Abstract
The GO-Cellular Component (GO-CC) ontology provides a controlled vocabulary for the consistent description of the subcellular compartments or macromolecular complexes where proteins may act. Current machine learning-based methods used for the automated GO-CC annotation of proteins suffer from the inconsistency of individual GO-CC term predictions. Here, we present FGGA-CC+, a class of hierarchical graph-based classifiers for the consistent GO-CC annotation of protein coding genes at the subcellular compartment or macromolecular complex levels. Aiming to boost the accuracy of GO-CC predictions, we make use of the protein localization knowledge in the GO-Biological Process (GO-BP) annotations to boost the accuracy of GO-CC prediction. As a result, FGGA-CC+ classifiers are built from annotation data in both the GO-CC and GO-BP ontologies. Due to their graph-based design, FGGA-CC+ classifiers are fully interpretable and their predictions amenable to expert analysis. Promising results on protein annotation data from five model organisms were obtained. Additionally, successful validation results in the annotation of a challenging subset of tandem duplicated genes in the tomato non-model organism were accomplished. Overall, these results suggest that FGGA-CC+ classifiers can indeed be useful for satisfying the huge demand of GO-CC annotation arising from ubiquitous high throughout sequencing and proteomic projects.
Similar content being viewed by others
Introduction
Eukaryotic cells are organized into a complex structure of subcellular compartments called organelles. Proteins synthesized in ribosomes can be trafficked to different organelles for the accomplishment of specific physiological functions. Hence, it is not surprising that unexpected protein subcellular localization often underlies the pathogenesis of many human diseases1,2,3. Proteins synthesized in ribosomes can also interact to form macromolecular complexes4 -naturally occurring machines inside cells- playing crucial roles in a variety of cellular processes5,6. The GO-CC ontology provides a controlled vocabulary for consistent description of both the subcellular structure or macromolecular complex location where proteins may act. Diverse experimental methods can be used to accurately determine the subcellular localization of proteins7, ranging from the identification of specific signals on cargo proteins8,9 to the use of advanced imaging techniques for revealing protein composition of organelles10,11,12. Similarly, a combination of chemical crosslinking13, mass spectrometry, and cryo-electron microscopy14 methods can be used to accurately determine the structure and function of macromolecular complexes. Although all these advanced experimental methods are beginning to bear fruits15,16, their time-consuming nature and elevated costs17,18 make then incompatible with current GO-CC protein annotation demands from ubiquitous large-scale sequencing and proteomic projects. In this scenario, in-silico methods for the automated GO-CC annotation of proteins, i.e., for predicting their localization, at the subcellular structure or macromolecular complex levels, become promising alternatives19,20,21,22. However, few studies have considered this problem as a whole, CELLO2GO23 and FFPred324 being two important exceptions. The CELLO2GO method entails a sequence-based approach for predicting the GO localization of proteins based on their homology to previously localized proteins, mostly belonging to model organisms. On the other hand, the FFPred3 method entails a machine learning-based approach for (separately) predicting all GO domains, including GO-CC, with main focus on divergent human protein chains for which homology-based methods can provide little aid. Since machine learning-based GO annotation methods can overcome the limitations of straightforward homology-based alternatives, they are particularly attractive for the annotation of proteins from non-model organisms.
Taking into account that the study of non-model organisms provides new opportunities for understanding the evolution of multicellular life and cell biological processes25, and that substantial reductions in the cost of DNA sequencing have recently burst their study, more efforts on the improvement of machine learning-based methods for GO protein localization are required. In this regard, it is worthy of note that only simplified versions of this problem have been mostly considered in literature. In particular, the prediction of a reduced set of subcellular localization (SCL) categories, often extracted from the SCL section of UniProt entries26, has been a frequently revisited problem considering single27,28,29 or multi-category30,31,32,33 prediction outputs. From a biological point of view, multi-category prediction methods are preferable since relevant proteins often show a ubiquitous character. In either case, protein SCL categories are predicted using the knowledge available at previously localized proteins by characterizing their sequences in terms of a fixed number of informative features. These features may range from the frequency of amino-acids to the existence of low-complexity regions, signal peptides, or trans-membrane helices34, among others. On the other hand, the unified prediction of highly-specific protein localization categories derived from ad-hoc ontologies like GO-CC has been occasionally considered.
Recalling that an ontology embodies a controlled vocabulary of terms and well-known relationships between them, in-silico methods for GO-CC protein localization can be further differentiated by the consistency of individual GO-CC term predictions. We note, however, that although admittedly important35, the consistency problem of ontology-based predictions has been rarely considered in literature. For example, GO-CC FFPred3 predictions are built from a predefined flat -unware of ontology relationships- set of 89 binary GO-CC term predictions; a final propagation step from selected leaf GO-CC terms to the root is then used to accomplish consistent GO-CC predictions. We note, however that consistent GO-CC predictions obtained this way may not be unique, and may not be optimal with respect to the minimization of the probability of erroneous GO-term predictions, since neither the prediction noise of flat GO-CC classifiers, nor the relationships between GO-CC terms are considered. In particular, false positive predictions will be always propagated to the root instead of attempting the prediction of less specific but easier terms, that could improve overall prediction accuracy.
In this paper, a graphical model-based machine learning approach for the automated and consistent GO-CC annotation of protein coding genes is presented. While the graphical component is used to specify the GO-CC ontology, the machine learning component is used to independently learn target GO-CC categories; both these components are then appropriately combined to infer consistent GO-CC annotations. Graphical models have been long used to provide intuitive visions and useful insights in a variety of biological problems at different levels of complexity, including the prediction of metabolic pathways36, the prediction of protein functions37, and the analysis of complicated drug metabolic systems38. Regarding our GO-CC annotation problem, we specifically rely on the power of factor graph models39 for obtaining a graphical and formal specification of the GO-CC ontology, for modeling the prediction noise of flat binary classifiers used to predict individual GO-CC categories, for graphically approximating A Posteriori Probabilities (APP) of individual GO-CC categories, for computing corresponding Maximum A Posteriori (MAP) estimates, and for downstream expert analysis of GO-CC predictions. Building upon these concepts, we present FGGA-CC+ classifiers, hierarchical ensembles of binary classifiers allowing the straightforward inference of consistent GO-CC annotations by means of the execution of the well-known Sum-Product algorithm39 in factor graphs.
Initial insights about FGGA-CC+ classifiers were obtained with the former introduction of FGGA classifiers40, hierarchical ensembles of binary classifiers defined over GO Molecular Function (GO-MF) compliant factor graphs designed to tackle the automated and consistent GO-MF annotation of protein coding genes. Aiming to deal with GO-CC annotations, FGGA classifiers are now extended into FGGA-CC+ counterparts. For this purpose, the factor graph modeling of the transitive is_a and part_of relationships between GO-CC terms is first considered. In addition, the factor graph modeling of the non-transitive occurs_in relationship between GO-CC and GO-BP terms, useful for specifying the subcellular location where a biological process occurs, is complimentary considered. As a result of the occurs_in modeling, relevant protein subcellular localization knowledge already available at the GO-BP subdomain can be formally exploited for the enrichment of GO-CC predictions.
Results
Characterization of protein sequences for their consistent GO-CC annotation with FGGA-CC+ classifiers
Automated GO-CC annotation of protein sequences with FGGA-CC+ classifiers requires their characterization in terms of a fixed number of informative protein features. Table 1 shows the average hierarchical Precision (HP), Recall (HR) and F-score (HF) results accomplished by native FGGA-CC classifiers, i.e., FGGA-CC+ classifiers without the GO-BP enrichment stage, on A. thaliana protein sequences. Firstly, a significant effect of characterization methods on hierarchical F-score results is revealed (p < 0.01; Friedman’s test). Secondly, significant differences (see Supplementary Tables S1 and S2) in favor of the Physicochemical+ characterization method are observed (p < 0.01; Wilcoxon test with Bonferroni correction).
GO-CC annotation of protein sequences with FGGA-CC+ classifiers
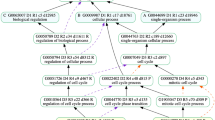
FGGA-CC+ classifiers were evaluated on protein sequences from five model organisms, D. rario, A. thaliana, S. cerevisiae, D. melanogaster and M. musculus, using a 5-fold cross-validation approach. In all cases, a Physicochemical+ characterization of protein sequences was used, and hierarchical Precision, Recall and F-score performance metrics were evaluated. A first insight into the benefits of requiring consistent GO-CC predictions can be appreciated in Fig. 1 where FGGA-CC+ processing over flat GO-CC predictions promotes consistency and reduces the number of false-positives.
GO-CC subgraphs induced in the annotation of the Q7ZVT3 protein in the D. rario model organism. Positive annotations are shown in light blue, negative ones in white, and erroneous ones with a crossline. (a) GO-CC annotations accomplished by a naive ensemble of SVM classifiers; erroneous/inconsistent annotations can be observed. (b) GO-CC annotations after FGGA-CC+ processing; consistent annotations, including just one false positive, can be observed.
A first round of evaluations was performed to evaluate the baseline annotation performance of FGGA-CC+ classifiers. For this purpose, native FGGA-CC classifiers were evaluated against naive ensembles of binary SVM classifiers trained to predict just individual GO-CC categories. FGGA-CC classifiers not only yielded better Area Under Curve (AUC) scores but did a particular good job at predicting more specific/deeper GO-CC terms (see Fig. 2 and Figure Supplementary S1).
Scatter plots of the average AUC scores attained by native FGGA-CC and baseline ensembles of SVM classifiers when performing the GO-CC annotation of protein sequences characterized by the Physicochemical+ method. As deeper GO-CC categories are considered, points in the scatter plot turn from yellow to red.
A second round of evaluations was performed to evaluate the actual benefits of introducing of SCL knowledge available in boundary GO-BP terms. For this purpose, GO-BP enriched FGGA-CC+ classifiers were evaluated against their FGGA-CC alternatives. As expected, FGGA-CC+ classifiers yielded higher AUC scores and did a particular good job at predicting even more specific/deeper GO-CC terms (see Fig. 3 and Figure Supplementary S2). Noteworthy, FGGA-CC+ classifiers noticeable increased the number of true positive GO-CC annotations (see Figure Supplementary S3). Furthermore, a graphical comparison of predicted GO-CC categories by the two classifiers revealed that these improvements came from positive FGGA-CC+ annotations to rather specific GO-CC nodes directly connected, or in the vicinity of, contributing boundary GO-BP nodes. This result makes sense since GO-BP contributing nodes are generally connected to GO-CC nodes located at certain depth. In addition, aiming to quantify observed differences between FGGA-CC+ and FGGA-CC classifiers taking into account consistency requirements, the hierarchical precision, recall and F-score performance metrics were evaluated. As expected, the advantages of FGGA-CC+ classifiers with respect to FGGA-CC alternatives were confirmed in all but the D. rario dataset (see Table 2), for which only one additional boundary GO-BP term was available.
Scatter plots of the average AUC scores attained by GO-BP enriched FGGA-CC+ and native FGGA-CC classifiers when performing the GO-CC annotation of protein sequences characterized by the Physicochemical+ method. As deeper GO-CC categories are considered, points in the scatter plot turn from yellow to red.
For the sake of completeness, a third round of evaluations was performed to evaluate the performance of FGGA-CC+ classifiers against two established methods for the automated GO-CC annotation of protein sequences, CELLO2GO and FFPred3. For this purpose, precision, recall and F-score performance metrics and corresponding hierarchical extensions were computed. The CELLO2GO method searches GO-CC annotated homologous proteins in the UniProtKB/TrEMBL database using the Blast algorithm. On the other hand, the FFPred3 method performs an extensive characterization of protein sequences and complementary feature selection before training a naive ensemble of SVM classifiers set to predict an empirically predefined set of 89 GO-CC categories. As a result, comparisons between methods were limited to the 89 predefined FFPred3 GO-CC categories and to the D. melanogaster model organism for which a precomputed FFPred3 characterization of protein sequences was publicly available. Based on these considerations, the Slim D. melanogaster dataset was first assembled (see Methods). A first insight on the annotation power of the three methods was assessed from ROC curves. Taking into account the natural imbalance between positively and negatively protein sequences annotated to each GO-CC category and the importance of positively annotated protein sequences, PR curves were complementary analyzed. Both ROC and PR curves showed promising comparative results for the FGGA-CC+ method (see Fig. 4). These results were further confirmed by precision, recall and F-score evaluations. In this regard, a significant effect (p < 0.01; Friedman’s test) of annotation methods on the F-score was first observed (see Table Supplementary S3); a significant difference in favor of the FGGA-CC+ method (p < 0.01; Wilcoxon test with Bonferroni correction) was afterwards observed (see Table 3, left). Finally, to shed further light on the actual comparative performance FGGA-CC+, FFPred3 and CELLO2GO methods taking into account consistency issues, hierarchical performance metrics were considered. For this end, predicted GO-CC categories by FFPred3 and CELLO2GO methods were first propagated to parent GO-CC terms. The advantages of the FGGA-CC + method could be then clearly observed (see Table 3, right).
FGGA-CC+ validation with the annotation of sHSPs in S. lycopersicum
In plants, fruit maturation and oxidative stress can induce small Heat Shock Proteins (sHSPs) synthesis to maintain cellular homeostasis. The diversity of the sHSP gene family is mostly supported by gene duplication events that result in genetic redundancy41. Protein SCL plays a key role in the functional diversification process of duplicated genes as follows from the differential distribution of their proteins across different subcellular compartments42. Current GO-CC annotation of the sHSP gene family in S. lycopersicum genome remains scarce, with less than 10 in a set of 33 gene family members having some GO-CC annotation. Here, we focus on the in-silico GO-CC annotation of three clusters (I, II and III) of tandem duplicated sHSP genes in S. lycopersicum. These three clusters involve a total of nine sHSP genes, six of them without a GO-CC annotation. In addition, at each of them, one representative sHSP gene with a GO-CC annotation supported by some experimental evidence is present (Solyc06g076520, Solyc08g062450, and Solyc08g078700). In this regard, we recall that a cytosolic SCL annotation has been reported for Solyc06g07652043 belonging to cluster I comprising four sHSP genes in chromosome 6, that a chloroplast SCL annotation has been reported for Solyc08g06245044 belonging to a cluster II comprising two sHSP genes in chromosome 8, and that a mitochondria SCL annotation has been reported for Solyc08g07870045 belonging to cluster III comprising three sHSP genes also in chromosome 8. Aiming to shed light on the GO-CC annotation of the six remaining sHSP genes without a GO-CC annotation, a FGGA-CC+ classifier trained on A. thaliana protein sequences was considered. Owing to the tandem constraint, we further expect that FGGA-CC+ annotations are consistent with the common ancestral origin of sHSP genes within each cluster, i.e., cytosolic, chloroplast, and mitochondrial related GO-CC annotations are respectively expected for sHSP genes in Clusters I, II, and III. Recalling that for hierarchical ensembles of classifiers like FGGA-CC+, a prediction is considered correct as long as the actual solution is contained in the predicted graph, the three positive GO-CC annotation controls were verified (see Table 4 and Figures Supplementary S4–S6). Expected GO-CC annotations within the three clusters were confirmed for all but one, with Cluster I and II retaining their ancestral cytosolic and chloroplastic localization respectively. Meanwhile, in Cluster III, the mitochondrial localization was only predicted for Solyc08g078700 and Solyc08g078720. For the remaining related gene Solyc08g078710, a nonspecific organelle localization was predicted, possible due to lack of some peptide signal, adding further evidence for its gene pseudogenization process.
Discussion and Conclusions
A graphical model-based machine learning approach for the automated and consistent GO-CC annotation of protein sequences has been presented. In this approach, a novel class of hierarchical classifiers, named FGGA-CC+, map the GO-CC protein annotation problem to that of discovering hidden nodes in factor graphs defined by, latent GO-CC and GO-BP categories, semantic relationships between categories, observable predictions of individual categories, and probability density functions modeling the prediction noise over individual categories. As a result, inconsistencies among observable GO-CC and GO-BP predictions -issued by binary SVM classifiers- can be transparently handled the well-known iterative Sum-Product algorithm in factor graphs. At the end of this leveraging process, a set of consistent GO-CC and GO-BP annotations are obtained. These computational modeling efforts are paid off when observing the improvement of AUC scores accomplished by native FGGA-CC classifiers with regard to naive ensembles of binary SVM classifiers. Similarly, they are pay-off when observing the improvement of AUC and hierarchical performance metrics of FGGA-CC+ classifiers with regard to their native FGGA-CC alternatives. Concerning mandatory comparisons of FGGA-CC+ classifiers with state of art GO-CC annotation methods like FFPred3 and CELLO2GO, flat performance metrics were first considered. In the former case, significant improvements on F-score results were observed. However, in terms of precision performance, the CELLO2GO method performed better, but at lower recall levels. We wonder if we could reach CELLO2GO precision levels (0.65) by raising the threshold of our decisions. In effect, we found that by raising the decision threshold from 0.5 to 0.9, an average precision of 0.65 with an average recall of 0.57 (F-score of 0.56) was accomplished (see Supplementary Table S4), suggesting that FGGA-CC+ classifiers can accommodate a wide spectrum of precision/recall requirements. To shed light on the actual comparative performance of the FGGA-CC+, FFPred3 and CELLO2GO methods, hierarchical performance metrics were then considered. In these evaluations, the advantages of the FGGA-CC+ method could be clearly observed, with comparable results in terms of the hierarchical precision and significant better results in terms of the hierarchical recall.
For purposes of FGGA-CC+ validation, the challenging annotation of nine sHSP genes of tandem duplication origin in the tomato genome was considered. Verification of three positive controls allowed us to tackle the in-silico annotation of the six remaining sHSP genes. Consistent GO-CC annotation results were mostly observed with the sole exception of Solyc08g078710, for which a nonspecific organelle prediction, instead of a mitochondrial one, was obtained. Posterior analysis of the sHSP characterization patterns used for FGGA-CC+ queries revealed that differently from its two tandem duplicated counterparts, Solyc08g078710 lacks of a key signal allowing its transport into the mitochondria. This signal is the Tom20 motif, a mitochondrial targeting signal expected at the N-terminal presequences that is recognized by the Tom20 import receptor at the outer mitochondrial membrane46. This finding points out the importance of using a comprehensive, GO-CC specific, characterization of protein sequences for their reliable GO-CC annotation.
Computational and biological concepts underlying the design of FGGA-CC+ classifiers, from techniques used for the characterization of protein sequences, to the factor graph modeling of target GO-CC subgraphs, including the integration of GO-BP knowledge and the modeling of GO-CC prediction noise at flat binary SVM classifiers, provide a systematic framework for designing a computational tool allowing the integral GO annotation of protein sequences. By characterizing a sufficient large collection of annotated proteins in the three GO subdomains, including those coming from orthologous protein coding genes, it should be possible to provide accurate and integral and precise GO annotations of protein coding genes in many non model organisms. Finally, as pointed out in47, and demonstrated in a series of recent publications33,48,49, user-friendly and publicly accessible web-servers represent the future direction for developing practically more useful prediction methods and computational tools. In this regard, we shall make efforts in our future work to provide a web-server for the method presented in this paper.
Methods
To develop really useful sequence-based statistical classifiers for a biological system, such as those reported in a series of recent publications33,48,50, one should observe the 5-step rule51. As a result, one should make the following five steps very clear: (i) how to construct a valid benchmark dataset to train and test the classifiers; (ii) how to characterize protein sequences so that they can reflect their intrinsic correlation with target categories; (iii) how to develop a powerful algorithm for predicting target categories; (iv) how to properly perform cross-validation tests to objectively evaluate the anticipated accuracy of classifiers; and (v) how to establish a user-friendly web-server for the classifiers that is accessible to the public. In what follows, we describe how to deal with these steps one-by-one for the specific case of FGGA-CC+ classifiers and the prediction of GO-CC categories.
Datasets
Benchmark datasets
GO-CC annotation data with experimental and computational evidence codes [http://geneontology.org/page/guide-go-evidence-codes] was first collected. Regarding experimental codes, Inferred from Experiment (EXP), inferred from Direct Assay (IDA), Inferred from Physical Interaction (IPI), Inferred from Mutant Phenotype (IMP), inferred from Genetic Interaction (IGI) and Inferred from Expression Pattern (IEP), were considered. Regarding computational evidence codes, inferred from Sequence or structural Similarity (ISS), inferred from Sequence Orthology (ISO), inferred from Sequence Alignment (ISA) and inferred from Sequence Model (ISM), were considered. In addition, annotation data was also collected for GO-BP boundary nodes, i.e., GO-BP terms connected to GO-CC terms through the non-transitive occurs_in relationship. For GO-BP boundary nodes, we considered annotation data with experimental, computational or Inferred from Electronic Annotation (IEA) evidence codes. This kind of soft annotation data policy for GO-BP boundary nodes aims to compensate the lack of sufficient protein sequences with experimental or computational evidence codes that may overshadow the actual power of GO-BP boundary nodes for the enhancement of GO-CC predictions. Regarding minimum requirements for learning individual GO terms with binary SVM classifiers, a minimum of 50 positively annotated protein sequences was considered. In addition, to assemble conveniently balanced binary training datasets52, positively annotated protein sequences were complemented with negative annotated protein counterparts using the inclusive separation policy53. As shown in Table 5, datasets comprising both positively and negatively GO-CC annotated protein sequences from five models organisms, D. rario54, A. thaliana55, S. cerevisiae56, D. melanogaster57 and M. musculus58, were finally assembled.
Slim D. melanogaster dataset
D. melanogaster protein sequences were collated from the UniProt database based on their annotation to any of the 89 GO-CC categories predefined by the FFPred3 method; to allow fair hierarchical comparisons of GO-CC predictions between methods, 22 ancestor GO-CC categories were also included. In addition, the same evidence codes of benchmark datasets were used. Taking into account the high computational overhead of FFPred3 GO-CC predictions, a reduced set of 270 protein sequences was finally considered (see Supplementary Data file 1).
S. lycopersicum (cv. Heinz 1706) sHSP dataset
Although the tomato reference genome was published in 2012, the functionality of sHSP genes in this model organism for fleshy fruit development remains mostly unknown. Using a transcriptomic (RNA-seq) and evolutionary genomic approach, a family of thirty-three sHSP genes in S. lycopersicum (cv. Heinz 1706) genome was recently established41. Here, the GO-CC annotation of this gene family is considered. We restrict our attention to tandem duplicated sHSP genes arranged into physical clusters with at least one of its members being up-regulated during fruit ripening to ensure functionality, and with a established SCL annotation to validate annotations at each cluster. As a result, the GO-CC annotation of nine tandem duplicated genes sHSP genes arranged into three physical clusters was tackled: a cluster of four members (Solyc06g076520, Solyc06g076540, Solyc06g076560, Solyc06g076570) in chromosome 6, a cluster of two members (Solyc08g078710, Solyc08g078720) in chromosome 8, and another cluster of three members (Solyc08g062340, Solyc08g062450, Solyc08g078700) also in chromosome 8.
Protein sequence representation
Protein sequences were characterized in terms of a fixed number features. As shown in Table 6, four characterization methods were analyzed: (i) Signal+, encoding features used by well-known TargetP27, SignalP59, Transmembrane Helices60, WoLF PSORT61, and MitoFates62 tools for the prediction of standard SCL categories, along with the presence/absence of localization signals collected in the LocSigDB63 database, (ii) PrositeBin encoding just the presence/absence of Prosite domains, (iii) Signal+ +, encoding features in the Signal+ and PrositeBin characterizations, and (iv) Physicochemical+, encoding features in the Signal+ characterization, together with of physicochemical and secondary structure properties64,65,66,67. Characterization methods were implemented with in-house R scripts. To assess the effect of characterization methods in the prediction power of FGGA-CC+ classifiers, the largest annotation dataset (A. thaliana) was considered.
Consistent GO-CC annotation of protein sequences with FGGA-CC+ classifiers
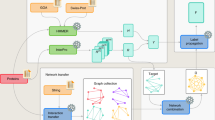
GO-CC annotation of protein sequences was first tackled with native FGGA-CC classifiers, hierarchical ensembles of binary SVM classifiers relying on the power of factor graph models for overcoming inconsistencies among flat SVM predictions of individual GO-CC categories (see Fig. 5). FGGA-CC classifiers arise as a natural extension of FGAA classifiers originally developed for the automated and consistent GO-MF annotation of protein coding genes. Both the GO-MF and GO-CC subontologies make extensive use of the fundamental transitive is-a relationship. In addition, the GO-CC subontology makes extensive use of the transitive part of relationship. Both relationships are considered by native FGGA-CC classifiers when performing GO-CC annotations. Aiming to accomplish more accurate GO-CC annotations, the integration of SCL knowledge from GO-BP boundary terms was additionally considered. As a result, native FGGA-CC classifiers were further extended into FGGA-CC+ classifiers. To accomplish this extension, the non-transitive occurs-in relationship between GO-CC and GO-BP terms was further considered. However, since non-transitive relationships may lead to non-transitive inference paths precluding the free propagation68,69 and consistency checking of GO-CC annotations in supporting factor graphs, a transitive closure screening process was introduced prior to factor graph modeling. After transitive closure processing, the resulting GO subgraph is ready to be transformed into a factor graph classification model using roughly the same methodology described in40.
GO-CC annotation of a protein sequences with FGGA-CC classifiers. A GO-CC subgraph defining the expected structure of GO-CC predictions is first converted to a factor graph (FG) model. Protein sequences of any length are characterized in terms of a fixed number of features. Flat binary SVM classifiers (SVM i ) predict individual GO-CC categories (GO:i) upon protein sequence queries. Flat, likely inconsistent, binary GO-CC predictions are leveraged by executing the Sum-Product algorithm on the FG model. At the end, a set of consistent GO-CC predictions is obtained.
Transitive Closure Screening of GO-CC subgraphs enriched with GO-BP boundary nodes
Given a GO-CC subgraph enriched with GO-BP boundary nodes, a transitive closure screening process is performed using a Depth-First Search (DFS) algorithm70 ignoring repeated nodes. Starting from a bottom-leaf node, a link between a child node and its parent node is accepted only if, for all grandparent nodes, the boolean function h is satisfied for all composite child-parent-grandparent relationships. If any of these h evaluations fail, the child-parent link is deleted. In the definition of h, all reasoning rules established for standard71 and experimental72 relationships, like the occurs_in, are considered (see Table 7).
For example, in Fig. 6a, the transitive closure of inference paths in a GO subgraph including non-transitive relationships is analyzed. In particular, the presence of the GO:7 → GO:6 link involving the non-transitive regulates relationship is evaluated. Since h is verified by composite path GO:7 → GO:6 → GO:4 but is rejected by composite path GO:7 → GO:6 → GO:5, the GO:7 → GO:6 link gets removed (see Fig. 6b).
(a) Original GO subgraph (b) After checking the graph transitive closure with the boolean function h, the link GO:7 → GO:6 is deleted (c) Factor graph model used for GO-CC predictions. Inside the dash lined box, a core factor graph contains binary variable nodes x i modeling GO terms, and boolean factor nodes f j modeling relationships between them. Outside the dash lined box, the core factor graph is enriched with observable, real-valued, variable nodes y i modeling independent GO-CC predictions, and probabilistic factor nodes g i modeling corresponding prediction noise.
Factor graph transformation and inference of GO-CC annotations
After transitive closure screening, the resulting GO subgraph is first transformed into a core factor graph (see Fig. 6c). For this purpose, GO terms are mapped to binary variable nodes and relationships between GO terms are mapped to logical factor nodes -logical functions- implementing the True Path Graph (TPG) constraint. Specifically, the TPG constraint ensures that if a child GO term is annotated positive, then its parent GO term(s) must also be annotated positive; on the other hand, if a parent GO term is annotated negative, then its children GO term(s) must also be annotated negative. The core factor graph is then enriched with observable variable nodes and probabilistic factor nodes. Observed variable nodes model practical binary SVM predictions over ideal, but hidden/latent, variable nodes in the core factor graph. On the other hand, probabilistic factor nodes model zero mean Gaussian distributions modeling the prediction noise of practical SVM classifiers.
For a given query protein sequence, GO-CC annotations are obtained by the execution of the iterative Sum-Product algorithm between nodes of the enriched factor graph. The algorithm starts from the observable but noisy predictions at leaf nodes of the factor graph. After a few number of iterations, approximated APPs on hidden variable nodes -target GO-CC categories- can be obtained40. We recall that only approximated APPs can be guaranteed since cycles73 are naturally expected in GO-CC compliant factor graphs. From these probabilities, corresponding MAP estimates -minimizing the probability of erroneous GO-CC predictions- are obtained; practically, a maximum of 50 iterations were allowed. Note that since our GO-CC predictions follow from MAP estimates, we do not expect they are able to optimize more elaborate performance metrics like the F-score. We note, however, that the design of optimal F-score classification algorithms remain a challenging computational problem even for the prediction of flat multiclass/multilabel categories74.
Soft-margin SVM classifiers with a radial basis function kernel and default parameters were used for the prediction of individual GO-CC and GO-BP categories. To fulfill the assumption of zero-mean Gaussian prediction noise, the margins of SVM classifier outputs were used. A complementary validation stage after the training of SVM classifiers was used to assess the standard deviation of Gaussian distributions modeling the prediction noise of individual GO-CC and GO-BP categories. Practically, SVMs classifiers were implemented with e-1071 R package75. In addition, the factor graph iterative Sum-Product algorithm was implemented with in-house R [https://cran.r-project.org/] scripts.
Performance evaluation
The prediction performance of native FGGA-CC and FGGA-CC+ classifiers was evaluated with 5-fold cross-validation tests. Taking into account the hierarchical relations among target GO-CC categories, both flat hierarchical classification performance metrics were considered76. Differently from their flat counterparts, hierarchical classification performance metrics appropriately recognize partially correct classifications and correspondingly penalize more distant or more superficial errors -prediction errors at upper levels of a hierarchy should be punished more severely that those at deeper levels77. In particular, the hierarchical precision (HP), the hierarchical recall (HR), and the hierarchical F-score (HF) measures introduced in78 were used. Below are their formulas:
where s is a protein sequence, G is GO subgraph, P G (s) ⊂ G is the predicted GO subgraph of s, C G (s) ⊂ G is the actual GO subgraph of s, l(P G (s)) is the set of leaves of the P G (s) and l(C G (s)) is the set of leaves of C G (s). In addition, ↑q is the set of ancestors of a node q belonging to P G (s), and ↑c is the set of ancestors of a node c belonging to C GO (s). Concerning fair comparisons of FGGA-CC+ classifiers against established, but not hierarchical, methods for the automated GO-CC annotation of protein coding genes, the average precision, the average recall and the average F-score performance metrics were used. Specifically, for each protein sequence s, the precision p(s) was calculated as \(\frac{tp(s)}{tp(s)+fp(s)}\), the recall r(s) as \(\frac{tp(s)}{tp(s)+fn(s)}\) and the F-score as \(\frac{2\cdot p(s)\cdot r(s)}{p(s)+r(s)}\), where tp is the number of GO-CC categories correctly predicted as positives (true positives), fp is the number of GO-CC categories incorrectly predicted as positives (false positives) and fn is the number of GO-CC categories incorrectly predicted as negatives (false negatives).
Evaluation protocol
Firstly, the annotation performance of native FGGA-CC classifiers was evaluated against that of flat ensembles of binary SVM classifiers. Secondly, the annotation performance of (GO-BP enriched) FGGA-CC+ classifiers was evaluated against that of native FGGA-CC counterparts. In the former case, average AUC scores79 at individual GO-CC categories were additionally computed using the facilities in the PerfMeas package80. FGGA-CC+ classifiers were also evaluated against two established methods for GO-CC annotation, FFPred3 and CELLO2GO; literature results on these methods have been reported only with flat performance metrics. Aiming fair comparisons, both flat and hierarchical performance metrics were used. For these evaluations, the Slim D. melanogaster dataset was used. In all cases, the statistical significance of observed differences was assessed with the Friedman and Wilcoxon rank sum tests at α = 0.01 significance level.
References
Chen, Y. et al. Aberrant subcellular localization of BRCA1 in breast cancer. Sci. 270, 789–791 (1995).
Hung, M. C. & Link, W. Protein localization in disease and therapy. J. Cell. Sci. 124, 3381–3392 (2011).
Pauty, J. et al. Cancer-causing mutations in the tumor suppressor PALB2 reveal a novel cancer mechanism using a hidden nuclear export signal in the WD40 repeat motif. Nucleic Acids Res. 45, 2644–2657 (2017).
Sali, A., Glaeser, R., Earnest, T. & Baumeister, W. From words to literature in structural proteomics. Nat. 422, 216–225 (2003).
Stroud, D. A. et al. Accessory subunits are integral for assembly and function of human mitochondrial complex I. Nat. 538, 123–126 (2016).
Compe, E. & Egly, J. M. TFIIH: when transcription met DNA repair. Nat. Rev. Mol. Cell Biol. 13, 343–354 (2012).
Bauer, N. C., Doetsch, P. W. & Corbett, A. H. Mechanisms Regulating Protein Localization. Traffic 16, 1039–1061 (2015).
Ozer, R. S. & Halpain, S. Phosphorylation-dependent localization of microtubule-associated protein MAP2c to the actin cytoskeleton. Mol. Biol. Cell 11, 3573–3587 (2000).
Nardozzi, J. D., Lott, K. & Cingolani, G. Phosphorylation meets nuclear import: a review. Cell Commun. Signal 8, 32 (2010).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Sci. 313, 1642–1645 (2006).
Huang, B., Babcock, H. & Zhuang, X. Breaking the diffraction barrier: super-resolution imaging of cells. Cell 143, 1047–1058 (2010).
Kremers, G.-J., Gilbert, S. G., Cranfill, P. J., Davidson, M. W. & Piston, D. W. Fluorescent proteins at a glance. J. Cell Sci. 124, 157–160 (2010).
Suchanek, M., Radzikowska, A. & Thiele, C. Photo-leucine and photo-methionine allow identification of protein-protein interactions in living cells. Nat. Methods 2, 261–267 (2005).
Fernandez-Leiro, R. & Scheres, S. H. Unravelling biological macromolecules with cryo-electron microscopy. Nat. 537, 339–346 (2016).
Cremer, C., Szczurek, A., Schock, F., Gourram, A. & Birk, U. Super-Resolution Microscopy Approaches to Nuclear Nanostructure Imaging. Methods (2017).
Schmidt, C. & Urlaub, H. Combining cryo-electron microscopy (cryo-em) and cross-linking mass spectrometry (cx-ms) for structural elucidation of large protein assemblies. Curr. Opin. Struct. Biol. 46(157), 168 (2017).
Lee, A., Tsekouras, K., Calderon, C., Bustamante, C. & Pressé, S. Unraveling the thousand word picture: An introduction to super-resolution data analysis. Chem. Rev. 117, 7276–7330 (2017).
Zybailov, B. L., Glazko, G. V., Jaiswal, M. & Raney, K. D. Large Scale Chemical Cross-linking Mass Spectrometry Perspectives. J Proteomics Bioinform 6, 001 (2013).
Chou, K.-C. & Elrod, D. W. Protein subcellular location prediction. Protein Eng. Des. Sel. 12, 107–118 (1999).
Chou, K.-C. & Cai, Y.-D. Using functional domain composition and support vector machines for prediction of protein subcellular location. J. Biol. Chem. 277, 45765–45769 (2002).
Zhou, G.-P. & Doctor, K. Subcellular location prediction of apoptosis proteins. Proteins: Struct. Funct. Bioinforma 50, 44–48 (2003).
Chou, K.-C. & Shen, H.-B. Recent progress in protein subcellular location prediction. Anal. Biochem. 370, 1–16 (2007).
Yu, C.-S. et al. Cello2go: A web server for protein subcellular localization prediction with functional gene ontology annotation. Plos One 9, 1–9 (2014).
Cozzetto, D., Minneci, F., Currant, H. & Jones, D. T. Ffpred 3: feature-based function prediction for all gene ontology domains. Sci. reports 6, 31865 (2016).
Russell, J. J. et al. Non-model model organisms. BMC Biol. 15, 55 (2017).
The UniProt Consortium. The universal protein resource (UniProt). Nucleic Acids Res. 36, D190–195 (2008).
Emanuelsson, O., Nielsen, H., Brunak, S. & von Heijne, G. Predicting subcellular localization of proteins based on their n-terminal amino acid sequence. J. Mol. Biol. 300, 1005–1016 (2000).
Hua, S. & Sun, Z. Support vector machine approach for protein subcellular localization prediction. Bioinforma. 17, 721 (2001).
Yu, N. Y. et al. Psortb 3.0: improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinforma. 26, 1608 (2010).
Chou, K.-C. & Shen, H.-B. Cell-PLoc: a package of Web servers for predicting subcellular localization of proteins in various organisms. Nat. Protoc. 3, 153–162 (2008).
Wang, X., Zhang, W., Zhang, Q. & Li, G.-Z. Multip-schlo: multi-label protein subchloroplast localization prediction with chou’s pseudo amino acid composition and a novel multi-label classifier. Bioinforma. 31, 2639 (2015).
Hasan, M. A. M., Ahmad, S. & Molla, M. K. I. Protein subcellular localization prediction using multiple kernel learning based support vector machine. Mol. BioSyst. 13, 785–795 (2017).
Cheng, X., Xiao, X. & Chou, K.-C. ploc-meuk: Predict subcellular localization of multi-label eukaryotic proteins by extracting the key go information into general pseaac. Genomics 110, 50–58 (2018).
Lobley, A., Swindells, M. B., Orengo, C. A. & Jones, D. T. Inferring function using patterns of native disorder in proteins. PLOS Comput. Biol. 3, 1–13 (2007).
Dolan, M. E., Ni, L., Camon, E. & Blake, J. A. A procedure for assessing go annotation consistency. Bioinforma. 21, i136 (2005).
Moriya, Y. et al. PathPred: an enzyme-catalyzed metabolic pathway prediction server. Nucleic Acids Res. 38, W138–W143 (2010).
Sharan, R., Ulitsky, I. & Shamir, R. Network-based prediction of protein function. Mol. Syst. Biol. 3 (2007).
Chou, K.-C. Graphic rule for drug metabolism systems. Curr. Drug Metab. 11, 369–378 (2010).
Kschischang, F. R., Frey, B. J. & Loeliger, H.-A. Factor graphs and the sum-product algorithm. IEEE Trans. Inf. Theor. 47, 498–519 (2001).
Spetale, F., Tapia, E., Krsticevic, F., Roda, F. & Bulacio, P. A factor graph approach to automated go annotation. Plos One 11, 1–16 (2016).
Krsticevic, F., Arce, D., Ezpeleta, J. & Tapia, E. Tandem Duplication Events in the Expansion of the Small Heat Shock Protein Gene Family in Solanum lycopersicum (cv. Heinz 1706). G3: Genes|Genomes|Genetics 6, 3027–3034 (2016).
Marques, A. C., Vinckenbosch, N., Brawand, D. & Kaessmann, H. Functional diversification of duplicate genes through subcellular adaptation of encoded proteins. Genome Biol. 9, R54 (2008).
Löw, D., Brändle, K., Nover, L. & Forreiter, C. Cytosolic heat-stress proteinshsp17.7 class i and hsp17.3 class ii of tomato act as molecular chaperones invivo. Planta 211, 575–582 (2000).
Neta-Sharir, I., Isaacson, T., Lurie, S. & Weiss, D. Dual role for tomato heat shock protein 21: Protecting photosystem ii from oxidative stress and promoting color changes during fruit maturation. The Plant Cell Online 17, 1829–1838 (2005).
Lui, J. & Shono, M. Characterization of mitochondria-located small heat shock protein from tomato (lycopersicon esculentum). Plant and Cell Physiol. 40, 1297–1304 (1999).
Duncan, O., Murcha, M. W. & Whelan, J. Unique components of the plant mitochondrial protein import apparatus. Biochimica et Biophysica Acta (BBA) – Mol. Cell Res. 1833, 304–313 (2013).
Chou, K.-C. & Shen, H. -B. Review: Recent advances in developing web-servers for predicting protein attributes. Nat. Sci. 63–92 (2009).
Liu, B., Yang, F., Huang, D.-S. & Chou, K.-C. ipromoter-2l: a two-layer predictor for identifying promoters and their types by multi-window-based pseknc. Bioinforma. 34, 33–40 (2018).
Chou, K.-C. An unprecedented revolution in medicinal chemistry driven by the progress of biological Science. Curr. Top. Medicinal Chem. 17, 2337–2358 (2017).
Asma, E., Khalid, M., Daanial, K. Y., Afzal, K. S. & Kuo-Chen, C. A Novel Modeling in Mathematical Biology for Classification of SignalPeptides. Sci. Reports 8, 1039 (2018).
Chou, K.-C. Some remarks on protein attribute prediction and pseudo amino acid composition. J. Theor. Biol. 273, 236–247 (2011).
Wei, Q. & Dunbrack, R. L. The role of balanced training and testing data sets for binary classifiers in bioinformatics. PloS one 8 (2013).
Eisner, R., Poulin, B., Szafron, D., Lu, P. & Greiner, R. Improving protein function prediction using the hierarchical structure of the gene ontology. In Proc. IEEE CIBCB, 1–10 (2005).
Carlson, M. Genome wide annotation for zebrafish http://bioconductor.org/packages/org.Dr.eg.db/. Version: 3.2.3, Accessed: 2017-07-06 (2016).
Carlson, M. Genome wide annotation for arabidopsis http://bioconductor.org/packages/org.At.tair.db. Version: 3.2.3, Accessed: 2017-07-06 (2017).
Carlson, M. Genome wide annotation for yeast http://bioconductor.org/packages/org.Sc.sgd.db. Version: 3.2.3, Accessed: 2017-07-06 (2017).
Carlson, M. Genome wide annotation for fly http://bioconductor.org/packages/org.Dm.eg.db. Version: 3.2.3, Accessed: 2017-07-06 (2017).
Carlson, M. Genome wide annotation for mouse http://bioconductor.org/packages/org.Mm.eg.db/. Version: 3.2.3, Accessed: 2017-07-06 (2016).
Nordahl, P. T., Soren, B., von Heijne, G. & Henrik, N. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Meth 8, 785–786 (2011).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden markov model: application to complete genomes11edited by f. cohen. J. Mol. Biol. 305, 567–580 (2001).
Horton, P. et al. WoLF PSORT: protein localization predictor. Nucleic Acids Res. 35, W585–W587 (2007).
Fukasawa, Y. et al. Mitofates: Improved prediction of mitochondrial targeting sequences and their cleavage sites. Mol. & Cell. Proteomics 14, 1113–1126 (2015).
Simarjeet, N., Pandey, S., Srinivasan, S., Mohammed, A. & Guda, C. LocSigDB: a database of protein localization signals. Database: The J. Biol. Databases and Curation 2015, bav003 (2015).
Lee, B., Shin, M., Oh, Y., Oh, H. & Ryu, K. Identification of protein functions using a machine-learning approach based on sequence-derived properties. Proteome Sci. 7, 27 (2009).
Lupas, A., Van Dyke, M. & Stock, J. Predicting coiled coils from protein sequences. Sci. 252, 1162–1164 (1991).
Chou, P. Y. & Fasman, G. D. Prediction of protein conformation. Biochem. 13, 222–245 (1974).
Chou, P. Y. & Fasman, G. D. Conformational parameters for amino acids in helical,β-sheet, and random coil regions calculated from proteins. Biochem. 13, 211–222 (1974).
Rhee, S. Y., Wood, V., Dolinski, K. & Draghici, S. Use and misuse of the gene ontology annotations. Nat. Rev. Genet. 9, 509–515 (2008).
Blondé, W. et al. Reasoning with bio-ontologies: using relational closure rules to enable practical querying. Bioinform. 27, 1562–1568 (2011).
Lawler, E. L. & Wood, D. E. Branch-And-Bound Methods: A Survey. Operations Research 14, 699–719 (1966).
Gene Ontology Consortium. Ontology relations http://geneontology.org/page/ontology-relations. Accessed: 2017-17-10 (2017).
Consortium, G. O. Annotation extension: Capturing cell and tissue types http://wiki.geneontology.org/index.php/Annotation_Extension:_Capturing_cell_and_tissue_types. Accessed: 2017-17-10 (2017).
Yedidia, J. S. Message-passing algorithms for inference and optimization. Journal of Statistical Physics 145, 860–890 (2011).
Pillai, I., Fumera, G. & Roli, F. Designing multi-label classifiers that maximize f measures: State of the art. Pattern Recognition 61, 394–404, http://www.sciencedirect.com/science/article/pii/S0031320316302217 (2017),
Meyer, D., Dimitriadou, E., Hornik, K., Weingessel, A. &Leisch, F. Misc functions of the department of statistics (e1071), tu wien http://cran.r-project.org/web/packages/e1071/index.html. Version: 1.6-8, Accessed: 2017-08-02 (2017).
Kiritchenko, S., Matwin, S. & Famili, A. F. Functional annotation of genes using hierarchical text categorization. In in Proc. of the BioLINK SIG: Linking Literature, Information and Knowledge for Biology held at ISMB-05 (2005).
Kosmopoulos, A., Partalas, I., Gaussier, E., Paliouras, G. & Androutsopoulos, I. Evaluation measures for hierarchical classification: A unified view and novel approaches. Data Min. Knowl. Discov. 29, 820–865 (2015).
Verspoor, K., Cohn, J., Mnizewski, S. & Joslyn, C. A categorization approach to automated ontological function annotation. Protein Sci. 15, 1544–1549 (2006).
Fawcett, T. An introduction to roc analysis. Pattern Recogn. Lett. 27, 861–874 (2006).
Valentini, G. & Re, M. Perfmeas: Performance measures for ranking and classification tasks http://cran.r-project.org/web/packages/PerfMeas/index.html. Version: 1.2-1, Accessed: 2017-08-02 (2015).
Acknowledgements
We thank the reviewers and editor for helpful suggestions and comments. The authors would like to thank the support of project INI 3600TC: “Anotación funcional GO basada en clasificación jerárquica supervisada”, Universidad Tecnológica Nacional; PICT-2016-4063: “Aprendizaje computacional para la anotación enriquecida de genes en ontologías vinculadas and PICT-2014-3181: “Optimización de procesos del mejoramiento genético de tomate mediante herramientas bioinformáticas”, National Agency for Science and Technology Promotion, Argentina.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: F.E.S., D.A. and E.T. Performed the experiments: F.E.S., P.B. and E.T. Analyzed the results: F.E.S., P.B. and E.T. Analyzed the biological data: D.A. and F.K. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Spetale, F., Arce, D., Krsticevic, F. et al. Consistent prediction of GO protein localization. Sci Rep 8, 7757 (2018). https://doi.org/10.1038/s41598-018-26041-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-26041-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.