Abstract
Coral reefs are significant ecosystems. The ecological success of coral reefs relies on not only coral-algal symbiosis but also coral-microbial partnership. However, microbiome assemblages in the South China Sea corals remain largely unexplored. Here, we compared the microbiome assemblages of reef-building corals Galaxea (G. fascicularis) and Montipora (M. venosa, M. peltiformis, M. monasteriata) collected from five different locations in the South China Sea using massively-parallel sequencing of 16S rRNA gene and multivariate analysis. The results indicated that microbiome assemblages for each coral species were unique regardless of location and were different from the corresponding seawater. Host type appeared to drive the coral microbiome assemblages rather than location and seawater. Network analysis was employed to explore coral microbiome co-occurrence patterns, which revealed 61 and 80 co-occurring microbial species assembling the Galaxea and Montipora microbiomes, respectively. Most of these co-occurring microbial species were commonly found in corals and were inferred to play potential roles in host nutrient metabolism; carbon, nitrogen, sulfur cycles; host detoxification; and climate change. These findings suggest that the co-occurring microbial species explored might be essential to maintain the critical coral-microbial partnership. The present study provides new insights into coral microbiome assemblages in the South China Sea.
Similar content being viewed by others
Introduction
Coral reef ecosystems are considered as the tropical rainforests of the sea, nurturing the highest biodiversity of marine life and providing vital ecosystem goods and services1,2. However, coral reefs around the world have suffered from declines and extinction risks largely due to bleaching events and emerging/reemerging diseases induced by climate change and anthropogenic disturbances3,4. The ecological success of coral reefs relies on coral-algal symbiosis, and recent studies using 16S rRNA gene amplicon pyrosequencing have revealed highly diverse and abundant microbes in individual coral colonies5,6,7,8,9. It is believed that some of these microbes can form partnerships with coral hosts and help them with possible access to those unavailable nutrients and metabolic pathways10. Compared with the well-understood coral-algal symbiosis, current understanding of coral-microbial partnership is rather limited. Although members of Endozoicomonas6 and Prosthecochloris11 have been shown to form potential symbioses with corals, no obligate coral-microbial symbiosis has been addressed to date. A recent advance in calcification for coral symbiotic algae demonstrated that free-living Symbiodinium in culture could form algal-microbial partnership, which facilitated the Symbiodinium calcification12. Taken these evidences, potentially complex coral-algal-microbial interactions in holobionts might facilitate the ecological success of coral reefs.
The coral mucus, tissue, and skeleton all contain large populations of associated microbes from three domains of life, including Eukarya, Archaea, and Bacteria, as well as many viruses13,14. The term coral microbiome is employed herein to collectively refer to the bacteria and archaea in coral holobionts. The characterization of coral core microbiomes reveals that two most abundant bacterial phylotypes affiliating to the genera Propionibacterium and Ralstonia are co-localized specifically with the host’s endosymbiotic algae which are likely to facilitate the success of coral-algal symbiosis10. Coral microbiomes are highly complex and dynamic, usually changing with environmental conditions, host types, and tempo-spatial gradients5,15. However, we assumed that there would be co-occurring microbial species assembling coral microbiomes which might be fundamental for coral holobionts to maintain the critical coral-microbial partnership. To explore these potential microbial species, co-occurrence network analysis was conducted in this study. Network-based approaches have been widely applied in microbial ecology studies16 such as soil17, coral18, human gut19, marine sediment20, stream biofilm21, activated sludge22, marine biofilm23, and other natural and man-made environments.
In recent years, there has been a growing interest in coral microbiome studies24,25,26,27,28, but very limited knowledge has been gained in understanding the microbiome assemblages for the South China Sea corals. We do not yet clearly understand how coral microbiomes are assembled in the South China Sea and potential co-occurring microbial species making up the coral microbiomes. The aims of this study were (i) to compare the microbiome assemblages using congeneric corals collected from different locations of the South China Sea, and (ii) to explore potential co-occurring microbial species through co-occurrence network analysis. We believe the present study facilitates our understanding of coral-microbial partnership in the South China Sea and serves as a useful reference for comparison with other regional coral microbiome assemblages.
Results
Microbiome assemblages and comparisons for Galaxea and Montipora at a high taxonomic level
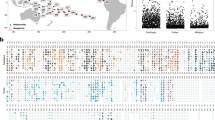
After filtering unqualified sequences, a total of 587,425 clean 16S tags were obtained for the 59 coral and seawater samples collected from the five locations LI, CB, LHT, SB, and DI (Table 1 and Fig. 1). Detailed information for each sample, including the sample ID, number of clean 16S tags, number of observed species, and Shannon index, is shown in Table S1. The three technical replicates for coral colony LHT-Mo4 exhibited similar microbiome assemblages (Figure S1). The seawater technical replicates (collected in parallel) for the five sampling locations also displayed similar profiles (Figure S1). Both coral and seawater technical replicates demonstrated the sequencing data had a good reproducibility. As shown in Fig. 2, coral microbiome assemblages displayed quite different profiles among host type, location, and seawater at the domain, phylum, or class level. Individual colony microbiome assemblage also displayed varied patterns, even from the same species and location (Figure S1). The phylum Proteobacteria dominated all of the coral and seawater microbiome assemblages with a relative abundance greater than 50%, among which Alphaproteobacteria or Gammaproteobacteria accounted for the largest proportion (Fig. 2). This finding is consistent with the results of other studies on coral-associated microbes7,8,18,29,30,31,32. Relative abundance of Deltaproteobacteria was much higher in corals than that in seawater (Fig. 2). Compared with Proteobacteria, the phyla Bacteroidetes, Cyanobacteria, Firmicutes, and Actinobacteria were less abundant (Fig. 2), but they were essential members assembling the coral microbiomes. The domain Archaea was detected in both coral and seawater samples with a low relative abundance (Fig. 2), but most of them were ecologically significant ammonia-oxidizing archaea Nitrosopumilus spp.33.
A geographic map of coral sampling locations in the South China Sea. The map showing dark gray, sandy, and blue indicates the land, shallow water, and deep water, respectively. The rectangles show the magnified areas of the red cycles. The red cycles and crosses indicate the sampling regions and locations, respectively. Galaxea and Montipora from three biogeographic regions in Hong Kong (Crescent Bay and Lamma Island, abbreviated as CB and LI), Sanya (Luhuitou and Sunny Bay, abbreviated as LHT and SB), and Sansha (Drummond Island, abbreviated as DI) were collected for this study. The map was plotted by Ocean Data View 4.7.2 (Schlitzer, R., Ocean Data View, odv.awi.de, 2017).
Coral microbiome assemblages at domain (only for Archaea), phylum, or class (only for Proteobacteria) level. Relative abundance for each taxon is plotted using a box chart statistically. The eight classified taxa Alphaproteobacteria, Gammaproteobacteria, Deltaproteobacteria, Archaea, Bacteroidetes, Cyanobacteria, Firmicutes, and Actinobacteria in Figure S1 were statistically analyzed. The detailed data are shown by stacked columns in Figure S1. CB, LI, LHT, SB, DI, Ga, Mo, and SW indicate Crescent Bay, Lamma Island, Luhuitou, Sunny Bay, Drummond Island, Galaxea, Montipora, and seawater, respectively.
Microbiome assemblages and comparisons for Galaxea and Montipora at a low taxonomic level
The recovered 97% OTU data were used for statistical analyses to examine the differences (PERMANOVA) and similarities (NMDS) in coral microbiome assemblages between host type, location, and seawater. One-way PERMANOVA revealed that the Galaxea microbiome assemblages were significantly different among locations CB, LHT, SB, and DI (P < 0.01, F = 4.18), and the Montipora microbiome assemblages were also significantly different among locations LI, CB, LHT, and SB (P < 0.01, F = 6.23). To further explore the differences in coral microbiome assemblages between host type, location, and seawater comprehensively, thirteen groups (i.e. LI-Mo, LI-SW, CB-Ga, CB-Mo, CB-SW, LHT-Ga, LHT-Mo, LHT-SW, SB-Ga, SB-Mo, SB-SW, DI-Ga, and DI-SW) were generated for pairwise comparisons using one-way PERMANOVA. As shown in Table S2, coral microbiome assemblages for each host type from each location was significantly different from any other host type and the corresponding seawater (P < 0.05, 1.96 < F < 22.44). To explore the similarities among these thirteen groups, NMDS (stress = 0.13) was conducted and shown in Fig. 3. Colonies for each coral group (i.e. biological replicates) and the three technical replicates for LHT-Mo4 exhibited different distances, revealing that individual differences for the same host type and location can be either very high or very low. The five seawater groups were closely clustered and distinct from all coral groups, which is consistent with the previous findings8,34. The eight coral groups were loosely clustered together with four Galaxea groups and four Montipora groups distributed at the top and bottom respectively, indicating that host type exhibited a consistent distribution pattern. However, there was no consistent geographic pattern in distribution for both Galaxea and Montipora groups, suggesting that location contributed less in driving the coral microbiome assemblages. According to the NMDS result, host type but not location showed a consistent distribution pattern for the coral microbiome assemblages and the coral and seawater microbiomes were assembled dissimilarly. Therefore host type is suggested to play an even more important role than location and seawater in driving coral microbiome assemblages.
NMDS ordination for 97% OTU data of all samples using Bray-Curtis distance. The similarities of microbiome assemblages at species level among host type, sampling locations, and corresponding seawater were compared appropriately. CB, LI, LHT, SB, DI, Ga, Mo, and SW indicate Crescent Bay, Lamma Island, Luhuitou, Sunny Bay, Drummond Island, Galaxea, Montipora, and seawater, respectively.
Co-occurrence patterns for Galaxea and Montipora microbiomes
In total, 122 and 134 OTUs derived from the Galaxea and Montipora microbiomes were used for co-occurrence network analysis after removal of the poorly represented OTUs, respectively. We first tested the two OTU datasets and observed non-random co-occurrence patterns, which is consistent with other studies17,19,20,22. A study even demonstrated that non-random community assemblage might be a common feature of all life domains35. As shown in Fig. 4, the generated networks had 61 and 80 nodes (i.e., 97% OTUs or microbial species) and 73 and 172 edges (i.e., links between every two nodes) for the Galaxea and Montipora microbiomes, respectively. The network topological parameters for the Galaxea microbiomes included an average node connectivity of 2.39, an average network distance of 2.18 edges, a network diameter of 6 edges, an average clustering coefficient of 0.37, and a modularity index of 0.73. While the network topological parameters for the Montipora microbiomes contained an average node connectivity of 4.30, an average network distance of 2.76 edges, a network diameter of 6 edges, an average clustering coefficient of 0.48, and a modularity index of 0.62. These topological properties were used to describe the general structures of co-occurrence networks17. Both networks of the Galaxea and Montipora microbiomes had modular structures because each modularity index value was higher than 0.4036. Based on the modularity class, each network could be further divided into several sub-networks, for example, those closely interacted nodes on the left side of each network (Fig. 4).
Co-occurrence patterns of microbial species assembling the Galaxea and Montipora microbiomes. Each connection indicates a strong and significant correlation, with the Spearman’s correlation coefficient higher than 0.6 and statistically significant (P < 0.01). Each node represents a microbial species, and its size is proportional to the node connectivity. Each edge represents a linkage between two co-occurring nodes, and its thickness is proportional to the Spearman’s correlation coefficients. All nodes are labeled with annotated species or unknown species, which are colored at the phylum level.
There were 114 OTUs in the co-occurrence networks of the Galaxea and Montipora microbiomes (Fig. 4), among which 27 OTUs were shared between the two corals (Figures S2 and S3). These OTUs were taxonomically annotated by BLAST online search against the GenBank 16S rRNA gene database using 94% similarity cutoff (Table S3). OTUs that exhibited less than 94% similarity or any ambiguous match to a specific genus were annotated as unknown species. There were a total of 78 annotated species and 36 unknown species (Figs 4 and S2), most of which were affiliated to Alphaproteobacteria and Gammaproteobacteria. To achieve a basic understanding of the unknown species, phylogenetic analysis was conducted using their closest relatives (Figure S4), and the result showed that 27 of the 36 unknown species were phylogenetically close to the previously reported coral-associated microbes, and the others were phylogenetically related to marine bacteria (Figure S4 and Table S3). Among the 78 annotated species, 64 were reported as coral-associated microbes, and the remainders were also known as marine bacteria (Table S3). Because the 114 OTUs derived from the co-occurrence networks were demonstrated to have statistically significant interactions, most of which were identified as coral-associated microbes in this and previous studies (see Table S3 for details). These OTUs are believed to serve as necessary co-occurring microbial species assembling the coral microbiomes.
Profiles of co-occurring microbial species of the Galaxea and Montipora microbiomes
As shown in Fig. 4, the 61 co-occurring microbial species making up the Galaxea microbiomes were assigned to Alphaproteobacteria (26), Betaproteobacteria (1), Gammaproteobacteria (23), Bacteroidetes (5), Cyanobacteria (3), Thaumarchaeota (2), and Euryarchaeota (1). While the 80 co-occurring microbial species assembling the Montipora microbiomes were classified into Alphaproteobacteria (39), Betaproteobacteria (2), Gammaproteobacteria (27), Bacteroidetes (1), Cyanobacteria (7), Thaumarchaeota (1), Nitrospirae (1), Chloroflexi (1), and Fusabacteria (1). The relative abundance of the 114 co-occurring microbial species and the corresponding seawater samples is shown using a heat map (Fig. 5). The 31 species in Fig. 5A showed similar profiles between coral and seawater samples, revealing that they might multiply well in both corals and seawater. While the 48 species in Fig. 5B exhibited higher relative abundance in corals than that in seawater, suggesting that they might benefit from coral hosts and proliferated more efficiently in corals than in seawater. The 35 species in Fig. 5C displayed unusual profiles between corals from different locations, indicating that some of them might be enriched in corals from certain location. Most of the 114 co-occurring microbial species are known as heterotrophs, which might utilize holobiont organic wastes for growth. The remainders are autotrophs, including photoautotrophs like cyanobacteria and chemoautotrophs such as ammonia-oxidizing archaea Nitrosopumilus and sulfur-oxidizing bacteria Thioalkalivibrio and Thioprofundum. Cluster analysis for the co-occurring microbial species explored were performed to reveal the similarities between corals from different locations and between corals and seawater (Figure S5). Corals from the same location clustered preferentially with individual colony differences and they were relatively distant to the seawater, which is similar to the NMDS result.
Profiles of co-occurring microbial species for the Galaxea and Montipora microbiomes. Relative abundance was log10-transformed for plotting. The scale bars ND, −2, −1, 0, and 1 indicate relative abundance of 0, 0.01%, 0.1%, 1%, and 10%, respectively. The bottom panel represents the sample IDs. The right panel shows the IDs of the co-occurring microbial species. CB, LI, LHT, SB, DI, Ga, Mo, SW, and TR indicate Crescent Bay, Lamma Island, Luhuitou, Sunny Bay, Drummond Island, Galaxea, Montipora, seawater, and technical replicate, respectively.
There were 27 shared co-occurring microbial species assembling the Galaxea and Montipora microbiomes (Figures S2 and S3). As shown in Figs 4 and S3, they belonged to Alphaproteobacteria (11), Betaproteobacteria (1), Gammaproteobacteria (13), Cyanobacteria (1), and Thaumarchaeota (1). These shared co-occurring microbial species included 7 unknown spp., 6 Acinetobacter spp., 3 Endozoicomonas spp., and a species from each of the following 11 genera: Brevundimonas, Brucella, Escherichia, Filomicrobium, Methyloceanibacter, Methylophilus, Nitrosopumilus, Pelagibius, Roseibium, Thalassomonas, and Thioprofundum. These shared co-occurring microbial species assembling the Galaxea and Montipora microbiomes might be prevalent in other corals from the South China Sea.
Discussion
In recent years, the concept of “core microbiome” has been introduced in coral microbial ecology studies10,26,37, while the counterpart “stable microbiome” has also been used in related studies26,37,38. According to the statistical methods used for data analysis, coral-associated microbes can be assigned into “core/stable microbiome” and “transient/sporadic microbiome”37. The “core microbiome” is described as the stable and consistent components across complex microbiome assemblages from similar habitats which can be determined with one of the five described variations, that is, a core based on shared presence, shared abundance, shared composition, phylogenetic information, or interaction37,39. Co-occurring microbial species explored here through co-occurrence network analysis are exactly equal to the “core microbiome” based on interaction, as illustrated in related studies37,39. In the present study, we collected coral samples with a large set of biological replicates from the five locations in the South China Sea, allowing us to compare the complex microbiome assemblages comprehensively and to explore co-occurrence patterns through network analysis. We further discussed the effects of host type and location in coral microbiome assemblages (the first section below) and potential roles of co-occurring microbial species explored including those functional microbes (the second section below) and those mostly found microbes (the third section below).
Effects of host type and location in coral microbiome assemblages
Corals harbor highly diverse microbes that might be assembled similarly or dissimilarly among host types, locations and other potential determinants. The study on Caribbean corals shows that different hosts harbor distinct host-specific microbes and that specificity varies by host type and location15. The study on Red Sea corals reveals that the microbiome assemblages vary largely with location and are shaped by host type5. Certain microbes are reported to form species-specific associations with corals40, suggesting the roles of host played in driving coral microbiome assemblages. The study on corals from the Great Barrier Reef shows that certain microbes are associated with corals specifically and that microbiome assemblages differ with the location but not the host type41. These findings support the present study that host type rather than location drives the coral microbiome assemblages, as shown in Fig. 3. There is a limitation that three species from the genus Montipora were collected for the present study because the shared species were not found in the selected locations except for Galaxea fascicularis. The study using three species of the genus Acropora found that they harbored conserved microbes and suggested that closely related corals of the same genus were assembled with similar microbes41. So it can be assumed that if one of the three studied Montipora species were shared and used for the present study, the general distribution pattern of microbiome assemblages for this species from different locations might be less likely affected in the NMDS ordination. The distribution pattern of the three Montipora species shown in Fig. 3 further supports that host type rather than location drives the coral microbiome assemblages because LHT-Mo and SB-Mo from the same host species Montipora monasteriata were clustered closely and were relatively distinct from LI-Mo and CB-Mo (from Montipora venosa and Montipora peltiformis respectively).
Potential roles of certain functional co-occurring microbial species
To achieve a better understanding of co-occurring microbial species, potential ecological roles and associations with coral hosts were further explored. As shown in Fig. 6, the listed co-occurring microbial species might be involved in several important biological and ecological processes: nutrient metabolism; carbon, nitrogen, and sulfur cycles; host detoxification; and climate change. Potential nitrogen-fixing bacteria such as Brevundimonas spp. and Phyllobacterium spp. might fix atmospheric nitrogen gas to ammonia. Potential ammonia-oxidizing archaea like Nitrosopumilus spp. might oxidize the holobiont ammonia to nitrite, and possible nitrite-oxidizing bacteria like Nitrospira sp. might oxidize the holobiont nitrite to nitrate. Finally, potential denitrifying bacteria such as Methylophilus sp. and Nitratireductor sp. might convert the holobiont nitrate to nitrogen gas. These co-occurring microbial species might thus be involved in the complete nitrogen cycle and changes in their population and activity might affect certain process of the nitrogen metabolism in coral holobionts42. It is suggested that they might serve as nitrogen regulators to keep the balance of holobiont nitrogen through providing sufficient bioavailable nitrogen and removing unneeded nitrogen. Carbon dioxide might be fixed into carbohydrates by Symbiodinium and Cyanobacteria through photosynthesis and by the co-occurring microbial species explored such as Nitrosopumilus spp., Thioalkalivibrio sp., and Thioprofundum spp. through chemosynthesis. Like Symbiodinium, the synthesized carbohydrates by the co-occurring microbial species might also serve as the host nutrients or food sources. The study has demonstrated that free-living Symbiodinium can calcify with the aid of microbial partners12, implying that potential co-occurring microbial species explored might also facilitate coral calcification directly or indirectly. Hydrogen sulfide is toxic to a wide range of eukaryotic organisms, including marine invertebrates like coral and sponge, by inhibiting cytochrome c oxidase and a type of catalase43,44. In addition, hydrogen sulfide generated by sulfate-reducing bacteria might lead to the initiation of coral black band disease, which has been consistently found in lab induction and field observation45,46. Co-occurring microbial species Thioalkalivibrio sp. and Thioprofundum spp., serving as potential sulfur-oxidizing bacteria, might oxidize holobiont accumulated hydrogen sulfide to sulfate, thus contributing to coral health through detoxification of reduced sulfur compounds. Both DMSP (dimethylsulfoniopropionate) and DMS (dimethylsulfide) are important compounds in the global sulfur cycle as they are closely related to cloud formation and global climate change47. Both reef-building corals and free-living Symbiodinium have been documented to be high producers of DMSP48,49. The generated DMSP might be degraded into climate-active gas DMS via the bacterial cleavage pathway by co-occurring microbial species explored such as Hoeflea sp., Loktanella sp., Phaeobacter sp., Roseovarius sp., Shewanella sp., and Vibrio sp.47. In summary, co-occurring microbial species explored might play potential roles in host nutrient metabolism; carbon, nitrogen, sulfur cycles; host detoxification; and climate change and might be essential to maintain the critical coral-microbial partnership. However, the real functions need to be tested in future studies. For example, microbial genome recovery using culture-independent methods such as genome binning and single-cell genomics is promising to validate the specific functions on a genome scale.
A schematic representation illustrating potential roles of certain co-occurring microbial species in corals. Potential roles might be played in host nutrient metabolism; carbon, nitrogen, sulfur cycles; host detoxification; and climate change. Those co-occurring microbial species shown in red were mapped onto the well-known ecological processes.
Potential roles of the mostly found co-occurring microbial species Acinetobacter spp. and Endozoicomonas spp
Among the 114 co-occurring microbial species assembling the Galaxea and Montipora microbiomes, 16 Acinetobacter spp. and 10 Endozoicomonas spp. were found, of which 6 Acinetobacter spp. and 3 Endozoicomonas spp. were shared between the two corals. This finding corroborates the high occurrence of Acinetobacter spp. and Endozoicomonas spp. in coral microbiomes found in different coral species and regions (Table S3). However, the roles of Endozoicomonas spp. and Acinetobacter spp. are poorly understood to date. One study demonstrated that Acinetobacter spp. were dominant in the microbiomes of bleached corals but were not detected in healthy corals50. Acinetobacter spp. are not likely to serve as bleaching causing agents because they can also be detected in high abundance in healthy corals, as observed in the present and the other studies7,8,15,34. It is important to note, however, that Acinetobacter baumannii is a highly troublesome human pathogen worldwide due to its resistance to antibiotics51. This implies that potential Acinetobacter sp. might be a threat to coral hosts. An early study demonstrated that Acinetobacter guillouiae strain 20B is a DMS oxidizer52, suggesting that Acinetobacter spp. might participate in holobiont DMS metabolism. However, more and more recent studies have demonstrated that Acinetobacter spp. have the denitrification function, for example, Acinetobacter baumannii H153, Acinetobacter johnonii DBP-354, Acinetobacter sp. HA255, Acinetobacter sp. YZS-X1–156, and Acinetobacter sp. SZ2857.
Besides the present study, Endozoicomonas spp. have been reported to be dominant in the microbiome assemblages of multiple coral species such as Porites astreoides15, Stylophora pistillata6, Acropora millepora31, Seriatopora hystrix32, and Coelastrea aspera58. Several Endozoicomonas species have been isolated from different marine invertebrates including a sponge59, a sea anemone60, a sea slug61, a comb pen shell62, and an octocoral63, and a stony coral64. Endozoicomonas spp. might play certain roles in their hosts because they are associated with a broad range of marine invertebrates. The roles of Endozoicomonas spp. in corals have been inferred in several studies31,32,65. Here PICRUSt prediction using 16S sequences assigned to Endozoicomonas spp. revealed an unreported function that they have a complete denitrification pathway including genes napAB, nirK, norBC, and nosZ responsible for converting nitrate to nitrite, nitrite to nitric oxide, nitric oxide to nitrous oxide, and nitrous oxide to nitrogen, respectively. However, further experiments are needed to elucidate their real roles in corals. In addition, Endozoicomonas spp. might be developed as coral health indicators because they are distributed extensively in corals and other marine invertebrates and no negative effects on corals have been reported to date.
Conclusions
In the present study, we examined the coral microbiome assemblages associated with Galaxea and Montipora collected from five locations in three biogeographic regions of the South China Sea. The highly dominant Proteobacteria and less abundant Bacteroidetes, Cyanobacteria, Firmicutes, and Actinobacteria are the main phyla in structuring the coral microbiomes. OTU-based multivariate statistical analysis demonstrated that each host species from each location was significantly different from any other host species and the corresponding seawater in the coral microbiome assemblages. Host type appeared to drive the coral microbiome assemblages rather than location and seawater. The network analysis explored a set of co-occurring microbial species assembling the coral microbiomes, which might play potential roles in host nutrient metabolism; carbon, nitrogen, sulfur cycles; host detoxification; and climate change. The findings of this study form a baseline for assessing coral microbiome development in the South China Sea, and for microbiome assemblage comparison across regions. The present study extends our knowledge of coral microbiome assemblages in the South China Sea and facilitates our understanding of coral-microbial partnership.
Methods
Sampling locations and corals
Three cities Hong Kong, Sanya, and Sansha, representing three different biogeographic regions of the South China Sea for hard coral growth, were selected for the present study (Fig. 1). Hong Kong’s climate is subtropical with seasonal sea surface temperature (SST) ranging from 14 °C in winter to 31 °C in summer66, which is a marginal environment for the development of coral communities. Hard corals of Hong Kong grow very slowly, and are unable to form real reefs. In contrast to Hong Kong, both Sanya and Sansha are tropical climates with a mean SST of 22.5 °C and 23.8 °C in winter and 30.0 °C and 29.8 °C in summer, respectively67. Sansha has a tropical climate for coral development, while Sanya is located in a transitional zone between tropical and subtropical climates. As shown in Fig. 1, two sampling locations were selected for Hong Kong (Lamma Island and Crescent Bay, abbreviated as LI and CB) and Sanya (Luhuitou and Sunny Bay, abbreviated as LHT and SB), but only one for Sansha (Drummond Island, abbreviated as DI) because of its simple and original environment. Lamma Island, close to the Pearl River, is a tough estuarine environment for coral survival. Crescent Bay, located in the Mirs Bay region, is a normal oceanic environment for coral growth. The coral cover of Luhuitou has significantly decreased for decades due to the frequent human activities and pollution, while Sunny Bay corals are well protected by the Sanya Coral Reef National Nature Reserve68. Galaxea and Montipora were used as the target corals of the present study because they are commonly distributed in Indo-Pacific region and they are also found in most of the selected sampling locations (Table 1). These conspecific and congeneric corals were thus collected from the three selected biogeographic regions, and in two of these regions with two locations of contrasting environmental conditions, which were used to explore the coral microbiome assemblages in the South China Sea, potential co-occurring microbial species and their roles in coral holobionts.
Coral collection
Information for each sampling location including location names, coordinates, sampling dates, coral species, and abbreviations is shown in Table 1. At each sampling, a small piece of coral tissue (~1 cm × 1 cm) from an apparently healthy colony was collected using a hammer & chisel and packaged into a tagged sterile bag. A total of six colonies representing biological replicates were collected for each coral species at each sampling location. After sampling, the coral specimens were immediately washed using sterile seawater to remove loosely attached microbes and fixed in 70% ethanol. All of the fixed coral specimens were kept with dry ice in an icebox. Seawater surrounding the sampled coral colonies was also collected in parallel for comparison with the corresponding coral samples. The microbes within 1 L seawater were concentrated by filtering the water through a 0.22-μm polycarbonate membrane using Pall Gelman 6-Place Aluminum Vacuum Filter Funnel Manifold #15403 and fixed with 50% ethanol. After fixation, all of the samples were stored at −30 °C until used for DNA extraction.
DNA extraction, PCR, and high-throughput sequencing
A small fraction (~0.5 cm × 0.5 cm) of each coral specimen fixed in ethanol was first rinsed with 1× PBSE (137 mM NaCl, 2.7 mM KCl, 4.3 mM Na2HPO4·7H2O, 1.4 mM KH2PO4 and 10 mM EDTA) and ground in 1× PBSE using a mortar & pestle. The resulting coral slurry was centrifuged at 12,000 g for 5 min to collect all pellets for total DNA extraction using the FastDNA® Spin Kit for Soil (MP Biomedicals, France). The DNA extracts were used as PCR templates after quality and purity determination through agarose gel electrophoresis and OD260/OD280 ratio measurement. To amplify and sequence the hypervariable V3 and V4 regions of the 16S rRNA gene, a universal primer set, 341 F (5′-CCTAYGGGRBGCASCAG-3′) and 802 R (5′-TACNVGGGTATCTAATCC-3′), was used for PCR amplification69,70. Unique barcodes of six nucleotides were modified at the 5′ terminus of the forward primer 341 F to allow multiplexed sequencing71. PCR was conducted using the following program: initial denaturation at 94 °C for 5 min, followed by 30 cycles at 94 °C for 0.5 min, 50 °C for 0.5 min, 72 °C for 1 min, and a final extension at 72 °C for 5 min. Each PCR reaction was pooled with ~50 ng DNA template, 200 nM of each primer, 25 μL 2× Premix Ex Taq solution (TaKaRa, China), and ddH2O up to 50 μL. To minimize potential PCR bias, three independent reactions were made for each sample and the products were pooled for purification using the PureLink® PCR Purification Kit (Invitrogen, USA). The purified PCR products were quantified using a Thermo NanoDrop 2000 UV-Vis Spectrophotometer and pooled at equivalent concentrations for subsequent multiplexed sequencing of 16S amplicons. The pooled DNA sample was sent to Novogene (Beijing, China) for sequencing on an Illumina MiSeq sequencer with a paired-end (PE) mode and a read length of 300 bp. Because the length of the 16S V3V4 fragment is approximately 450 bp, paired-end 300 bp sequencing is sufficient to form overlaps for full-length amplicon assembly72,73. The sequencing datasets have been deposited in the NCBI Sequence Read Archive under accession number SRP066229. Totally, there were 59 samples sequenced for the five locations LI, CB, LHT, SB, and DI (Table 1 and Fig. 1): 6 samples of Galaxea for each location of CB, LHT, SB, and DI; 4–6 samples of Montipora for locations of LI (6), CB (5), LHT (4), and SB (5); 2 additional Montipora technical replicates for the location of LHT, and 2–3 technical replicates of seawater samples for locations of LI (2), CB (2), LHT (3), SB (3), and DI (3). There were four Montipora samples failed in the DNA extraction or PCR amplification, that is, each one from the locations of CB and SB and two from the location of LHT.
Data trimming and bioinformatics analysis
The raw data were first trimmed to remove sequencing adaptors, short reads (<300 bp), and low quality reads (average quality score <20). To obtain the full-length of 16S V3V4 sequences, the PEAR (paired-end read merger) tool74 was employed to merge the overlapping PE reads into 16S tags using default parameters. The QIIME (quantitative insights into microbial ecology) platform75 was used to demultiplex 16S tags into individual samples identified by unique barcodes. Potential PCR chimeras were detected and removed using the ChimeraSlayer76 command in the QIIME. After strict data trimming, the clean 16S tags were recovered and normalized for the downstream analysis. OTUs (operational taxonomic units) at 97% similarity (equal to the species level) were clustered and annotated using the QIIME pipeline under default settings. Relative abundance data for community structures were automatically generated from phylum to genus. A 97% OTU matrix was recovered from the generated OTU table of biom file format, and the data were used for species-level permutational multivariate analysis of variance (PERMANOVA), non-metric multidimensional scaling (NMDS), and network analysis. Those poorly represented OTUs (mean relative abundance lower than 0.1%) were removed before the network analysis as previously described17. Briefly, microbial species-level co-occurrence networks were conducted in the R77 environment using the vegan78, igraph79, and Hmisc80 packages. Positive co-occurrence events were considered if the OTUs demonstrated the Spearman’s correlation coefficient higher than 0.6 and statistically significant (P < 0.01)17,21. The constructed networks were further optimized by the interactive visualization platform Gephi81. The R software was also employed for the heat map plotting to visualize profiles of the co-occurring microbial species in each sample. MEGA 6.0682 was used to construct the phylogenetic tree of 16S rRNA gene. PAST 3.1083 was used for PERMANOVA, NMDS and cluster analysis based on the Bray-Curtis distance. OriginPro 9.0 was employed for interactive scientific graphing. PICRUSt (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States) prediction was conducted following the established pipeline84.
References
Moberg, F. & Folke, C. Ecological goods and services of coral reef ecosystems. Ecol. Econ. 29, 215–233, https://doi.org/10.1016/S0921-8009(99)00009-9 (1999).
Polidoro, B. & Carpenter, K. Dynamics of coral reef recovery. Science 340, 34–35, https://doi.org/10.1126/science.1236833 (2013).
Hughes, T. P. et al. Climate change, human impacts, and the resilience of coral reefs. Science 301, 929–933, https://doi.org/10.1126/science.1085046 (2003).
Carpenter, K. E. et al. One-third of reef-building corals face elevated extinction risk from climate change and local impacts. Science 321, 560–563, https://doi.org/10.1126/science.1159196 (2008).
Lee, O. O. et al. Spatial and species variations in bacterial communities associated with corals from the Red Sea as revealed by pyrosequencing. Appl. Environ. Microbiol. 78, 7173–7184, https://doi.org/10.1128/AEM.01111-12 (2012).
Bayer, T. et al. The microbiome of the Red Sea coral Stylophora pistillata is dominated by tissue-associated Endozoicomonas Bacteria. Appl. Environ. Microbiol. 79, 4759–4762, https://doi.org/10.1128/Aem.00695-13 (2013).
Carlos, C., Torres, T. T. & Ottoboni, L. M. Bacterial communities and species-specific associations with the mucus of Brazilian coral species. Sci. Rep. 3, 1624, https://doi.org/10.1038/srep01624 (2013).
Li, J. et al. Bacterial dynamics within the mucus, tissue and skeleton of the coral Porites lutea during different seasons. Sci. Rep. 4, 7320, https://doi.org/10.1038/srep07320 (2014).
Fernando, S. C. et al. Microbiota of the major South Atlantic reef building coral Mussismilia. Microb. Ecol. 69, 267–280, https://doi.org/10.1007/s00248-014-0474-6 (2015).
Ainsworth, T. D. et al. The coral core microbiome identifies rare bacterial taxa as ubiquitous endosymbionts. ISME J. 9, 2261–2274, https://doi.org/10.1038/ismej.2015.39 (2015).
Cai, L. et al. Metagenomic analysis reveals a green sulfur bacterium as a potential coral symbiont. Sci. Rep. 7, 9320, https://doi.org/10.1038/s41598-017-09032-4 (2017).
Frommlet, J. C. et al. Coral symbiotic algae calcify ex hospite in partnership with bacteria. P. Natl. Acad. Sci. USA 112, 6158–6163, https://doi.org/10.1073/pnas.1420991112 (2015).
Rosenberg, E., Koren, O., Reshef, L., Efrony, R. & Zilber-Rosenberg, I. The role of microorganisms in coral health, disease and evolution. Nat. Rev. Microbiol. 5, 355–362, https://doi.org/10.1038/nrmicro1635 (2007).
Sweet, M. J., Croquer, A. & Bythell, J. C. Bacterial assemblages differ between compartments within the coral holobiont. Coral Reefs 30, 39–52, https://doi.org/10.1007/s00338-010-0695-1 (2011).
Morrow, K. M., Moss, A. G., Chadwick, N. E. & Liles, M. R. Bacterial associates of two Caribbean coral species reveal species-specific distribution and geographic variability. Appl. Environ. Microbiol. 78, 6438–6449, https://doi.org/10.1128/AEM.01162-12 (2012).
Faust, K. & Raes, J. Microbial interactions: from networks to models. Nat. Rev. Microbiol. 10, 538–550, https://doi.org/10.1038/nrmicro2832 (2012).
Barberan, A., Bates, S. T., Casamayor, E. O. & Fierer, N. Using network analysis to explore co-occurrence patterns in soil microbial communities. ISME J. 6, 343–351, https://doi.org/10.1038/ismej.2011.119 (2012).
Rodriguez-Lanetty, M., Granados-Cifuentes, C., Barberan, A., Bellantuono, A. J. & Bastidas, C. Ecological inferences from a deep screening of the complex bacterial consortia associated with the coral Porites astreoides. Mol. Ecol. 22, 4349–4362, https://doi.org/10.1111/mec.12392 (2013).
Zhang, Z. G. et al. Spatial heterogeneity and co-occurrence patterns of human mucosal-associated intestinal microbiota. ISME J. 8, 881–893, https://doi.org/10.1038/ismej.2013.185 (2014).
Muller, A. L. et al. Endospores of thermophilic bacteria as tracers of microbial dispersal by ocean currents. ISME J. 8, 1153–1165, https://doi.org/10.1038/ismej.2013.225 (2014).
Widder, S. et al. Fluvial network organization imprints on microbial co-occurrence networks. P. Natl. Acad. Sci. USA 111, 12799–12804, https://doi.org/10.1073/pnas.1411723111 (2014).
Ju, F., Xia, Y., Guo, F., Wang, Z. P. & Zhang, T. Taxonomic relatedness shapes bacterial assembly in activated sludge of globally distributed wastewater treatment plants. Environ. Microbiol. 16, 2421–2432, https://doi.org/10.1111/1462-2920.12355 (2014).
Lawes, J. C., Neilan, B. A., Brown, M. V., Clark, G. F. & Johnston, E. L. Elevated nutrients change bacterial community composition and connectivity: high throughput sequencing of young marine biofilms. Biofouling 32, 57–69, https://doi.org/10.1080/08927014.2015.1126581 (2016).
Hernandez-Agreda, A., Leggat, W., Bongaerts, P. & Ainsworth, T. D. The microbial signature provides insight into the mechanistic basis of coral success across reef habitats. mB io 7, e00560-16, https://doi.org/10.1128/mBio.00560-16 (2016).
Glasl, B., Herndl, G. J. & Frade, P. R. The microbiome of coral surface mucus has a key role in mediating holobiont health and survival upon disturbance. ISME J. 10, 2280–2292, https://doi.org/10.1038/ismej.2016.9 (2016).
Sweet, M. J., Brown, B. E., Dunne, R. P., Singleton, I. & Bulling, M. Evidence for rapid, tide-related shifts in the microbiome of the coral Coelastrea aspera. Coral Reefs 36, 815–828, https://doi.org/10.1007/s00338-017-1572-y (2017).
Bourne, D. G. et al. Coral reef invertebrate microbiomes correlate with the presence of photosymbionts. ISME J. 7, 1452–1458, https://doi.org/10.1038/ismej.2012.172 (2013).
Welsh, R. M. et al. Bacterial predation in a marine host-associated microbiome. ISME J. 10, 1540–1544, https://doi.org/10.1038/ismej.2015.219 (2016).
Kimes, N. E. et al. The Montastraea faveolata microbiome: ecological and temporal influences on a Caribbean reef-building coral in decline. Environ. Microbiol. 15, 2082–2094, https://doi.org/10.1111/1462-2920.12130 (2013).
Lema, K. A., Bourne, D. G. & Willis, B. L. Onset and establishment of diazotrophs and other bacterial associates in the early life history stages of the coral Acropora millepora. Mol. Ecol. 23, 4682–4695, https://doi.org/10.1111/mec.12899 (2014).
Lema, K. A., Willis, B. L. & Bourne, D. G. Amplicon pyrosequencing reveals spatial and temporal consistency in diazotroph assemblages of the Acropora millepora microbiome. Environ. Microbiol. 16, 3345–3359, https://doi.org/10.1111/1462-2920.12366 (2014).
Pantos, O., Bongaerts, P., Dennis, P. G., Tyson, G. W. & Hoegh-Guldberg, O. Habitat-specific environmental conditions primarily control the microbiomes of the coral Seriatopora hystrix. ISME J. 9, 1916–1927, https://doi.org/10.1038/ismej.2015.3 (2015).
Stahl, D. A. & de la Torre, J. R. Physiology and diversity of ammonia-oxidizing archaea. Annu. Rev. Microbiol. 66, 83–101, https://doi.org/10.1146/annurev-micro-092611-150128 (2012).
Chen, C. P., Tseng, C. H., Chen, C. A. & Tang, S. L. The dynamics of microbial partnerships in the coral Isopora palifera. ISME J. 5, 728–740, https://doi.org/10.1038/ismej.2010.151 (2011).
Horner-Devine, M. C. et al. A comparison of taxon co-occurrence patterns for macro- and microorganisms. Ecology 88, 1345–1353, https://doi.org/10.1890/06-0286 (2007).
Newman, M. E. J. Modularity and community structure in networks. P. Natl. Acad. Sci. USA 103, 8577–8582, https://doi.org/10.1073/pnas.0601602103 (2006).
Sweet, M. J. & Bulling, M. T. On the importance of the microbiome and pathobiome in coral health and disease. Front. Mar. Sci. 4, 9, https://doi.org/10.3389/fmars.2017.00009 (2017).
Hester, E. R., Barott, K. L., Nulton, J., Vermeij, M. J. & Rohwer, F. L. Stable and sporadic symbiotic communities of coral and algal holobionts. ISME J. 10, 1157–1169, https://doi.org/10.1038/ismej.2015.190 (2016).
Shade, A. & Handelsman, J. Beyond the Venn diagram: the hunt for a core microbiome. Environ. Microbiol. 14, 4–12, https://doi.org/10.1111/j.1462-2920.2011.02585.x (2012).
Rohwer, F., Seguritan, V., Azam, F. & Knowlton, N. Diversity and distribution of coral-associated bacteria. Mar. Ecol. Prog. Ser. 243, 1–10, https://doi.org/10.3354/meps243001 (2002).
Littman, R. A., Willis, B. L., Pfeffer, C. & Bourne, D. G. Diversities of coral-associated bacteria differ with location, but not species, for three acroporid corals on the Great Barrier Reef. FEMS Microbiol. Ecol. 68, 152–163, https://doi.org/10.1111/j.1574-6941.2009.00666.x (2009).
Rädecker, N., Pogoreutz, C., Voolstra, C. R., Wiedenmann, J. & Wild, C. Nitrogen cycling in corals: the key to understanding holobiont functioning? Trends Microbiol. 23, 490–497 (2015).
Oeschger, R. & Vetter, R. D. Sulfide detoxification and tolerance in Halicryptus spinulosus (Priapulida): a multiple strategy. Mar. Ecol. Prog. Ser. 86, 167–179 (1992).
Downs, C. A., Fauth, J. E., Downs, V. D. & Ostrander, G. K. In vitro cell-toxicity screening as an alternative animal model for coral toxicology: effects of heat stress, sulfide, rotenone, cyanide, and cuprous oxide on cell viability and mitochondrial function. Ecotoxicology 19, 171–184, https://doi.org/10.1007/s10646-009-0403-5 (2010).
Richardson, L. L. et al. Sulfide, microcystin, and the etiology of black band disease. Dis. Aquat. Organ. 87, 79–90, https://doi.org/10.3354/dao02083 (2009).
Miller, A. W. & Richardson, L. L. Fine structure analysis of black band disease (BBD) infected coral and coral exposed to the BBD toxins microcystin and sulfide. J. Invertebr. Pathol. 109, 27–33, https://doi.org/10.1016/j.jip.2011.09.007 (2012).
Raina, J. B., Dinsdale, E. A., Willis, B. L. & Bourne, D. G. Do the organic sulfur compounds DMSP and DMS drive coral microbial associations? Trends Microbiol. 18, 101–108, https://doi.org/10.1016/j.tim.2009.12.002 (2010).
Broadbent, A. D., Jones, G. B. & Jones, R. J. DMSP in corals and benthic algae from the Great Barrier Reef. Estuar. Coast. Shelf S. 55, 547–555, https://doi.org/10.1006/ecss.2002.1021 (2002).
Raina, J. B. et al. DMSP biosynthesis by an animal and its role in coral thermal stress response. Nature 502, 677–680, https://doi.org/10.1038/nature12677 (2013).
Koren, O. & Rosenberg, E. Bacteria associated with the bleached and cave coral Oculina patagonica. Microb. Ecol. 55, 523–529, https://doi.org/10.1007/s00248-007-9297-z (2008).
Peleg, A. Y., Seifert, H. & Paterson, D. L. Acinetobacter baumannii: emergence of a successful pathogen. Clin. Microbiol. Rev. 21, 538–582, https://doi.org/10.1128/CMR.00058-07 (2008).
Horinouchi, M., Kasuga, K., Nojiri, H., Yamane, H. & Omori, T. Cloning and characterization of genes encoding an enzyme which oxidizes dimethyl sulfide in Acinetobacter sp. strain 20B. FEMS Microbiol. Lett. 155, 99–105, https://doi.org/10.1111/j.1574-6968.1997.tb12692.x (1997).
Cao, H. P. et al. Isolation and characterization of a denitrifying Acinetobacter baumannii H1 using NO2 −-N as nitrogen source from shrimp farming ponds. Afr. J. Microbiol. Res. 6, 2258–2264, https://doi.org/10.5897/AJMR11.814 (2012).
Li, M. T., Liu, J. H., Zhao, S. J., Wang, Z. X. & Hao, L. L. The characteristics of nitrate removal by the psychrotolerant denitrifying bacterium Acinetobacter johnonii DBP-3, isolated from a low-temperature eutrophic body of water. J. Environ. Sci. Heal. B. 48, 885–892, https://doi.org/10.1080/03601234.2013.796836 (2013).
Yao, S., Ni, J. R., Ma, T. & Li, C. Heterotrophic nitrification and aerobic denitrification at low temperature by a newly isolated bacterium, Acinetobacter sp. HA2. Bioresour. Technol. 139, 80–86, https://doi.org/10.1016/j.biortech.2013.03.189 (2013).
Zhang, H., Li, X., Zhang, B. & Liu, C. Draft genome sequence of Acinetobacter sp. strain YZS-X1-1, a denitrifying bacterium isolated from freshwater pond sludge in China. Genome Announc. 3, e01579–01514, https://doi.org/10.1128/genomeA.01579-14 (2015).
Su, J. et al. Characterization of the anaerobic denitrification bacterium Acinetobacter sp. SZ28 and its application for groundwater treatment. Bioresour. Technol. 192, 654–659, https://doi.org/10.1016/j.biortech.2015.06.020 (2015).
Williams, A. D., Brown, B. E., Putchim, L. & Sweet, M. J. Age-related shifts in bacterial diversity in a reef coral. PLoS ONE 10, https://doi.org/10.1371/journal.pone.0144902 (2015).
Nishijima, M., Adachi, K., Katsuta, A., Shizuri, Y. & Yamasato, K. Endozoicomonas numazuensis sp. nov., a gammaproteobacterium isolated from marine sponges, and emended description of the genus Endozoicomonas Kurahashi and Yokota 2007. Int. J. Syst. Evol. Microbiol. 63, 709–714, https://doi.org/10.1099/ijs.0.042077-0 (2013).
Du, Z. J., Zhang, W. Y., Xia, H. J., Lu, G. Q. & Chen, G. J. Isolation and diversity analysis of heterotrophic bacteria associated with sea anemones. Acta Oceanol. Sin. 29, 62–69, https://doi.org/10.1007/s13131-010-0023-1 (2010).
Kurahashi, M. & Yokota, A. Endozoicomonas elysicola gen. nov., sp. nov., a gamma-proteobacterium isolated from the sea slug Elysia ornata. Syst. Appl. Microbiol. 30, 202–206, https://doi.org/10.1016/j.syapm.2006.07.003 (2007).
Hyun, D. W. et al. Endozoicomonas atrinae sp. nov., isolated from the intestine of a comb pen shell Atrina pectinata. Int. J. Syst. Evol. Microbiol. 64, 2312–2318, https://doi.org/10.1099/ijs.0.060780-0 (2014).
Pike, R. E., Haltli, B. & Kerr, R. G. Description of Endozoicomonas euniceicola sp. nov. and Endozoicomonas gorgoniicola sp. nov., bacteria isolated from the octocorals Eunicea fusca and Plexaura sp., and an emended description of the genus. Endozoicomonas. Int. J. Syst. Evol. Microbiol. 63, 4294–4302, https://doi.org/10.1099/ijs.0.051490-0 (2013).
Yang, C. S. et al. Endozoicomonas montiporae sp. nov., isolated from the encrusting pore coral Montipora aequituberculata. Int. J. Syst. Evol. Microbiol. 60, 1158–1162, https://doi.org/10.1099/ijs.0.014357-0 (2010).
Morrow, K. M. et al. Natural volcanic CO2 seeps reveal future trajectories for host-microbial associations in corals and sponges. ISME J. 9, 894–908, https://doi.org/10.1038/ismej.2014.188 (2015).
Goodkin, N. F. et al. Coral communities of Hong Kong: long-lived corals in a marginal reef environment. Mar. Ecol. Prog. Ser. 426, 185–196, https://doi.org/10.3354/meps09019 (2011).
Liu, Y. et al. The decline of winter monsoon velocity in the South China Sea through the 20th century: Evidence from the Sr/Ca records in corals. Global Planet. Change 63, 79–85, https://doi.org/10.1016/j.gloplacha.2008.05.003 (2008).
Li, X. B. et al. Spatial and temporal variations in sediment accumulation and their impacts on coral communities in the Sanya Coral Reef Reserve, Hainan, China. Deep-Sea Res. Pt. II 96, 88–96, https://doi.org/10.1016/j.dsr2.2013.04.015 (2013).
Behrendt, L. et al. Endolithic chlorophyll d-containing phototrophs. ISME J. 5, 1072–1076, https://doi.org/10.1038/ismej.2010.195 (2011).
Cai, L., Ye, L., Tong, A. H., Lok, S. & Zhang, T. Biased diversity metrics revealed by bacterial 16S pyrotags derived from different primer sets. PLoS ONE 8, e53649, https://doi.org/10.1371/journal.pone.0053649 (2013).
Bartram, A. K., Lynch, M. D., Stearns, J. C., Moreno-Hagelsieb, G. & Neufeld, J. D. Generation of multimillion-sequence 16S rRNA gene libraries from complex microbial communities by assembling paired-end illumina reads. Appl. Environ. Microbiol. 77, 3846–3852, https://doi.org/10.1128/AEM.02772-10 (2011).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 79, 5112–5120, https://doi.org/10.1128/AEM.01043-13 (2013).
Fadrosh, D. W. et al. An improved dual-indexing approach for multiplexed 16S rRNA gene sequencing on the Illumina MiSeq platform. Microbiome 2, 6, https://doi.org/10.1186/2049-2618-2-6 (2014).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620, https://doi.org/10.1093/bioinformatics/btt593 (2014).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336, https://doi.org/10.1038/nmeth.f.303 (2010).
Haas, B. J. et al. Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res. 21, 494–504, https://doi.org/10.1101/gr.112730.110 (2011).
Ihaka, R. & Gentleman, R. R: A language for data analysis and graphics. J. Comp. Graph. Stat. 5, 299–314, https://doi.org/10.2307/1390807 (1996).
Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 14, 927–930, https://doi.org/10.1111/j.1654-1103.2003.tb02228.x (2003).
Csardi, G. & Nepusz, T. The igraph software package for complex network research. InterJ. Complex Syst. 1695 (2006).
Harrell, F. E. Hmisc: harrell miscellaneous. R package version (2014).
Bastian, M., Heymann, S. & Jacomy, M. Gephi: an open source software for exploring and manipulating networks. International AAAI conference on weblogs and social media: San Jose, California (2009).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729, https://doi.org/10.1093/molbev/mst197 (2013).
Hammer, Ø., Harper, D. A. T. & Ryan, P. D. PAST: paleontological statistics software package for education and data analysis. Palaeontologia Electronica 4, 9pp (2001).
Langille, M. G. I. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 31, 814–821, https://doi.org/10.1038/nbt.2676 (2013).
Acknowledgements
This study was supported by the NSFC-Guangdong Joint Fund (U1301232).
Author information
Authors and Affiliations
Contributions
H.H. and P.Y.Q. designed this study. L.C., R.M.T., G.Z., H.T., Y.H.W., W.Z., A.P.Y.C., J.Y.X., J.W.Q., P.O.A., and S.L. performed the research and data analysis. L.C. drafted the paper. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cai, L., Tian, RM., Zhou, G. et al. Exploring coral microbiome assemblages in the South China Sea. Sci Rep 8, 2428 (2018). https://doi.org/10.1038/s41598-018-20515-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-20515-w
This article is cited by
-
A genome-centric view of the role of the Acropora kenti microbiome in coral health and resilience
Nature Communications (2024)
-
The microbiomes of five temperate soft corals declining in the Sea of Marmara
Marine Biodiversity (2024)
-
Algae-coral symbiosis: fragility owing to anthropogenic activities and adaptive response to changing climatic trends
Environment, Development and Sustainability (2024)
-
Genomic exploration of coral-associated bacteria: identifying probiotic candidates to increase coral bleaching resilience in Galaxea fascicularis
Microbiome (2023)
-
Deficiency of exchange protein directly activated by cAMP (EPAC)-1 in mice augments glucose intolerance, inflammation, and gut dysbiosis associated with Western diet
Microbiome (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.