Abstract
Trachoma, the leading infectious cause of blindness, is caused by Chlamydia trachomatis (Ct), a bacterium of the phylum Chlamydiae. Recent investigations revealed the existence of additional families within the phylum Chlamydiae, also termed Chlamydia-like organisms (CLOs). In this study, the frequency of Ct and CLOs was examined in the eyes of healthy Sudanese (control) participants and those with trachoma (case). We tested 96 children (54 cases and 42 controls) and 93 adults (51 cases and 42 controls) using broad-range Chlamydiae and Ct-specific (omcB) real-time PCR. Samples positive by broad-range Chlamydiae testing were subjected to DNA sequencing. Overall Chlamydiae prevalence was 36%. Sequences corresponded to unclassified and classified Chlamydiae. Ct infection rate was significantly higher in children (31.5%) compared to adults (0%) with trachoma (p < 0.0001). In general, 21.5% of adults and 4.2% of children tested positive for CLOs (p = 0.0003). Our findings are consistent with previous investigations describing the central role of Ct in trachoma among children. This is the first study examining human eyes for the presence of CLOs. We found an age-dependent distribution of CLO DNA in human eyes with significantly higher positivity in adults. Further studies are needed to understand the impact of CLOs in trachoma pathogenicity and/or protection.
Similar content being viewed by others
Introduction
Trachoma constitutes the leading infectious cause of blindness worldwide1. It is considered a public health problem in 41 countries and is responsible for the visual impairment of approximately 1.9 million people, of whom 0.450 million are blind2. Trachoma is caused by repeated infection of the conjunctiva with Chlamydia trachomatis, a bacterium of the phylum Chlamydiae1,3. The majority of infections heal without sequelae, whereas subgroups of infected patients develop chronic inflammation and scarring of the ocular surface. However, the factors responsible for the different outcomes are not yet fully understood.
The phylum Chlamydiae was originally considered to consist of only a single family, the Chlamydiaceae4. Since the mid-1990s, advances in molecular techniques have uncovered new Chlamydiae, termed Chlamydia-like organisms (CLOs) or environmental Chlamydiae, belonging to novel families in this phylum; i.e., the Parachlamydiaceae, Simkaniaceae, Rhabdochlamydiaceae, Waddliaceae, Candidatus Piscichlamydiaceae, Candidatus Parilichlamydiaceae, Candidatus Amphibiichlamydiaceae, Candidatus Clavichlamydiaceae, and Criblamydiaceae5,6,7,8,9. There is also molecular evidence for an even greater diversity within the phylum, suggesting the existence of more than 180 families therein10. Chlamydiae are obligate intracellular bacteria with a biphasic developmental cycle that includes an extra-cellular elementary body and an intracellular dividing reticulate body11,12,13. CLOs have been identified from various environmental sources and hosts such as humans, warm-blooded terrestrial vertebrates, fish, reptiles, amphibians, arthropods, and eukaryotic microorganisms such as amoebae6,7,8,9,14,15. Parachlamydia acanthamoebae and Neochlamydia hartmannellae were found in eyes of cats with keratitis, conjunctivitis, and other ocular diseases16,17. In a study on the eyes of sheep with conjunctivitis, DNA of uncultured CLOs was found in two-thirds (26/32) of the samples18. The presence of P. acanthamoebae and other CLOs have also been reported in symptomatic eyes of guinea pigs and in the one-day disposal contact lenses of their owner19. Additionally, results from several studies on humans and animals have shown the presence of CLOs in urogenital sites, the skin, respiratory tract and eye20,21,22. Furthermore, to date, Parachlamydiaceae species (spp.), Simkania negevensis and Waddlia chondrophila have been linked to human respiratory diseases and adverse pregnancy outcomes with various levels of evidence5,23,24,25,26,27.
Increasing evidence suggests an association between the presence of non-C. trachomatis bacteria and trachoma. Dean et al. reported a high prevalence of Chlamydia psittaci, Chlamydia suis, Chlamydia pecorum, and Chlamydia pneumoniae in eyes of patients with trachoma from Nepal28,29. Results from two separate studies in Tanzania, one, a cross-sectional study on children30 and the other a case-control study on adults31, revealed a strong association between non-chlamydial bacterial infections (such as Streptococcus pneumoniae and Haemophilus influenza) and clinical signs of trachoma in children and adults. These findings were independently supported by two studies on Gambian children and adults with signs of trachoma32,33. In addition, a study in Ethiopia suggested that conjunctival colonization with pathogenic bacterial species is more likely in patients with trachomatous trichiasis34. Trichiasis, the introversion of the eyelashes, has been described repeatedly as a direct route for conjunctival colonization with pathogens in the eyes of patients with trachoma34,35.
Altogether, such surveys provide insights into the possible role of various chlamydial species on ocular surface pathogenicity. Several similarities have been demonstrated between different CLOs and the Chlamydiaceae concerning their developmental cycle, virulence factors, and invasion mechanisms to the host cells5,10,20,36,37. Considering the importance of C. trachomatis in trachoma and the status of CLOs as highly prevalent microorganisms in the environment, it is necessary to examine the frequency of CLOs on the ocular surface and their possible association with trachoma. The main aim of this case-control study was therefore to investigate the frequency of C. trachomatis and other Chlamydiae in ocular samples of children and adults from Al Qadarif region in Sudan (Fig. 1) with trachoma and healthy participants.
Results
Study population
The distribution of participants by age, sex, and grade of trachoma is shown in Table 1. The study population comprised 189 individuals, of whom 96 (children) were aged 1–9 years and 93 (adults) aged 15–88 years. The median ages of the children and adults were 5 and 56.3 years, respectively.
Broad-range Chlamydiae real-time polymerase chain reaction and sequence classification
Among 189 swab samples in total, 68 (36%) were positive for Chlamydiae DNA. All the nucleic acid extraction negative controls and real-time polymerase chain reaction (PCR) negative controls remained negative when tested with Chlamydiae real-time PCR. Although no swab has been taken as a field control, the risk of contamination can be ruled out because of existing differences in the distribution of positive samples for Chlamydiae real-time PCR and various profiles of sequences corresponding to different Chlamydiae families among children and adults as well as cases and controls. There were no statistically significant differences in positivity for Chlamydiae or C. trachomatis between male and female participants (p > 0.05). The average threshold cycle (CT) value for Chlamydiae real-time PCR was 34 ± 9.4. A standard curve was prepared to evaluate the efficiency of real-time PCR and to achieve a comparative concentration (inclusion forming units (IFUs)) for the samples based on their CT values (Supplementary Fig. S1). Among the 68 positive samples, 55 (81%) yielded assembled forward and reverse sequences (~200 bp) that were used for BLAST-n analysis, Naïve Bayesian classification in RDP, and phylogenetic analysis (Table 2, Figs 2 and 3) (Supplementary Tables S1 and S2). We were not able to assemble the sequences of the 13 remaining samples owing to multiple peaks in chromatogram lanes, which either might suggest the presence of more than one member of Chlamydiae in the ocular swab, or low amounts of DNA in the sample, which can cause PCR-generated artefactual species variation38,39,40,41.
Phylogenetic affiliation of 16S rRNA gene sequences detected in this study. A Bayesian tree built from full-length chlamydial sequences is shown, to which the partial sequences generated in this study were added using Parsimony. Only sequences closely related to the sequences found in this study are depicted. The full dendrogram including all used sequences is available as Supplementary Fig. S2. Branches belonging to the Chlamydiaceae and CLOs are marked (light and dark purple, respectively). Sequences of cases (purple) and controls (dark purple) of children (cTF and cC, respectively), and cases (dark blue) and controls (cyan) of adults (aTT and aC, respectively), are marked.
The maximum likelihood phylogenetic tree derived from the partial 16 S rRNA sequences detected in this study. Household IDs of samples are shown on left. Children cases are marked in purple (cTF), children controls (cC) in red, adults cases in dark blue (aTT), adults controls in cyan (aC), and the reference strains corresponding to species for which there was at least one best BLAST hit in black. The best BLAST hit identity (%) for each sample stands on the right side of the lables.
BLAST-n results for sequences were categorized based on the first (Table 2) and second BLAST-hit identification (excluding uncultured bacteria) (Supplementary Table S2) and closest 16 S ribosomal RNA (16S rRNA) classification match (Table 2). Of 16 samples that were detected as Chlamydiales bacterium by first BLAST-hit identification, nine sequences had highest similarity to Chlamydiales bacterium CRIB 32 and CRIB 33, six to Chlamydiales bacterium NS11, and one to both Chlamydiales bacterium CRIB 32/NS11 (Table 2). Sequences for Chlamydiales bacterium CRIB 32/33 and Chlamydiales bacterium NS11 were previously identified in domestic water and human nasal samples, respectively42,43.
Based on this 16S rRNA classification, four sequences corresponded to new species levels, 21 to new genus levels, and one to a new family level of the phylum Chlamydiae. Altogether, 54 out of 55 assembled sequences were identified as the nearest neighbor to four families including Chlamydiaceae (55.6%), Parachlamydiaceae (29.6%), Simkaniaceae (13%), and Criblamydiaceae (1.9%) (Table 2). Additionally, the retrieved 16S rRNA gene sequences were classified using the Naïve Bayesian Classifier of RDP, by which the majority of the sequences were also placed within Chlamydiaceae (52.7%) and Parachlamydiaceae (27.3%). Owing to length, the remaining sequences could only be assigned as unclassified Chlamydiales (20%) (Table 2). In general, the prevalence of Chlamydiaceae sequences was significantly higher in children than in adults (p = 0.0069) and CLO sequences were more frequent in adults than in children (p = 0.0003) (Figs 4 and 5).
Abundance of 16 S rRNA gene sequences in the ocular samples taken from Al Qadarif-Sudan classified at the family-level in the Chlamydiae based on the closest BLAST hit. (a) Distribution of sequences obtained from children, assigned to members of chlamydial families. (b) Distribution of sequences obtained from adults, assigned to members of chlamydial families. The statistical significance is indicated as follows: **p < 0.01, and ****p < 0.0001.
Abundance of 16 S rRNA gene sequences in the ocular samples taken from Al Qadarif-Sudan at the family-level in the Chlamydiae based on phylogenetic analysis. (a) Sequences derived from samples of children. (b) Sequences derived from samples of adults. The statistical significance is indicated as follows: *p < 0.05, **p < 0.01, and ****p < 0.0001.
Phylogenetic analysis of sequences
Phylogenetic analyses were performed with two different datasets to obtain better insights into the taxonomic placement of the 16S rRNA sequences from this study. The sequences obtained in this study were added to a Bayesian inference tree of full-length chlamydial 16S rRNA gene sequences representing the major known chlamydial lineages using Parsimony (Fig. 2) (Supplementary Table S1 and Supplementary Fig. S2). Similar to BLAST-n results, the majority of sequences were assigned to the Chlamydiaceae (52.7%). The sequences affiliating with CLOs grouped together with sequences of members of the Parachlamydiaceae (9.1%) and Criblamydiaceae (1.8%). A total of 36.3% of the sequences were closely related to as yet unclassified chlamydial sequences and reflect the high diversity within the Chlamydiae. A maximum likelihood phylogenetic tree (Fig. 3) revealed the phylogenetic relationship of the obtained 16S rRNA gene sequences to their closest BLAST-hit including these sequences along with some reference sequences.
Prevalence of Chlamydiae DNA in children
Of 96 samples from children (54 in the trachomatous inflammation-follicular (cTF) group and 42 in the control (cC) group), 32 (33.3%) yielded positive results for Chlamydiae (25 in the cTF group and 7 in the cC group). The prevalence of Chlamydiae DNA in the cTF group (45.3%) was significantly higher than that in the cC group (16.7%) (p = 0.0032). The average CT values in the cTF and cC groups tested using Chlamydiae real-time PCR were 30.8 ± 9.2 and 35.7 ± 0.7, respectively (p = 0.0154) (Fig. 6a).
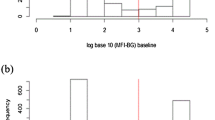
Average real-time PCR threshold cycle (CT) values for amplification of the Chlamydiae 16 S rRNA gene. (a) Recorded CT values for positive samples among children and adults. The children case group showed significantly lower CT values compared to the children control group and adults case and control groups. (b) Recorded CT values for 16 S rRNA sequences classified as families in the phylum Chlamydiae, C. trachomatis (CT), and non-trachomatis Chlamydiaceae (NTC). CT values were significantly lower for the Chlamydiaceae family compared to Parachlamydiaceae, Simkaniaceae, and Criblamydiaceae families. CT values for positive samples for C. trachomatis were recorded as significantly lower than those for NTC positive samples. (c) Recorded CT values for 16 S rRNA sequences classified as families in the phylum Chlamydiae, CT, and NTC among children. (d) Recorded CT values for 16 S rRNA sequences classified as families in the phylum Chlamydiae, CT, and NTC among adults. The statistical significance is indicated as follows: *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001.
Among the 32 positive samples, 27 sequences were assembled. Sequences from 26 samples corresponded to various families: 22 (84.6%) were positive for Chlamydiaceae, three (11.5%) for Parachlamydiaceae, and one for (3.8%) Criblamydiaceae (Table 2 and Fig. 4a). In addition, one sample (cTF092) exhibited 87% similarity to the order Chlamydiales and appeared to be phylogenetically close to unclassified Chlamydiales (Table 2 and Fig. 2).
In the cTF group, among the 19 sequences classified as Chlamydiaceae, 17 (89.5%) showed 99–100% homology to C. trachomatis (p < 0.0001). These results were confirmed using omcB real-time PCR. In general, C. trachomatis sequences had a frequency of 31.5% (17/54) among the cTF group. The remaining two positive samples showed sequence homology with Chlamydia caviae/Chlamydia felis and Chlamydia gallinacea and remained negative when tested with omcB real-time PCR. Among three assembled sequences in the cC group, two revealed highest similarity to both C. caviae/C. felis and one to only C. caviae (Table 2). None of the samples in the cC group exhibited similarities to C. trachomatis or had positive testing results using omcB real-time PCR. Moreover, among the three sequences classified as Parachlamydiaceae in the cTF group, two sequences showed highest similarity to Parachlamydia and one to Neochlamydia. In addition, one sample (cTF051) was classified as Criblamydiaceae (Table 2). This is also reflected in the phylogenetic trees (Figs 2 and 3).
Altogether, C. trachomatis comprised the majority of 16S rRNA sequences compared to non-trachomatis Chlamydiaceae (NTC) and all CLOs among children (C. trachomatis vs. NTC and CLOs, p = 0.0005 and p = 0.0001, respectively) (Fig. 7a). The frequency of Chlamydiaceae sequences was significantly higher than that of all classified or unclassified CLOs among the children samples (p < 0.0001) (Figs 4a and 5a). No significant differences were found between cTF and cC groups with respect to the affiliation to different CLO families (p > 0.05) (Fig. 7a).
Distribution of 16 S rRNA gene sequences assigned to families in the phylum Chlamydiae, C. trachomatis (CT), and non-trachomatis Chlamydiaceae (NTC) among (a) children and (b) adults. Prevalence of classified sequences as CT in the children case group was significantly higher than that in the children control group. The statistical significance is indicated as follows: *p < 0.05, ***p < 0.001, and ****p < 0.0001.
Prevalence of Chlamydiae DNA in adults
Overall, 36 out of 93 (38.7%) swabs taken from adults tested positive for Chlamydiae DNA. Prevalence of Chlamydiae in the adult control (aC) group (20/42) was higher than that in adults with trachomatous trichiasis (aTT) (16/51); however, this difference was not significant (p > 0.05). The average CT values in the aTT and aC groups tested using Chlamydiae real-time PCR were 36.0 ± 0.45 and 36 ± 0.8, respectively (p > 0.05) (Fig. 6a). The majority of sequences from adults were identified by BLAST search as being similar to the Parachlamydiaceae family (13 sequences, 46.4%), eight (28.6%) similar to the Chlamydiaceae and seven (25%) to the Simkaniaceae family (Table 2 and Fig. 4b). Phylogenetic analysis of the sequences from adults, including many lineages of yet uncultured chlamydial species, revealed three main groups including unclassified Chlamydiales (64.3%), Chlamydiaceae (25%), and Parachlamydiaceae (10.7%) (Fig. 5b).
The distribution of Chlamydiaceae sequences within the aTT group consisted of one sequence that showed highest similarity to genus Chlamydia, one to C. pecorum, and three that showed equivalent identification as either C. caviae or C. felis. With regard to the three controls positive for the Chlamydiaceae family, two sequences revealed highest similarity to C. trachomatis and also tested positive in omcB real-time PCR. The remaining sample exhibited equivalent similarity to C. caviae/C. felis (Table 2).
In general, the number of sequences classified as CLOs within the adult group was significantly higher than that classified as Chlamydiaceae (p = 0.0137) (Figs 4b and 5b). Nevertheless, no significant differences were found between aTT and aC groups with respect to the affiliation to different Chlamydiae families (p > 0.05) (Fig. 7b).
Discussion
This study provided evidence for the presence of Chlamydiaceae and CLO DNA in human ocular samples of patients with trachoma and healthy controls in Sudan. Overall, 36% of all samples tested positive for Chlamydiae DNA. Whereas the prevalence of C. trachomatis was significantly higher in children than in adults, a significantly higher distribution of CLO DNA was detected in adult samples compared to those from children. However, based on differences in the occurrence of identified CLO sequences among the patients with trachoma and healthy controls in children and adults, a pathogenic role for these bacteria in trachoma is not evident.
To our knowledge, this study is the first to examine human ocular samples for the presence of all members of the Chlamydiae. Whereas 22.9% of children’ and 8.6% of adults’ samples corresponded to Chlamydiaceae sequences, amplicons of 21.5% of adults’ samples and 4.2% of children samples corresponded to sequences originating from different CLOs. These results are comparable with a prior study on cats with and without (controls) ocular diseases16. Specifically, C. felis, the known pathogen for feline chlamydial conjunctivitis44, was found among the majority of diseased eyes in animals under the age of five. The distribution of positive samples for non-C. felis Chlamydiales increased from 30% for cats under the age of one to 54% for cats older than 10 years in animals with and without ocular diseases16. Furthermore, in two separate studies, in nasopharyngeal samples from healthy children and adults with and without pneumonia, 11.4% of children samples and 36% of samples from adults tested positive for members of various families in the phylum Chlamydiae43,45. It is not clear why the distribution of CLOs in adults was more than five times higher than that in children. Although this difference might constitute a bias resulting from the small sample size in each group, this does not appear likely.
In the present study, we found a higher diversity of CLOs in adults than in children. Some of the analyzed sequences belong to members of the Parachlamydiaceae, which have previously been detected in other mammalian tissues46. Parachlamydia spp. have been found directly and indirectly in human corneal samples and disposable contact lenses, respectively19,47. To our knowledge, this is the first report of Criblamydiaceae in human eyes. The majority of sequences obtained, however, were affiliated with various as yet uncharacterized chlamydial lineages. This is unexpected, because these lineages are generally to date represented only by environmental clone sequences, with members of such lineages not having been considered to be also associated with humans. Although the sequences belonging to these divergent CLOs have mainly been detected in healthy participants and may also originate from other sources, it is still noteworthy to find such a diversity of CLOs sequences in human eyes. Considering that similar profiles of CLOs were obtained among the aTT and aC groups, we cannot suggest any association of these bacteria with trachoma in adults. In children, the distribution of CLOs did not significantly differ between the cTF and cC groups.
Several studies have reported the pathogenic impacts of Parachlamydia spp. in respiratory disorders6,20,21,26 and S. negevenesis in patients with bronchiolitis, although S. negevenesis pathogenic potential remains controversial6,20,21,48,49,50. Notably, the possible role of some CLOs in respiratory tract infections was mainly suspected in pediatric populations; similarly, in the present study, the only limited evidence suggesting a role of Parachlamydiaceae and Criblamydiaceae in trachoma pathogenesis was documented in children.
In our study, no strong evidence of the pathogenic impact of CLOs on adult human eyes was obtained, which supports previous results by Vidgen et al. suggesting CLOs as commensals of the ocular microbiota in koala51. Estimated loads of CLOs in this study are comparable with the reported load of the bona fide microbiome: 100 to 1000 colony-forming unit (CFU) per mL in human tear fluid52. These results are in line with several studies reporting a non-significant distribution of Parachlamydiaceae and other CLOs among diseased and healthy eyes of sheep, pigs, and cats16,17,18,53. Moreover, CLOs have been detected in different samples such as skin, nose, and cervicovaginal samples of healthy individuals22,38,43,45.
The high prevalence of Chlamydiae DNA in the present study is comparable with findings of human skin and nasal samples, which support the idea that ocular microbiota colonizes in parallel with microbiota of the upper respiratory system and skin22,43. This high prevalence compared to results from human respiratory or cervicovaginal samples can be explained through the frequent contact of the ocular surface directly or indirectly with a source of bacteria in the environment, infected animals, and humans. In particular, the majority of unclassified chlamydial sequences in this study demonstrated highest homology to previously isolated sequences from water. Additionally, members of Parachlamydiaceae are known to naturally infect amoeba, which they utilize as a replicative niche23,42,54,55,56. Moreover, Simkania and Criblamydia have been detected repeatedly in domestic waters14,42,54,55,57,58, which highlights the importance of water as a rich source of CLOs5 and possibly a main source of ocular infection with Chlamydiae. These results also emphasize the validity of environmental improvement as an essential factor of the SAFE strategy (Surgery for trichiasis, Antibiotics to clear ocular infection, promotion of Facial cleanliness, and Environmental improvement).
The profile of the Chlamydiaceae community present in ocular mucosa differed significantly between adults and children. Whereas C. trachomatis was only detected in two (2.1%) adult samples, 17 samples (31.5%) of children with TF were found to be positive for C. trachomatis. These results are consistent with previous data suggesting a central role for C. trachomatis in active trachoma among children1,59,60. Notably, the only two samples from adults that yielded positive results for C. trachomatis infection in the present study were in the aC group. The absence of infection with C. trachomatis in adults with trachoma can be explained through: [1] the effects of non-C. trachomatis bacteria and mechanical irritation on the eyes30,31,32,34,61,62; [2] the development of acquired immunity to C. trachomatis and its impact on the duration of ocular infection, which decreases significantly with age28,34,59,61,63; and [3] the kinetics of the disease, which can be explained by the recovery phase after repeated infections with C. trachomatis60. During this time, patients can be diagnosed positive for trachoma although Chlamydia is no longer present in the eyes60.
Classification of the Chlamydiaceae positive sequences into C. trachomatis and NTC provided an opportunity to investigate the prevalence of NTC bacteria. Overall, we found no association between NTC species and trachoma in this population. Although NTC species in the aTT group (9.8%) compared to the aC group (2.4%) were more frequent, this difference was not statistically significant. This is in contrast to previous studies, which found a high prevalence of NTC species in trachoma samples from Nepal28,29. Differences in NTC prevalence between present data and previous studies by Dean et al.28,29 could be due to the small sample size in all of these studies and regional variations.
Unfortunately, BLAST results for the majority (8/11, 72.7%) of sequences positive for NTC species could not differentiate between C. caviae and C. felis as the same identification score was obtained for both species. Moreover, the non-significant distribution of sequences positive for C. caviae/C. felis between cases compared to controls was unable to provide a clear indication of their role in this study. C. caviae and C. felis cause keratoconjunctivitis in guinea pigs and cats, respectively16,19,28,64,65. Previous studies have also reported the presence of C. caviae and C. felis in human eyes19,66,67. In our study, one sequence obtained in the aTT group and one in the cTF group demonstrated highest homology to C. pecorum and C. gallinacea, respectively. The presence of C. pecorum in an ocular sample from a patient with trachoma has also been previously reported29. To the best of our knowledge, this is the first molecular evidence of the presence of C. gallinacea DNA in humans, a species recently added to the Chlamydiaceae that was reported for the first time in cattle by Li et al.68 and subsequently described as a causative agent of avian chlamydiosis with zoonotic potential69,70,71.
CT values revealed important information about the bacterial load in each sample. The average CT value for children infected with C. trachomatis corresponds to 1.000 to 10.000 IFU per swab (Fig. 6c), whereas those for NTC and CLO positive samples correspond to 10 to 100 IFU per swab (Fig. 6b–d). These data support prior findings that active infection (higher loads) of C. trachomatis is associated with active trachoma61,72,73,74. Furthermore, consistent with previous data, we found higher loads of bacteria in samples from children59,72,73, suggesting this age group as the major source of C. trachomatis infection59. Higher CT values, which are corresponding to lower loads of bacteria in positive samples for CLOs might be due to the persistent form of infection in these samples as it has been described earlier74,75. Nevertheless, high CT values may also be the consequence of an early infection, the presence of residual DNA from a previous infection, or inoculation of the eye with dead microorganisms28,61.
Our study also provides evidence that the presence of CLOs and C. trachomatis in human eyes was influenced by age. Previously, 16S rRNA gene sequencing revealed a highly diverse bacterial community in human eyes76,77,78. Moreover, age-dependent changes in the ocular microbiota have been reported in prior studies79,80. It has been shown that the ocular microbiome of individuals with healthy conjunctiva has a significantly different bacterial abundance and diversity in children aged ≤10 compared to older participants80. In addition, recent studies have uncovered strong evidence regarding the impacts of ocular commensal microbiota on regulating local immune responses and developing defense mechanisms against colonization of pathogens in the eyes52,79,81,82. The absence of CLOs and C. trachomatis in children and adults can thus be partially explained owing to age-dependent changes in the microbiota of eyes and its impacts on the development of different local immune responses. This may in turn influence the presence of Chlamydiae and multiplicity of infection. For example, a study on the cervical microbiota of women infected with C. trachomatis and healthy controls have shown a different diversity of microbiota in those with asymptomatic infection of C. trachomatis83. Additionally, the abundance of bacterial taxa was different in the cervical microbiota of women infected with C. trachomatis compared to that in healthy controls83. Vidgen et al. have also demonstrated a correlation between urogenital microbiota of koalas and infection with C. pecorum51. Furthermore, previous results have shown that co-infection of C. trachomatis serovar E and another intracellular pathogen, Toxoplasma gondii, can lead to a stress-induced persistent growth of C. trachomatis in the host cells owing to better ability of T. gondii in scavenging nutrients inside the cells84. The suggestion of CLOs as comprising part of the commensal microbiota of the eye thus raises the question of whether the presence of CLOs can influence successful infection of C. trachomatis in human eyes.
This study has several limitations. As the sample size was relatively small, the use of a larger sample size within this study might have minimized the bias in biodiversity of detected CLOs in each group and provided a better understanding of the possible associations of CLOs and NTC species with eye health or disease. In addition, the length of the amplicons used for sequencing was not sufficient to resolve the species (i.e. C. felis/C. caviae) or genus level in every case, due to the sequence identity or low representation of many chlamydial lineages in public databases, respectively.
In summary, C. trachomatis is associated with active trachoma in Sudanese children. We presented evidence demonstrating an age-dependent distribution of divergent groups of CLOs in human eyes. However, the biology of the CLOs identified in this study remains unknown. Our findings do not support a significant association between the presence of CLOs and NTC species and active trachoma in children or TT in adults. Comparative studies with a larger sample size on the prevalence of CLOs in human eyes from different geographic regions may provide insight into the possible role of these microorganisms in pathogenicity and/or protection. Further studies are needed to understand the impacts of ocular microbiota and local immune responses on the presence of C. trachomatis and CLOs in the eyes.
Material and Methods
Ethical approval
The study was conducted in accordance with the Declaration of Helsinki. The National Ethics Authorities in Sudan (Sudanese National Research Ethics Review Committee (NRERC) (No. 174-8-12)) and the ethical committee of the Medical University of Vienna approved the study and all the included procedures. Written informed consent was obtained from all adult individuals at the time of sample collection. For those participants aged <18 years that wished to take part in the study, consent was obtained from a parent/guardian. All samples were anonymized.
Trachoma grading
Clinical phenotypes were assessed in the field by an experienced ophthalmologist trained in trachoma grading using the World Health Organization simplified grading system85. Individuals presenting with follicular trachoma were classified as TF, individuals with pronounced papillary hypertrophy and inflammatory thickening of tarsal conjunctiva were classified as “Trachomatous Inflammation − Intense (TI)” and individuals with tarsal conjunctival scarring as “Trachomatous scarring (TS)”. Individuals with inturned eyelashes were graded as TT. Individuals with no clinical signs of follicles, papillary hypertrophy, or conjunctival scarring were classed as healthy controls. Cases and controls were matched by age and gender.
Study population and sampling
Participants for this study were recruited in trachoma endemic areas in Sudan86. For adults with TT, sample collection took place in a Field Surgery Clinic in the Al Qadarif region (Fig. 1). A total of 51 aTT scheduled for trichiasis surgery and 42 matched aC without history and present signs of trachoma were enrolled. In addition, 54 cTF and 42 cC subjects aged 1 to 9 years were recruited at two Quran Schools in the Al Qadarif region. Conjunctival samples were taken from the upper tarsal conjunctiva from both eyes using polyester flocked swabs (UTM-RT collection kits, Copan USA, Murrieta, CA, USA) using standard methodology87,88. The sampling ophthalmologist wore gloves that were changed after each individual. Swabs were stored in universal transport medium and frozen immediately in liquid nitrogen cryogenic shipping containers. In Austria, samples were subsequently stored at −80 °C.
DNA preparation
Genomic DNA was extracted from the conjunctival swab samples taken from left eyes. As a nucleic acid extraction negative control, one empty swab was included during each DNA extraction run. Each swab was vigorously vortexed for 5 min inside the universal transport medium, then the entire transport medium was transferred to a new tube. Cells in the swab transport medium were pelleted by centrifugation at 17,000 × g for 30 min. DNA was extracted from the resuspended pellet in ATL buffer of the QIAamp DNA Investigator Kit (QIAGEN GmbH, Hilden, Germany), according to manufacturer instruction, and DNA was stored at −20 °C until further use.
Broad-range Chlamydiae real-time PCR and sequencing
A Chlamydiae-specific real-time PCR was performed on a final reaction volume of 20 µL targeting a fragment of approximately 207 to 215 bp of the 16S rRNA gene as described by Lienard et al.45. Cycling conditions were 5 min at 95 °C, followed by 50 repetitions of 3-step cycles of 15 s at 95 °C, 20 s at 67 °C, and 15 s at 72 °C, all carried out in a PikoReal real-time PCR System (Thermo Fisher Scientific). All samples were tested in duplicate. After optimization of the PCR, a CT value of ≤37 has been suggested as a reliable cutoff to provide reproducible results and samples with this CT value were considered as positive samples. In each run, 2 wells were dedicated to negative controls (distilled water) and 2 wells were used as nucleic acid extraction negative controls. A standard curve was generated for each real-time PCR run using decimal serial dilutions of extracted genomic DNA from 1 × 106 to 1 × 101 IFUs of C. trachomatis (Supplementary Fig. S1).
Purification of amplicons for positive samples was performed using the QIAquick PCR Purification Kit (QIAGEN GmbH) according to the manufacturer’s protocol. Sanger sequencing of purified PCR products was performed by Eurofins Genomics AT (Vienna, Austria). Inner primers (panFseq and panRseq) resulted in an approximately 200 bp amplicon as explained previously by Lienard et al.45.
Specific TaqMan real-time PCR for the omcB gene of C. trachomatis
To assess C. trachomatis prevalence in particular, a specific real-time PCR was carried out on all samples to target the omcB gene of C. trachomatis. The reaction was performed in a final volume of 20 µL using iTaq Supermix (BioRad, Reinach, Switzerland), 0.3 µM concentration of each primer, 0.1 µM concentration of probe, and 5 µL sample DNA89. The program was set at 95 °C for 5 min, followed by 45 cycles of 15 s at 95 °C and 30 s at 60 °C, all carried out in a PikoReal real-time PCR System. All samples were tested in duplicate. Samples with a CT value of ≤37 were considered as positive. Distilled water (a negative control) and C. trachomatis DNA (a positive control) were included in each experiment.
Data analysis
The obtained partial forward and reverse sequences of 16S rRNA gene were trimmed, aligned and assembled in the Geneious R10.2.2 software package. Of the resulting 55 sequences, 50 (90.9%) were of high quality with average phred quality scores above 30 and 5 (9.1%) of the sequences had average phred quality scores above 23 (cTF-025, aC-259, aC-261, aC-268 and aC-285). All consensus sequences were checked for the presence of chimera with DECIPHER90 and compared with sequences available in the GenBank database of the National Center for Biotechnology Information via the BLAST server (https://www.ncbi.nlm.nih.gov/blast/). 16S rRNA gene identification cut-offs of 97, 95, and 90% were applied for the achieved sequences to approximately classify members of the phylum Chlamydiae to the species, genus, and family levels, respectively91. In addition, the 16S rRNA sequences were classified with the Naïve Bayesian classifier of RDP using a 70% confidence interval92. Obtained nucleotide sequences have been submitted to the NCBI database (https://www.ncbi.nlm.nih.gov/genbank/) under accession numbers (MH119764-99).
Based on the results from sequencing of 16S rRNA gene sequences and C. trachomatis omcB real-time PCR, samples assigned to the Chlamydiaceae were divided into two groups: C. trachomatis and NTC.
Phylogenetic analysis
Phylogenetic analyses were carried out with two datasets. First, a dataset contained full-length 16S rRNA gene sequences of Chlamydiae, which were downloaded from GenBank and aligned to the SILVA SSU Ref database containing preconfigured high-quality full-length sequences in ARB93,94. Bayesian inference analysis of full-length 16S rRNA gene sequences was carried out with MrBayes 3.2.6 using standard settings via the CIPRES Science Gateway95,96. The partial 16S rRNA gene sequences obtained in this study were added subsequently to the Bayesian tree using the Quick-Add Parsimony option in ARB93. The phylogenetic tree was visualized with iTOL and FigTree97,98 (Fig. 2).
Second, all obtained consensus sequences as well as the 16S rRNA gene sequences of C. trachomatis A/Har-13T (A_E17344.1), C. caviae strain: Gp/Ic (=ATCC VR813) (D85708.1), N. hartmannellae strain A1Hsp (NR_025037.1), Criblamydia sequanensis strain CRIB-18 (NR_115696.1), S. negevensis strain Z (NR_029194.1), and P. acanthamoebae strain CRIB43 (FJ532291.1) were used. Sequences were aligned using MAFFT v7.30999. Jalview v2.10.1 was used to trim the alignment and to look for reverse complemented sequences100. Maximum likelihood-based phylogenetic analyses were performed using FastTree 2.1 software (with parameters “-nt -gtr -spr 4 -mlacc 2 -slownni”)101. The phylogenetic representation was rerooted at the midpoint using Archaeopterix v0.9920102 and displayed with FigTree v1.4.298 (Fig. 3).
Statistics
Variables were compared by t test. A p-value < 0.05 was taken as statistically significant. Data analysis was performed using GraphPad Prism 6.0 (GraphPad Inc., La Jolla, CA, USA) software.
References
Taylor, H. R., Burton, M. J., Haddad, D., West, S. & Wright, H. Trachoma. Lancet 384, 2142–2152, https://doi.org/10.1016/S0140-6736(13)62182-0 (2014).
WHO. Trachoma: Fact sheet, http://www.who.int/mediacentre/factsheets/fs382/en/ (2016).
Pillonel, T., Bertelli, C., Salamin, N. & Greub, G. Taxogenomics of the order Chlamydiales. Int J Syst Evol Micr 65, 1381–1393, https://doi.org/10.1099/ijs.0.000090 (2015).
Kuo, C.-C., Stephens, R. S., Bavoil, P. M. & Kaltenboeck, B. Chlamydia. Bergey’s Manual of Systematics of Archaea and Bacteria., 1–28. (2015.).
Taylor-Brown, A., Vaughan, L., Greub, G., Timms, P. & Polkinghorne, A. Twenty years of research into Chlamydia-like organisms: a revolution in our understanding of the biology and pathogenicity of members of the phylum Chlamydiae. Pathog Dis 73, 1–15, https://doi.org/10.1093/femspd/ftu009 (2015).
Taylor-Brown, A. & Polkinghorne, A. New and emerging chlamydial infections of creatures great and small. New Microbes New Infect 18, 28–33, https://doi.org/10.1016/j.nmni.2017.04.004 (2017).
Pawlikowska-Warych, M. & Deptula, W. Characteristics of chlamydia-like organisms pathogenic to fish. Journal of Applied Genetics 57, 135–141, https://doi.org/10.1007/s13353-015-0303-8 (2016).
Horn, M. Chlamydiae as symbionts in eukaryotes. Annu Rev Microbiol 62, 113–131, https://doi.org/10.1146/annurev.micro.62.081307.162818 (2008).
Thomas, V., Casson, N. & Greub, G. Criblamydia sequanensis, a new intracellular Chlamydiales isolated from Seine river water using amoebal co-culture. Environmental Microbiology 8, 2125–2135, https://doi.org/10.1111/j.1462/2920.2006.01094.x (2006).
Lagkouvardos, I. et al. Integrating metagenomic and amplicon databases to resolve the phylogenetic and ecological diversity of the Chlamydiae. Isme J 8, 115–125, https://doi.org/10.1038/ismej.2013.142 (2014).
Abdelrahman, Y. M. & Belland, R. J. The chlamydial developmental cycle. FEMS Microbiol Rev 29, 949–959, https://doi.org/10.1016/j.femsre.2005.03.002 (2005).
Hybiske, K. & Stephens, R. S. Mechanisms of host cell exit by the intracellular bacterium Chlamydia. Proc Natl Acad Sci USA 104, 11430–11435, https://doi.org/10.1073/pnas.0703218104 (2007).
Greub, G. & Raoult, D. Crescent bodies of Parachlamydia acanthamoeba and its life cycle within Acanthamoeba polyphaga: an electron micrograph study. Appl Environ Microbiol 68, 3076–3084 (2002).
Kahane, S., Greenberg, D., Newman, N., Dvoskin, B. & Friedman, M. G. Domestic water supplies as a possible source of infection with Simkania. J Infection 54, 75–81, https://doi.org/10.1016/j.jinf.2006.01.011 (2007).
Lienard, J., Croxatto, A., Prod’hom, G. & Greub, G. Estrella lausannensis, a new star in the Chlamydiales order. Microbes Infect 13, 1232–1241, https://doi.org/10.1016/j.micinf.2011.07.003 (2011).
von Bomhard, W. et al. Detection of novel chlamydiae in cats with ocular disease. Am J Vet Res 64, 1421–1428 (2003).
Richter, M. et al. Parachlamydia acanthamoebae in domestic cats with and without corneal disease. Vet Ophthalmol 13, 235–237, https://doi.org/10.1111/j.1463-5224.2010.00789.x (2010).
Polkinghorne, A. et al. Molecular evidence for chlamydial infections in the eyes of sheep. Vet Microbiol 135, 142–146, https://doi.org/10.1016/j.vetmic.2008.09.034 (2009).
Lutz-Wohlgroth, L. et al. Chlamydiales in guinea-pigs and their zoonotic potential. J Vet Med A Physiol Pathol Clin Med 53, 185–193, https://doi.org/10.1111/j.1439-0442.2006.00819.x (2006).
Corsaro, D. & Greub, G. Pathogenic potential of novel Chlamydiae and diagnostic approaches to infections due to these obligate intracellular bacteria. Clin Microbiol Rev 19, 283–297, https://doi.org/10.1128/CMR.19.2.283-297.2006 (2006).
Haider, S., Collingro, A., Walochnik, J., Wagner, M. & Horn, M. Chlamydia-like bacteria in respiratory samples of community-acquired pneumonia patients. Fems Microbiol Lett 281, 198–202, https://doi.org/10.1111/j.1574-6968.2008.01099.x (2008).
Hokynar, K. et al. Chlamydia-Like Organisms (CLOs) in Finnish Ixodes ricinus Ticks and Human Skin. Microorganisms 4, https://doi.org/10.3390/microorganisms4030028 (2016).
Baud, D., Regan, L. & Greub, G. Emerging role of Chlamydia and Chlamydia-like organisms in adverse pregnancy outcomes. Curr Opin Infect Dis 21, 70–76, https://doi.org/10.1097/QCO.0b013e3282f3e6a5 (2008).
Ammerdorffer, A., Stojanov, M., Greub, G. & Baud, D. Chlamydia trachomatis and chlamydia-like bacteria: new enemies of human pregnancies. Curr Opin Infect Dis 30, 289–296, https://doi.org/10.1097/QCO.0000000000000369 (2017).
Casson, N., Michel, R., Muller, K. D., Aubert, J. D. & Greub, G. Protochlamydia naegleriophila as etiologic agent of pneumonia. Emerging Infectious Diseases 14, 168–172, https://doi.org/10.3201/eid1401.070980 (2008).
Lamoth, F., Jaton, K., Vaudaux, B. & Greub, G. Parachlamydia and Rhabdochlamydia: emerging agents of community-acquired respiratory infections in children. Clin Infect Dis 53, 500–501, https://doi.org/10.1093/cid/cir420 (2011).
Niemi, S., Greub, G. & Puolakkainen, M. Chlamydia-related bacteria in respiratory samples in Finland. Microbes Infect 13, 824–827, https://doi.org/10.1016/j.micinf.2011.04.012 (2011).
Dean, D., Kandel, R. P., Adhikari, H. K. & Hessel, T. Multiple Chlamydiaceae species in trachoma: implications for disease pathogenesis and control. PLoS Med 5, e14, https://doi.org/10.1371/journal.pmed.0050014 (2008).
Dean, D., Rothschild, J., Ruettger, A., Kandel, R. P. & Sachse, K. Zoonotic Chlamydiaceae species associated with trachoma, Nepal. Emerg Infect Dis 19, 1948–1955, https://doi.org/10.3201/eid1912.130656 (2013).
Burton, M. J. et al. What is causing active trachoma? The role of nonchlamydial bacterial pathogens in a low prevalence setting. Invest Ophthalmol Vis Sci 52, 6012–6017, https://doi.org/10.1167/iovs.11-7326 (2011).
Hu, V. H. et al. Bacterial infection in scarring trachoma. Invest Ophthalmol Vis Sci 52, 2181–2186, https://doi.org/10.1167/iovs.10-5829 (2011).
Burr, S. E. et al. Association between ocular bacterial carriage and follicular trachoma following mass azithromycin distribution in The Gambia. PLoS Negl Trop Dis 7, e2347, https://doi.org/10.1371/journal.pntd.0002347 (2013).
Burton, M. J. et al. Bacterial infection and trachoma in the gambia: A case-control study. Invest Ophth Vis Sci 48, 4440–4444, https://doi.org/10.1167/iovs.07-0315 (2007).
Cevallos, V. et al. Association of conjunctival bacterial infection and female sex in cicatricial trachoma. Invest Ophthalmol Vis Sci 53, 5208–5212, https://doi.org/10.1167/iovs.12-9984 (2012).
Cox, J. T. et al. Trachomatous Scarring and Infection With Non-Chlamydia Trachomatis Bacteria in Women in Kongwa, Tanzania. Invest Ophth Vis Sci 58, 3249–3253, https://doi.org/10.1167/iovs.17-21519 (2017).
Collingro, A. et al. Unity in variety–the pan-genome of the Chlamydiae. Mol Biol Evol 28, 3253–3270, https://doi.org/10.1093/molbev/msr161 (2011).
Konig, L. et al. Biphasic Metabolism and Host Interaction of a Chlamydial Symbiont. mSystems 2, https://doi.org/10.1128/mSystems.00202-16 (2017).
Reid, F., Oakeshott, P., Kerry, S. R., Hay, P. E. & Jensen, J. S. Chlamydia related bacteria (Chlamydiales) in early pregnancy: community-based cohort study. Clin Microbiol Infect 23, 119 e119–119 e114, https://doi.org/10.1016/j.cmi.2016.10.011 (2017).
Burnard, D. et al. Novel Chlamydiales genotypes identified in ticks from Australian wildlife. Parasite Vector 10, https://doi.org/10.1186/s13071-017-1994-y (2017).
Pilloux, L. et al. The High Prevalence and Diversity of Chlamydiales DNA within Ixodes ricinus Ticks Suggest a Role for Ticks as Reservoirs and Vectors of Chlamydia-Related Bacteria. Appl Environ Microb 81, 8177–8182, https://doi.org/10.1128/Aem.02183-15 (2015).
Burnard, D. et al. Molecular evidence of Chlamydia pecorum and arthropod-associated Chlamydiae in an expanded range of marsupials. Sci Rep 7, 12844, https://doi.org/10.1038/s41598-017-13164-y (2017).
Corsaro, D. et al. Novel Chlamydiales strains isolated from a water treatment plant. Environ Microbiol 11, 188–200, https://doi.org/10.1111/j.1462-2920.2008.01752.x (2009).
Corsaro, D. & Venditti, D. Detection of novel Chlamydiae and Legionellales from human nasal samples of healthy volunteers. Folia Microbiol (Praha) 60, 325–334, https://doi.org/10.1007/s12223-015-0378-y (2015).
Ohya, K. et al. Chlamydophila felis CF0218 is a novel TMH family protein with potential as a diagnostic antigen for diagnosis of C. felis infection. Clin Vaccine Immunol 15, 1606–1615, https://doi.org/10.1128/CVI.00134-08 (2008).
Lienard, J. et al. Development of a new chlamydiales-specific real-time PCR and its application to respiratory clinical samples. J Clin Microbiol 49, 2637–2642, https://doi.org/10.1128/JCM.00114-11 (2011).
Wheelhouse, N. et al. Involvement of Parachlamydia in bovine abortions in Scotland. Vet J 193, 586–588, https://doi.org/10.1016/j.tvjl.2012.01.008 (2012).
Fritsche, T. R., Gautom, R. K., Seyedirashti, S., Bergeron, D. L. & Lindquist, T. D. Occurrence of bacterial endosymbionts in Acanthamoeba spp. isolated from corneal and environmental specimens and contact lenses. J Clin Microbiol 31, 1122–1126 (1993).
Vouga, M., Baud, D. & Greub, G. Simkania negevensis, an insight into the biology and clinical importance of a novel member of the Chlamydiales order. Crit Rev Microbiol 43, 62–80, https://doi.org/10.3109/1040841X.2016.1165650 (2017).
Friedman, M. G., Kahane, S., Dvoskin, B. & Hartley, J. W. Detection of Simkania negevensis by culture, PCR, and serology in respiratory tract infection in Cornwall, UK. J Clin Pathol 59, 331–333, https://doi.org/10.1136/jcp.2004.025601 (2006).
Kahane, S., Greenberg, D., Friedman, M. G., Haikin, H. & Dagan, R. High prevalence of “Simkania Z,” a novel Chlamydia-like bacterium, in infants with acute bronchiolitis. J Infect Dis 177, 1425–1429 (1998).
Vidgen, M. E., Hanger, J. & Timms, P. Microbiota composition of the koala (Phascolarctos cinereus) ocular and urogenital sites, and their association with Chlamydia infection and disease. Sci Rep 7, 5239, https://doi.org/10.1038/s41598-017-05454-2 (2017).
Reyes, N. J. & Saban, D. R. A Commencement for Eye Commensals. Immunity 47, 6–8, https://doi.org/10.1016/j.immuni.2017.07.006 (2017).
Becker, A. et al. Intensively kept pigs pre-disposed to chlamydial associated conjunctivitis. J Vet Med A Physiol Pathol Clin Med 54, 307–313, https://doi.org/10.1111/j.1439-0442.2007.00963.x (2007).
Lienard, J. et al. Prevalence and diversity of Chlamydiales and other amoeba-resisting bacteria in domestic drinking water systems. New Microbes New Infect 15, 107–116, https://doi.org/10.1016/j.nmni.2016.10.003 (2017).
Horn, M. & Wagner, M. Evidence for additional genus-level diversity of Chlamydiales in the environment. Fems Microbiol Lett 204, 71–74, https://doi.org/10.1111/j.1574-6968.2001.tb10865.x (2001).
Birtles, R. J., Rowbotham, T. J., Storey, C., Marrie, T. J. & Raoult, D. Chlamydia-like obligate parasite of free-living amoebae. Lancet 349, 925–926 (1997).
Kahane, S., Platzner, N., Dvoskin, B., Itzhaki, A. & Friedman, M. G. Evidence for the presence of Simkania negevensis in drinking water and in reclaimed wastewater in Israel. Appl Environ Microb 70, 3346–3351, https://doi.org/10.1128/Aem.70.6.3346-3351.2004 (2004).
Perez, L. M. et al. Searching Simkania negevensis in environmental waters. Folia Microbiol 57, 11–14, https://doi.org/10.1007/s12223-011-0086-1 (2012).
Solomon, A. W. et al. Strategies for control of trachoma: observational study with quantitative PCR. Lancet 362, 198-204, doi:S0140-6736(03)13909-8 [pii] 10.1016/S0140-6736(03)13909-8 (2003).
Robinson, E. et al. Trachoma rapid assessments in Unity and Northern Bahr-el-Ghazal States, Southern Sudan. PLoS One 5, https://doi.org/10.1371/journal.pone.0013138 (2010).
Grassly, N. C., Ward, M. E., Ferris, S., Mabey, D. C. & Bailey, R. L. The natural history of trachoma infection and disease in a Gambian cohort with frequent follow-up. PLoS Negl Trop Dis 2, e341, https://doi.org/10.1371/journal.pntd.0000341 (2008).
Macleod, C. K. et al. Low Prevalence of Ocular Chlamydia trachomatis Infection and Active Trachoma in the Western Division of Fiji. PLoS Negl Trop Dis 10, e0004798, https://doi.org/10.1371/journal.pntd.0004798 (2016).
Bailey, R., Duong, T., Carpenter, R., Whittle, H. & Mabey, D. The duration of human ocular Chlamydia trachomatis infection is age dependent. Epidemiol Infect 123, 479–486 (1999).
Rohde, G., Straube, E., Essig, A., Reinhold, P. & Sachse, K. Chlamydial zoonoses. Dtsch Arztebl Int 107, 174–180, https://doi.org/10.3238/arztebl.2010.0174 (2010).
Zoonotic Chlamydiae Maintained in Mammals. Chlamydiosis. 1–14 (Institute for International Cooperation in AnimalBiologics, Iowa State University, Iowa, 2017).
Hartley, J. C. et al. Conjunctivitis due to Chlamydophila felis (Chlamydia psittaci feline pneumonitis agent) acquired from a cat: case report with molecular characterization of isolates from the patient and cat. J Infect 43, 7–11, https://doi.org/10.1053/jinf.2001.0845 (2001).
Wons, J., Meiller, R., Bergua, A., Bogdan, C. & Geissdorfer, W. Follicular Conjunctivitis due to Chlamydia felis-Case Report, Review of the Literature and Improved Molecular Diagnostics. Front Med (Lausanne) 4, 105, https://doi.org/10.3389/fmed.2017.00105 (2017).
Li, J. et al. Chlamydia pecorum is the endemic intestinal species in cattle while C. gallinacea, C. psittaci and C. pneumoniae associate with sporadic systemic infection. Vet Microbiol 193, 93–99, https://doi.org/10.1016/j.vetmic.2016.08.008 (2016).
Laroucau, K. et al. Outbreak of psittacosis in a group of women exposed to Chlamydia psittaci-infected chickens. Euro Surveill 20 (2015).
Hulin, V. et al. Host preference and zoonotic potential of Chlamydia psittaci and C. gallinacea in poultry. Pathog Dis 73, 1–11, https://doi.org/10.1093/femspd/ftv005 (2015).
Guo, W. et al. Chlamydia gallinacea, not C. psittaci, is the endemic chlamydial species in chicken (Gallus gallus). Sci Rep 6, 19638, https://doi.org/10.1038/srep19638 (2016).
Burton, M. J. et al. Which members of a community need antibiotics to control trachoma? Conjunctival Chlamydia trachomatis infection load in Gambian villages. Invest Ophth Vis Sci 44, 4215–4222, https://doi.org/10.1167/iovs.03-0107 (2003).
Burton, M. J., Holland, M. J., Jeffries, D., Mabey, D. C. W. & Bailey, R. L. Conjunctival chlamydial 16S ribosomal RNA expression in trachoma: Is chlamydial metabolic activity required for disease to develop? Clinical Infectious Diseases 42, 463–470, https://doi.org/10.1086/499814 (2006).
Prusty, B. K. et al. Chlamydia trachomatis Infection Induces Replication of Latent HHV-6. Plos One 8, https://doi.org/10.1371/journal.pone.0061400 (2013).
Dirks, J. A. M. C. et al. Chlamydia trachomatis Load in Population-Based Screening and STI-Clinics: Implications for Screening Policy. Plos One 10, https://doi.org/10.1371/journal.pone.0121433 (2015).
Schabereiter-Gurtner, C. et al. Investigation of an anaerobic microbial community associated with a corneal ulcer by denaturing gradient gel electrophoresis and 16S rDNA sequence analysis. Diagn Microbiol Infect Dis 43, 193–199 (2002).
Schabereiter-Gurtner, C. et al. 16S rDNA-based identification of bacteria from conjunctival swabs by PCR and DGGE fingerprinting. Invest Ophthalmol Vis Sci 42, 1164–1171 (2001).
Willcox, M. D. Characterization of the normal microbiota of the ocular surface. Exp Eye Res 117, 99–105, https://doi.org/10.1016/j.exer.2013.06.003 (2013).
Miller, D. & Iovieno, A. The role of microbial flora on the ocular surface. Curr Opin Allergy Clin Immunol 9, 466–470, https://doi.org/10.1097/ACI.0b013e3283303e1b (2009).
Zhou, Y. et al. The conjunctival microbiome in health and trachomatous disease: a case control study. Genome Med 6, 99, https://doi.org/10.1186/s13073-014-0099-x (2014).
St Leger, A. J. et al. An Ocular Commensal Protects against Corneal Infection by Driving an Interleukin-17 Response from Mucosal gammadelta T Cells. Immunity 47, 148–158 e145, https://doi.org/10.1016/j.immuni.2017.06.014 (2017).
Kugadas, A. et al. Impact of Microbiota on Resistance to Ocular Pseudomonas aeruginosa-Induced Keratitis. Plos Pathog 12, https://doi.org/10.1371/journal.ppat.1005855 (2016).
Filardo, S. et al. Diversity of Cervical Microbiota in Asymptomatic Chlamydia trachomatis Genital Infection: A Pilot Study. Front Cell Infect Microbiol 7, 321, https://doi.org/10.3389/fcimb.2017.00321 (2017).
Romano, J. D. et al. A novel co-infection model with Toxoplasma and Chlamydia trachomatis highlights the importance of host cell manipulation for nutrient scavenging. Cell Microbiol 15, 619–646, https://doi.org/10.1111/cmi.12060 (2013).
WHO simplified trachoma grading system. Community Eye Health 17, 68 (2004).
Solomon, A. W. & Kurylo, E. The global trachoma mapping project. Community Eye Health 27, 18 (2014).
Keenan, J. D. et al. Clinical activity and polymerase chain reaction evidence of chlamydial infection after repeated mass antibiotic treatments for trachoma. Am J Trop Med Hyg 82, 482–487, https://doi.org/10.4269/ajtmh.2010.09-0315 (2010).
Last, A. R. et al. Risk factors for active trachoma and ocular Chlamydia trachomatis infection in treatment-naive trachoma-hyperendemic communities of the Bijagos Archipelago, Guinea Bissau. PLoS Negl Trop Dis 8, e2900, https://doi.org/10.1371/journal.pntd.0002900 (2014).
Pickett, M. A., Everson, J. S., Pead, P. J. & Clarke, I. N. The plasmids of Chlamydia trachomatis and Chlamydophila pneumoniae (N16): accurate determination of copy number and the paradoxical effect of plasmid-curing agents. Microbiol-Sgm 151, 893–903, https://doi.org/10.1099/mic.0.27625-0 (2005).
Wright, E. S., Yilmaz, L. S. & Noguera, D. R. DECIPHER, a search-based approach to chimera identification for 16S rRNA sequences. Appl Environ Microbiol 78, 717–725, https://doi.org/10.1128/AEM.06516-11 (2012).
Everett, K. D. E., Bush, R. M. & Andersen, A. A. Emended description of the order Chlamydiales, proposal of Parachlamydiaceae fam. nov. and Simkaniaceae fam. nov., each containing one monotypic genus, revised taxonomy of the family Chlamydiaceae, including a new genus and five new species, and standards for the identification of organisms. Int J Syst Bacteriol 49, 415–440 (1999).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73, 5261–5267, https://doi.org/10.1128/AEM.00062-07 (2007).
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Research 32, 1363–1371, https://doi.org/10.1093/nar/gkh293 (2004).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41, D590–596, https://doi.org/10.1093/nar/gks1219 (2013).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61, 539–542, https://doi.org/10.1093/sysbio/sys029 (2012).
Miller, M. A., Pfeiffer, W. & Schwartz, T. In Creating the CIPRES Science Gateway for inference of large phylogenetic trees. Proc Gatew Comput Environ Workshop GCE 1–8 (2010).
Letunic, I. & Bork, P. Interactive tree of life (iTOL)v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res 44, W242–245, https://doi.org/10.1093/nar/gkw290 (2016).
Rambaut, A. F, a graphical viewer of phylogenetic trees. Available from: http://tree.bio.ed.ac.uk/software/figtree (2014).
Katoh, K. & Standley, D. M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol Biol Evol 30, 772–780, https://doi.org/10.1093/molbev/mst010 (2013).
Waterhouse, A. M., Procter, J. B., Martin, D. M., Clamp, M. & Barton, G. J. Jalview Version 2–a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191, https://doi.org/10.1093/bioinformatics/btp033 (2009).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2-Approximately Maximum-Likelihood Trees for Large Alignments. Plos One 5, https://doi.org/10.1371/journal.pone.0009490 (2010).
Zmasek, C. Archaeopteryx. Available from: http://tree.bio.ed.ac.uk/software/figtree (2015).
Acknowledgements
We are immensely grateful to Prof. Matthias Horn for his invaluable advice and suggestions that greatly improved the manuscript. We thank Dr. Hanna Marti for helpful discussions and Bozidar Jokanovic for assistance in graphical design. This research was funded by the “Laura Bassi Centers of Expertise” program of the Austrian Federal Ministry of Economy through the Austrian Research Promotion Agency (FFG Project Number: 822768).
Author information
Authors and Affiliations
Contributions
E.G., A.I.K. and T.B.A. conceived and designed the study. E.G. performed the experiments and wrote the manuscript. E.G., A.C., F.T. and D.K. analyzed the data. H.A., B.E.B. and T.B.A. provided the samples. E.G., A.I.K., A.C., F.T., E.S., N.S., D.K., N.B., G.G. and T.B.A. reviewed and edited the manuscript. T.B.A. supervised the study.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ghasemian, E., Inic-Kanada, A., Collingro, A. et al. Detection of Chlamydiaceae and Chlamydia-like organisms on the ocular surface of children and adults from a trachoma-endemic region. Sci Rep 8, 7432 (2018). https://doi.org/10.1038/s41598-018-23887-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-23887-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.