Abstract
The blue coral, Heliopora coerulea, is a reef-building octocoral that prefers shallow water and exhibits optimal growth at a temperature close to that which causes bleaching in scleractinian corals. To better understand the molecular mechanisms underlying its biology and ecology, we generated a reference transcriptome for H. coerulea using next-generation sequencing. Metatranscriptome assembly yielded 90,817 sequences of which 71% (64,610) could be annotated by comparison to public databases. The assembly included transcript sequences from both the coral host and its symbionts, which are related to the thermotolerant C3-Gulf ITS2 type Symbiodinium. Analysis of the blue coral transcriptome revealed enrichment of genes involved in stress response, including heat-shock proteins and antioxidants, as well as genes participating in signal transduction and stimulus response. Furthermore, the blue coral possesses homologs of biomineralization genes found in other corals and may use a biomineralization strategy similar to that of scleractinians to build its massive aragonite skeleton. These findings thus offer insights into the ecology of H. coerulea and suggest gene networks that may govern its interactions with its environment.
Similar content being viewed by others
Introduction
Octocorals are the most diverse coral group and can be found in various marine environments, including shallow tropical reefs, deep seamounts, and submarine canyons1. They reportedly exhibit greater resistance and resilience to environmental perturbations, particularly to thermal stress, compared to scleractinians2,3,4. They are also suitable indicators of ecological disturbances, as they are generally fast-growing and have the ability to colonize open spaces. Furthermore, octocorals provide habitats for molluscs, polychaetes, fish, and other marine fauna5. Octocorals contribute to the structure of reef communities through the production of allelopathic compounds that drive interactions with other organisms6,7.
The Octocorallia comprises approximately 3,500 extant species and is traditionally divided into three genetically distinct clades – the order Alcyonacea (soft corals and gorgonians), the Pennatulacea (sea pens), and the Helioporacea (blue corals)8. The Helioporacea is unique, as its members produce calcified skeletons of crystalline aragonite, a convergent feature shared with scleractinians. The blue coral, Heliopora coerulea, is the sole extant member of the family Helioporidae. Its fibrous aragonite skeleton is characterized by a distinctive and permanent blue pigmentation resulting from iron salts9. The blue coral is a reef-building coral that superficially resembles many scleractinian corals in its gross morphology. Its colonies exhibit considerable morphological plasticity and can be arborescent or plate-like. In very calm water it can form slender columns or in more turbulent areas, it develops stout trunks and branches. H. coerulea frequently co-exists with massive stony corals like Porites and Montipora10.
Based on fossil records, the genus Heliopora was widely distributed in shallow waters of all oceans10. However, climatic changes, such as glacial periods of lowered temperatures, may have resulted in contraction of its distribution. Presently, H. coerulea is found in equatorial waters between 25°N and 25°S in the Indo-Western Pacific region, Red Sea, American Samoa, southern Japan, and Australia. Although it is uncommon throughout its range, it is the dominant species on some reefs10. The distribution of H. coerulea may be determined by its optimal growth temperature, short larval duration, and other environmental factors. H. coerulea thrives in waters with a mean annual minimum temperature above 22 °C, which is considerably higher than the 18 °C marginal isotherm for many corals10. Its shallow depth range and higher optimal growth temperature are close to conditions that cause bleaching in scleractinian corals11.
Scleractinians, or hard corals, are the primary bioconstructors of reef ecosystems. However, in recent years, heat stress due to global warming and increasing intensity of El Niño Southern Oscillation (ENSO) events have worsened the bleaching of hard corals worldwide. Such recurrent disturbances have caused mass die-offs of many hard coral species, reducing their abundance and total cover12. Modification of reef substrata may allow opportunistic proliferation of non-scleractinian taxa, such as octocorals12. In fact, in some ecosystems, octocoral abundance is reported to be increasing coincidentally with scleractinian decline, following severe bleaching events13,14. More specifically, a shift in the dominant taxa from branching scleractinian corals to H. coerulea have been observed on reefs in Ishigaki island, Okinawa, Japan and Bolinao, Pangasinan, Philippines15,16.
To date, very few studies have examined octocorals and available transcriptomic data mostly represent corals of the order Alcyonacea17,18. Sequencing of the first octocoral transcriptome from the Carribean sea fan, Gorgonia ventalina, revealed mechanisms of sea fan immunity in response to pathogen exposure18, while a study of the red coral, Corallium rubrum, further highlighted the molecular basis of local adaptation to thermal stress17. Thus, while a few studies have begun to explain the stress tolerance and adaptive potential of some octocorals to environmental perturbations, transcriptome sequencing of other octocoral species will prove valuable in understanding the potential impacts of changing environments on members of this diverse coral group.
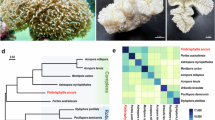
In this study, we describe a reference transcriptome of the non-scleractinian reef-building coral, H. coerulea (Fig. 1a). Gene content analysis and comparison with other cnidarian sequences potentially reveal the molecular basis for unique characteristics of this coral, its symbiotic interactions, and responses to the environment. This transcriptome resource provides a platform for future investigations into the gene complement and gene expression dynamics of H. coerulea.
De novo transcriptome assembly of the blue coral, Heliopora coerulea. (a) H. coerulea contributes to reef structure through production of an aragonite skeleton that forms colonies with digitate (white arrowhead) or laminar (black arrowhead) morphologies. (b) Classification of H. coerulea transcripts into those originating from the coral host (Hcoe-coral) or the symbiont (Hcoe-symbiont). Unclassified transcripts are potentially Heliopora-specific (Hcoe-specific) with about 56% aligning to metazoan sequences. (c) Percent GC of transcript sets classified by origin. (d) Gene ontology (GO) analysis for H. coerulea transcripts classified by origin. Enrichment p-values for selected terms are shown. Only GO terms with a p-value < 0.05 (Fisher’s exact test) were considered significantly enriched. (e–f) Top 30 enriched PFAM protein domains in (e) Hcoe-coral and (f) Hcoe-symbiont peptides compared to other organisms. Heatmaps were generated using pheatmap in R. Species names are abbreviated as follows: H. coerulea coral host (Hcoe), G. ventalina (Gven), N. vectensis (Nvec), A. digitifera (Adig), A. pallida (Apal), H. coerulea symbionts (Hcoe_s), C. owczarzaki (Cowc), S. kawagutii (Skaw), S. microadriaticum (Smic), and S. minutum (Smin).
Results
RNA sequencing and transcriptome assembly
Sequencing of cDNA libraries from three individual colonies of H. coerulea yielded a total of 108.7 million paired-end reads (Supplementary Tables 1, 2), of which 82.8 million remained after adapter trimming and quality filtering. De novo transcriptome assembly yielded 231,515 transcripts (>300 bp), with an N50 of 1,106 bp. 155,859 (67%) transcripts were translated into peptides and 90,817 (39%) were retained after clustering similar sequences with CDHIT (Supplementary Table 2).
Classification of transcripts
Like most scleractinian corals, the blue coral is a holobiont consisting of a cnidarian host and endosymbiotic dinoflagellates of the genus Symbiodinium. To identify sequences in the H. coerulea transcriptome that originate from either the coral host or its symbionts, transcripts were aligned to cnidarian and Symbiodinium databases using blastn (Supplementary Table 3). This recovered 22,174 (24%) sequences originating from the coral host (Hcoe-coral) and 26,085 (29%) originating from symbionts (Hcoe-symbiont) (Fig. 1b, Supplementary Table 4). About 47% (42,558 sequences) of the transcripts could not be classified, as they did not match any sequences in the cnidarian and symbiont databases. The latter set of sequences may include transcripts specific to the H. coerulea holobiont (Hcoe-specific).
Transcripts classified as originating from the coral (Hcoe-coral) exhibited a GC content of about 43% (Fig. 1c). This is similar to the GC content of other anthozoans, such as Acropora digitifera (39%), Porites australiensis (40%), and Nematostella vectensis (47%)19,20,21. On the other hand, transcripts classified as originating from the symbiont (Hcoe-symbiont) had a GC content of approximately 54%, which is similar to the GC content reported for Symbiodinium microadriaticum (50.5%) and considerably higher than the coding GC content of S. minutum (43.5%) and S. kawagutii (45.5%)22. The GC content of unclassified (Hcoe-specific) transcripts also peaked at around 45% supporting the idea that most of these originated from the blue coral host or other organisms associated with the coral holobiont.
Transcriptome annotation
In total, 64,610 (71%) of the 90,817 H. coerulea transcriptome sequences were annotated using public databases (NCBI RefSeq, UniProt-SwissProt, PFAM, and KEGG) (Supplementary Table 5). Specifically, blastp alignments revealed that 61,922 (68%) and 49,796 (55%) had significant matches to proteins in the NCBI-RefSeq and UniProt-SwissProt databases, respectively. A total of 21,573 (97%) Hcoe-coral and 13,025 (50%) Hcoe-symbiont peptide sequences were homologous to NCBI-RefSeq proteins. Among peptides from the Hcoe-specific set, 27,324 (64%) had blastp matches to NCBI-RefSeq proteins, the majority (56%) of which were from metazoans (Fig. 1b, Supplementary Table 6). Comparison of predicted peptides against the OrthoMCL database23 revealed that most of the transcriptome sequences cluster with eukaryote protein families (Supplementary Table 7).
Gene ontology analysis revealed that Hcoe-coral transcripts are enriched for typical metazoan cellular processes, such as cell communication, signal transduction, cellular response to stimuli, and regulation of gene expression (Fig. 1d)24. On the other hand, Hcoe-symbiont transcripts are enriched for metabolic processes, including carbohydrate and amino acid synthesis, and photosynthesis. Hcoe-specific transcripts are enriched for exocytosis and secretion-related functions.
PFAM domains were identified in 48,834 (54%) of predicted peptides from the H. coerulea transcriptome. Global comparisons of PFAM domain composition of various taxa revealed the greatest similiarity of the domain complement of predicted peptides from Hcoe-coral transcripts with that of other cnidarians (Supplementary Figure 1). On the other hand, the PFAM domain composition of peptides from Hcoe-symbiont transcripts was more similar to that of other Symbiodinium species. PFAM domains that are most abundant in the Hcoe-coral peptides include those involved in signaling (14-3-3, Ras, Arf, GTP, G-alpha), cytoskeleton and transport (adaptin, microtub_bd, CH), scavenger receptors (SRCR), extracellular matrix (EGF_2), heat shock proteins, and antioxidants (Cpn60_TCP1, HSP90, thioredoxin, redoxin) (Fig. 1e). Abundant PFAM domains in Hcoe-symbiont peptides include antioxidants (glutaredoxin, thioredoxin), methyltransferases, and calcium-binding domains (EF_hand, SPARC_Ca_bdg) (Fig. 1f).
Divergence time estimation
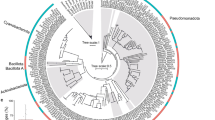
Phylogenetic analysis was conducted based on alignment of 62,390 amino acid positions from peptides predicted from transcripts originating from the coral host. This estimated the divergence of the blue coral from the sea fan, G. ventalina, at about 370 MYA (Carboniferous period) (Fig. 2a). The octocorals diverged from the hexacorals around 595 MYA (Precambrian period). These findings support divergence estimates based on mitochondrial protein-coding genes25,26.
Comparison of the H. coerulea gene complement with those of other corals. (a) Divergence time estimates for H. coerulea and other cnidarians. Phylogenetic relationships among taxa were reconstructed using PhyML with the sponge, A. queenslandica, as an outgroup. Red circles at the nodes represent 100% support in PhyML analysis. Divergence was estimated using the Reltime method in MEGA 7. Approximate divergence times of each lineage are shown at the nodes. Blue horizontal bars at each node represent 95% confidence intervals. (b) Numbers of orthologous protein groups in H. coerulea (Hcoe-coral and Hcoe-specific peptides only), G. ventalina, and scleractinians, as determined by OrthoMCL analysis. (c) Gene ontology (GO) analysis for orthologous protein families found in both scleractinians and octocorals (Scle-Octo), octocorals only (Octo), and H. coerulea only (Hcoe). Enrichment p-values for selected terms are shown. Only GO terms with a p-value < 0.05 (Fisher’s exact test) were considered significantly enriched.
Gene functions in orthologous groups
OrthoMCL analysis was conducted to determine the number of shared and unique orthologous protein families in H. coerulea, G. ventalina, and scleractinians. Using OrthoMCL, 10,407 ortholog groups were identified from 69,757 coral proteins. Of these, 879 groups were common to the octocorals, H. coerula and G. ventalina, and the scleractinians (Fig. 2b). 3,926 gene groups were shared only between the two octocorals. 375 protein families were common only to scleractinian corals and H. coerulea. 2,369 gene groups were unique to H. coerulea.
Ortholog groups shared by scleractinians and octocorals included gene families associated with cellular component organization, metabolic processes, chromatin modification, gene expression, cell cyle, and transport (Fig. 2c). A similar set of gene ontology terms was enriched among the octocoral-specific ortholog groups. Gene families associated with cell communication, signal transduction, response to stimulus, and development were enriched in ortholog groups unique to H. coerulea. In particular, signaling pathways that are regulated by mitogen-activated protein kinases (MAPKs) and small GTPases were overrepresented.
ITS2 identification and metabolic pathway analysis
The Symbiodinium hosted in the tissues of H. coerulea from Bolinao, Philippines was found to be most closely related to the thermally tolerant S. thermophilum C3-Gulf ITS2 type. Two ITS2 sequences were recovered from H. coerulea by DGGE. One sequence was 99.5% identical to the partial S. thermophilum ITS2 sequence and the other was 92.6% identical. Both ITS2 sequences contained an 8-bp insertion similar to C3-Gulf ITS227 (Supplementary Figure 2). No other ITS2 types were identified in the coral using DGGE and we did not detect assembled sequences of nuclear rDNA in the transcriptome dataset.
Identification of enzymes within KEGG orthology (KO) pathways that were present in either member of the holobiont or in both revealed biochemical processes that may be performed principally by the host coral or by the symbiont. Examples of modules that are represented mainly in the host are cell signaling and RNA processing (Fig. 3). Enzymes for metabolism of glycans and glycosaminoglycans, which are key extracellular matrix components, are also more highly represented in the host. On the other hand, modules with more enzymes represented only in symbiont transcripts include terpenoid biosynthesis and photosynthesis, as well as pathways for metabolism of amino acids such as lysine, histidine, serine, and threonine.
Metabolic complementation analysis for the H. coerulea metatranscriptome. Percentages of enzymes in selected KEGG pathways that are represented by transcripts originating from the coral symbiont (Hcoe-symbiont, yellow), transcripts originating from the coral host (Hcoe-coral, light blue), transcripts specific to the H. coerulea holobiont (Hcoe-specific, dark blue), and transcripts present in both host and symbiont (Both, grey).
Key enzymes for amino acid biosynthesis, including diaminopimelate decarboxylase and aspartate kinase for lysine synthesis, glutamine amidotransferase, ATP phosphoribosyltransferase, and histidinol dehydrogenase for histidine synthesis, homoserine kinase and threonine aldolase for serine and threonine synthesis, 5-methyltetrahydrofolate-homocysteine methyltransferase for methionine synthesis, and acetolactate synthase for branched chain amino acid synthesis, were represented in symbiont-derived transcripts but not in host coral-derived transcripts (Supplementary Table 8).
Biomineralization-related genes
Coral biomineralization requires sources of inorganic carbon and calcium. Hydration of inorganic carbon to bicarbonate is mediated by carbonic anhydrases. Secreted or membrane-bound, cytosolic, and mitchondrial alpha-carbonic anhydrase (α-CA) types are present in the H. coerulea transcriptome and in the transcriptomes of other octocorals, G. ventalina and C. rubrum (Fig. 4a). Phylogenetic analysis revealed that α-CAs from these three octocorals cluster together (Fig. 4b). Representatives of α-CA isoforms, CruCA4 and CruCA6 of C. rubrum, were identified in the H. coerulea transcriptome. CruCA4 is the top gene candidate involved in calcification in C. rubrum while CruCA6 is expressed at lower levels in tissues of this soft coral28. No homolog of α-CA isoform CruCA5, the most ubiquitous isoform in C. rubrum, was identified in the H. coerulea transcriptome. Interestingly, four of the six α-CA homologs detected in the H. coerulea transcriptome are derived from the Hcoe-specific set of transcripts.
Biomineralization genes in H. coerulea. (a) Biomineralization-related genes that are represented in the H. coerulea transcriptome and those of other anthozoans (present, colored boxes; not detected, white boxes). (b) Phylogenetic analyses of biomineralization-related genes present in H. coerulea. The unrooted trees were derived from PhyML analysis. Numbers on selected branches represent maximum likelihood bootstrap values.
The concentration of bicarbonate anions at the site of calcification is mediated by bicarbonate transporters29, such as SLC4 and SLC26. SLC4γ, which is reported to be a key scleractinian-specific gene for calcification30, was not detected in H. coerulea (Fig. 4b). However, the H. coerulea transcriptome included a homolog of SLC4δ, which is present in many cnidarians, as well as SLC26γ, a protein that is highly expressed in the calicoblastic ectoderm of the scleractinian coral, Stylophora pistillata30. Octocorals possess additional SLC4 and SLC26 homologs that do not cluster closely with scleractinian genes.
Precipitation of calcium carbonate or aragonite crystals is facilitated by organic matrix proteins such as coral acid-rich proteins (CARPs) and skeletal acidic proteins (SAPs). Homologs of CARPs are present in the H. coerulea transcriptome (Fig. 4a, Supplementary Figure 3). An acid-rich peptide with low percent identity (40%) to A. millepora SAP2 was also identified in the blue coral transcriptome (Fig. 4a). This may be a putative homolog, although SAP proteins were previously reported to be restricted to acroporids31. Organization of aragonite crystals into the stable macrostructure that characterizes the coral skeleton is mediated by other organic matrix proteins, such as adhesion proteins (collagen triple helix and zona pellucida domain-containing proteins), lectin-containing proteins (lectin, MAM, and fibronectin-containing proteins), and proteins associated with calcium ion-binding (calmodulin, SPARC_Ca_bdg, EGF, and laminin G domain-containing proteins) all of which were detected in H. coerulea (Fig. 4a). In addition, the H. coerulea transcriptome was found to contain a homolog of galaxin, a component of the skeletal organic matrix protein first identified in the coral, Galaxea fascicularis32. The H. coerulea galaxin homolog clusters with galaxins from the hard corals, Favia sp. and S. pistillata (Fig. 4b). Furthermore, a homolog of scleritin, a secreted protein in the sclerite organic matrix of the soft coral, C. rubrum, was detected in the H. coerulea transcriptome (Fig. 4a).
Discussion
The blue coral, H. coerulea, is unique among octocorals for its ability to produce a massive skeleton of crystalline aragonite. It has a shallow depth range and a higher optimal growth temperature that suggests greater resilience to warming environments than scleractinian corals. These features make this coral an interesting model organism for understanding environmental responses, adaptation to elevated temperature, and biomineralization.
H. coerulea has been described as a living fossil because its skeletal features remain unchanged from the oldest representative of the genus, Heliopora japonica, of the Lower Cretaceous33. Octocorals are thought to have originated from a Precambrian ancestor that diverged 696 MYA, with an approximately 400 MY gap until the appearance of the ancestor of modern octocoral species that emerged around 241 MYA25. Our divergence estimation based on transcriptome data shows that the divergence between H. coerulea and G. ventalina occurred around 370 MYA. This suggests that octocorals may have originated earlier than what was estimated based on mitochondrial protein-coding genes and the fossil record25.
The H. coerulea metatranscriptome includes transcripts originating from the coral animal or its symbiotic dinoflagellate, Symbiodinium. These transcript sets were enriched for signature functions of animals and photosynthetic organisms, respectively. Comparison of orthologous protein families further revealed that, while the blue coral transcriptome includes many functions that are shared with scleractinians and other octocorals, gene families that are unique to H. coerulea are enriched for potential roles in signal transduction. This is also reflected in the abundance of protein domains associated with signaling in the blue coral. The expanded repertoire of signaling genes may contribute to its ability to sense and respond to various stimuli in its environment. In addition, the enrichment of transcripts with functions related to cytoskeleton, extracellular matrix, and cell cycle in the blue coral transcriptome suggests a capacity for tissue growth and remodeling that may partly explain the morphological plasticity of this species34 and its ability to compete for space on the reef.
The H. coerulea transcriptome contains representatives of gene families that are commonly associated with stress responses. This includes an abundance of transcripts with heat shock protein and antioxidant domains that may be important for maintenance of cellular homeostasis. Similar genes are differentially expressed in scleractinians, suggesting a link between expression levels of these genes and bleaching resistance of corals35,36,37. Expression of protective genes in scleractinians is rapid and highly coordinated, indicating a strong transcriptional response to thermal stress in the coral host38. Whether this is also the case in H. coerulea remains to be determined.
Octocoral species exhibit various degrees of thermotolerance, depending on the presence or absence of symbionts and on the specificity of symbiosis39,40. In symbiotic octocorals, the coral hosts are more adapted to higher seawater temperatures than their zooxanthellae4. The ability of the holobiont to withstand environmental stressors may be related to the degree of interdependency between the coral host and the symbionts20,21. Many coral species are capable of hosting multiple types of Symbiodinium that vary in their susceptibility to bleaching. Changes in the endosymbiont community composition to include more thermotolerant clades may increase resistance to future bleaching41. H. coerulea from the Andaman Sea, in the northeastern Indian Ocean, hosts thermally tolerant symbionts of ITS2 type D2-4-542, while the same species in Guam hosts generalist symbionts of type C343. The present study revealed that H. coerulea from Bolinao, Pangasinan, host symbionts of the C3-Gulf ITS2 type, a thermotolerant lineage previously detected in corals from the Persian Gulf, where temperatures can reach high seasonal maxima. This study provides the first evidence for this ITS2 variant in a coral outside of the Persian/Arabian Gulf region. This finding suggests that the high optimal growth temperature of H. coerulea may be supported by the presence of thermally-tolerant symbiont types in its tissues.
Co-evolution of host and symbionts results in genomic changes in both symbiotic partners. In the coral holobiont, this is evidenced by metabolic complementation between the coral and Symbiodinium, with about 80% of the daily photosynthate transferred from the symbiont to the scleractinian coral host. In octocorals, the degree of metabolic exchange with symbionts varies depending on colony morphology, depth range, and symbiont specificity40. Transcriptome analysis revealed that the blue coral lacks enzymes for key steps in amino acid metabolic pathways and probably relies on its symbionts for amino acids that it is unable to produce. This is similar to what has been shown in A. digitifera, P. australiensis, and G. fascicularis20,21,44. It should be noted, however, that the inability to detect representatives for certain enzymes involved in metabolic pathways may not necessarily indicate their absence from the coral or symbiont genome. It may simply be that they were not captured in the tissues used for transcriptome sequencing. In addition to acquiring photosynthates from their symbionts, corals can augment their nutrient supply through heterotrophic feeding45. Octocorals, in particular, are known to exhibit mixed phototrophic and heterotrophic nutrition to sustain their metabolic needs40,46,47. Heterotrophy may reduce host dependency on endosymbionts thereby allowing them to allocate energy toward repair and recovery from stress. However, it remains to be determined whether the blue coral exhibits extended heterotrophic compensation as an adaptive strategy to survive adverse conditions, as has been reported in other corals40,48.
Evolution of calcification has been linked to the presence of bicarbonate transport proteins, expansion of the secreted and membrane-associated carbonic anhydrase repertoire, and recruitment of extracellular matrix-derived genes to regulate mineralization30,49. In scleractinians, precipitation of aragonite, which occurs under the calicoblastic ectodermal layer, is mediated by the concentration of inorganic carbon and calcium ions and proteins that catalyze the nucleation process and crystallization29,50,51. The blue coral skeleton, like that of scleractinians, is composed of fibrous aragonite instead of the typical calcitic sclerite found in soft corals. An in situ microRaman spectral mapping study revealed the regular cyclic secretion of organic and mineral components in biomineralization zones of the blue coral52. This suggests that H. coerulea skeleton fiber growth is induced by the organic matrix. Manual curation of the H. coerulea transcriptome revealed coral biomineralization-related genes, including carbonic anhydrases, bicarbonate transporters, calcium-binding proteins, and skeletal organic matrix proteins (SOMPs). Representative SOMPs involved in various stages of skeleton formation in A. digitifera31 were identified in the H. coerulea transcriptome, along with other extracellular matrix proteins and proteins involved in secretion and exocytosis. The H. coerulea biomineralization gene repertoire shares similarities with both scleractinians and other octocorallians, although some key scleractinian biomineralization genes were not detected in the transcriptome. Genome sequencing of H. coerulea, as well as other octocorals, would provide a definitive answer as to whether the genes are indeed absent in these taxa. The presence of homologs of biomineralization enzymes, transporters, and organic matrix proteins, such as galaxin, that are typically found in scleractinian corals may underlie the ability of the blue coral to form massive aragonite skeletons. However, differences in the activities and biochemical characteristics of these proteins may result in the production of skeletons with different properties compared to those of scleractinian corals. Further studies are needed to understand the roles of these genes in H. coerulea biomineralization.
The H. coerulea gene complement shares many similarities with both scleractinians and other octocorals. The presence of genes that function within known pathways or networks highlight genetic mechanisms that potentially govern interactions of the blue coral with its environment. Additional experiments need to be conducted to determine how this gene complement responds to changing ocean conditions. The transcriptome resource presented in this study thus serves as a springboard for further investigations into the physiology of the blue coral and its environmental responses. These studies will contribute to a deeper understanding of the phylogeny, character evolution, and key innovations within the Octocorallia.
Methods
Coral collection
Blue coral colonies were collected at a depth of 10–15 meters by scuba diving in Lucero, Bolinao, Pangasinan, Philippines (N 16°2441 E 119°5420) in January 2015 (Supplementary Table 1). H. coerulea exists in several forms, but only the digitate form was used in this study. Collections were conducted with permission from the Philippines Department of Agriculture Bureau of Fisheries and Aquatic Resources (DA-BFAR GP-0097-15). Colonies were temporarily placed in tanks with flowing 10 um-filtered seawater. To avoid excessive mucus production during fragment collection, care was taken to minimize air exposure of the colonies. Coral fragments were immediately flash-frozen in liquid nitrogen after collection, prior to transport and subsequent molecular analyses.
RNA extraction and mRNA sequencing
Total RNA from coral fragments originating from three biologically independent coral colonies was extracted using Trizol (Invitrogen) following the manufacturer’s protocol. Coral fragments (~2.5 cm long) from each coral colony were manually homogenized using a mortar and pestle. Contaminating genomic DNA in the RNA extracts was removed using a Turbo DNA-free kit (Life Technologies) followed by ethanol precipitation. Nucleic acid concentrations were quantified using a Qubit 3.0 Fluorometer (Life Technologies). RNA integrity was assessed by electrophoresis on native agarose gels with denaturing loading dye and using an Agilent Bioanalyzer 2100 (Supplementary Table 2). RNA samples were sent to the Beijing Genomic Institute (BGI), Hong Kong, for preparation of barcoded libraries using the Illumina TruSeq RNA Sample Prep Kit V2 protocol. mRNA was polyA-selected from total RNA and fragmented followed by first- and second-strand synthesis to produce products ready for library construction. mRNA-enriched libraries were sequenced on the Illumina HiSeq 2000 platform generating 100 bp paired-end reads with all cDNA libraries pooled in a single lane.
Transcriptome assembly and post-assembly filtering
To detect a greater number of genes expressed by the coral holobiont, reads from three biologically independent coral colonies were pooled and assembled. Raw sequence reads were filtered to remove adapter sequences and low-quality reads. Trimmomatic 0.36 was used to trim the first 10 bases of the reads53. Reads were scanned using a 4-base sliding window, deleting bases when the average quality dropped below 20. Bases with quality scores below 30 at leading and trailing ends were removed. Only reads that passed the quality filters and were longer than 30 bases were retained for further analysis. De novo assembly was carried out using Trinity software V.2.1.154. Transcripts shorter than 300 bp were removed as these were potentially misassembled sequences or did not encode proteins. Coding regions within the transcripts were identified using TransDecoder V.2.0.1. Only the longest ORF (>100 amino acids) from each transcript was included in the H. coeruela transcriptome reference peptide set. CDHIT was then implemented using default parameters to reduce redundancy among proteins (95% similarity and word size of 10)55.
Transcriptome annotation
The H. coerulea transcriptome was aligned with proteins in the UniProt-SwissProt and NCBI-RefSeq databases using blastp with an e-value cutoff of 1 × 10−5. The top hit from the NCBI-RefSeq blastp alignment was used as input for Blast2GO to retrieve gene ontology terms. Protein domains were identified by mapping predicted peptides against the PFAM 28.0 database using HMMER v3.1b1.
Classification of coral and symbiont transcripts
To identify transcripts originating from the coral host or the symbiont, the H. coerulea reference assembly was aligned by blastn to customized databases. The cnidarian database included the genomes of A. digitifera (scleractinian)21, Aiptasia (sea anemone)56, Hydra magnipapillata57, Discosoma sp.58, and Amplexidiscus fenestrafer58 while the Symbiodinium database consisted of the S. minutum59, S. kawagutii60, and S. microadriaticum22 genomes, as well as transcriptomes of Symbiodinium cultures (clades A1, A2, B2, C and D) (Supplementary Table 3)61,62. Transcripts that aligned only to cnidarian polynucleotides were classified as cnidarian sequences (Hcoe-coral) while those that aligned only to Symbiodinium polynucleotides were classified as symbiont sequences (Hcoe-symbiont). Sequences that had hits to both databases were further segregated using the psytrans script (https://github.com/sylvainforet/psytrans). Unclassified sequences that did not align to sequences in either database, were regarded as Hcoe-specific. Different e-value cutoffs (1 × 10−4,, 1 × 10−5, 1 × 10−8, 1 × 10−10) were examined and the cutoff that resulted in a minimum number of unclassified transcripts was chosen (Supplementary Table 4).
GO enrichment analysis for binned transcripts was performed using the topGO package in R. Only GO terms with a p-value < 0.05 were considered significantly enriched. Enriched PFAM domains in Hcoe-coral and Hcoe-symbiont sequences were also identified. Log-transformed, normalized PFAM domain counts were visualized as heatmaps using pheatmap in R.
Divergence time estimation
Peptides from six taxa (H. coerulea, G. ventalina, A. digitifera, H. magnipapillata, N. vectensis, and A. queenslandica) were assigned into ortholog groups (OGs) using a hidden Markov model-based search with HaMStR. Only Hcoe-coral peptides were used for this analysis and only OGs shared by all six taxa were selected. Each OG was aligned using MAFFT v7.220 (parameters:–localpair–maxiterate 1000), and then alignments were trimmed using TrimAl v1.2 with a gap threshold of 0.9, a similarity threshold of 0.001, and a window size of 6. Trimmed alignments shorter than 30 amino acids were discarded. Ortholog alignments were concatenated into a supermatrix using FASconCAT-G. There were a total of 60,627 positions in the final dataset. The best substitution model was determined using ProtTest (v3.4). Maximum-likelihood analysis was implemented on PhyML 3.062 with 1,000 bootstrap replicates. Reltime ML using the JTT matrix-based model was run on MEGA 7 to calculate the divergence time with A. queenslandica (Porifera) as the outgroup. The reference divergence times of H. magnipapillata, N. vectensis, G. ventalina, and A. digitifera were obtained from TimeTree website.
OrthoMCL analysis and orthologous group enrichment analysis
Orthologous gene groups were identified using OrthoMCL with default settings (e-value cutoff 1 × 10−5, protein identity 50%, and MCL inflation of 1.5). The G. ventalina and scleractinian data sets contained 31,334 and 16,249 peptides, respectively. The H. coerulea dataset included 64,732 peptides from the Hcoe-coral and Hcoe-specific clusters. Only ortholog groups with three or more members were considered. The scleractinian ortholog data set, which includes sequences from 20 hard coral species, was downloaded from comparative.reefgenomics.org63. Enrichment analysis for species and lineage-specific gene groups was performed using the topGO package in R. Only GO terms with a p-value < 0.05 were considered significantly enriched.
Denaturing Gradient Gel Electrophoresis
Total DNA from the coral holobiont was extracted using a modified CTAB method. Briefly, small coral fragments from three biologically independent coral colonies were homogenized with a mortar and pestle in CTAB extraction buffer (100 mM TrisCl, pH 8.0, 20 mM EDTA, 2% CTAB, 1.4 M NaCl, 2.5 mg/mL lysozyme) and incubated at 37 °C for 40 mins. After addition of 0.2% β-mercaptoethanol and 0.1 mg/mL proteinase K, samples were incubated at 60 °C for 1 hr, followed by chloroform fractionation and isopropanol precipitation64. The DNA pellet was washed with 70% ethanol and dried at room temperature. DNA was dissolved in 1× TE buffer and stored at −20 °C. PCR amplification of the internal transcribed spacer 2 (ITS2) region was performed using the primers ITSinfor2 and ITS2CLAMP65. Fragments were resolved on a 30–60% linear gradient of urea and formamide with 8% acrylamide66. Gels were run in 1× TAE at 60 V for 16 hours at 60 °C in a DGGE apparatus (C.B.S. Scientific). After electrophoresis, the gel was stained with SYBR Gold nucleic acid stain in 1× TAE buffer, rinsed, and photographed. Distinct bands were excised from the gel and placed in 30 uL of nuclease-free water overnight to diffuse the DNA. Eluted DNA was re-amplified using ITSinfor2 and ITSrev65. Reaction products were sent to First Base, Malaysia for direct sequencing.
Metabolic complementation analysis
Sequences in the Hcoe-coral, Hcoe-symbiont, and Hcoe-specific transcript sets were compared to enzyme sequences in the curated Kyoto Encyclopedia of Genes and Genomes (KEGG) to identify homologs of enzymes in different metabolic pathways. For each KEGG pathway, the percentage of enzymes represented by coral or symbiont transcripts was calculated using a custom Perl script.
Phylogenetic analyses
Protein sequences were aligned using Clustal Omega67 and trimmed with Gblocks68. The best substitution model for each alignment was determined using ProtTest (v3.4). Maximum-likelihood analysis was implemented on PhyML 3.062 with 1,000 bootstrap replicates. Genes used for phylogenetic comparisons were identifed through reciprocal blastp.
Accession numbers
Raw sequence reads were deposited in the NCBI Short Read Archive database (PRJNA397991). The Transcriptome Shotgun Assembly project has been deposited at DDBJ/ENA/GenBank under the accession GFVH00000000. The version described in this paper is the first version, GFVH00000000.
References
McFadden, C. S., Sanchez, J. A. & France, S. C. Molecular phylogenetic insights into the evolution of Octocorallia: a review. Integr Comp Biol 50, 389–410, https://doi.org/10.1093/icb/icq056 (2010).
Netherton, S. E., Scheer, D. M., Morrison, P. R., Parrin, A. P. & Blackstone, N. W. Physiological correlates of symbiont migration during bleaching of two octocoral species. J Exp Biol 217, 1469–1477, https://doi.org/10.1242/jeb.095414 (2014).
Ateweberhan, M. et al. Climate change impacts on coral reefs: synergies with local effects, possibilities for acclimation, and management implications. Mar Pollut Bull 74, 526–539, https://doi.org/10.1016/j.marpolbul.2013.06.011 (2013).
Sammarco, P. W. & Strychar, K. B. Responses to high seawater temperatures in zooxanthellate octocorals. PLoS One 8, e54989, https://doi.org/10.1371/journal.pone.0054989 (2013).
Reijnen, B. T., van der Meij, S. E. & van Ofwegen, L. P. Fish, fans and hydroids: host species of pygmy seahorses. Zookeys, 1–26, https://doi.org/10.3897/zookeys.103.953 (2011).
Sammarco, P. W., Coll, J. C. & Labarre, S. Competitive Strategies of Soft Corals (Coelenterata, Octocorallia) .2. Variable Defensive Responses and Susceptibility to Scleractinian Corals. Journal of Experimental Marine Biology and Ecology 91, 199–215, https://doi.org/10.1016/0022-0981(85)90176-5 (1985).
Fleury, B. G., Lages, B. G., Barbosa, J. P., Kaiser, C. R. & Pinto, A. C. New hemiketal steroid from the introduced soft coral Chromonephthea braziliensis is a chemical defense against predatory fishes. J Chem Ecol 34, 987–993, https://doi.org/10.1007/s10886-008-9499-y (2008).
McFadden, C. S., France, S. C., Sanchez, J. A. & Alderslade, P. A molecular phylogenetic analysis of the Octocorallia (Cnidaria: Anthozoa) based on mitochondrial protein-coding sequences. Mol Phylogenet Evol 41, 513–527, https://doi.org/10.1016/j.ympev.2006.06.010 (2006).
Hill, D. Possible intermediates between Alcyonaria and Tabulata, Tabulata and Rugosa and Rugosa and Hexacoralla. Rept. Intern. Geol. Congress, XXI Sess., Copenhagen, Pt. 22, 51–58 (1960).
Zann, L. P. & Bolton, L. The Distribution, Abundance and Ecology of the Blue Coral Heliopora-Coerulea (Pallas) in the Pacific. Coral Reefs 4, 125–134, https://doi.org/10.1007/Bf00300871 (1985).
Kayanne, H., Harii, S., Ide, Y. & Akimoto, F. Recovery of coral populations after the 1998 bleaching on Shiraho Reef, in the southern Ryukyus, NW Pacific. Marine Ecology Progress Series 239, 93–103, https://doi.org/10.3354/meps239093 (2002).
Hoegh-Guldberg, O. et al. Coral reefs under rapidclimate change and ocean acidification. Science 318, 1737–1742, https://doi.org/10.1126/science.1152509 (2007).
Lenz, E. A., Bramanti, L., Lasker, H. R. & Edmunds, P. J. Long-term variation of octocoral populations in St. John, US Virgin Islands. Coral Reefs 34, 1099–1109, https://doi.org/10.1007/s00338-015-1315-x (2015).
Ruzicka, R. R. et al. Temporal changes in benthic assemblages on Florida Keys reefs 11 years after the 1997/1998 El Nino. Marine Ecology Progress Series 489, 125–141, https://doi.org/10.3354/meps10427 (2013).
Harii, S., Hongo, C., Ishihara, M., Ide, Y. & Kayanne, H. Impacts of multiple disturbances on coral communities at Ishigaki Island, Okinawa, Japan, during a 15 year survey. Marine Ecology Progress Series 509, 171-+, https://doi.org/10.3354/meps10890 (2014).
Atrigenio, M., Porfirio, A. & Conaco, C. Influence of the Blue Coral Heliopora coerulea on Scleractinian Coral Larval Recruitment. Journal of Marine Biology 2017, 1–5, https://doi.org/10.1155/2017/6015143 (2017).
Pratlong, M. et al. The red coral (Corallium rubrum) transcriptome: a new resource for population genetics and local adaptation studies. Mol Ecol Resour 15, 1205–1215, https://doi.org/10.1111/1755-0998.12383 (2015).
Burge, C. A., Mouchka, M. E., Harvell, C. D. & Roberts, S. Immune response of the Caribbean sea fan, Gorgonia ventalina, exposed to an Aplanochytrium parasite as revealed by transcriptome sequencing. Front Physiol 4, 180, https://doi.org/10.3389/fphys.2013.00180 (2013).
Putnam, N. H. et al. Sea anemone genome reveals ancestral eumetazoan gene repertoire and genomic organization. Science 317, 86–94, https://doi.org/10.1126/science.1139158 (2007).
Shinzato, C., Inoue, M. & Kusakabe, M. A snapshot of a coral “holobiont”: a transcriptome assembly of the scleractinian coral, porites, captures a wide variety of genes from both the host and symbiotic zooxanthellae. PLoS One 9, e85182, https://doi.org/10.1371/journal.pone.0085182 (2014).
Shinzato, C. et al. Using the Acropora digitifera genome to understand coral responses to environmental change. Nature 476, 320–323, https://doi.org/10.1038/nature10249 (2011).
Aranda, M. et al. Genomes of coral dinoflagellate symbionts highlight evolutionary adaptations conducive to a symbiotic lifestyle. Sci Rep 6, 39734, https://doi.org/10.1038/srep39734 (2016).
Chen, F., Mackey, A. J., Stoeckert, C. J. Jr. & Roos, D. S. OrthoMCL-DB: querying a comprehensive multi-species collection of ortholog groups. Nucleic Acids Res 34, D363–368, https://doi.org/10.1093/nar/gkj123 (2006).
Srivastava, M. et al. The Amphimedon queenslandica genome and the evolution of animal complexity. Nature 466, 720–726, https://doi.org/10.1038/nature09201 (2010).
Park, E. et al. Estimation of divergence times in cnidarian evolution based on mitochondrial protein-coding genes and the fossil record. Mol Phylogenet Evol 62, 329–345, https://doi.org/10.1016/j.ympev.2011.10.008 (2012).
Figueroa, D. F. & Baco, A. R. Octocoral mitochondrial genomes provide insights into the phylogenetic history of gene order rearrangements, order reversals, and cnidarian phylogenetics. Genome Biol Evol 7, 391–409, https://doi.org/10.1093/gbe/evu286 (2014).
Hume, B. C. et al. Symbiodinium thermophilum sp. nov., a thermotolerant symbiotic alga prevalent in corals of the world’s hottest sea, the Persian/Arabian Gulf. Sci Rep 5, 8562, https://doi.org/10.1038/srep08562 (2015).
Le Goff, C. et al. Carbonic Anhydrases in Cnidarians: Novel Perspectives from the Octocorallian Corallium rubrum. PLoS One 11, e0160368, https://doi.org/10.1371/journal.pone.0160368 (2016).
Bhattacharya, D. et al. Comparative genomics explains the evolutionary success of reef-forming corals. Elife 5, https://doi.org/10.7554/eLife.13288 (2016).
Zoccola, D. et al. Bicarbonate transporters in corals point towards a key step in the evolution of cnidarian calcification. Sci Rep 5, 9983, https://doi.org/10.1038/srep09983 (2015).
Takeuchi, T., Yamada, L., Shinzato, C., Sawada, H. & Satoh, N. Stepwise Evolution of Coral Biomineralization Revealed with Genome-Wide Proteomics and Transcriptomics. PLoS One 11, e0156424, https://doi.org/10.1371/journal.pone.0156424 (2016).
Fukuda, I. et al. Molecular cloning of a cDNA encoding a soluble protein in the coral exoskeleton. Biochemical and Biophysical Research Communications 304, 11–17, https://doi.org/10.1016/s0006-291x(03)00527-8 (2003).
Eguchi, M. Fossil Helioporidae from Japan and the South Sea Islands. Journal of Paleontology (1948).
Villanueva, R. D. Cryptic speciation in the stony octocoral Heliopora coerulea: temporal reproductive isolation between two growth forms. Marine Biodiversity 46, 503–507, https://doi.org/10.1007/s12526-015-0376-y (2015).
Ruiz-Jones, L. J. & Palumbi, S. R. Tidal heat pulses on a reef trigger a fine-tuned transcriptional response in corals to maintain homeostasis. Sci Adv 3, e1601298, https://doi.org/10.1126/sciadv.1601298 (2017).
Paxton, C. W., Davy, S. K. & Weis, V. M. Stress and death of cnidarian host cells play a role in cnidarian bleaching. J Exp Biol 216, 2813–2820, https://doi.org/10.1242/jeb.087858 (2013).
Bellantuono, A. J., Hoegh-Guldberg, O. & Rodriguez-Lanetty, M. Resistance to thermal stress in corals without changes in symbiont composition. Proc Biol Sci 279, 1100–1107, https://doi.org/10.1098/rspb.2011.1780 (2012).
Lutz, A., Raina, J. B., Motti, C. A., Miller, D. J. & van Oppen, M. J. Host Coenzyme Q Redox State Is an Early Biomarker of Thermal Stress in the Coral Acropora millepora. PLoS One 10, e0139290, https://doi.org/10.1371/journal.pone.0139290 (2015).
Torrents, O., Tambutte, E., Caminiti, N. & Garrabou, J. Upper thermal thresholds of shallow vs. deep populations of the precious Mediterranean red coral Corallium rubrum (L.): Assessing the potential effects of warming in the NW Mediterranean. Journal of Experimental Marine Biology and Ecology 357, 7–19, https://doi.org/10.1016/j.jembe.2007.12.006 (2008).
Baker, D. M. et al. Productivity links morphology, symbiont specificity and bleaching in the evolution of Caribbean octocoral symbioses. ISME J 9, 2620–2629, https://doi.org/10.1038/ismej.2015.71 (2015).
Berkelmans, R. & van Oppen, M. J. The role of zooxanthellae in the thermal tolerance of corals: a ‘nugget of hope’ for coral reefs in an era of climate change. Proc Biol Sci 273, 2305–2312, https://doi.org/10.1098/rspb.2006.3567 (2006).
LaJeunesse, T. C. et al. Long-standing environmental conditions, geographic isolation and host-symbiont specificity influence the relative ecological dominance and genetic diversification of coral endosymbionts in the genus Symbiodinium. J Biogeogr 37, 785–800, https://doi.org/10.1111/j.1365-2699.2010.02273.x (2010).
Pochon, X. et al. Comparison of Endosymbiotic and Free-Living Symbiodinium (Dinophyceae) Diversity in a Hawaiian Reef Environment1. Journal of Phycology 46, 53–65, https://doi.org/10.1111/j.1529-8817.2009.00797.x (2010).
Lin, Z. et al. Transcriptome profiling of Galaxea fascicularis and its endosymbiont Symbiodinium reveals chronic eutrophication tolerance pathways and metabolic mutualism between partners. Sci Rep 7, 42100, https://doi.org/10.1038/srep42100 (2017).
Houlbreque, F. & Ferrier-Pages, C. Heterotrophy in tropical scleractinian corals. Biol Rev Camb Philos Soc 84, 1–17, https://doi.org/10.1111/j.1469-185X.2008.00058.x (2009).
Fabricius, K. E. & Klumpp, D. W. Widespread mixotrophy in reef-inhabiting soft corals: the influence of depth, and colony expansion and contraction on photosynthesis. Marine Ecology Progress Series 125, 195–204 (1995).
van de Water, J., Allemand, D. & Ferrier-Pages, C. Host-microbe interactions in octocoral holobionts - recent advances and perspectives. Microbiome 6, 64, https://doi.org/10.1186/s40168-018-0431-6 (2018).
Hughes, A. D. & Grottoli, A. G. Heterotrophic compensation: a possible mechanism for resilience of coral reefs to global warming or a sign of prolonged stress? PLoS One 8, e81172, https://doi.org/10.1371/journal.pone.0081172 (2013).
Lin, M. F. et al. Analyses of corallimorpharian transcriptomes provide new perspectives on the evolution of calcification in the Scleractinia (corals). Genome Biol Evol. https://doi.org/10.1093/gbe/evw297 (2017).
Moya, A. et al. Whole transcriptome analysis of the coral Acropora millepora reveals complex responses to CO(2)-driven acidification during the initiation of calcification. Mol Ecol 21, 2440–2454, https://doi.org/10.1111/j.1365-294X.2012.05554.x (2012).
Watanabe, T., Fukuda, I., China, K. & Isa, Y. Molecular analyses of protein components of the organic matrix in the exoskeleton of two scleractinian coral species. Comp Biochem Physiol B Biochem Mol Biol 136, 767–774, https://doi.org/10.1016/s1096-4959(03)00177-5 (2003).
Zhang, F., Cai, W., Zhu, J., Sun, Z. & Zhang, J. In situ raman spectral mapping study on the microscale fibers in blue coral (Heliopora coerulea) skeletons. Anal Chem 83, 7870–7875, https://doi.org/10.1021/ac2017663 (2011).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120, https://doi.org/10.1093/bioinformatics/btu170 (2014).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat Protoc 8, 1494–1512, https://doi.org/10.1038/nprot.2013.084 (2013).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659, https://doi.org/10.1093/bioinformatics/btl158 (2006).
Baumgarten, S. et al. The genome of Aiptasia, a sea anemone model for coral symbiosis. Proc Natl Acad Sci USA 112, 11893–11898, https://doi.org/10.1073/pnas.1513318112 (2015).
Chapman, J. A. et al. The dynamic genome of Hydra. Nature 464, 592–596, https://doi.org/10.1038/nature08830 (2010).
Wang, X. et al. Draft genomes of the corallimorpharians Amplexidiscus fenestrafer and Discosoma sp. Mol Ecol Resour 17, e187–e195, https://doi.org/10.1111/1755-0998.12680 (2017).
Shoguchi, E. et al. Draft assembly of the Symbiodinium minutum nuclear genome reveals dinoflagellate gene structure. Curr Biol 23, 1399–1408, https://doi.org/10.1016/j.cub.2013.05.062 (2013).
Lin, S. et al. The Symbiodinium kawagutii genome illuminates dinoflagellate gene expression and coral symbiosis. Science 350, 691–694, https://doi.org/10.1126/science.aad0408 (2015).
Baumgarten, S. et al. Integrating microRNA and mRNA expression profiling in Symbiodinium microadriaticum, a dinoflagellate symbiont of reef-building corals. BMC Genomics 14, 704, https://doi.org/10.1186/1471-2164-14-704 (2013).
Rosic, N. et al. Unfolding the secrets of coral-algal symbiosis. ISME J 9, 844–856, https://doi.org/10.1038/ismej.2014.182 (2015).
Liew, Y. J., Aranda, M. & Voolstra, C. R. Reefgenomics.Org - a repository for marine genomics data. Database (Oxford) 2016, https://doi.org/10.1093/database/baw152 (2016).
Winnepenninckx, B., Backeljau, T. & De Wachter, R. Extraction of high molecular weight DNA from molluscs. Trends Genet 9, 407 (1993).
Lajeunesse, T. C. & Trench, R. K. Biogeography of two species of Symbiodinium (Freudenthal) inhabiting the intertidal sea anemone Anthopleura elegantissima (Brandt). Biol Bull 199, 126–134, https://doi.org/10.2307/1542872 (2000).
Sampayo, E. M., Dove, S. & Lajeunesse, T. C. Cohesive molecular genetic data delineate species diversity in the dinoflagellate genus Symbiodinium. Mol Ecol 18, 500–519, https://doi.org/10.1111/j.1365-294X.2008.04037.x (2009).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7, 539, https://doi.org/10.1038/msb.2011.75 (2011).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17, 540–552 (2000).
Acknowledgements
We thank the OIST High Performance Computing facility for server usage and the Bolinao Marine Laboratory and Michael Atrigenio for assistance with coral collections. We also thank Steven D. Aird for editing the manuscript. This study was funded by the University of the Philippines System Emerging Interdisciplinary Research Grant (OVPAA-EIDR-C04-005) to CC.
Author information
Authors and Affiliations
Contributions
C.C. designed the study; C.G. performed the experiments; C.G., C.S., T.M.L. and C.C. analyzed the data. C.G., C.S. and C.C. wrote the manuscript. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guzman, C., Shinzato, C., Lu, TM. et al. Transcriptome analysis of the reef-building octocoral, Heliopora coerulea. Sci Rep 8, 8397 (2018). https://doi.org/10.1038/s41598-018-26718-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-26718-5
This article is cited by
-
Application of phylogenomic tools to unravel anthozoan evolution
Coral Reefs (2022)
-
The skeletome of the red coral Corallium rubrum indicates an independent evolution of biomineralization process in octocorals
BMC Ecology and Evolution (2021)
-
De novo transcriptome assembly from the gonads of a scleractinian coral, Euphyllia ancora: molecular mechanisms underlying scleractinian gametogenesis
BMC Genomics (2020)
-
Integrated evidence reveals a new species in the ancient blue coral genus Heliopora (Octocorallia)
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.