). In the EWSR1 interactome, multiple ALS-causative proteins are present (color coded red, Fig. 2). A top interactor of TAF15 is PABPC1, which is known to be mislocalized in ALS patient motor neurons29. A top hit in both the FUS and TAF15 interactomes is HNRNPR, which interacts with SMN1 in the axons of motor neurons30,31. HNRNPR is also found in the EWSR1 and MATR3 interactomes (ranked 34 and 19, respectively in Table S1). The observation that HNRNPR is a common interactor of ALS proteins and SMN1 reveals a new molecular link between ALS and SMA. Among the top hits in the MATR3 interactome are numerous proteins that function in the DNA damage response, including NUMA1, CAD, RUVBL1, RUVBL2, HNRNPK and DDB1. Consistent with these results, MATR3 itself is involved in the DNA damage response32, which has emerged as a pathway disrupted in multiple types of ALS and SMA33,34,35,36,37,38,39. Moreover, both RUVBL1 and RUVBL2 are components of the HSP90/R2TP chaperone complex, which interacts with the SMN complex and functions in facilitating snRNP assembly40,41, suggesting an involvement of MATR3 in snRNP assembly.
To identify complexes in each of the interactomes, we analyzed the data in Table S1 using the STRING database (https://string-db.org). This analysis showed that each of the interactomes contain numerous distinct complexes. Several well-known complexes were not separated into clusters by STRING. Thus, we manually clustered these proteins (Figs 3–6). These data revealed complexes in common among the ALS protein interactomes so we next analyzed the proteins shared by all four ALS interactomes (Fig. S1). These interactomes share three dead box helicases (DHX9, DDX5 and DDX17), which, similar to the ALS proteins, are DNA/RNA binding proteins with roles in transcription and splicing. The DBIRD complex, reported to function in coupling transcription to alternative splicing42, as well as hnRNP proteins, are also shared by the four interactomes. The factors that are shared by the 4 ALS protein interactomes are good candidates for being disease-relevant. Consistent with this possibility, several ALS-causative proteins are also shared among the 4 interactomes, including HNRNPA1, HNRNPA2B1, FUS and MATR3.
Protein-protein interaction network of the FUS interactome. The network of the FUS interactome constructed using STRING database (confidence score >0.7) is shown. Disconnected nodes are omitted. Protein complexes and clusters of functionally related proteins are indicated by yellow circles and blue circles, respectively. The ALS and SMA-causative proteins are in red.
Protein-protein interaction network of the EWSR1 interactome. Same as Fig. 3, except that the network of the EWSR1 interactome is shown. The ALS and SMA causative proteins are in red.
Protein-protein interaction network of the TAF15 interactome. Same as Fig. 3, except that the network of the TAF15 interactome is shown. The ALS causative proteins are in red.
Protein-protein interaction network of the MATR3 interactome. Same as Fig. 3, except that the network of the MATR3 interactome is shown. The ALS causative proteins are in red.
The data in Figs 3–6 reveal complexes/factors unique to one or more of the ALS protein interactomes. FUS and TAF15 interactomes contain the SMN complex, thereby linking both of these ALS proteins to the SMA-causative SMN1 protein (Figs 3 and 5). The FUS and TAF15 interactomes also contain the transcription elongation factor P-TEFb, and the FUS interactome uniquely contains many other factors/complexes that function in transcription, including subunits of RNAP II, and BAF, PAF and integrator complex components (Fig. 3). The TAF15 interactome also contains the TREX mRNA export complex, which has been tied to ALS previously43,44,45,46. In addition, ASAP RNA processing/apoptosis complex is present in the TAF15 interactome, and apoptosis is a pathway associated with ALS (Fig. 5)47,48. Finally, the MATR3 interactome contains CUL1 and CUL2, factors that are components of the ubiquitin proteasome pathway. This pathway is known to be important in ALS via genes such as UBQLN2, which is mutated to cause the disease (Fig. 6)49,50. We conclude that the four ALS-causative proteins have multiple interaction partners, many of which are linked to different pathways involved in ALS/SMA, and these interaction partners are new candidates for factors involved in pathogenesis via these known pathways. Moreover, on a more basic science note, our data reveal that, despite the structural similarities and their common association with U1 snRNP, the four ALS proteins have many unique interaction partners that likely explain the distinct roles that these proteins have been reported to play in such processes as transcription and splicing.
The U1 snRNP machinery overlaps extensively with the RNAP II machinery
In light of the observation that FUS, EWSR1, TAF15 and MATR3 all associate with U1 snRNP, we next investigated the interactome of this machinery, identifying 226 proteins within it. These proteins and their functions are color coded in Table S2. As expected, the top two hits are core components (SNRNP70 and SNRPA) of U1 snRNP and the other core component (SNRPC), which is low molecular weight, is 48th on the list. Numerous SRSFs and the snRNP core proteins (SNRPs) that are known U1 snRNP components are also in the interactome. We next investigated the U1 interactome using STRING (Fig. 7). Unexpectedly, this analysis revealed numerous complexes not typically associated with the role of U1 snRNP as a canonical splicing factor. Indeed, the U1 snRNP interactome contained many complexes that we recently identified in the interactome of the RNAP II machinery. The latter machinery contains 274 proteins (BC et al., submitted). Thus, we next directly compared the U1 snRNP and RNAP II machineries to one another. Remarkably, as shown in the Venn diagram, we found that virtually the entire U1 snRNP machinery interactome (>90%) overlaps with the RNAP II machinery interactome (Fig. 8a). The extent of the overlap between the U1 snRNP and RNAP II machineries is exemplified by the observation that core U1 snRNP components (SNRNP70, SNRPC, SNRNA) and SNRP core proteins are among the most abundant components of both the U1 snRNP and RNAP II machineries (Table S2 and BC et al., submitted). The abundance of the U1 snRNP components is readily apparent on a Commassie stained gel in which high levels of these components can be seen in both the U1 snRNP and RNAP II machineries (Fig. S2). Moreover, consistent with our observation that that the two machineries overlap, we found that RNAP II elutes in the same fractions as U1 snRNP components (SNRPA and SNRPC) in the high molecular weight region of a gel filtration column (Fig. 8b).
Protein-protein interaction network of the U1 snRNP machinery. Same as Fig. 3, except that the network of the U1 snRNP interactome is shown.
The U1 snRNP machinery overlaps with the RNAP II machinery. (a) Venn diagram showing overlap of the U1 snRNP and RNAP II machineries. (b) HeLa cell nuclear extract was separated on a Sephacryl-S500 column. The indicated fractions were used for Western analyses with antibodies against RNAP II and U1 snRNP components (SNRPA and SNRPC). Fraction 25 is the void volume and 69 is the included volume. (c) IPs were carried out from nuclear extract using an antibody to the POLR2A subunit of the RNAP II (left panel) or an antibody to the SNRPC subunit of the U1 snRNP (right panel) as well as a negative control antibody (EIF4A3) followed by Westerns with antibodies to the DEAD box helicases (DHX9, DDX5 and DDX17). (d) IPs were carried out with the indicated antibodies from nuclear extract treated with a U1 snRNA AMO or a negative control AMO followed by Western using the SNRPC antibody.
To validate the association of components of the U1 snRNP machinery with the RNAP II machinery, we carried out IP/Westerns. As shown in Fig. 8c, three DEAD box helicases, DHX9, DDX5, and DDX17, which are shared by both machineries all co-IP with both RNAP II and U1 snRNP, but not with the negative control EIF4A3. In addition, reciprocal IPs showed that all three proteins co-IP with U1 snRNP, and this association is specific as it is completely disrupted by the U1 AMO (Fig. 8d). Although the U1 snRNP machinery is highly abundant in RNAP II IPs, we do not observe reciprocal IPs of the RNAP II machinery using antibodies against U1 snRNP. One possible explanation for this is that the epitope on U1 snRNP that is recognized by the antibody is buried within the RNAP II machinery. We note that antibodies against the two other U1 snRNP core components also do not co-IP the RNAP machinery.
In addition to the U1 snRNP components and the DEAD box helicases, the two machineries have numerous complexes in common. Included among these are transcription complexes (P-TEFb, the BAF complex, and the ASC-1 complex), the SMN complex, the tRNA ligase complex, the NONO-SFPQ complex, the DNA-PK complex and the ASAP complex. The RNAP II and U1 snRNP machineries also share numerous proteins that are ALS or SMA causative. Both the ASC-1 complex and SMN complex contain SMA-causative proteins (ASCC1 and TRIP4, and SMN1, respectively). The other shared SMN-causative proteins are HSPB1 and EXOSC8. Multiple ALS-causative proteins are also in common, including FUS, EWSR1, TAF15, MATR3, TIA1, HNRNPA1, and HNRNPA2B1. Thus, the RNAP II/U1 snRNP machinery is clearly extensively associated with motor neuron disease-causative proteins.
There are 24 proteins unique to the U1 snRNP interactome. Among these are key components of the 3’ end formation machinery, including CPSF1, which binds to the AAUAAA polyadenylation signal and CPSF3, which is the endonuclease that cleaves the transcript prior to addition of the polyA tail51,52,53. Previous studies showed that the binding of U1 snRNP to 5’ splice sites blocks 3’ end formation at cryptic polyA sites in a process known as telescripting54. The mechanisms behind this are not understood. Our observation that critical 3’ end formation factors associate with U1 snRNP raises the possibility that these factors are the targets for U1 snRNP during telescripting.
70 proteins are specific to the RNAP II machinery, including the subunits of RNAP II itself as well as general transcription factors (e.g. TFIIF and NELF complex). It is not clear why some transcription factors are shared by the U1 snRNP and RNAP II machineries, whereas others are specific to RNAP II. One possibility is that the shared transcription factors are involved in coupling between transcription and splicing. It is well known that transcription by RNAP II potently enhances splicing, and our previous work indicated that this enhancement is due to the association of U1 snRNP with the RNAP II machinery, which allows efficient recruitment of U1 snRNP to 5’ splice sites during pre-mRNA synthesis28,55,56. There is also evidence that reciprocal coupling occurs in which splicing enhances transcription, but the mechanisms involved in this coupling are less well understood57. Our observation that the U1 snRNP machinery associates extensively with transcription factors and with the RNAP II machinery suggests that these interactions are involved in the reciprocal coupling. In particular, HTATSF1, which associates with p-TEFb, was previously identified as a factor involved in the reciprocal coupling57, and both factors are present in the U1 snRNP and RNAP II machinery interactomes. Thus, the association between the RNAP II and U1 snRNP machinery may be the molecular mechanism for bi-directional coupling between transcription and splicing.
As mentioned above, the RNAP II/U1 snRNP machinery contains numerous motor-neuron disease causative proteins and thus the processes in which the RNAP II/U1 snRNP machinery functions are candidates for pathways involved in the pathogenesis of motor neuron disease. These pathways include transcription, splicing, reciprocal coupling of transcription and splicing, snRNP biogenesis and DNA repair. In addition, the unexpected association of other factors, such as the ASAP complex and the tRNA ligase complex with the RNAP II/U1 snRNP machinery raises the possibility that other pathways are involved in the pathogenesis of ALS/SMA.
Material and Methods
Plasmids and Antibodies
The monoclonal antibodies used in this study were 8WG16 (against POLR2A, the large subunit of RNAP II) from Biolegend (cat # 920102), SNRPC from Sigma (cat # SAB4200188), DHX9 from Abcam (cat # ab26271), SNRPA (cat # sc-101149), DDX5 (cat # sc-166167), DDX17 (cat # sc-86409) from Santa Cruz. The polyclonal antibodies were FUS (cat # A300–293A), EWSR1 (cat # A300–418A) and MATR3 (cat # A300–591A) from Bethyl, TAF15 from Novus (cat # NB100–567), DDX17 (cat # sc-86409) from Santa Cruz, and DHX9 (cat # ab 26271) from Abcam. Our rabbit polyclonal antibodies to THOC2, EIF4A3, and DDX39B have been described58,59.
Immunoprecipitations (IPs)
For IPs, antibodies were coupled to Protein A Sepharose beads (GE healthcare) and covalently cross-linked using dimethylpimelimidate (Sigma). Reaction mixtures (1 ml) contains 300 μl of HeLa nuclear extract60, 300 μl of SDB (20 mM HEPES, pH 7.9, 100 mM KCl), 500 μM ATP, 3.2 mM MgCl2 and 20 mM creatine phosphate. The mixtures were incubated for 30 min at 30 °C to turn over endogenous complexes in the nuclear extract56. Reaction mixtures were then added to 500 μl of buffer A (1X PBS, 0.1% Triton, 0.2 mM PMSF, protease inhibitor EDTA-free [Roche]) and 40 μl of antibody-crosslinked beads. The IPs were carried out overnight at 4 °C. After five washes with buffer A, proteins were eluted at room temperature using 80 μl of protein gel loading buffer (125 mM Tris, 5% SDS, 20% glycerol, 0.005% Bromophenol blue). After elution, DTT was added to a final concentration of 40 mM, and 15 μl of each eluate was analyzed on a 4–12% SDS-PAGE gradient gel (Life technologies). AMO treatment was performed by adding control AMO (5′-CCTCTTACCTCAGTTACAATTTATA-3′) or U1 AMO (5′-GGTATCTCCCCTGCCAGGTAAGTAT-3′)27,28 to HeLa nuclear extract to a final concentration of 12 μM before IP. For analysis of total RNAs in the IPs, beads were treated with proteinase K for 10 min at 37 °C, and RNAs were recovered by phenol/chloroform extraction and ethanol precipitation. RNAs were run on 8% denaturing polyacrylamide gels and stained with Ethidium Bromide.
Mass Spectrometry
To identify the interactomes of FUS, EWSR1, TAF15 and MATR3, the IP samples were trichloroacetic acid (TCA) precipitated and subjected to shotgun mass spectrometry. The total peptide number of the antigen in each IP was set as 100 and the relative peptide numbers of each interactors are shown in Table S1. Abundant cytoplasmic proteins, ribosomal proteins, proteins greater than 250 kDa, and proteins for which the relative total peptide number is smaller than 5 were omitted. For mass spectrometry of the U1 snRNP machinery, the IP was TCA precipitated and the digested peptides were labeled by tandem mass tag61 for MS3 analysis on an Orbitrap Fusion mass spectrometer coupled to a Proxeon EASY-nLC 1000 liquid chromatography (LC) pump (Thermo Scientific). Abundant cytoplasmic proteins, ribosomal proteins, proteins greater than 200 kDa with less than 10 spectral counts, proteins greater than 70 kDa with less than 4 spectral counts, and proteins with one spectral count were not included in Table S2. The proteins in Tables S1 and S2 were annotated with functions using the Gene Cards database (www.genecards.org)62 and/or searching the literature. To compare the U1 snRNP machinery and the RNAP II machinery, the quantitative mass spectrometry data of the WT RNAP II machinery (BC et al., submitted) was filtered using the same criteria used for the U1 snRNP machinery as mentioned above.
Gel filtration
A reaction mixture containing 300 μl of HeLa nuclear extract60, 300 μl of SDB (20 mM HEPES, pH 7.9, 100 mM KCl), 500 μM ATP, 3.2 mM MgCl2 and 20 mM creatine phosphate was incubated for 30 min at 30 °C. After incubation, the mixture was separated on a Sephacryl S500 (GE Healthcare) gel filtration column. The gel filtration column buffer contains 20 mM HEPES, 60 mM KCl, 2.5 mM EDTA and 0.1% Triton X-100.
Data availability statement
The materials and datasets generated during and/or analyzed during the current study are available from the corresponding author upon request.
References
Peters, O. M., Ghasemi, M. & Brown, R. H. Jr. Emerging mechanisms of molecular pathology in ALS. The Journal of clinical investigation 125, 2548, https://doi.org/10.1172/JCI82693 (2015).
Jovicic, A., Paul, J. W., III & Gitler, A. D. Nuclear transport dysfunction: a common theme in amyotrophic lateral sclerosis and frontotemporal dementia. Journal of neurochemistry, https://doi.org/10.1111/jnc.13642 (2016).
Gao, F. B., Almeida, S. & Lopez-Gonzalez, R. Dysregulated molecular pathways in amyotrophic lateral sclerosis-frontotemporal dementia spectrum disorder. EMBO J, https://doi.org/10.15252/embj.201797568 (2017).
Paez-Colasante, X., Figueroa-Romero, C., Sakowski, S. A., Goutman, S. A. & Feldman, E. L. Amyotrophic lateral sclerosis: mechanisms and therapeutics in the epigenomic era. Nat Rev Neurol 11, 266–279, https://doi.org/10.1038/nrneurol.2015.57 (2015).
Taylor, J. P., Brown, R. H. Jr. & Cleveland, D. W. Decoding ALS: from genes to mechanism. Nature 539, 197–206, https://doi.org/10.1038/nature20413 (2016).
Li, Y. R., King, O. D., Shorter, J. & Gitler, A. D. Stress granules as crucibles of ALS pathogenesis. The Journal of cell biology 201, 361–372, https://doi.org/10.1083/jcb.201302044 (2013).
Kwiatkowski, T. J. Jr. et al. Mutations in the FUS/TLS gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science 323, 1205–1208, https://doi.org/10.1126/science.1166066 (2009).
Vance, C. et al. Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323, 1208–1211, https://doi.org/10.1126/science.1165942 (2009).
Sreedharan, J. et al. TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 319, 1668–1672, https://doi.org/10.1126/science.1154584 (2008).
Van Deerlin, V. M. et al. TARDBP mutations in amyotrophic lateral sclerosis with TDP-43 neuropathology: a genetic and histopathological analysis. Lancet neurology 7, 409–416, https://doi.org/10.1016/S1474-4422(08)70071-1 (2008).
Ratti, A. & Buratti, E. Physiological Functions and Pathobiology of TDP-43 and FUS/TLS proteins. Journal of neurochemistry, https://doi.org/10.1111/jnc.13625 (2016).
Ticozzi, N. et al. Mutational analysis reveals the FUS homolog TAF15 as a candidate gene for familial amyotrophic lateral sclerosis. Am J Med Genet B Neuropsychiatr Genet 156B, 285–290, https://doi.org/10.1002/ajmg.b.31158 (2011).
Couthouis, J. et al. Feature Article: From the Cover: A yeast functional screen predicts new candidate ALS disease genes. Proc Natl Acad Sci USA 108, 20881–20890, https://doi.org/10.1073/pnas.1109434108 (2011).
Couthouis, J. et al. Evaluating the role of the FUS/TLS-related gene EWSR1 in amyotrophic lateral sclerosis. Hum Mol Genet 21, 2899–2911, https://doi.org/10.1093/hmg/dds116 (2012).
Johnson, J. O. et al. Mutations in the Matrin 3 gene cause familial amyotrophic lateral sclerosis. Nat Neurosci 17, 664–666, https://doi.org/10.1038/nn.3688 (2014).
Mackenzie, I. R. et al. TIA1 Mutations in Amyotrophic Lateral Sclerosis and Frontotemporal Dementia Promote Phase Separation and Alter Stress Granule Dynamics. Neuron 95, 808–816 e809, https://doi.org/10.1016/j.neuron.2017.07.025 (2017).
Tan, A. Y. & Manley, J. L. The TET family of proteins: functions and roles in disease. J Mol Cell Biol 1, 82–92, https://doi.org/10.1093/jmcb/mjp025 (2009).
Schwartz, J. C., Cech, T. R. & Parker, R. R. Biochemical Properties and Biological Functions of FET Proteins. Annual review of biochemistry 84, 355–379, https://doi.org/10.1146/annurev-biochem-060614-034325 (2015).
Coelho, M. B. et al. Nuclear matrix protein Matrin3 regulates alternative splicing and forms overlapping regulatory networks with PTB. EMBO J 34, 653–668, https://doi.org/10.15252/embj.201489852 (2015).
Kim, H. J. et al. Mutations in prion-like domains in hnRNPA2B1 and hnRNPA1 cause multisystem proteinopathy and ALS. Nature 495, 467–473, https://doi.org/10.1038/nature11922 (2013).
Johnson, J. O. et al. Exome sequencing reveals VCP mutations as a cause of familial ALS. Neuron 68, 857–864, https://doi.org/10.1016/j.neuron.2010.11.036 (2010).
Boczonadi, V. et al. EXOSC8 mutations alter mRNA metabolism and cause hypomyelination with spinal muscular atrophy and cerebellar hypoplasia. Nature communications 5, 4287, https://doi.org/10.1038/ncomms5287 (2014).
Evgrafov, O. V. et al. Mutant small heat-shock protein 27 causes axonal Charcot-Marie-Tooth disease and distal hereditary motor neuropathy. Nature genetics 36, 602–606, https://doi.org/10.1038/ng1354 (2004).
Ackerley, S. et al. A mutation in the small heat-shock protein HSPB1 leading to distal hereditary motor neuronopathy disrupts neurofilament assembly and the axonal transport of specific cellular cargoes. Hum Mol Genet 15, 347–354, https://doi.org/10.1093/hmg/ddi452 (2006).
Knierim, E. et al. Mutations in Subunits of the Activating Signal Cointegrator 1 Complex Are Associated with Prenatal Spinal Muscular Atrophy and Congenital Bone Fractures. Am J Hum Genet 98, 473–489, https://doi.org/10.1016/j.ajhg.2016.01.006 (2016).
Oliveira, J., Martins, M., Pinto Leite, R., Sousa, M. & Santos, R. The new neuromuscular disease related with defects in the ASC-1 complex: report of a second case confirms ASCC1 involvement. Clinical genetics. https://doi.org/10.1111/cge.12997 (2017).
Kaida, D. et al. U1 snRNP protects pre-mRNAs from premature cleavage and polyadenylation. Nature 468, 664–668, https://doi.org/10.1038/nature09479 (2010).
Yu, Y. & Reed, R. FUS functions in coupling transcription to splicing by mediating an interaction between RNAP II and U1 snRNP. Proc Natl Acad Sci USA 112, 8608–8613, https://doi.org/10.1073/pnas.1506282112 (2015).
Prpar Mihevc, S., Baralle, M., Buratti, E. & Rogelj, B. TDP-43 aggregation mirrors TDP-43 knockdown, affecting the expression levels of a common set of proteins. Scientific reports 6, 33996, https://doi.org/10.1038/srep33996 (2016).
Rossoll, W. et al. Specific interaction of Smn, the spinal muscular atrophy determining gene product, with hnRNP-R and gry-rbp/hnRNP-Q: a role for Smn in RNA processing in motor axons? Hum Mol Genet 11, 93–105 (2002).
Mourelatos, Z., Abel, L., Yong, J., Kataoka, N. & Dreyfuss, G. SMN interacts with a novel family of hnRNP and spliceosomal proteins. EMBO J 20, 5443–5452, https://doi.org/10.1093/emboj/20.19.5443 (2001).
Salton, M., Lerenthal, Y., Wang, S. Y., Chen, D. J. & Shiloh, Y. Involvement of Matrin 3 and SFPQ/NONO in the DNA damage response. Cell Cycle 9, 1568–1576, https://doi.org/10.4161/cc.9.8.11298 (2010).
Lopez-Gonzalez, R. et al. Poly(GR) in C9ORF72-Related ALS/FTD Compromises Mitochondrial Function and Increases Oxidative Stress and DNA Damage in iPSC-Derived Motor Neurons. Neuron 92, 383–391, https://doi.org/10.1016/j.neuron.2016.09.015 (2016).
Kenna, K. P. et al. NEK1 variants confer susceptibility to amyotrophic lateral sclerosis. Nature genetics 48, 1037–1042, https://doi.org/10.1038/ng.3626 (2016).
Hill, S. J. et al. Two familial ALS proteins function in prevention/repair of transcription-associated DNA damage. Proc Natl Acad Sci USA 113, E7701–E7709, https://doi.org/10.1073/pnas.1611673113 (2016).
Higelin, J. et al. FUS Mislocalization and Vulnerability to DNA Damage in ALS Patients Derived hiPSCs and Aging Motoneurons. Frontiers in cellular neuroscience 10, 290, https://doi.org/10.3389/fncel.2016.00290 (2016).
Farg, M. A., Konopka, A., Soo, K. Y., Ito, D. & Atkin, J. D. The DNA damage response (DDR) is induced by the C9orf72 repeat expansion in amyotrophic lateral sclerosis. Hum Mol Genet 26, 2882–2896, https://doi.org/10.1093/hmg/ddx170 (2017).
Walker, C. et al. C9orf72 expansion disrupts ATM-mediated chromosomal break repair. Nat Neurosci 20, 1225–1235, https://doi.org/10.1038/nn.4604 (2017).
Jangi, M. et al. SMN deficiency in severe models of spinal muscular atrophy causes widespread intron retention and DNA damage. Proc Natl Acad Sci USA 114, E2347–E2356, https://doi.org/10.1073/pnas.1613181114 (2017).
Bizarro, J. et al. NUFIP and the HSP90/R2TP chaperone bind the SMN complex and facilitate assembly of U4-specific proteins. Nucleic Acids Res 43, 8973–8989, https://doi.org/10.1093/nar/gkv809 (2015).
Cloutier, P. et al. R2TP/Prefoldin-like component RUVBL1/RUVBL2 directly interacts with ZNHIT2 to regulate assembly of U5 small nuclear ribonucleoprotein. Nature communications 8, 15615, https://doi.org/10.1038/ncomms15615 (2017).
Close, P. et al. DBIRD complex integrates alternative mRNA splicing with RNA polymerase II transcript elongation. Nature 484, 386–389, https://doi.org/10.1038/nature10925 (2012).
Shi, K. Y. et al. Toxic PRn poly-dipeptides encoded by the C9orf72 repeat expansion block nuclear import and export. Proc Natl Acad Sci USA 114, E1111–E1117, https://doi.org/10.1073/pnas.1620293114 (2017).
Jovicic, A. et al. Modifiers of C9orf72 dipeptide repeat toxicity connect nucleocytoplasmic transport defects to FTD/ALS. Nat Neurosci 18, 1226–1229, https://doi.org/10.1038/nn.4085 (2015).
Zhang, K. et al. The C9orf72 repeat expansion disrupts nucleocytoplasmic transport. Nature 525, 56–61, https://doi.org/10.1038/nature14973 (2015).
Boehringer, A. et al. ALS Associated Mutations in Matrin 3 Alter Protein-Protein Interactions and Impede mRNA Nuclear Export. Scientific reports 7, 14529, https://doi.org/10.1038/s41598-017-14924-6 (2017).
Dangond, F. et al. Molecular signature of late-stage human ALS revealed by expression profiling of postmortem spinal cord gray matter. Physiol Genomics 16, 229–239, https://doi.org/10.1152/physiolgenomics.00087.2001 (2004).
Sathasivam, S., Ince, P. G. & Shaw, P. J. Apoptosis in amyotrophic lateral sclerosis: a review of the evidence. Neuropathology and applied neurobiology 27, 257–274 (2001).
Deng, H. X. et al. Mutations in UBQLN2 cause dominant X-linked juvenile and adult-onset ALS and ALS/dementia. Nature 477, 211–215, https://doi.org/10.1038/nature10353 (2011).
Teyssou, E. et al. Novel UBQLN2 mutations linked to amyotrophic lateral sclerosis and atypical hereditary spastic paraplegia phenotype through defective HSP70-mediated proteolysis. Neurobiol Aging 58, 239 e211–239 e220, https://doi.org/10.1016/j.neurobiolaging.2017.06.018 (2017).
Keller, W., Bienroth, S., Lang, K. M. & Christofori, G. Cleavage and polyadenylation factor CPF specifically interacts with the pre-mRNA 3’ processing signal AAUAAA. EMBO J 10, 4241–4249 (1991).
Murthy, K. G. & Manley, J. L. The 160-kD subunit of human cleavage-polyadenylation specificity factor coordinates pre-mRNA 3′-end formation. Genes & development 9, 2672–2683 (1995).
Mandel, C. R. et al. Polyadenylation factor CPSF-73 is the pre-mRNA 3′-end-processing endonuclease. Nature 444, 953–956, https://doi.org/10.1038/nature05363 (2006).
Berg, M. G. et al. U1 snRNP determines mRNA length and regulates isoform expression. Cell 150, 53–64, https://doi.org/10.1016/j.cell.2012.05.029 (2012).
Das, R. et al. Functional coupling of RNAP II transcription to spliceosome assembly. Genes & development 20, 1100–1109 (2006).
Das, R. et al. SR Proteins Function in Coupling RNAP II Transcription to Pre-mRNA Splicing. Molecular cell 26, 867–881, https://doi.org/10.1016/j.molcel.2007.05.036 (2007).
Fong, Y. W. & Zhou, Q. Stimulatory effect of splicing factors on transcriptional elongation. Nature 414, 929–933 (2001).
Masuda, S. et al. Recruitment of the human TREX complex to mRNA during splicing. Genes & development 19, 1512–1517 (2005).
Cheng, H. et al. Human mRNA export machinery recruited to the 5′ end of mRNA. Cell 127, 1389–1400 (2006).
Dignam, J. D., Lebovitz, R. M. & Roeder, R. G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res 11, 1475–1489 (1983).
McAlister, G. C. et al. Increasing the multiplexing capacity of TMTs using reporter ion isotopologues with isobaric masses. Analytical chemistry 84, 7469–7478, https://doi.org/10.1021/ac301572t (2012).
Stelzer, G. et al. The GeneCards Suite: From Gene Data Mining to Disease Genome Sequence Analyses. Curr Protoc Bioinformatics 54(1), 30 31–31 30 33, https://doi.org/10.1002/cpbi.5 (2016).
Acknowledgements
We are grateful to lab members for useful discussions and critical reading of the manuscript. This work was supported by NIH grants R01GM043375 and R35GM122524 to R.R.
Author information
Authors and Affiliations
Contributions
B.C. and R.R. conceived the project. The experiments were carried out by B.C. and T.Y. Mass spectrometry was performed by J.D.O. The data were analyzed by B.C., J.D.O., S.P.G. and R.R. with assistance from J.G. The manuscript was written by B.C. and R.R. with input from all authors.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chi, B., O’Connell, J.D., Yamazaki, T. et al. Interactome analyses revealed that the U1 snRNP machinery overlaps extensively with the RNAP II machinery and contains multiple ALS/SMA-causative proteins. Sci Rep 8, 8755 (2018). https://doi.org/10.1038/s41598-018-27136-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-27136-3
This article is cited by
-
TASOR epigenetic repressor cooperates with a CNOT1 RNA degradation pathway to repress HIV
Nature Communications (2022)
-
The upstream 5′ splice site remains associated to the transcription machinery during intron synthesis
Nature Communications (2021)
-
DDX17 is involved in DNA damage repair and modifies FUS toxicity in an RGG-domain dependent manner
Acta Neuropathologica (2021)
-
RNA dependent suppression of C9orf72 ALS/FTD associated neurodegeneration by Matrin-3
Acta Neuropathologica Communications (2020)
-
RNA-recognition motif in Matrin-3 mediates neurodegeneration through interaction with hnRNPM
Acta Neuropathologica Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.