Abstract
The circadian clock in plants is the intrinsic rhythmic expression of thousands of genes in a 24 h period, which is set by the day-night cycles in the environment. The study of the circadian clock often requires expression profiling of genes over a large number of samples for which RT-qPCR is invariably used. Reliability of the results depends largely on the house-keeping genes, which serve as control and thus should be chosen carefully to prevent erroneous results. In this study, ten house-keeping genes were chosen from rice for stability analysis with 48 tissue samples harvested from plants subjected to diurnal/circadian cycles. Although, all the genes were found to be stable, however, six of them showed cyclic expression patterns and caused major changes in the expression profiles of the target genes when used to normalize their expression data, thereby making them poor candidates for diurnal/circadian studies. In conclusion, reference genes need to be selected for diurnal/circadian studies with extra caution as more than 80% of transcriptome in plants undergoes cycling, which remains undetected by the gene stability assessment software and can severely affect the RT-qPCR based gene expression profiling. The geometric mean of two or more most stable reference genes is hence recommended for diurnal/circadian studies in plants.
Similar content being viewed by others
Introduction
Plants perceive their environment as a daily change in light and temperature conditions known as photocycles and thermocycles, respectively that vary with different seasons and regions (latitude) on the planet. These sessile organisms, like all animals, have evolved a 24 h period circadian clock that synchronizes their internal physiology and metabolism to the correct time of the day with the daily photo- and thermo-cycle. This is an important means of their assessment of the time of day and season1,2,3. Circadian clock benefits the plants with increased fitness and vigour and provides an adaptive advantage as they are able to anticipate daily and seasonal environmental variations and make appropriate changes in their metabolism4,5. It has been found that up to 89% of genes in Arabidopsis, when entrained in a day-night cycle, show daily cyclic changes (oscillate) in their transcript levels with a peak at some point of the day; such genes are diurnally controlled. About 10–30% genes are such that will continue to oscillate with the same rhythm, for some days, even when transferred to a free running or constant condition, i.e. when there is no day-night cycle, and such genes are called circadian genes1,2. These genes continue to drive the circadian rhythms of plants, at least for some days, even in the absence of external environmental cues. Nearly all the genes associated with hormone biosynthesis and signaling and the integrated stress signaling pathway are regulated by the circadian clock that synchronizes them for better growth and development and improved fitness6,7,8.
Study of the involvement of a gene in the functioning of the circadian clock oscillator, or if the gene is controlled by the circadian clock itself, frequently requires gene expression profiling to be carried out of the gene itself and its effect on the signature output pathway genes in diurnal as well as circadian conditions (free running). It becomes a mammoth task as the sample number is invariably high in studies dealing with circadian gene expression and the precision required is also high. Real-time quantitative PCR (RT-qPCR) has been widely accepted and employed as a method for gene expression quantification in plant science research9. It is the method of choice for high throughput, precise and speedy quantification of native genes in diurnally entrained and/or free running (circadian) tissue samples for studying cyclic expression patterns. The technique has become all the more powerful with the introduction of nano-litre plates that can analyze 3072 samples at the same time10 and the microfluidic technology where gene expression is measured using dynamic arrays, empowering the users to analyze 9216 samples in the same run11. Having said this, it would be appropriate to talk about the shortcomings and careful considerations that need to be addressed for a correct and meaningful analysis using RT-qPCR, as the technique is prone to both human and technical errors. After careful tissue sampling, RNA extraction and purification, and cDNA synthesis with all possible quality and quantity checks and minimized pipetting errors, the most important factor to be considered is the selection of one, or more appropriately, more than one most suitable house-keeping genes (also known as internal control or reference genes) for normalizing the data12,13. The most important criteria for selection of house-keeping genes for an experiment is that its expression level should remain stable across all the samples to be analysed12,14,15,16. These genes were initially thought to play central roles in house-keeping activities or basic cellular processes in the cell and hence were presumed to be constantly expressed and unregulated. Later, with the growing popularity of this technique, it was found that these house-keeping genes were also regulated under certain conditions17,18, hence their preference as control genes vary for different sample categories19. In general, there are no fixed internal control genes that can be used for all RT-qPCR analyses and the choice of these genes differ with experiment type to be performed, such as exposure to abiotic stresses, hormonal treatments, developmental stages, diurnal studies and other treatments. An internal control best suited for one experimental set may vary in another set of treated tissues and hence may be rendered unsuitable, which is the case for most of the known house-keeping genes in plants. Choosing an inappropriate internal control can lead to anonymous errors and eventually to serious misleading interpretations of results and conclusions12,13,14,16,19.
Rice is not only a major food crop in several Asian countries, feeding millions of people, but is also a model system for genetic and genomic studies in monocots as its genome is fully sequenced to finished quality and very well annotated. It is the simplest among all the major cereals, such as wheat and maize, and shows good genetic correlations, making the studies with other cereal crops easier. Over the past decade, several house-keeping genes have been identified across cereals, legumes, shrubs and woody perennials and evaluated for their stability under different conditions using freely available statistical algorithms, such as GeNORM12,20, BestKeeper21, NormFinder22, ∆Ct23, RefFinder24, etc. Many new house-keeping genes have been identified in Arabidopsis25 and in rice26 for a wide variety of sample types. These studies are sufficient to assess the possible best internal control genes for some common set of experiments across many other species but there is a continuous need for expanding the list of experiments and sample types. Until now, to the best of our knowledge, there has been no proper validation of house-keeping genes for diurnal and circadian studies in dicots or monocots.
In this study, we have attempted to evaluate 10 commonly used house-keeping genes, earlier validated for rice in our laboratory27, for their utility as best suitable internal controls for diurnal and circadian expression profiling in rice for normalizing RT-qPCR data and also reinforce the importance of using multiple genes as superior internal controls. Also, the reliability of the results was confirmed by normalizing the RT-qPCR data of 5 known circadian genes against the chosen 10 reference genes and analyzing their expression profiles. We have shown that the expression of some of the popular house-keeping genes in rice oscillates with a circadian rhythm and hence are unsuitable for normalizing RT-qPCR data in diurnal/circadian studies. Furthermore, we conclude that the geometric mean of three reference genes is best for normalizing circadian RT-qPCR data.
Results
Selection of candidate house-keeping genes, primer designing, amplification efficiency and specificity
In this study, a total of 10 house-keeping genes from rice were considered for analysis based on a previously published study from our laboratory27. We had chosen some of the most commonly used house-keeping genes at that time, when the rice genome was sequenced and made publically available (IRGSP 2005)28 and was being actively annotated; our group also actively participated in this initiative. These genes are still very popular amongst the researchers across the globe and numerous studies have been carried out utilizing these genes for normalization of the RT-qPCR data. In fact, utility of these genes has been already demonstrated in a variety of plant tissues (developmental stages) as well as treated samples by many groups. The PCR efficiencies for all the primer pairs tested in our study ranged between 93% (eEF-1α) and 107% (eIF-4a) (Table 1). The melting curve analysis revealed single peaks for each primer pair indicating the specificity of the PCR reactions (Fig. 1A), which was also confirmed by analyzing the final PCR products on 2% agarose gel that showed presence of a single band of expected sizes (Fig. 1B). The primers, thus used, were efficient and specific enough for accepting them for routine RT-qPCR analysis.
Specificity of the primers used in RT-qPCR amplification. (A) Melting curves of all the 10 house-keeping genes showing single peaks obtained from the three technical replicates of the first day samples (12 cDNA samples). (B) 2% agarose gel picture showing amplification of specific PCR products of expected sizes by the primer pairs of each of the house-keeping genes used in this study.
Expression profiling of the house-keeping genes
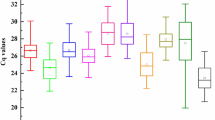
The amplification plots of all the house-keeping genes are given in Supplemental Fig. S1 that shows efficient amplification rates for all the primer pairs. The mean of the three technical replicates of the two biological replicates (six Ct values in total) were averaged and used for all further analyses; these values have been presented in Supplemental Table S1. The mean Ct values for the 10 reference genes across the 48 diurnal and circadian samples ranged from 9.34 for 25S rRNA to 27.34 for UBC (Fig. 2). While the Ct values for UBQ5 (22.79 ± 0.39 SD, 1.71% CV), UBQ10 (21.54 ± 0.49 SD, 2.3% CV), eEF-1α (22.67 ± 0.45 SD, 2% CV), ACT11 (24.11 ± 0.56 SD, 2.34% CV), GAPDH (23.20 ± 0.70 SD, 3.02% CV), β-TUB (23.52 ± 0.42 SD, 1.8% CV), and eIF-4a (24.49 ± 0.42 SD, 1.74% CV) lied mostly between 20 and 25 and hence they were moderately expressed, the Ct values for UBC (26.11 ± 0.53 SD, 2.04% CV) were higher and ranged between 25 and 27, which makes it the lowest expressing, amongst the 10 genes selected for this study. The 18S rRNA (10.56 ± 0.59 SD, 5.57% CV) and 25S rRNA (10.05 ± 0.50 SD, 5.03% CV) transcripts were found to be extremely abundant and their Ct values ranged between 9 and 11. The coefficient of variation for all the genes ranged from 1.71%–5.57%. The expression variability across all the samples for all the house-keeping genes (Fig. 3) was higher for GAPDH, ACT11, UBQ10, UBC, 18S rRNA and 25S rRNA and lower for UBQ5, eEF-1α, β-TUB and eIF-4a. Moreover, this expression variability was least when the geometric mean of UBQ5, eEF-1α and eIF-4a (Ct 23.3 ± 0.39 SD, 1.69% CV) was taken for normalization.
The mean Ct values of all the house-keeping genes. The graph shows the overall variation in the Ct values for each of the house-keeping genes across all the 48 samples. The Ct values of 18S rRNA and 25S rRNA were very low indicating extremely high expression levels. The error bars represent standard error. The average Ct values for all the house-keeping genes have been given in Supplemental Table S1.
Variation in Ct values of the house-keeping genes across all the cDNA samples. Box plot graph showing variation in Ct values for each of the house-keeping genes across all the 48 diurnal/circadian samples. The median values are represented as lines across the box. The lower and the upper boxes represent the 25th and 75th percentile, respectively. Whiskers represent the maximum and minimum values.
Expression stability analysis of the house-keeping genes
The mean Ct values were directly imported to the four algorithms used in this study, i.e. GeNORM, NormFinder, BestKeeper and ∆Ct that calculate the stability of the genes across the entire sample set in an experiment. In general, all the house-keeping genes appeared to be stable (Fig. 2) with overall less variation in their expression values but their suitability for diurnal and circadian studies needed further assessment and confirmation.
GeNorm measures the stability of a reference gene based on the principle that the expression ratio between two ideal reference genes should remain the same under all experimental conditions and in the tissues being analyzed. Any deviation from this ratio will reflect either one or both reference genes being not constantly expressed and any increase in this ratio will mean a corresponding decrease in gene stability. The algorithm assigns a gene stability value M to all the reference genes, which is the average pairwise variation of a gene with all other reference genes included in the study. The genes with least M value have the most stable expression and stepwise exclusion of genes with highest M-values will result in a combination of two genes with highest stability12. eIF-4a, eEF-1α and UBQ5 were predicted to be the most stable genes by GeNORM with least M-values of 0.139, 0.184 and 0.219, respectively (Fig. 4A). GAPDH, ACT11 and 18S rRNA were found to be least stable with the M-values of 0.445, 0.418 and 0.385, respectively. Nevertheless, all the genes analyzed showed good overall stability with their M values much lower than the set threshold (M < 1.5) by Vandesompele et al.12. The BestKeeper algorithm determines the best stable reference genes out of a set of at least 10 genes for an experimental set, based on the principle that the expression of good reference genes are highly correlated and constructs a correlative index out of it, which is then compared with the target genes to reach a conclusion whether or not the reference genes are regulated21. UBQ5, β-TUB and eIF-4a were predicted to be most stable by the BestKeeper algorithm with the stability values (standard deviation) of 0.299, 0.314 and 0.331, respectively, followed by eEF-1α (stability value 0.348). GAPDH (0.577), 18S rRNA (0.487) and ACT11 (0.443) were the least stable genes (Fig. 4B). The cut-off value for this stability measure is 1, below which all the genes are deemed to be stably expressed; this was the case for all the genes investigated in this study. NormFinder is based on the principle that any house-keeping gene will show some extent of variation across the entire set of samples to be studied in an experiment. Hence, to improve the stability measure of the reference genes, it takes into account this overall variation and, in addition, it also considers the variation between different subgroups of samples in the experiment, such as control and treated samples22. NormFinder predicted UBC, eIF-4a and UBQ5 to be the most stable genes with stability values of 0.0064, 0.011 and 0.013, respectively, followed by eEF-1α (0.0135), whereas 18S rRNA, 25S rRNA, ACT11, and GAPDH were the least stable with stability values of 0.447, 0.329, 0.0195 and 0.0191, respectively (Fig. 4C). The ∆Ct approach compares the relative expression of pairs of genes for each sample in an experiment and ranks the reference genes for their stability according to repeatability of the expression difference across all the samples23. According to the ∆Ct approach, eIF-4a (SD = 0.373), 25S rRNA (SD = 0.389) and UBQ5 (SD = 0.4) were the most stable genes followed by eEF-1α (SD = 0.403), whereas GAPDH (SD = 0.567), 18S rRNA (SD = 0.546) and ACT11 (SD = 0.541) were the least stable (Fig. 4D). Taking together, the results suggest eIF-4a, UBQ5 and eEF-1α to be the most stable genes as all four algorithms have given them high ranking for their stability. In comparison, GAPDH, 18S rRNA and ACT11 were given the least rankings by all the algorithms and hence were the least stable.
Expression stability analysis and ranking of the house-keeping genes by the four algorithms used in this study. Gene expression stability plots and subsequent ranking of the house-keeping genes by (A) GeNORM, based on average stability value (M), (B) BestKeeper, based on the standard deviation values, (C) NormFinder, based on the stability values calculated by the software, and (D) ∆Ct, based on the standard deviation values. The most stable genes are on the right hand side and the least stable genes are arranged on the left. The stability values, for each gene, by all the four programs are indicated on the curves.
Optimization of the number of reference genes required for RT-qPCR analysis
It is also important to know the optimum number of reference genes that are required to normalize RT-qPCR data for the given samples in an experiment. Based on the stability of expression, GeNorm is also programmed to evaluate the number of house-keeping genes required to normalize RT-qPCR data. For this purpose, the three most stable genes (with least M-values) are used to calculate the normalization factor (NFn, n = 3) and stepwise inclusion of one more reference gene until the inclusion of (n + 1)th gene has no significant effect on the newly arrived normalization factor (NFn+1). Pairwise variation, Vn/n+1, calculated between the sequential normalization factors, NFn and NFn+1, if large, means that the added gene has a significant contribution for a reliable normalizing factor and should be included in the analysis. If this variation is negligible or low, it means the newly added gene has no significant contribution towards the normalizing factor and is virtually unnecessary12. Taking into consideration, the recommended pairwise variation cutoff value (V < 0.15), the geometric mean of two most stable reference genes, i.e. eIF-4a/UBQ5, eIF-4a/eEF-1α or eEF-1α/UBQ5, is sufficient to normalize diurnal/circadian RT-qPCR data in rice, and no third gene is required, as the pairwise variation was far less than the set cut-off (0.15) for all the genes (Fig. 5). Keeping in mind that certain genes show very weak cyclic patterns or have a very low transcript abundance, and can be badly affected by the reference gene selection, it is recommended that the geometric mean of three most stable reference genes (eIF-4a/UBQ5/eEF-1α) be taken for normalization of diurnal/circadian RT-qPCR data in rice. Although, there was no significant improvement in the stability value of geometric mean of the three genes (M = 0.139, GeNorm; SD = 0.298, BestKeeper; stability value = 0.001, NormFinder and SD = 0.348, ∆CT), it has been ranked number one by GeNorm, BestKeeper and ∆Ct methods and number two by NormFinder, making the combination most stable.
Pairwise variation (V) analysis of the house-keeping genes by GeNORM for finding out the optimal number of genes for data normalization. The pairwise variation (Vn/Vn+1) was analyzed between normalization factors NFn and NFn+1. The default cut-off value was 0.15, below which all gene pairs were considered stable with no need to add the third gene for normalization.
Validation of the house-keeping genes for normalizing diurnal/circadian RT-qPCR data in rice
Firstly, the geometric mean of eIF-4a/UBQ5/eEF-1α was used to analyze the expression patterns of all the 10 reference genes. Secondly, in order to find out the effects of the individual house-keeping genes and the geometric means of the three most and least stable house-keeping genes on the expression patterns of known diurnal/circadian genes, the expression data of five known diurnal/circadian genes, viz. OsLHY/CCA1, OsTOC1, OsFKF1, OsRVE1 and OsRVE8, were normalized and analyzed. Thirdly, the reference genes were interchangeably used to normalize the expression of the remaining reference genes to see their individual effects on the expression patterns of non-cyclic genes and genes with weak cyclic patterns.
When the reference genes were studied for their expression profiles (Fig. 6), eIF-4a, UBQ5 and eEF-1α showed no cyclic pattern when their expression was normalized with respect to the geometric mean of eIF-4a/UBQ5/eEF-1α and were very stable. The UBQ10, although did not show any cyclic pattern, was highly unstable and showed irregular peaks and troughs. UBC had a morning expression pattern with the peak forming two hours after dawn, ZT 2, in a circadian fashion, although the peaks were very shallow and the expression appeared stable. ACT11 and GAPDH were clearly morning expressing genes, peaking immediately at dawn (ZT 0) and two hours after dawn (ZT 2), respectively. GAPDH showed sharp peaks whereas ACT11 exhibited relatively shallow peaks and both the genes oscillated in a circadian fashion. β-TUB, on the other hand, showed evening expression pattern, also circadian in nature, and peaked at ZT 8–12, although the peaks were broad and shallow. 18S rRNA and 25S rRNA had expressions manifolds higher than the remaining reference genes and they even displayed morning cyclic patterns of expression (ZT 4), although 18S rRNA appeared diurnal and 25S rRNA cycled in a circadian fashion.
Expression profiles of all the house-keeping genes analysed in this study normalized with the geometric mean of the best three stable house-keeping genes. Relative expression profiles of all the house-keeping genes were obtained after normalization with the geometric mean of the three best stable genes, eIF-4a, UBQ5 and eEF-1α, as determined by the four computer algorithms used in this study. The house-keeping genes have been categorized into three sub-groups; (A) genes that did not show cyclic expression patterns, viz. eIF-4a, UBQ5, eEF-1α and UBQ10; (B) genes that showed cyclic expression patterns, viz. UBC, ACT 11 and GAPDH, that displayed morning peaked circadian patterns of expression, and β-TUB, that exhibited evening peaked circadian expression pattern; and finally (C) genes that showed extremely high expression levels, viz. 18S rRNA and 25S rRNA. The error bars represent standard error.
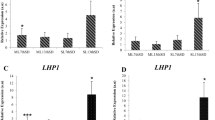
The reference genes, when studied individually, had significant effect on the expression curves of the five known diurnal/circadian genes (Fig. S2). OsLHY/CCA1, OsRVE1 and OsRVE8 are morning expressing genes, forming sharp peaks at ZT 0, ZT 22 and ZT 22, respectively. OsLHY/CCA1 and OsRVE1 show clear circadian expression patterns whereas OsRVE8 is a diurnal gene and its cycling gets disrupted in the free running conditions. OsFKF1 and OsTOC1 are evening expression circadian genes, showing peaks at ZT 8–10 and ZT 8–12, respectively. The cyclic expression profiles obtained were in accordance to the previously published microarray studies and are freely available in the public database www.ricearray.org (E-MEXP-2506 and E-MTAB-275). The minor differences, if any, may be due to small differences in the entrainment strategy. The 18S rRNA and 25S rRNA genes, due to their exceptionally high abundance levels diminished the cycling patterns of all the genes tested, including the robustly cycling OsLHY/CCA1, and thus should be grossly avoided for any diurnal/circadian RT-qPCR studies in rice. UBQ10 also had considerably dwarfed the peaks of the test genes that could be attributed to its relatively higher expression levels. UBC, on the other hand, due to its lowest expression level among all the house-keeping genes, showed highly amplified cycling patterns of the test genes, but was still a good reference gene. Rest all the other house-keeping genes generated good cyclic expression patterns of the test genes with moderate peak amplitudes and somewhat similar diurnal/circadian patterns, i.e. broadening, narrowing or shifting the peaks by a couple of hours. Similar were the results when the geometric means of the three most stable genes, i.e. eIF-4a/UBQ5/eEF-1α, and the three least stable ones, i.e. GAPDH/ACT11/UBQ10, were used for normalization of the test genes (Fig. 7). There were only minor differences in the peak amplitudes or the cycling patterns of the test genes, when either of the two geometric means was used.
Expression profiling of the five circadian genes using the geometric means of the three most stable and three least stable house-keeping genes for normalization. The same cDNA samples for all the 48 time points, that were used to study the house-keeping genes, were used to generate the expression profiles of the 5 known circadian genes, viz. LHY/CCA1, TOC1, FKF1, RVE1 and RVE8. The expression data of the circadian genes were normalized with the geometric mean of the three most stable house-keeping genes, (A) i.e. eIF-4a, eEF-1α and UBQ5, and the three least stable house-keeping genes, (B) i.e. GAPDH, ACT11 and UBQ10, identified by the gene stability analyses.
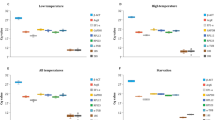
The real test of these house-keeping genes was when dealing with test genes showing no cyclic patterns or weak cyclic patterns because the choice of house-keeping genes can dramatically affect their expression patterns (Fig. 8). For this purpose, the house-keeping genes showing cyclic patterns, i.e. UBC, ACT11, GAPDH and β-TUB, were interchangeably used to normalize expression of non-cyclic genes, eIF-4a, UBQ5 and eEF-1α, and genes with weaker cyclic patterns, i.e. UBC, ACT11, GAPDH and β-TUB, all of them now treated as test genes. The morning expressing house-keeping genes, UBC, ACT11 and GAPDH, when used individually to normalize the RT-qPCR data of the remaining house-keeping genes, now treated as test genes, converted the non-cyclic genes eIF-4a, UBQ5 and eEF-1α to a cyclic pattern, circadian in nature, where they developed very broad but conspicuous expression peaks spanning mid-day, evening and mid-night and dropped sharply at dawn, the peak time for UBC, ACT11 and GAPDH. The circadian rhythms of the morning expressing genes (UBC, ACT11 and GAPDH) were disturbed dramatically as the peaks broadened, shifted or were completely diminished. The evening expressing gene β-TUB showed narrower evening peaks when normalized with UBC and ACT11, but developed an additional mid-night peak along with a narrower evening peak with GAPDH. When the evening expressing β-TUB was used for normalization, the non-cyclic eIF-4a, UBQ5 and eEF-1α genes once again showed circadian patterns of expression with broad peaks spanning evening, midnight and morning time-points. Their peaks dropped in the mid-day to evening transition phase, the peak time for β-TUB. The morning circadian peaks of UBC, ACT11 and GAPDH were not affected when normalized with β-TUB but their mid-day and evening expression levels were reduced making their morning peaks appear sharper and steeper.
Effect of cyclic house-keeping genes on the expression profiles of non-cyclic genes and cyclic genes with weak cyclic expression patterns. The averaged Ct. values of the four cyclic house-keeping genes, i.e. the morning expressing genes UBC (A), ACT11 (B) and GAPDH (C), and the evening expressing β-TUB (D) were used individually to normalize the relative expression data of rest of the house-keeping genes which were now treated as test genes. In all cases, the non-cyclic genes, viz. eIF-4a, UBQ5 and eEF-1α developed cyclic patterns of expression when their relative expression profiles were normalized against the Ct. values of the cyclic house-keeping genes. The expression patterns of the cyclic genes, viz. UBC, ACT 11, GAPDH and β-TUB were greatly disturbed when their relative expression profiles were normalized interchangeably with each other. The error bars represent standard error.
Discussion
The circadian clock controls the entire physiological and molecular functioning of plants, conferring them better adaptability in their ambient environment and enhanced vigor4. The clock functions by directing the expression of thousands of genes to peak at particular times in a 24-h day, when their products are needed for performing different physiological processes, thereby enabling better utilization of the plant’s energy and resources1,2. It is a routine practice to quantify gene expression levels throughout the day or even days together, when studying circadian clock functioning, which is an interplay of transcript abundance of thousands of genes in a 24 h period. RT-qPCR becomes the obvious choice for gene quantification in diurnal/circadian studies as it provides the opportunity of high-throughput and precise measurements in least possible time. Many genes have low expression levels and some genes show weak cyclic expression patterns, which tends to get easily maneuvered by selection of a not so suitable reference gene. This necessitates the need for standardization of some house-keeping genes for RT-qPCR analysis with diurnal/circadian samples, to facilitate reliable gene expression quantification. Several validation studies have been conducted in many plant species with different experimental samples where different house-keeping genes were deemed stable. Species to species variations are very common and, moreover, within a species, a gene which is stable for one set of samples may be unstable for another set of samples19,29. Keeping in mind that even the validated genes may show some variations across different samples, we began analyzing 10 house-keeping genes with a set of 48 diurnal and circadian samples, spread over four days. To the best of our knowledge, this work is the first of its kind detailed study of house-keeping genes for diurnal and circadian gene expression studies in plants. The genes chosen have already been studied previously with developmental, abiotic stress and hormonal treatment samples in rice by our group27. After confirming the primer efficiencies and specificities of the genes, the raw Ct values from the RT-qPCR data were averaged and analyzed for stability using four algorithms, viz. GeNORM, BestKeeper, NormFinder and ∆Ct. These four algorithms are based on different principles and independently calculate and rank the genes for their stability. eIF-4a, eEF-1α and UBQ5 were predicted to be the most stable genes by GeNORM; UBQ5, β-TUB, eIF-4a and eEF-1α by BestKeeper; UBC, eIF-4a, UBQ5 and eEF-1α according to NormFinder; and eIF-4a, 25S rRNA, UBQ5 and eEF-1α were the most stable as calculated by the ∆Ct method. On the other hand, GAPDH, ACT11 and 18S rRNA were the least stable according to GeNORM; GAPDH, 18S rRNA and ACT11 according to BestKeeper; 18S rRNA, 25S rRNA, ACT11 and GAPDH according to NormFinder; and GAPDH, 18S rRNA and ACT11 were the least stable according to ∆Ct method. These results suggest that eIF-4a, eEF-1α and UBQ5 are the most stable genes and GAPDH, 18S rRNA and ACT11 are the least stable, as predicted by all four programs. The minor variations observed in the gene rankings across all the four programs are obvious as they use different statistical approaches30,31,32,33. All the 10 genes, under study, showed good overall stability as their stability values were much below the default cut-off values set by the four programs. In the initial study by Jain et al.27 eEF-1α and UBQ5 were found to be the most stable while carrying out RT-qPCR studies with rice samples of various developmental stages and abiotic stress and hormone treated tissues, and UBQ10 was found to be the least stable, although ACT11 and β-TUB were also ranked low in terms of stability, which supports our results. The only validation study, so far, for diurnal samples in lettuce by Sgamma et al.34 however, showed GAPDH to be relatively stable but Actin genes had relatively lower stability values. This may be partly due to different set of reference genes taken for analysis. eEF-1α and eIF-4a have been found to be very stable in samples representing various developmental stages and those subjected to diverse treatments in numerous plant species, such as switchgrass, Brachiaria, Lolium, perennial ryegrass, flax, potato, chickpea, cucumber, Pyrus etc30,35,36,37,38,39,40,41,42. GAPDH showed good stability in coffee, flax, sugarcane and pigeonpea for developmental tissues and certain stress treated samples31,32,38,43,44, but was expressed highly in some treated tissues in case of Lolium36. To make things all the more complicated GAPDH exhibited inter-cultivar differences of several folds with the same samples in rice and Petunia45,46. Gimeno et al.30 found Actin and GAPDH genes to be less stable under stress conditions in switchgrass. Actin genes were found to be unstable under salinity stress in potato, Lolium and cucumber36,39,41, but were considered stable under drought conditions in barley, only at specific developmental stages, thereby adding another dimension to the complexity in selecting reference genes47. The deviations observed in the otherwise stable expression levels of the reference genes, under certain conditions, may be due to some specific roles these genes might be playing apart from their main house-keeping function. For example, GAPDH is also involved in tRNA binding and age specific apoptosis in animal system48,49, which is not its usual role in glycolysis.
A general notion has developed over the past few years that the use of a single reference gene for normalization of an RT-qPCR data can introduce errors in the results and thus may be unreliable. The use of geometric mean of two to three best stable reference genes was therefore suggested by Vandesompele et al.12 for normalizing the data, which is getting much acceptance, worldwide. Hence, pairwise variation study by GeNORM was performed which predicted the geometric mean of only the two best genes to be sufficient as reference but we suggest the geometric mean of the best three genes, i.e. eIF-4a, eEF-1α and UBQ5, to be taken as reference for all diurnal/circadian studies in rice as it was concluded by all the four programs to be the most stable. Similar conclusions were given by Sgamma et al.34 where the geometric mean of the three best genes were recommended for diurnal samples in lettuce.
The reference genes showing the least stability should be grossly avoided as they tend to change the cyclic expression patterns of low expressing genes. In order to validate the stability results and to find out if these genes were good candidates for use in normalizing diurnal/circadian RT-qPCR data, the cycling patterns of five well known diurnal/circadian genes, viz. OsLHY/CCA1, OsTOC1, OsFKF1, OsRVE1 and OsRVE8, were analyzed using, (1) each of the 10 reference genes alone and, (2) geometric mean of the 3 best stable and 3 least stable genes. The results indicated that all the reference genes were stable and did not cause any major shifts in the peaks of the test genes, viz. at ZT-0, ZT-22, ZT-22, ZT-8–10 and ZT-8–12 for OsLHY/CCA1, OsRVE1, OsRVE8, OsFKF1 and OsTOC1, respectively, except for minor broadening of peaks, although the peak amplitudes were severely affected by the choice of the reference genes. The 18S rRNA and 25S rRNA transcripts levels were exceptionally high with the Ct values reaching in the order of 9–11 and almost completely diminished the peak expression levels of even the known test genes with sharp peaks, when used as reference. The 18S rRNA and 25S rRNA genes are therefore not recommended to be used for reference purpose with diurnal/circadian samples. Similar results were obtained by Jain et al.27 and Sinha et al.31,32. Their use as reference genes has been viewed with skepticism as its expression has been found to show variations, not only in different samples across different species, but also inter-cultivar variations were found29. Kim et al.45 however, found 18S rRNA to be very stable for studying gene expression across various developmental stages in rice.
The geometric mean of the three most stable genes was used to study the expression patterns of the reference genes. eIF-4a, UBQ5 and eEF-1α did not exhibit any cyclic patterns but, to our surprise, GAPDH, ACT11 and UBC showed clear morning circadian rhythms whereas β-TUB showed evening circadian rhythms. A reference gene, which is itself undergoing cycling, cannot be used for normalizing diurnal/circadian RT-qPCR data. This became apparent when genes with no cyclic patterns or weak cyclic patterns were studied. When normalized with the morning peaked reference genes the non cyclic test genes developed midday to evening peaked cycling pattern and with evening peaked β-TUB, the test genes developed midnight to morning peaks. When the genes with weak cyclic patterns were studied with cyclic reference genes, the cycling patterns of the test genes were greatly distorted, to the extent of becoming arrhythmic in some cases. GAPDH and ACT11 had scored badly in the stability rankings, but UBC and β-TUB were in the middle or high-up in those rankings, still none of the programs could detect the unsuitability of the four genes for diurnal/circadian studies. This reinforces the notion that the choice of house-keeping genes is experiment dependent and cannot be deemed best or universal for all sample types and plant species.
Methods
Plant material, day-night entrainment and sample harvesting
Rice seeds (Oryza sativa L. ssp. Indica, var. PB1) were obtained from the Indian Agricultural Research Institute, New Delhi. The seeds were dehusked and surface sterilized by washing with 70% ethanol for 1 min followed by two washes of 10 min each, with 0.1% HgCl2, 1% bavistin solution to which were added a few drops of teepol (detergent), with vigorous shaking. The seeds were subsequently rinsed, several times, with sterile reverse osmosis water and left overnight for soaking. Next day, seeds were blotted dry on sterile filter paper and inoculated on ½ MS salts supplemented with 1% sucrose and 0.4% phytagel (Sigma) for gelling, in magenta boxes. These boxes were transferred to a growth chamber (Conviron, Canada) under a daily photoperiodic cycle of 14 h light and 10 h dark. The temperatures were set at 35 °C in the light phase and 25 °C in the dark phase. After one week, the healthy seedlings were washed under tap water and transferred to liquid rice growth medium50 in culture tubes. After another two weeks in the same entrainment conditions, the healthy plants were harvested, in two biological replicates, every two hours for four consecutive days, beginning from the onset of the light period. During the third and the fourth day, the plants were given a free-running treatment of continuous light and hot condition, beginning the dawn of the third day. The harvesting of the plants was continued every two hours. The details of the light and temperature conditions and the sampling layout from the first to the fourth day have been summarized in supplemental Table S3.
RNA isolation, quality checks and cDNA preparation
The total RNA was isolated independently from the two biological sets of samples, 48 samples in each set, using RNeasy Mini kit (Qiagen, USA) according to the manufacturer’s instructions. The isolated RNA was checked for its quality and quantity using Nanovue® spectrophotometer (GE Healthcare, UK). The samples had an A260/280 ratio between 1.9 and 2.1 and the A260/230 ratio between 2 and 2.3. The integrity of the RNA samples was also checked on a denaturing 1.5% agarose gel, which were then processed for cDNA synthesis. First-strand cDNA synthesis was done using the High Capacity cDNA Reverse Transcription kit (Applied Biosystems, USA) as per the manufacturer’s instructions, using 2 µg total RNA in a 40 µl reaction volume.
Selection of the candidate house-keeping genes, primer designing and RT-qPCR analysis
In one of the first reference genes validation reports in rice by Jain et al.27 some of the commonly used genes were analyzed in various categories, such as abiotic stresses, hormonal treatments and developmental stages. In that study, some genes and gene pairs were established as good house-keeping gene candidates for all three categories. With the aim of extending that research for diurnal and circadian studies, the same genes and primer sets were used in this study also without any modification, details of which have been given in Table 1; see also Jain et al.27. The RT-qPCR analysis was carried out using 2 × Roche SYBR Green I master mix on a Roche Light Cycler 480 II instrument (Roche, Switzerland). The 18 µl reaction mix consisted of 6.25 µl PCR grade water, 9 µl 2 × Roche SYBR Green I master mix, 1 µl cDNA and 1.75 µl of 5 µM dilution of forward and reverse primers. Three technical replicates (5 µl each) from the above 18 µl reaction were subjected to the recommended thermal cycling profile of initial denaturation of 10 min at 95 °C and 45 cycles of 10 s at 95 °C, 20 s at 60 °C and 20 s at 72 °C. The expression values were obtained in the form of Ct (cycle threshold) values in RT-qPCR analysis, which is broadly defined as the cycle number at which the fluorescence levels in an amplification reaction clearly crosses the background noise. The fluorescence threshold levels were automatically determined by software of the RT-qPCR instrument. These Ct values were processed further mathematically to get the relative expression values for a gene. The completion of each run was followed by a melting curve analysis which denotes the specificity of the amplification. As the temperature rises from 60 °C to 95 °C, with an increment of 0.5 °C each time, the fluorescence levels are lost as the amplicons in the reaction continue to denature at increasing temperatures and the temperature at which half of the fluorescence is lost is denoted by Tm. This Tm is specific for each species of template in the reaction mix and the presence of more than one peak at the end of melting curve analysis shows non-specific amplifications. The amplification efficiencies in the RT-qPCR reactions for each primer pairs were calculated by making serial 10-fold dilutions of the pooled cDNA. The amplification efficiency is the rate of DNA amplification in any PCR reaction, which theoretically should be 100% implicating exponential doubling of the template DNA per PCR cycle. The final amplification products were also resolved on a 2% agarose gel to check for their specificity.
Analysis of the house-keeping gene stability
For ranking the stability of the chosen house-keeping genes, the mean Ct (cycle threshold) values of the technical replicates from the two biological replicates were averaged, which were then directly imported to the four statistical algorithms, used in this study, that measures the stability value of the house-keeping genes, viz. GeNorm in the qBase Plus software version 2.5 (http://www.biogazelle.com/qbaseplus), NormFinder (http://www.moma.dk/normfinder-software), BestKeeper (http://www.gene-quantification.com/bestkeeper.html) and ∆Ct. The GeNorm programme also calculates the pairwise variation to identify the number of house-keeping genes that should be taken for analysis, for each experiment set.
Validation of the reference gene stability
For validating the reference genes stability results, the expression profiles of 5 well known circadian genes, viz. OsLHY/CCA1 (LOC_Os08g06110), OsTOC1 (LOC_Os02g40510), OsFKF1 (LOC_Os11g34460), OsRVE1 (LOC_Os02g46030) and OsRVE8 (LOC_Os02g45670) were investigated by performing RT-qPCR analysis of cDNA samples for all the 48 time points, in the same way as stated above. The expression values were normalized against each of the 10 selected reference genes, separately, as well as against the geometric mean of the three most stable and least stable reference genes. The primer details of these target circadian genes have been given in the Supplemental Table S2.
References
Michael, T. P. et al. Network discovery pipeline elucidates conserved time-of-day-specific cis-regulatory modules. PLoS Genet 4, e14, https://doi.org/10.1371/journal.pgen.0040014 (2008).
Filichkin, S. A. et al. Global profiling of rice and poplar transcriptomes highlights key conserved circadian-controlled pathways and cis-regulatory modules. PLoS One 6, e16907, https://doi.org/10.1371/journal.pone.0016907 (2011).
Nagel, D. H. & Kay, S. A. Complexity in the wiring and regulation of plant circadian networks. Curr Biol 22, R648–657, https://doi.org/10.1016/j.cub.2012.07.025 (2012).
Dodd, A. N. et al. Plant circadian clocks increase photosynthesis, growth, survival, and competitive advantage. Science 309, 630–633, https://doi.org/10.1126/science.1115581 (2005).
Ni, Z. et al. Altered circadian rhythms regulate growth vigour in hybrids and allopolyploids. Nature 457, 327–331, https://doi.org/10.1038/nature07523 (2009).
Covington, M. F., Maloof, J. N., Straume, M., Kay, S. A. & Harmer, S. L. Global transcriptome analysis reveals circadian regulation of key pathways in plant growth and development. Genome Biol 9, R130, https://doi.org/10.1186/gb-2008-9-8-r130 (2008).
Nozue, K. & Maloof, J. N. Diurnal regulation of plant growth. Plant Cell Environ 29, 396–408, https://doi.org/10.1111/j.1365-3040.2005.01489.x (2006).
Green, R. M., Tingay, S., Wang, Z. Y. & Tobin, E. M. Circadian rhythms confer a higher level of fitness to Arabidopsis plants. Plant Physiol 129, 576–584, https://doi.org/10.1104/pp.004374 (2002).
Gachon, C., Mingam, A. & Charrier, B. Real-time PCR: what relevance to plant studies? J Exp Bot 55, 1445–1454, https://doi.org/10.1093/jxb/erh181 (2004).
Morrison, T. et al. Nanoliter high throughput quantitative PCR. Nucleic Acids Res 34, e123, https://doi.org/10.1093/nar/gkl639 (2006).
Spurgeon, S. L., Jones, R. C. & Ramakrishnan, R. High throughput gene expression measurement with real time PCR in a microfluidic dynamic array. PLoS One 3, e1662, https://doi.org/10.1371/journal.pone.0001662 (2008).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3, research0034, https://doi.org/10.1186/gb-2002-3-7-research0034 (2002).
Bustin, S. A. et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55, 611–622, https://doi.org/10.1373/clinchem.2008.112797 (2009).
Udvardi, M. K., Czechowski, T. & Scheible, W. R. Eleven golden rules of quantitative RT-PCR. Plant Cell 20, 1736–1737, https://doi.org/10.1105/tpc.108.061143 (2008).
VanGuilder, H. D., Vrana, K. E. & Freeman, W. M. Twenty-five years of quantitative PCR for gene expression analysis. Biotechniques 44, 619–626, https://doi.org/10.2144/000112776 (2008).
Radonić, A. et al. Guideline to reference gene selection for quantitative real-time PCR. Biochem Biophys Res Commun 313, 856–862, https://doi.org/10.1016/j.bbrc.2003.11.177 (2004).
Schmittgen, T. D. & Zakrajsek, B. A. Effect of experimental treatment on housekeeping gene expression: validation by real-time, quantitative RT-PCR. J Biochem Biophys Methods 46, 69–81, https://doi.org/10.1016/S0165-022X(00)00129-9 (2000).
Greer, S., Honeywell, R., Geletu, M., Arulanandam, R. & Raptis, L. Housekeeping genes; expression levels may change with density of cultured cells. J Immunol Methods 355, 76–79, https://doi.org/10.1016/j.jim.2010.02.006 (2010).
Gutierrez, L. et al. The lack of a systematic validation of reference genes: a serious pitfall undervalued in reverse transcription-polymerase chain reaction (RT-PCR) analysis in plants. Plant Biotechnol J 6, 609–618, https://doi.org/10.1111/j.1467-7652.2008.00346.x (2008).
Hellemans, J., Mortier, G., De Paepe, A., Speleman, F. & Vandesompele, J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol 8, R19, https://doi.org/10.1186/gb-2007-8-2-r19 (2007).
Pfaffl, M. W., Tichopad, A., Prgomet, C. & Neuvians, T. P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper–Excel-based tool using pair-wise correlations. Biotechnol Lett 26, 509–515, https://doi.org/10.1023/B:BILE.0000019559.84305.47 (2004).
Andersen, C. L., Jensen, J. L. & Ørntoft, T. F. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64, 5245–5250, https://doi.org/10.1158/0008-5472.CAN-04-0496 (2004).
Silver, N., Best, S., Jiang, J. & Thein, S. L. Selection of housekeeping genes for gene expression studies in human reticulocytes using real-time PCR. BMC Mol Biol 7, 33, https://doi.org/10.1186/1471-2199-7-33 (2006).
Xie, F., Xiao, P., Chen, D., Xu, L. & Zhang, B. miRDeepFinder: a miRNA analysis tool for deep sequencing of plant small RNAs. Plant Mol Biol, https://doi.org/10.1007/s11103-012-9885-2 (2012).
Czechowski, T., Stitt, M., Altmann, T., Udvardi, M. K. & Scheible, W. R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol 139, 5–17, https://doi.org/10.1104/pp.105.063743 (2005).
Jain, M. Genome-wide identification of novel internal control genes for normalization of gene expression during various stages of development in rice. Plant Science 176, 702–706, https://doi.org/10.1016/j.plantsci.2009.02.001 (2009).
Jain, M., Nijhawan, A., Tyagi, A. K. & Khurana, J. P. Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR. Biochem Biophys Res Commun 345, 646–651, https://doi.org/10.1016/j.bbrc.2006.04.140 (2006).
Project, I. R. G. S. The map-based sequence of the rice genome. Nature 436, 793–800, https://doi.org/10.1038/nature03895 (2005).
Kozera, B. & Rapacz, M. Reference genes in real-time PCR. J Appl Genet 54, 391–406, https://doi.org/10.1007/s13353-013-0173-x (2013).
Gimeno, J., Eattock, N., Van Deynze, A. & Blumwald, E. Selection and validation of reference genes for gene expression analysis in switchgrass (Panicum virgatum) using quantitative real-time RT-PCR. PLoS One 9, e91474, https://doi.org/10.1371/journal.pone.0091474 (2014).
Sinha, P., Saxena, R. K., Singh, V. K., Krishnamurthy, L. & Varshney, R. K. Selection and Validation of Housekeeping Genes as Reference for Gene Expression Studies in Pigeonpea (Cajanus cajan) under Heat and Salt Stress Conditions. Front Plant Sci 6, 1071, https://doi.org/10.3389/fpls.2015.01071 (2015).
Sinha, P. et al. Evaluation and validation of housekeeping genes as reference for gene expression studies in pigeonpea (Cajanus cajan) under drought stress conditions. PLoS One 10, e0122847, https://doi.org/10.1371/journal.pone.0122847 (2015).
Yang, Q. et al. Reference gene selection for qRT-PCR in Caragana korshinskii Kom. under different stress conditions. Mol Biol Rep 41, 2325–2334, https://doi.org/10.1007/s11033-014-3086-9 (2014).
Sgamma, T., Pape, J., Massiah, A. & Jackson, S. Selection of reference genes for diurnal and developmental time-course real-time PCR expression analyses in lettuce. Plant Methods 12, 21, https://doi.org/10.1186/s13007-016-0121-y (2016).
Silveira, E. D., Alves-Ferreira, M., Guimarães, L. A., da Silva, F. R. & Carneiro, V. T. Selection of reference genes for quantitative real-time PCR expression studies in the apomictic and sexual grass Brachiaria brizantha. BMC Plant Biol 9, 84, https://doi.org/10.1186/1471-2229-9-84 (2009).
Dombrowski, J. E. & Martin, R. C. Evaluation of reference genes for quantitative RT-PCR in Lolium temulentum under abiotic stress. Plant Science 176, 390–396, https://doi.org/10.1016/j.plantsci.2008.12.005 (2009).
Lee, J. M., Roche, J. R., Donaghy, D. J., Thrush, A. & Sathish, P. Validation of reference genes for quantitative RT-PCR studies of gene expression in perennial ryegrass (Lolium perenne L.). BMC Mol Biol 11, 8, https://doi.org/10.1186/1471-2199-11-8 (2010).
Huis, R., Hawkins, S. & Neutelings, G. Selection of reference genes for quantitative gene expression normalization in flax (Linum usitatissimum L.). BMC Plant Biol 10, 71, https://doi.org/10.1186/1471-2229-10-71 (2010).
Nicot, N., Hausman, J. F., Hoffmann, L. & Evers, D. Housekeeping gene selection for real-time RT-PCR normalization in potato during biotic and abiotic stress. J Exp Bot 56, 2907–2914, https://doi.org/10.1093/jxb/eri285 (2005).
Garg, R., Sahoo, A., Tyagi, A. K. & Jain, M. Validation of internal control genes for quantitative gene expression studies in chickpea (Cicer arietinum L.). Biochem Biophys Res Commun 396, 283–288, https://doi.org/10.1016/j.bbrc.2010.04.079 (2010).
Wan, H. et al. Selection of appropriate reference genes for gene expression studies by quantitative real-time polymerase chain reaction in cucumber. Anal Biochem 399, 257–261, https://doi.org/10.1016/j.ab.2009.12.008 (2010).
Imai, T., Ubi, B. E., Saito, T. & Moriguchi, T. Evaluation of reference genes for accurate normalization of gene expression for real time-quantitative PCR in Pyrus pyrifolia using different tissue samples and seasonal conditions. PLoS One 9, e86492, https://doi.org/10.1371/journal.pone.0086492 (2014).
Barsalobres-Cavallari, C. F., Severino, F. E., Maluf, M. P. & Maia, I. G. Identification of suitable internal control genes for expression studies in Coffea arabica under different experimental conditions. BMC Mol Biol 10, 1, https://doi.org/10.1186/1471-2199-10-1 (2009).
Iskandar, H. M. et al. Comparison of reference genes for quantitative real-time polymerase chain reaction analysis of gene expression in sugarcane. Plant Molecular Biology Reporter 22, 325–337, https://doi.org/10.1007/BF02772676 (2004).
Kim, B. R., Nam, H. Y., Kim, S. U., Kim, S. I. & Chang, Y. J. Normalization of reverse transcription quantitative-PCR with housekeeping genes in rice. Biotechnol Lett 25, 1869–1872, https://doi.org/10.1023/A:1026298032009 (2003).
Mallona, I., Lischewski, S., Weiss, J., Hause, B. & Egea-Cortines, M. Validation of reference genes for quantitative real-time PCR during leaf and flower development in Petunia hybrida. BMC Plant Biol 10, 4, https://doi.org/10.1186/1471-2229-10-4 (2010).
Rapacz, M., Stępień, A. & Skorupa, K. Internal standards for quantitative RT-PCR studies of gene expression under drought treatment in barley (Hordeum vulgare L.): the effects of developmental stage and leaf age. Acta Physiologiae Plantarum 34, 1723–1733, https://doi.org/10.1007/s11738-012-0967-1 (2012).
Singh, R. & Green, M. R. Sequence-specific binding of transfer RNA by glyceraldehyde-3-phosphate dehydrogenase. Science 259, 365–368, https://doi.org/10.1126/science.8420004 (1993).
Ishitani, R. et al. Evidence that glyceraldehyde-3-phosphate dehydrogenase is involved in age-induced apoptosis in mature cerebellar neurons in culture. J Neurochem 66, 928–935, https://doi.org/10.1046/j.1471-4159.1996.66030928.x (1996).
Yoshida, S., Forno, D. A. Cock, J. H. & Gomez, K. A. Routine procedures for growing rice plants in culture solution in Laboratory manual for physiological studies of rice (eds Yoshida, S., Forno, D. A. Cock, J. H. & Gomez, K. A.) 61 (International Rice Research Institute, 1976).
Acknowledgements
This study was supported by a research grant received from the Department of Biotechnology, Government of India (BT/PR12394/AGIII/103 891/2014). The University Grants Commission, New Delhi, and the Department of Science and Technology, New Delhi, provided infrastructure support. NJ and SV acknowledge the award of Research Fellowships from the University Grants Commission and the Indian Council of Medical Research, New Delhi, respectively. We thank Dr. Mukesh Jain, Jawaharlal Nehru University, New Delhi, for useful suggestions. JPK also thanks SERB, New Delhi, for the financial support provided through the J.C. Bose Fellowship.
Author information
Authors and Affiliations
Contributions
N.J., S.V. and J.P.K. conceived and designed the experiments. N.J. and S.V. performed the experiments. N.J., S.V. and J.P.K. analyzed the data. N.J. and S.V. wrote the manuscript. J.P.K. supervised the study and critically revised the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jain, N., Vergish, S. & Khurana, J.P. Validation of house-keeping genes for normalization of gene expression data during diurnal/circadian studies in rice by RT-qPCR. Sci Rep 8, 3203 (2018). https://doi.org/10.1038/s41598-018-21374-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-21374-1
This article is cited by
-
Transgenic rice expressing Trichoderma spp. endochitinase gene exhibits moderate resistance to sheath blight
Plant Cell, Tissue and Organ Culture (PCTOC) (2023)
-
Un-biased housekeeping gene panel selection for high-validity gene expression analysis
Scientific Reports (2022)
-
Conversion of sheath blight susceptible indica and japonica rice cultivars into moderately resistant through expression of antifungal β-1,3-glucanase transgene from Trichoderma spp.
Transgenic Research (2022)
-
Stable reference gene selection for quantitative real-time PCR normalization in passion fruit (Passiflora edulis Sims.)
Molecular Biology Reports (2022)
-
An optimized protocol for stepwise optimization of real-time RT-PCR analysis
Horticulture Research (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.