Abstract
Antimicrobial resistance against colistin has emerged worldwide and is threatening the efficacy of colistin treatment of multi-resistant Gram-negative bacteria. In this study, PCRs were used to detect mcr genes (mcr-1, mcr-2, mcr-3) in 213 anal and 1,339 nasal swabs from pigs (n = 1,454) in nine provinces of China, and 1,696 cloacal and 1,647 oropharyngeal samples from poultry (n = 1,836) at live-bird markets in 24 provinces. The mcr-1 prevalences in pigs (79.2%) and geese (71.7%) were significantly higher than in chickens (31.8%), ducks (34.6%) and pigeons (13.1%). The mcr-2 prevalence in pigs was 56.3%, significantly higher than in chickens (5.5%), ducks (2.3%), geese (5.5%) and pigeons (0%). The mcr-3 prevalences in pigs (18.7%), ducks (13.8%) and geese (11.9%) were significantly higher than in chickens (5.2%) and pigeons (5.1%). In total, 173 pigs and three chickens were positive for all three mcr genes. The prevalences of the mcr were significantly higher in nasal/oropharyngeal swabs than in the anal /cloacal swabs. Phylogenetic studies identified 33 new mcr-2 variants and 12 new mcr-3 variants. This study demonstrates high prevalences of mcr in pigs and poultry in China, and indicates there is need for more thorough surveillance and control programs to prevent further selection of colistin resistance.
Similar content being viewed by others
Introduction
Antimicrobial resistance is recognized as one of the most serious global health threats with the ESKAPE group of pathogens being a large problem1. Few treatment options are left and this has triggered the reintroduction of the older and less user-friendly antibiotic colistin2. However, the usefulness of colistin as a last resort antimicrobial is now compromised by the presence of an increasing number of mobile colistin resistance genes (mcr). To date, five different mcr and their variants have been described, mainly mcr-1 (11 variants)3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21, mcr-2 (three variants)11,12,13,14,15,16,17,18,21,22, mcr-3 (ten variants)19,20,23,24,25,26, mcr-427 and mcr-528. The mcr-1, mcr-2 and mcr-3 were originally described on plasmids in Enterobacteriaceae but have recently been identified on the chromosomes of Moraxella spp. and Aeromonas veronii21,24,29,30. mcr-4 and mcr-5 have only been described very recently, after we carried out our study in which we used short amplicon PCRs to determine the prevalence of the mcr-1, mcr-2 and mcr-3 in swabs from the initial and terminal alimentary system of pigs and poultry in China. Positive samples were tested with long amplicon PCRs for the three mcr genes that were used for sequencing and comparative studies.
Results
PCRs for mcr
The detection limit for real-time PCRs with short amplicons was one gene copy in a 20 µL reaction mixture, and the detection limit was 50 copies per reaction for conventional long amplicon PCRs. The established PCRs for mcr-1, mcr-2 and mcr-3 in this study amplified only the intended mcr target and not the other mcr31.
Prevalence of mcr-1
The mcr-1 was identified from pigs sampled in all nine provinces, and poultry in 21 of the 24 provinces of China we studied (Fig. 1). The mcr-1 specific PCR identified the gene in 83.6% of anal (178/213) and 79.0% of nasal swabs (1,058/1,139) from pigs, and 25.7% of cloacal (436/1,696) and 28.8% of oropharyngeal swabs (475/1,647) from poultry (1,498 chickens; 130 ducks; 109 geese; 99 pigeons) (Table 1, Tables S1–S5). Overall, the mcr-1 prevalence in chickens and ducks was significantly lower than in pigs and geese, but significantly higher than in pigeons (Fig. 2).
Distribution of mcr-1, mcr-2 and mcr-3 in pigs and poultry. The mcr were detected in specimens from 25 provinces/municipalities of China (mcr-1: 2,147/4,895; mcr-2: 926/4,895; mcr-3: 409/4,895). The prevalences of the mcr were much higher in pigs (mcr-1: 79.2%, 1,152/1,454; mcr-2: 58.4%, 818/1454; mcr-3: 17.3%, 272/1454) than in poultry (mcr-1: 33.0%, 606/1,836; mcr-2: 4.8 88/1,836; mcr-3: 5.7%, 105/1,836). The colors (dark, red, blue), shapes (circle, rectangle, triangle) and positions of filled and empty graphics indicate the mcr positivity, species sampled and the cities involved, respectively. The Adobe Illustrator CS 11.0.0 was used to create the map in this study.
Prevalences of mcr-1, mcr-2 and mcr-3 in swabs from pigs and poultry. The prevalences of mcr-1 in chickens (31.8%) and ducks (34.6%) were significantly lower than those in pigs (79.2%) and geese (71.7%), but significantly higher than in pigeons (13.1%). The mcr-2 prevalence in pigs was 56.3%, significantly higher than in chickens (5.5%), ducks (2.3%), geese (5.5%) and pigeons (0%). The prevalences of the mcr-3 in pigs (18.7%), ducks (13.8%) and geese (11.9%) were significantly higher than in chickens (5.2%) and pigeons (5.1%). Different letters within bars indicate statistical significance across species, as determined by multiple Pearson Chi-Square test via comparing proportions between two species with Bonferroni adjusted p-values.
Geese had the highest mcr-1 prevalence (65.1%, 71/109 of cloacal swabs; 45.0%, 49/109 of oropharyngeal swabs), followed by chickens (23.4%, 323/1,383 of cloacal swabs; 28.6%, 386/1,350 of oropharyngeal swabs), ducks (24.6%, 30/122 of cloacal swabs; 21.3%, 26/122 of oropharyngeal swabs) and pigeons (14.6%, 12/82 of cloacal swabs; 21.2%, 14/66 of oropharyngeal swabs).
Prevalence of mcr-2
The mcr-2 was identified in pigs sampled from all nine provinces, and poultry in 19 of the 24 provinces of China we studied (Fig. 1). The prevalence of mcr-2 in pigs was 56.3% (nasal: 58.4%, 782/1,339; anal: 23.0%, 49/213), being significantly higher than in poultry (oropharyngeal: 3.6%, 60/1,647; cloacal: 2.1%, 35/1,696) (Table 1, Fig. 3, Tables S1–S5). The mcr-2 gene was identified in chickens (5.5%, 82/1,498), ducks (2.3%, 3/130) and geese (5.5%, 6/109), but not in pigeons (0/99).
Prevalence of mcr-1, mcr-2 and mcr-3 in the initial and terminal alimentary system of pigs and poultry. Swabs from both initial and terminal alimentary systems were collected from 98 pigs and 1,507 poultry in this study. For all three mcr genes in swabs from pigs and mcr-1 and mcr-2 in swabs from poultry, the prevalences in nasal/oropharyngeal were significantly higher than in the anal /cloacal swabs.
Prevalence of mcr-3
Overall, the mcr-3 was detected by PCR in 8.4% of samples (409/4,895) including anal (23%, 49/213) and nasal swabs (17.3%, 232/1,339) from pigs, and oropharyngeal (4.9%, 81/1,647) and cloacal swabs (2.8%, 47/1,696) from poultry (Table 1, Fig. 3, Tables S1–S5). Positive mcr-3 PCRs were obtained from pigs in all nine provinces, and poultry sampled in 17 of the 24 provinces (chicken: 5.2%, 78/1,498; duck: 13.8%, 18/130; goose: 11.9%, 13/109; pigeons: 5.1%, 5/99) (Figs 1, 2).
Co-occurrence of mcr-1, mcr-2 and mcr-3
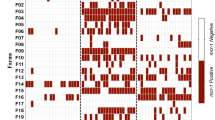
One-hundred and seventy three PCRs on anal and nasal swabs from 1,454 pigs were positive for all three colistin resistance genes (Table 2). Dual positivity was identified in 730 pigs for mcr-1 and mcr-2, 267 pigs for mcr-1 and mcr-3, and 177 pigs for mcr-2 and mcr-3.
Three chickens were positive for mcr-1, mcr-2 and mcr-3. Both of the mcr-1 and mcr-2 were identified in 37 birds (31 chickens; 2 ducks; 4 geese) while 66 birds (48 chickens, 7 ducks, 9 gooses, 2 pigeons) were positive for mcr-1 and mcr-3 and 4 chickens were positive for both mcr-2 and mcr-3 (Table 2).
Comparison of mcr between samples from initial and terminal alimentary systems
Swabs from both of the initial and terminal alimentary systems were collected from 98 pigs and 1,507 poultry. In general, the mcr prevalences in nasal/oropharyngeal swabs were significantly higher than in anal/cloacal swabs (Fig. 3).
For the 98 pigs with both anal and nasal swabs available, 84 were mcr-1 positive in both samples, three had only positive anal swabs, and 11 only had positive nasal swabs. For the mcr-2, only 13 pigs were positive in both swabs, 11 were positive for anal swabs only and 41 for cloacal swabs only. Nine pigs were positive for mcr-3 in both swabs with 12 having only nasal swabs positive and 18 having only anal swabs positive (Fig. 3, Tables 1, 2, Table S1).
Both oropharyngeal and cloacal swabs were positive for mcr-1 in 285 poultry (230 chickens, 11 ducks, 42 geese, 2 pigeons), for mcr-2 in four poultry (3 chickens, 1 goose) and for mcr-3 in 14 poultry (9 chickens, 3 ducks, 2 geese). For the mcr-1, only the cloacal swab was positive for 123 birds (75 chickens, 19 ducks, 29 geese) and only the oropharyngeal swab was positive for 186 birds (152 chickens, 15 ducks, 7 geese, 12 pigeons). For the mcr-2, 23 birds only had positive cloacal swabs (20 chickens, 1 duck, 2 geese) while 42 only had positive oropharyngeal swabs (37 chickens, 2 ducks, 3 geese). Cloacal swabs of 23 birds (18 chickens, 2 ducks, 2 geese, 1 pigeon) were positive for mcr-3, while oropharyngeal swabs of 61 birds (36 chickens, 13 ducks, 9 geese, 3 pigeons) were positive for mcr-3 (Fig. 3, Table 1, Tables 2, S2–S5).
Phylogenetic comparison
We successfully sequenced long amplicons (1,497-bp) of 66 mcr-1 PCRs (31 pigs, 23 chickens, 6 ducks, 3 geese, 3 pigeons), 33 mcr-2 PCRs (28 pigs, 4 chickens, 1 duck) and 25 mcr-3 PCRs (14 pigs, 2 chickens, 2 ducks, 7 geese). The nucleotide sequences of the mcr-1 amplified in this study were identical to those of the mcr-1 previously reported in bacteria from flies (MF598564)31 and Escherichia coli from pigs in China (KP347127)3.
The nucleotide sequences obtained for the mcr-2 revealed 33 new variants of mcr-2 (mcr-2.3 to mcr-2.35) which had high levels of similarity (95.9–99.9%). They also had high similarity (95.8% and 98%) with the mcr-2 from E. coli KP37 isolated from pigs and cattle in Belgium (NG_051171). The other two variants we found had lower similarity: variant mcr-2.1 (MF176239) with 95.4% to 97.5% and variant mcr-2.2 (MF176240) with 87.0% to 88.4% (Fig. 4). The deduced amino acid sequences (347-aa) of the mcr-2.3 to mcr-2.35 were 98.6%-100% identical to those of the E. coli KP37 strain from Belgium (Figure S1).
Phylogenetic analysis of colistin resistance gene mcr-2. The nucleotide sequences of mcr-2 gene (1,042-bp) identified in this study are compared with representative sequences from NCBI (in bold font) using the Neighbor-Joining method. The optimal tree with the sum of branch length (0.419) is shown. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (500 replicates) are shown next to the branches. The evolutionary distances were computed using the Kimura 2-parameter method and are in the units of the number of base substitutions per site. All positions containing gaps and missing data were eliminated. Evolutionary analyses were conducted in MEGA6.
We also identified 12 new variants of the mcr-3 (mcr-3.11 to mcr-3.22) which formed three potential clusters with phylogenetic tree analysis (Fig. 5). The variants in the first cluster (cluster 1) had 0–2 nucleotides mismatches and comprised five from this study (mcr-3.11 to mcr-3.15) and three reported before (mcr-3.1, NG_055505; mcr-3.2, NG_055523, mcr-3.5 (NG_055782). Those in cluster 2 had 2–21 nucleotide mismatches and consisted of seven variants from this study (mcr-3.16 to mcr-3.22) and two described before (mcr-3.3, NG_055783; mcr-3.10, NG_055799). The remaining four variants (mcr-3.6, MF598076; mcr-3.7, NG_055661; mcr-3.8, NG_055662; mcr-3.9, NG_055663) were polymorphic with the other mcr-3 (Fig. 5). Not all sequence mutations lead to differences at the amino acid level with the variants in cluster 1 having 0–2 different amino acids (Figure S2) and those in cluster 2 having 0–8 amino acid changes (Figure S2).
Evolutionary relationships of mcr-3 sequences obtained in this study. The evolutionary history of mcr-3 nucleotides (1,064-bp) identified in this study and NCBI (in bold font) was inferred using the Neighbor-Joining method. The optimal tree with the sum of branch length (0.0372) is shown. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (500 replicates) are shown next to the branches. The evolutionary distances were computed using the Kimura 2-parameter method and are in the units of the number of base substitutions per site. After eliminating the positions containing gaps and missing data, there were a total of 1,063 positions in the final dataset. Evolutionary analyses were conducted in MEGA6.
All the nucleotide sequences were submitted to GenBank with accession numbers MG017397 to MG017429 for mcr-2, and MG017385 to MG017396 for mcr-3.
Discussion
While colistin is a last-line antibiotic used to treat multidrug resistant Gram-negative bacteria, its efficacy is being compromised by the recently detected mobile colistin resistance genes, mcr-13, and subsequently mcr-2, mcr-3, mcr-4, mcr-511,24,27,28. Our study has shown that the mcr-1, mcr-2 and mcr-3 occurred widely in pigs and poultry from China (Table 1, Tables S1–S5). The prevalences we found were considerably higher than those previously reported in China and elsewhere in the world3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,29,30 which were based on the presence of the mcr in bacterial isolates. Bacterial isolation, however, is cumbersome, costly, and time-consuming, and it is also selective in that samples must be appropriately collected, transported and stored to maximize the number of viable bacteria with the mcr that can be selectively cultured. Directly testing samples with the sensitive and specific mcr-PCRs we developed meant that we avoided the limitations of isolation and the resultant underestimation of the mcr genes that the technique involves. The phenomenon whereby direct sample testing gives far higher prevalences than testing of isolates from samples has been termed the ‘phantom resistome’32 and indicates that the existing data on the prevalence of the mcr-1, mcr-2 and mcr-3 in China greatly underestimates the real situation. We note, however, that both methods are needed to understand the epidemiology of resistance. While direct PCR testing is ideal for the rapid estimation of risk and risk analysis, it does not readily enable investigations of movement of resistance between bacteria of the same or different species. As we did not base the study on the isolation of the bacteria, we are not sure which bacteria carry the resistance genes we identified, or the mobile genetic elements on which the genes are carried. Previous studies have shown the mcr-1, mcr-2 and mcr-3 can be on plasmids in Enterobacteriaceae and chromosomes in Moraxella spp.21,29,30.
The high levels of the mcr-1 we found in the pigs and the poultry we studied from 21 of the 24 provinces in China (Table 1, Tables S1–S5) is consistent with previous reports from China20,31,32,33 and is likely associated with the prolonged and widespread use of colistin as a growth promoter in food animals.
We also found a high prevalence of the mcr-2 in both pigs and poultry which is surprising as this gene has only been found in a limited number of other studies11,12,14,15,17,18,22,27. This is likely because our PCR for mcr is more sensitive in detecting the resistance gene than the conventional methods relying on bacterial isolation as discussed above. The PCR, however, did not enable us to determine the prevalences of mcr-2 in the different bacterial populations in our samples and it is possible that only a small part of the bacterial population carries this resistance gene. In studies on selectively isolated bacteria however, its presence may be masked if only low numbers of mcr-2 positive strains are present and higher numbers of strains with mcr-1 or mutational mediated resistance strains.
The mcr-3 gene was first characterized on a IncHI2-type plasmid pWJ1 from porcine E. coli in China23 and later shown to be present in bacteria isolated from humans from Denmark, chickens from China, cattle from Spain19,20,24,25,26, and flies in China31. Our findings indicate that the gene is quite widespread, as we found it in samples from pigs and poultry from many provinces in China, albeit at low prevalence in some cases. Further studies in other countries and from distinct sources should be performed to determine how this gene is spread and its overall role in colistin resistance.
The mcr-1 sequences were highly conserved in the pigs and various poultry we studied and identical to sequences described before3. However, the nucleotide sequences of the mcr-2 we found varied and we identified 33 variants which were very similar (21–44 nucleotide mismatches) to those originally reported by Xavier BB et al.22. Although highly polymorphic at the nucleotide level, there were only few differences at the amino acid level (0–5 mismatches). It remains unclear whether these mutations influence the level of colistin resistance. Furthermore, these data are difficult to interpret, as we do not know in which bacterial species the gene resides. Differences in codon usage of the different bacterial species might have been the cause of the high variability found here. Further studies are warranted to explore the influence of mcr-2 mutation on colistin resistance.
Alignment of mcr-3 nucleotide sequences obtained in this study (mcr-3.11 to mcr-3.22) and those reported before (mcr-3.1 to mcr-3.10) revealed five clusters. Those in cluster 1 were obtained from pigs and people, except for the mcr-3.12 which was obtained from a goose in our study. All the sequences in clusters 2, 3 and 4 were derived from poultry, while sequences of cluster 5 were from fish and turkeys. The host specificity of the mcr-3 was demonstrated at sites 188, 286, 302, 321, 326, 330 and 347 of the amino acid sequences. Further studies are needed to investigate the possible interrelationship between the host-specificity of nucleotide and amino acid sequences and the epitopes of colistin resistance.
Our comparison of the prevalence of the mcr genes in different ecosystems on the same animal, that is both nasal/oropharyngeal swabs and anal/cloacal swabs from pigs and chickens, respectively, showed that most positive animals had mcr containing bacteria at both sites. In the remaining animals, more were only mcr positive in nasal/ oropharyngeal swabs than only positive in anal/cloacal swabs. These results are consistent with bacteria in these ecosystems being exposed to colistin during ingestion of food containing the additive, and also during digestion of this food. It should also be noted, however, that there can be substantial concentration of colistin in dust34 and this might also be a source of respiratory mucosa exposure. Whereas feces are most commonly considered as the major factor in the transfer and spread of the mcr genes, our data showing significantly higher mcr prevalences in nasal/oropharyngeal swabs than in anal/cloacal swabs suggests that both saliva and respiratory secretions might also play important roles. The identification of mcr variants in Moraxella species (mcr-1 and mcr-2)21,29,30 and Aeromonas veronii (mcr-3)24 indicates there are organisms outside the Enterobacteriaceae that contribute to colistin resistance and these might be responsible for the differences in prevalences that we found.
In summary, our study indicates that the mcr-1, mcr-2 and mcr-3 are relatively common and widespread in food producing animals of China. The high sequence variability of some of the genes indicates there is ongoing evolution, probably in different bacterial species. Future studies should focus on the bacterial species carrying these genes and the localization of these genes in the microorganisms.
Materials and Methods
Ethics statement
This study was reviewed and approved by the Institutional Animal Care and Use Committee of Yangzhou University College of Veterinary Medicine, and was performed in accordance with the relevant guidelines and regulations.
Swab samples from pigs and poultry
In 2016, 232 anal and 1,339 nasal swabs from apparently healthy pigs (n = 1,552) in nine provinces were collected. In addition, 1,690 cloacal and 1,628 oropharyngeal samples from poultry (n = 1,836) at 38 live-bird markets in 24 different provinces in China were collected in 2014 (Fig. 1)35. Both of the swabs from the initial and terminal alimentary systems were collected from 98 pigs and 1,507 poultry in this study.
To collect an oropharyngeal swab sample from poultry, the swab was introduced into the bird’s mouth and rubbed around the tracheal opening and up along the choanae. To collect the cloacal swabs from poultry, the swab was gently introduced into the cloaca through the vent and gently twirled at an appropriate depth to ensure contact with the mucous membranes.
To collect the porcine nasal swabs, the nose was wiped with a dry piece of paper and the swab inserted into the ventral nasal passage and rotated through ninety degrees for three seconds. To collect anal swabs from pigs, the swab was inserted one cm into the rectum while being rotated.
Following sampling, the swabs were immersed in 400 μl DNA/RNA Stabilization Buffer (Roche Molecular Biochemicals, IN, USA) in sterile tubes and frozen at −80 °C until DNA was extracted.
DNA extraction
Swabs were centrifuged in the DNA/RNA Stabilization Buffer (3,000×g, 4 °C for 5 min) and DNA was extracted from the supernatants using either the High Pure PCR Template Preparation Kit (Roche Diagnostic, USA) following the manufacturer’s instructions for the oropharyngeal and nasal swabs or the QIAamp DNA Stool Mini Kit (Qiagen, USA) for the cloacal and anal swabs31.
PCR assays
In this study, we used previously described PCRs31. These PCRs do not amplify the chromosomal mcr genes described in Moraxella hydrophila and Aeromonas veronii. The PCRs for the mcr-1, mcr-2 and mcr-3 genes were performed on a Roche Light-Cycler 480II PCR instrument31. The short amplicon PCRs for mcr-1(342-bp), mcr-2 (282-bp) and mcr-3 (267-bp) were used to establish prevalence data and positive samples were tested with the long amplicon PCRs for mcr-1 (1,497-bp), mcr-2 (1,042-bp) and mcr-3 (1,063-bp). Positive long amplicon PCR products were verified by gel electrophoresis and sequenced using upstream and downstream primers (BGI, Shanghai, China). Sequences obtained were compared with those published in the NCBI database (www.ncbi.nlm.nih.gov) using the Clustal Multiple Alignment Algorithm. Negative controls consisting of sterile molecular grade water were used to detect cross-contamination during DNA purification and PCR processing.
Phylogenetic analysis
The sequences from this study and those from GenBank for the mcr-1, mcr-2 and mcr-3 were aligned using the MEGA 6.0 software. Based on these alignments, phylogenetic trees were constructed by the neighbor-joining method using the Kimura 2-parameter model with MEGA 6.0. Bootstrap values were calculated using 500 replicates.
Statistical analysis
Multiple Pearson Chi-Square test was used for comparing differences between animal species as well as between anal/oropharyngeal and nasal/oral swabs with Bonferroni adjusted p-values.
References
Pogue, J. M., Kaye, K. S., Cohen, D. A. & Marchaim, D. Appropriate antimicrobial therapy in the era of multidrug-resistant human pathogens. Clin. Microbiol. Infect. 21, 302–312 (2015).
Temkin, E., Adler, A., Lerner, A. & Carmeli, Y. Carbapenem-resistant Enterobacteriaceae: biology, epidemiology, and management. Ann. N. Y. Acad. Sci. 1323, 22–42 (2014).
Liu, Y. Y. et al. Emergence of plasmid-mediated colistin resistance mechanism mcr-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect. Dis. 16, 161–168 (2016).
Doumith, M. et al. Detection of the plasmid-mediated mcr-1 gene conferring colistin resistance in human and food isolates of Salmonella enterica and Escherichia coli in England and Wales. J. Antimicrob. Chemother. 71, 2300–2305 (2016).
El Garch, F. et al. mcr-1 is borne by highly diverse Escherichia coli isolates since 2004 in food-producing animals in Europe. Clin. Microbiol. Infect. 23, 51.e1–51.e4 (2017).
Haenni, M., Metayer, V., Gay, E. & Madec, J. Y. Increasing trends in mcr-1 prevalence among ESBL-producing E. coli in French calves despite decreasing exposure to colistin. Antimicrob. Agents Chemother. 60, 6433–6434 (2016).
Hasman, H. et al. Detection of mcr-1 encoding plasmid-mediated colistin-resistant Escherichia coli isolates from human bloodstream infection and imported chicken meat, Denmark 2015. Euro Surveill. 20, pii:30085 (2015).
Lei, L. et al. mcr-1 in Enterobacteriaceae from companion animals, Beijing, China, 2012–2016. Emerg. Infect. Dis. 23, 710–711 (2017).
Veldman, K. et al. Location of colistin resistance gene mcr-1 in Enterobacteriaceae from livestock and meat. J. Antimicrob. Chemother. 71, 2340–2342 (2016).
Nordmann, P., Assouvie, L., Prod’Hom, G., Poirel, L. & Greub, G. Screening of plasmid-mediated mcr-1 colistin-resistance from bacteremia. Eur. J. Clin. Microbiol. Infect. Dis. 35, 1891–1892 (2016).
Kawanishi, M. et al. Prevalence of colistin resistance gene mcr-1 and absence of mcr-2 in Escherichia coli isolated from healthy food-producing animals in Japan. Antimicrob. Agents Chemother. 61, pii:e02057–16 (2016).
Pragasam, A. K. et al. Molecular mechanisms of colistin resistance in Klebsiella pneumoniae causing bacteremia from India-a first report. Front. Microbiol. 7, 2135, https://doi.org/10.3389/fmicb.2016.02135 (2017).
Prim, N. et al. Prevalence of colistin resistance in clinical isolates of Enterobacteriaceae: A four-year cross-sectional study. J. Infect. 75, 493–498 (2017).
Roschanski, N. et al. Retrospective survey of mcr-1 and mcr-2 in German pig-fattening farms, 2011-2012. Int. J. Antimicrob. Agents. 50, 266–271 (2017).
Sari, A. N. et al. Results of a multicenter study investigating plasmid mediated colistin resistance genes (mcr-1 and mcr-2) in clinical Enterobacteriaceae isolates from Turkey. Mikrobiyol. Bul. 51, 299–303 (2017).
Venditti, C., Nisii, C., D’Arezzo, S., Vulcano, A., Di Caro, A. Letter to the Editor: Surveillance of mcr-1 and mcr-2 genes in Carbapenem-resistant Klebsiella pneumoniae strains from an Italian Hospital. Euro Surveill. 22, pii = 30604 (2017).
Wang, X. et al. Molecular epidemiology of colistin-resistant Enterobacteriaceae in inpatient and avian isolates from China: high prevalence of mcr-negative Klebsiella pneumoniae. Int. J. Antimicrob. Agents. 50, 536–541 (2017).
Zurfluh, K. et al. Screening for fecal carriage of MCR-producing Enterobacteriaceae in healthy humans and primary care patients. Antimicrob. Resist. Infect. Control. 6, 28, https://doi.org/10.1186/s13756-017-0186-z (2017).
Hernández, M. et al. Co-occurrence of colistin-resistance genes mcr-1 and mcr-3 among multidrug-resistant Escherichia coli isolated from cattle, Spain, September 2015. Euro Surveill. 22, pii30586 (2017).
Liu, L., Feng, Y., Zhang, X., McNally, A. & Zong, Z. A new variant of mcr-3 in an extensively drug-resistant Escherichia coli clinical isolate carrying mcr-1 and blaNDM-5. Antimicrob. Agents Chemother. 61, pii:AAC.01757-17 (2017).
AbuOun, M. et al. mcr-1 and mcr-2 variant genes identified in Moraxella species isolated from pigs in Great Britain from 2014 to 2015. J. Antimicrob. Chemother. 72, 2745–2749 (2017).
Xavier, B. B. et al. Identification of a novel plasmid-mediated colistin-resistance gene, mcr-2, in Escherichia coli, Belgium, June 2016. Euro Surveill. 21, pii:30280 (2016).
Yin, W. et al. Novel plasmid-mediated colistin resistance gene mcr-3 in Escherichia coli. MBio. 8, pii:e01166-17 (2017).
Ling, Z. et al. Chromosome-mediated mcr-3 variants in Aeromonas veronii from chicken meat. Antimicrob. Agents Chemother. 61, pii:e01272–17 (2017).
Litrup, E. et al. Plasmid-borne colistin resistance gene mcr-3 in Salmonella isolates from human infections, Denmark, 2009-17. Euro Surveill. 22, pii:30587 (2017).
Roer, L. et al. Novel mcr-3 variant, encoding mobile colistin resistance, in an ST131 Escherichia coli isolate from bloodstream infection, Denmark, 2014. Euro Surveill. 22, pii:30584 (2017).
Carattoli, A. et al. Novel plasmid-mediated colistin resistance mcr-4 gene in Salmonella and Escherichia coli, Italy 2013, Spain and Belgium, 2015 to 2016. Euro Surveill. 22, pii:30589 (2017).
Borowiak, M. et al. Identification of a novel transposon-associated phosphoethanolamine transferase gene, mcr-5, conferring colistin resistance in d-tartrate fermenting Salmonella enterica subsp. enterica serovar Paratyphi B. J. Antimicrob. Chemother. 72, 3317–3324 (2017).
Kieffer, N., Nordmann, P. & Poirel, L. Moraxella species as potential sources of MCR-like polymyxin resistance determinants. Antimicrob Agents Chemother. 61, pii: e00129–17 (2017).
Poirel, L. et al. MCR-2-mediated plasmid-borne polymyxin resistance most likely originates from Moraxella pluranimalium. J. Antimicrob. Chemother. 72, 2947–2949 (2017).
Zhang, J. et al. Housefly (Musca domestica) and blow fly (Protophormia terraenovae) as vectors of colistin resistance genes-carrying bacteria. Appl. Environ. Microbiol. 84, pii: e01736-17 (2017).
Wang, Y. et al. Comprehensive resistome analysis reveals the prevalence of NDM and MCR-1 in Chinese poultry production. Nat. Microbiol. 2, 16260, https://doi.org/10.1038/nmicrobiol.2016.260 (2017).
Shen, Z., Wang, Y., Shen, Y., Shen, J. & Wu, C. Early emergence of mcr-1 in Escherichia coli from food-producing animals. Lancet Infect. Dis. 16, 293 (2016).
Schulz, J. et al. Antimicrobial-resistant Escherichia coli survived in dust samples for more than 20 years. Front. Microbiol. 7, 866, https://doi.org/10.3389/fmicb.2016.00866 (2016).
Guo, W. et al. Chlamydia gallinacea, not C. psittaci, is the endemic chlamydial species in chicken (Gallus gallus). Sci. Rep. 6, 19638, https://doi.org/10.1038/srep19638 (2016).
Acknowledgements
This work was supported by a grant from the National Key R & D Program of China (2016YFD0500804), a grant from the National Natural Science Foundation of China (NO: 31472225), the Priority Academic Program Development of Jiangsu Higher Education Institutions, and the Jiangsu Co-Innovation Center for Prevention and Control of Important Animal Infectious Diseases and Zoonoses.
Author information
Authors and Affiliations
Contributions
C.W., J.Z. and L.C. conceived the study, J.Z., L.C., A.K.Y., J.G., W.G., J.L., M.L., F.Y., Z.F., P.J., C.S., Y.W., J.Y., and Y.Y. collected the samples and performed the experiments. J.L., L.C., C.W., P.B., P.K., S.P., K.Q. and Y.K. collected and analyzed the data. C.W., J.Z., L.C., P.B., P.K. and S.P. wrote the first draft of this work. All authors contributed toward data analysis and revising the paper and agree to be accountable for all aspects of the work.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, J., Chen, L., Wang, J. et al. Molecular detection of colistin resistance genes (mcr-1, mcr-2 and mcr-3) in nasal/oropharyngeal and anal/cloacal swabs from pigs and poultry. Sci Rep 8, 3705 (2018). https://doi.org/10.1038/s41598-018-22084-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-22084-4
This article is cited by
-
Population structure and antibiotic resistance of swine extraintestinal pathogenic Escherichia coli from China
Nature Communications (2024)
-
Plasmid-mediated colistin resistance and ESBL production in Escherichia coli from clinically healthy and sick pigs
Scientific Reports (2022)
-
Molecular survey of mcr1 and mcr2 plasmid mediated colistin resistance genes in Escherichia coli isolates of animal origin in Iran
BMC Research Notes (2021)
-
Phenotypic and genotypic characterization of mcr-1-positive multidrug-resistant Escherichia coli ST93, ST117, ST156, ST10, and ST744 isolated from poultry in Poland
Brazilian Journal of Microbiology (2021)
-
Rapid detection of human origin colistin-resistance genes mcr-1, mcr-3, mcr-8, mcr-10 in clinical fecal samples
Archives of Microbiology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.