Abstract
Jumonji (JmjC) domain proteins are known regulators of gene expression and chromatin organization by way of histone demethylation. Chromatin modification and remodeling provides a means to modulate the activity of large numbers of genes, but the importance of this class of predicted histone-modifying enzymes for different aspects of post-developmental processes remains poorly understood. Here we test the function of all 11 non-lethal members in the regulation of circadian rhythms and sleep. We find loss of every Drosophila JmjC gene affects different aspects of circadian behavior and sleep in a specific manner. Together these findings suggest that the majority of JmjC proteins function as regulators of behavior, rather than controlling essential developmental programs.
Similar content being viewed by others
Introduction
All living things, ranging from bacteria to eukaryotes, possess an innate need to respond to environmental conditions that undergo predictable daily oscillations such as light and temperature1,2. The circadian clock confers this ability by allowing animals to anticipate these changes and adapt appropriately by adjusting metabolism3, hormone secretion4 and immunity5. In particular, sleep is an essential biological process whose timing is regulated by the circadian clock6,7,8,9,10. Circadian regulation of these processes requires parallel, rhythmic fine-tuning of thousands of genes11,12,13 and this coordinated regulation results in an optimal synchronization of physiological processes and behaviors to the time of day. Disruption of circadian rhythms contributes to common metabolic disorders such as obesity and diabetes, and other disorders such as cardiovascular diseases, anxiety, sleep disorders and cancer14,15,16,17,18.
Transcriptional control at the level of chromatin is emerging as an important mechanism in the regulation of sleep and circadian rhythms19,20,21,22,23. Several studies have shown that histone modifications at specific sites throughout the genome oscillate during the circadian cycle23,24,25,26, and mutations in chromatin remodeling enzymes such as Brahma or the histone methyltransferase MLL1 alter circadian rhythms27,28,29. The Jumonji (JmjC) protein family includes histone demethylases that act to remove methyl marks off of specific lysine residues on histone proteins, which has a direct impact on chromatin organization and gene expression programs30,31. By targeting methyl marks on histone proteins, different JmjC proteins can positively or negatively influence transcription and are expected to serve as key regulators of gene expression in a broad number of contexts. How these family members influence different biological processes remains under investigation.
We hypothesized that fine-tuning of chromatin organization and gene expression programs may critically regulate post-developmental processes such as behavior. To systematically test this model, we took advantage of an established a toolkit, which includes molecularly defined null alleles and tagged transgenic lines for every JmjC family member encoded by the Drosophila genome32. We tested these JmjC mutants in two reproducibly quantifiable behaviors: sleep and circadian rhythms. All 11 mutants tested displayed significant and some distinct changes in circadian rhythms and/or sleep/wake activity. These data indicate that most members of the Drosophila JmjC gene family act to fine-tune behavior, many of them in a non-overlapping manner.
Materials and Methods
Fly Stocks
All flies were maintained on a standard cornmeal/molasses diet at 25 °C and 75% humidity on a 12-hour day and 12-hour night cycle. The following lines were acquired from the Bloomington Stock Center: lid10424 (BL# 12367) and lidk06801 (BL# 10403). PSRFM1 was provided by Kristin White (Massachusetts General Hospital, Charlestown, MA). The generation and verification of the knockout lines and tagged transgenes is described in Shalaby et al.32. Flies were outcrossed in groups of >20 for at least 5 generations to the w* Berlin genetic background for behavioral analyses.
Immunohistochemistry, microscopy and image processing
Adult brains were dissected in 1 × PBS and fixed for 30 min in 4% formaldehyde, washed 3 times, 10 min each, in PBT (1 × PBS, 0.3% Triton-X-100, 0.5% BSA), and incubated in primary antibody diluted in PBT overnight at 4 °C. Next day, samples were washed 3 times, 10 min each, in PBT and incubated in secondary antibodies diluted in PBT at RT for 4 hrs in the dark. Samples were then washed twice in PBT and once in 1 × PBS. TOTO3 was used to stain nuclei. Brains were mounted on a slide with a drop of Vectashield. The following antibodies were used: rat anti-HA 3F10 (Roche, 1:100), Fluorescence-conjugated secondary antibody Alexa488 (Molecular Probes) was used at 1:200. Images were taken using Leica SP8, processed and quantified using Amira 5.3 (Indeed, Berlin, Germany) and Adobe Photoshop CS6.
Behavioral analysis
Sleep and circadian rhythms were assayed as described before33. In brief, 2–7 day old flies were entrained for at least 3 days to a 12 hr light: 12 hr dark regimen (LD), and Drosophila Activity Monitors (DAM; TriKinetics, Waltham, MA) were used, for circadian rhythm and sleep/activity studies filled with standard food (see above), and for starvation assays filled with 0.7% agar. For sleep/activity studies, we collected data in 2 min bins for 3 days in 12:12 LD cycles, and analyzed them with a custom-written Excel spreadsheet for total sleep time in light and dark, with sleep defined as 6 minutes of uninterrupted inactivity. For circadian rhythms, flies were monitored for 6 days in complete darkness (DD). The data was collected in 30 min bins, and analyzed for rhythm length and strength (peak height above Chi square) using the FaasX software34. To determine starvation-induced hyperactivity35, which suppresses sleep36, flies were placed in DAM on 0.7% agar to provide water. The total cumulative starvation-induced hyperactivity was determined using a custom-written Excel spreadsheet and was defined as the longest continuous activity bout without any interruption by sleep. This normally occurred just prior to the flies’ death (data not shown and35). For the primary screen, and for Fig. 1, we assayed each mutant in groups of 16, and repeated the assay at least one time on different days (median n = 31, quartiles: 26–41). Because of its semi-lethality, lid10424/lidk06801 transheterozygous mutants had the lowest n of 16 for some assays. For the follow-up behavioral experiments (Figs 2–4), repeated on separate days, we assayed n = 32–64 mutant and rescue flies. The tables in Fig. 5B and the Supplemental Information was derived from combined primary screen and follow up data.
Circadian rhythm phenotypes of JmjC genes. (A) KDM2KO and JMJD5KO flies showed mildly reduced period lengths (estimated effect sizes compared to wild type at −0.74 ± 0.39 95% confidence interval for KDM2KO, and −0.40 ± 0.33 for JMJD5KO respectively). Note, these effect sizes were estimated assuming normal distributions, which these data were not. Thus, all bar graphs shown in this and following Figures are medians with 95% confidence interval error bars. (Kruskal-Wallis test for multiple comparisons with Dunn’s post hoc adjustment to detect significant differences). (B) KDM2KO and JMJD5KO flies also showed increased rhythm power compared to w Berlin control flies. (*p < 0.05, **p < 0.01, ***p < 0.001; effect sizes = 0.89 ± 0.40 for KDM2KO, 0.77 ± 0.34 for JMJD5KO).
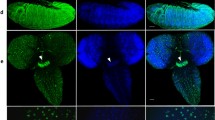
Sleep (and activity) phenotypes of JMJD5KO and JMJD7KO flies. (A,B) JMJD5KO showed moderately reduced daytime sleep (effect size = −0.68 ± 0.40) concomitant with (B) strongly increased daytime activity (effect size = 0.97 ± 40), while nighttime levels of sleep and activity were unaffected (n > 28). In this, and the next two Figures, data from w Berlin controls are in grey. The daytime phenotypes are rescued to wild-type levels for sleep, and beyond wild type for activity. (C) The JMJD5::HA rescue construct (right two brain hemispheres) was expressed in neurons close to the dorsal lateral neurons (arrow). Brains were stained with anti-HA (white) and a negative control is shown at the left. (D) JMJD7KO strongly reduced daytime sleep (effect size = −1.45 ± 0.33 confidence interval), which was partially rescued towards wild-type by expression of the JMJD7::HA transgene (n > 61). The mutant also mildly affected nighttime sleep (effect size = 0.55 ± 0.30). (E) The JMJD7KO mutants also displayed a moderate nighttime activity loss phenotype (effect size = −0.78 ± 0.31). (F) The expression pattern (green = anti-HA) of JMJD7::HA genomic rescue construct is shown (red = nuclei). Arrowheads point to the pars intercerebralis and arrows point to the fan shaped body, magnified in inset.
NO66KO hyperactivity phenotype. (A) NO66KO flies showed a reduction in sleep, strong for the daytime (effect size = 1.10 ± 0.40) and moderate for the night (effect size = 0.66 ± 0.39). This was completely rescued to wild type by expression of the NO66::HA genomic transgene (n > 31). (B,C) Flies also showed a very strong increase in activity, shown for one example fly (B; individual flies shown in this, and the next Figure have values within half a standard deviation from the mean of that genotypic cohort) and the whole cohort. (C) The effect size was 1.69 ± 0.43 for daytime activity, and 1.15 ± 0.40 for the night. These phenotypes were also completely rescued by NO66::HA expression. (D) The hyperactivity phenotype was in part driven by a strong increase in the activity per waking hour (effect size = 1.10 ± 0.40 for daytime waking activity, and 0.66 ± 0.39 at night). The phenotype in daytime waking activity was rescued by NO66::HA expression, but rescue flies were no different from mutants or wild type for nighttime waking activity. (E–G) The hyperactivity phenotype was also evident in the total starvation activity, which was very strongly enhanced (effect size = 1.81 ± 0.35) and partly rescued towards wild type (n > 47). (E,F) show a single fly example, and (G) the whole cohort. (H) This increased hyperactivity does not cause premature death; NO66KO flies even showed a mild delay in starvation-induced death (effect size = 0.41 ± 0.32), which was also rescued by the NO66::HA transgene (n > 31). (I) The expression pattern (green = anti-HA) of NO66::HA genomic rescue construct is shown (red = nuclei). Arrowheads point to the pars intercerebralis and arrows point to the fan shaped body, magnified in inset.
KDM4BKO sleep phenotype. (A–H) Shown are mutant (light blue) and rescue (dark) medians with 95% confidence interval. Averages of double-plotted daily sleep of one representative fly (per genotype, A) and the two cohorts (B) show a strong increase in sleep time in KDM4BKO (effect size = 1.89 ± 0.46 for daytime sleep, effect size = 0.78 ± 0.42 for night sleep). Average daily activity of one fly each (C) and the two cohorts (D) indicates a concomitant strong reduction in activity (effect size = −1.79 ± 0.45 for daytime activity, effect size = −0.82 ± 0.42 for night sleep). The daytime sleep and activity phenotypes were rescued to wild type by expressing KDM4B::HA, while nighttime sleep and activity were rescued beyond wild-type values. (E) The waking activity was not affected in KDM4BKO mutants compared to wild type (effect sizes = −0.14 ± 0.41 day, −0.41 ± 0.42 night). Individual fly actograms (F,G) show starvation-induced hyperactivity on non-nutritious agar in rescue flies (G), but not in KDM4BKO mutants (F). As a group (H), KDM4BKO flies showed absence of starvation hyperactivity (effect size = −1.22 ± 0.39) but no effect on death (I). Note that in numerous assays shown here, KDM4B::HA rescue flies showed opposite phenotypes from KDM4BKO mutants (as in B,D,H,I; n >31). (J) The expression pattern of KDM4B::HA genomic rescue construct is shown in (green = anti-HA, red = nuclei) and includes the pars intercerebralis (arrowheads) and fan shaped body (arrows).
Summary of circadian behavior and sleep phenotypes of JmjCKO mutants. (A) Circadian rhythm, and sleep phenotype strengths of JmjCKO strains. Phenotype strength (Herge’s g effect size) was determined assuming normal distributions and incorporating data from the primary screen and from follow-ups (Figs 2–4). The phenotype strength is shown as the z-score and percentile in an idealized wild-type distribution (color coded, as in Fig. 5B). Note that the means and standard deviations were calculated for each measure in a large w Berlin population (see Supplemental Table 1), but they are idealized, as some measures were not normally distributed (eg. w Berlin total starvation activity). The point is to illustrate the strength of each JmjC mutant’s strongest phenotype, and that few phenotypes are outside the 10th, or 90th percentiles, corresponding to an effect size of >1.3. This is a reflection of i) the lack of “strong” phenotypes, and ii) the considerable variability of the wild-type behavior. (B) Phenotype table indicating the behaviors of JmjC mutants significantly different from wild type (*p < 0.05, **p < 0.01, ***p < 0.001; Kruskal-Wallis test with Dunn’s post hoc multiple comparison). The strength of those phenotypes is color coded (according to the percentile within the wild-type distribution; see A). Hierarchical clustering dendrograms of these scores are depicted on the left, grouping JmjCKO mutants, and at the bottom, grouping phenotypic measures. (A/min stands for waking activity per minute; tau for the circadian period length; Pwr. for the power of the rhythm; AR/R for the frequency of arrhythmic flies; TSA for total starvation-induced hyperactivity; and TOD for time of death starting from removal from food).
Statistics
Data were analyzed in Prism (GraphPad Software, La Jolla, CA). Data from arrhythmicity determinations were dualistic by definition (rhythmic, or not), and were compared by Fisher’s exact test with Bonferroni correction. Half of the period length measures were not normally distributed, while 94/108 other measures passed normality tests (D’Agostino & Pearson normaility test with Bonferroni correction). To achieve highest specificity of our results, we compared all parametric measures by Kruskal-Wallis test with Dunn’s post hoc multiple comparison correction, which does not assume normal distribution. We also analyzed the effect size of the mutants’ phenotypes. Since most distributions were normal, we assumed normal distribution for these effect size estimates, chiefly, in order to determine 95% confidence intervals (for which there is no consensus definition with non-normal data). We determined effect sizes with an Excel spreadsheet (Durham University, UK; http://www.cem.org/effect-size-calculator) that determines Herge’s g, a bias-corrected form of Cohen’s d, which is essentially the z-score in a normal distribution (Fig. 5; where d = 1 means the mutant is one standard deviation away from the wild-type mean), as well as the 95% confidence intervals. Classically, an effect size of >0.5 is considered medium, and >0.8 large. In our experience with behavioral mutants, we consider an effect size of >0.8 a moderate phenotype, >1.3 a strong one, >1.65 a very strong one, and >2.03 an exceptionally strong one; these cutoffs correspond to standing at the following percentiles in a wild-type normal distribution: 20/80, 10/90, 5/95, 2/98. Note that because we can easily assay many individual flies, we have great statistical power (assuming normal distribution). For example, we found 5 genotypes with significantly changed waking activity at night, but only one of those was even of moderate effect size ( > 0.8). With an n of 64, which is easily obtained using just 2 DAM monitors, we have the power to detect effect sizes of 0.5 (alpha = 0.05, beta = 0.8). This explains why we found many significant phenotypes, but considerably fewer strong ones (9/46). Hierarchical clustering was done using the Ward method (www.wessa.net).
ERG Recordings
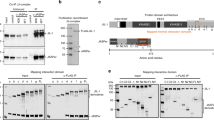
ERGs were performed as described in37 with the following modifications: Flies were fixed using Elmer’s non-toxic Glue-All. 2 M NaCl was used in the recording and reference electrodes. Electrode voltage was amplified by a Digidata 1440 A, filtered through a Warner IE-210, and recorded using Clampex 10.1 by Axon Instruments. Light stimulus was provided in 1 sec pulses by a computer-controlled white LED system (Schott MC1500). For quantification of depolarization and ‘on’ transients all experiments were carried out in triplicate with at least 10 recordings for each genotype.
Results
Several JmjC mutants differentially modulate circadian rhythms
A recently established collection of viable JmjC knockouts in Drosophila, generated using ends-out homologous recombination using an eyeless-driven RFP cassette that replaces the open reading frame to create a molecularly-defined null allele32, allowed us to test the effect of JmjC-dependent modulation on organism function and behavior in intact animals. Because of the known importance of cyclic transcription in circadian rhythms, we first assayed JmjC mutants’ circadian rhythms (the ability of organisms to anticipate and respond to predictable daily oscillations in light and temperature) and sleep/wake activity. First, we examined circadian rhythms in constant darkness to assess endogenous circadian period lengths. Of the 11 viable JmjC mutants, seven showed no significant changes, while two exhibited high levels of arrhythmicity (lid and KDM3KO), and two others exhibited a subtle shortening of the circadian period length (KDM2KO and JMJD5KO; Fig. 1A). The latter two mutants also displayed a significant increase in the rhythm power (strength of the rhythm; Fig. 1B).
KDM3KO mutants showed a high incidence of arrhythmic flies (20 of 78). We were able to rescue the high incidence of arrhythmia (to 7 out of 75 flies; p = 0.01, Fisher Exact test) by expressing a tagged genomic KDM3::HA transgene generated as part of the JmjC toolkit32. Anti-HA staining revealed KDM3::HA expression in numerous nuclei (Supplemental Fig. 1), including some close to the dorsal lateral neurons involved in circadian rhythms38. Surviving transheterozygotes of previously described lid loss-of-function mutants showed a high incidence of arrhythmicity (11/28 of lidk06801/lid10424 transheterozygotes had no discernible rhythms). This phenotype was similar to a previously reported high incidence of arrhythmicity, albeit in lid10424/ + heterozygotes25.
Several JmjC mutants differentially modulate sleep and activity levels
We next measured sleep and activity levels in 12hr:12hr light:dark (LD) conditions, to assess sleep duration and locomotion activity. In addition, we assayed mutants’ reaction to starvation on non-nutritious, but hydrating agar. Measures of starvation included both total counts of starvation-induced hyperactivity (and concomitant sleep suppression–as part of a food-searching strategy), as well as time to death35,36. Many of the JmjC mutants significantly affected sleep and activity levels in LD (see below). We analyzed some of the stronger phenotypes in more detail and rescued them with the genomic tagged JmjC::HA rescue transgenes (Figs 2–4). JMJD5KO showed a reduction in daytime sleep, as well as an increase in daytime activity, both of which could be rescued towards wild-type levels (Fig. 2A,B). JMJD5::HA from the rescue transgene was also expressed in numerous nuclei (Fig. 2C), including some close to the dorsal lateral neurons involved in circadian rhythms38.
JMJD7KO exhibited a reduction in daytime sleep, that could be rescued towards wild type with the genomic-tagged transgene (Fig. 2D,E). The JMJD7:HA transgene showed expression in the pars intercerebralis and fan-shaped body (of the adult fly brain), regions known to affect sleep time (Fig. 2 F;39,40).
NO66KO flies exhibited a reduced sleep phenotype and increased activity phenotype (Fig. 3A–D). This was in part driven by an increase in waking activity, i.e. activity per waking minute (Fig. 3D). Upon removal from food, these mutants also showed a strong increase in cumulative starvation-induced hyperactivity (Fig. 3E–G). The enhanced starvation activity did not cause premature death, however. Rather, NO66KO flies showed a slight delay in starvation-induced death (Fig. 3H). Almost all of these phenotypes were rescued by expressing the genomic NO66::HA transgene (Fig. 3A–H), and anti-HA staining showed expression in the pars intercerebralis and parts of the fan-shaped body in the adult fly brain (Fig. 3I).
Lastly, KDM4BKO exhibited a strong increase in daytime sleep and also significantly more nighttime sleep (Fig. 4A,B), in concert with decreased activity (Fig. 4C,D). Waking activity was not affected, arguing against KDM4BKO flies being lethargic, or sickly (Fig. 4E). Remarkably, the increased sleep phenotype of KDM4BKO also extended into their starvation response, and the mutants showed no sign of starvation-induced hyperactivity and arousal from sleep (Fig. 4F–H). A number of these phenotypes could be rescued to wild-type levels by expressing a KDM4B::HA transgene, but in some instances, we “rescued” beyond wild type, suggesting possible overexpression effects. The rescuing genomic-tagged transgene was also expressed in the pars intercerebralis and weakly in the fan-shaped body of the adult fly brain (Fig. 4J).
All JmjC non-lethal mutants affect sleep and circadian rhythm
As shown in Fig. 5, all 11 viable JmjC mutants tested exhibited significant differences (stars in Fig. 5B) in specific aspects of circadian behaviors, which are described in detail above. Most of the phenotypes observed, however, showed modest z-scores of less than 1.3 (standard deviations away from the mean of wild type; see Fig. 5A and depth of color in Fig. 5B), and effect sizes (Herge’s g; Supplemental Table 1). The phenotypes are unlikely to be caused by defective vision, as electroretinogram recordings of all JmjC mutants showed normal neurotransmission (Supplemental Fig. 2). However, we identified mutants that specifically affected overall activity (NO66KO), starvation-induced hyperactivity (KDM3KO, KDM4BKO) daytime sleep (JMJD5KO, JMJD7KO and KDM4BKO), nighttime sleep (lid, PSR, KDM4AKO and HSPBAP1KO), rhythm power (JMJD5KO and KDM2KO), period length (JMJD5KO, KDM2KO, KDM3KO) and overall rhythmicity (KDM3KO, lid; Fig. 5B). A hierarchical clustering analysis of this phenotype by genotype space revealed behavioral measures that are affected in concert. This includes, day- (or night-) time sleep and activity: as sleep increases, total activity decreases, we therefore expected these phenotypes to be affected together.
Buoyed by the validity of this clustering along the behavioral axis, we also examined the clustering along the genotype axis, which grouped the JmjC genes into two distinct categories, largely correlating with increased, versus decreased amounts of sleep. Mutants of KDM3, KDM4A, and KDM4B all fell into the same category of long-sleepers, and as mentioned above, KDM4A and KDM4B are known to demethylate H3K36me2/3 and all three members are predicted to affect H3K9me2/3 methylation31. Furthermore, five out of six genes in this category also affected chromatin32. In the second category of short-sleepers, four out of five genes belong to the JmjC domain-only subgroup and all four have recently been shown to act as protein hydroxylases41,42.
Taken together, our data indicate that Drosophila JmjC genes are involved in the modulation of circadian rhythms and sleep. All non-lethal JmjC mutants affect circadian rhythms and/or sleep, from very subtly (JMJD4) to very strongly (KDM4B). Importantly, none of the measures we assayed were affected by all of the mutants, and no single mutant affected all the measures, implying a certain degree of behavioral specificity for these genes. This is supported by our finding that no mutant tested exhibited the same set of phenotypes as another mutant.
Discussion
Most JmjC genes regulate circadian rhythms and/or sleep
Using sensitized genetic backgrounds for different signaling pathways, we identified genetic modulation as a mode of action for JmjC mutants32. To further uncover whether these mutants are required for evolutionarily relevant behaviors, we tested all 11 non-lethal mutants in behavioral assays (sleep, activity and circadian rhythm) that are quantitative and reproducible. To our surprise, we found that all eleven JmjC mutants tested significantly altered one or more measures of rhythm and sleep. It had been previously shown that histone modifications at specific sites throughout the genome oscillate during the circadian cycle23,24,25,26. In addition, a few JmjC proteins have been found to regulate circadian measures/behaviors in various organisms: KDM2A knock down in human cells led to a shortening of the period length of cells (as measured by an oscillating luciferase reporter;43. We show that Drosophila KDM2KO also caused a shortening of the period (Fig. 1A). Similarly, loss of JMJD5/JMJ30, a JmjC-domain only KDM, caused period-shortening in Arabidopsis and in cultured human cells, and the plant and mammalian genes are conserved enough to rescue the phenotype in the reciprocal system44,45. We found that Drosophila JMJD5KO mutation also caused a significantly shortened period-length (Fig. 1A), and in addition reduced daytime sleep (Fig. 2A). Lastly, mammalian Jarid1a, and its Drosophila homolog Lid, regulate the expression of the Period central clock gene by inhibiting HDAC1 activity, and Drosophila lid mutants have been described as highly arrhythmic25. We observed similar phenotypes (Fig. 5B), and overall our data indicate that JmjC gene function in circadian rhythms is highly conserved from plants to flies to human cells. Additionally, we found that KDM3KO mutants also have a high incidence of arrhythmicity (Fig. 5B). We therefore newly implicate this histone demethylase, which is involved in gene activation32,46, in the regulation of the core clock mechanism.
Much less is known about the involvement of histone modifications and JmjC genes in the regulation of sleep. We found that eight of eleven JmjC mutants significantly affected the amounts of day- and/or night-time sleep. Half of the phenotypes were less than 1.3 standard deviation (sd) from the mean, yet still highly significant, mainly because of the large number (>30) of individual flies that can be easily assayed. Four mutants showed strong sleep/activity phenotypes (>1.3 sd): lid, KDM4AKO, and HSPBAP1KO all showed increased night time sleep, while KDM4BKO increased daytime sleep. This latter mutant is particularly intriguing, because not only did it display a very strong phenotype with increased daytime sleep compared to the rescued control (while leaving relative activity per waking minute largely unaffected; Fig. 4A–E), but this increased sleep phenotype also extended into food deprivation. Wild-type flies react to food deprivation with a long bout of starvation-induced hyperactivity, including sleep-suppression, a conserved phenomenon thought to represent foraging for food35,47. KDM4BKO flies lacked starvation-induced arousal, and with it hyperactivity (Fig. 4F–H). This suggests that KDM4B may be generally involved in arousal. Supporting this hypothesis is the finding that our genomic tagged transgene and rescue construct, KDM4B::HA, is expressed in the dorsal fan-shaped body (Fig. 4I) strongly resembling a layer receiving dopaminergic input for arousal40. It will be interesting to see whether KDM4B mutants show disrupted expression of genes involved in dopaminergic transmission, or regulation of Rho-family GTPases, which are also required in the dorsal fan-shaped body for normal sleep48.
Contrary to our initial hypothesis, the majority of Drosophila JmjC genes are not essential during development32. Rather, many of them modulate changes in chromatin organization and gene expression programs32. Here we show that many Drosophila JmjC genes also modulate behavior, specifically circadian rhythms and sleep. It remains to be determined, whether the phenotypes we observed are a consequence of developmental abnormalities in the nervous system, or whether these genes acutely participate in (cyclic) neuronal function and transcription. We have found that a subset of these JmjC genes is also involved in modulating behavioral responses to ethanol, and that lid, KDM3 and NO66 are required in the nervous system for normal reactions to ethanol49. Together, these data imply that JmjC proteins help to modulate a variety of processes in this organism, including behavior, such as circadian rhythms and sleep.
References
Cirelli, C. & Bushey, D. Sleep and wakefulness in Drosophila melanogaster. Annals of the New York Academy of Sciences 1129, 323–329, https://doi.org/10.1196/annals.1417.017 (2008).
Masri, S., Cervantes, M. & Sassone-Corsi, P. The circadian clock and cell cycle: interconnected biological circuits. Curr Opin Cell Biol 25, 730–734, https://doi.org/10.1016/j.ceb.2013.07.013 (2013).
Eckel-Mahan, K. & Sassone-Corsi, P. Metabolism and the circadian clock converge. Physiol Rev 93, 107–135, https://doi.org/10.1152/physrev.00016.2012 (2013).
Gamble, K. L., Berry, R., Frank, S. J. & Young, M. E. Circadian clock control of endocrine factors. Nat Rev Endocrinol 10, 466–475, https://doi.org/10.1038/nrendo.2014.78 (2014).
Dumbell, R., Matveeva, O. & Oster, H. Circadian Clocks, Stress, and Immunity. Frontiers in endocrinology 7, 37, https://doi.org/10.3389/fendo.2016.00037 (2016).
Moore, R. Y. Circadian rhythms: basic neurobiology and clinical applications. Annual review of medicine 48, 253–266, https://doi.org/10.1146/annurev.med.48.1.253 (1997).
Borbely, A. A. & Achermann, P. Sleep homeostasis and models of sleep regulation. J Biol Rhythms 14, 557–568 (1999).
Toh, K. L. et al. An hPer2 phosphorylation site mutation in familial advanced sleep phase syndrome. Science 291, 1040–1043, https://doi.org/10.1126/science.1057499 (2001).
Xu, Y. et al. Functional consequences of a CKIdelta mutation causing familial advanced sleep phase syndrome. Nature 434, 640–644, https://doi.org/10.1038/nature03453 (2005).
Lowrey, P. L. & Takahashi, J. S. Genetics of circadian rhythms in Mammalian model organisms. Adv Genet 74, 175–230, https://doi.org/10.1016/B978-0-12-387690-4.00006-4 (2011).
Lu, S. X. & Tobin, E. M. Chromatin remodeling and the circadian clock: Jumonji C-domain containing proteins. Plant signaling & behavior 6, 810–814, https://doi.org/10.4161/psb.6.6.15171 (2011).
Aguilar-Arnal, L. & Sassone-Corsi, P. Chromatin landscape and circadian dynamics: Spatial and temporal organization of clock transcription. Proceedings of the National Academy of Sciences of the United States of America 112, 6863–6870, https://doi.org/10.1073/pnas.1411264111 (2015).
Partch, C. L., Green, C. B. & Takahashi, J. S. Molecular architecture of the mammalian circadian clock. Trends Cell Biol 24, 90–99, https://doi.org/10.1016/j.tcb.2013.07.002 (2014).
Asher, G. & Sassone-Corsi, P. Time for food: the intimate interplay between nutrition, metabolism, and the circadian clock. Cell 161, 84–92, https://doi.org/10.1016/j.cell.2015.03.015 (2015).
Masri, S., Kinouchi, K. & Sassone-Corsi, P. Circadian clocks, epigenetics, and cancer. Curr Opin Oncol 27, 50–56, https://doi.org/10.1097/CCO.0000000000000153 (2015).
Stevens, R. G., Brainard, G. C., Blask, D. E., Lockley, S. W. & Motta, M. E. Breast cancer and circadian disruption from electric lighting in the modern world. CA: a cancer journal for clinicians 64, 207–218, https://doi.org/10.3322/caac.21218 (2014).
Lucassen, E. A. et al. Environmental 24-hr Cycles Are Essential for Health. Current biology: CB. https://doi.org/10.1016/j.cub.2016.05.038 (2016).
Liu, S. et al. WIDE AWAKE mediates the circadian timing of sleep onset. Neuron 82, 151–166, https://doi.org/10.1016/j.neuron.2014.01.040 (2014).
Curtis, A. M. et al. Histone acetyltransferase-dependent chromatin remodeling and the vascular clock. The Journal of biological chemistry 279, 7091–7097, https://doi.org/10.1074/jbc.M311973200 (2004).
Doi, M., Hirayama, J. & Sassone-Corsi, P. Circadian regulator CLOCK is a histone acetyltransferase. Cell 125, 497–508, https://doi.org/10.1016/j.cell.2006.03.033 (2006).
Etchegaray, J. P., Yang, X. & DeBruyne, J. P. The polycomb group protein EZH2 is required for mammalian circadian clock function. Journal of Biological …, https://doi.org/10.1074/jbc.M603722200 (2006).
Naruse, Y., Oh-hashi, K., Iijima, N. & Naruse, M. Circadian and light-induced transcription of clock gene Per1 depends on histone acetylation and deacetylation. and cellular biology, https://doi.org/10.1128/MCB.24.14.6278-6287.2004 (2004).
Valekunja, U. K. & Edgar, R. S. Histone methyltransferase MLL3 contributes to genome-scale circadian transcription. Proceedings of the …, https://doi.org/10.1073/pnas.1214168110 (2013).
Etchegaray, J.-P. P., Lee, C., Wade, P. A. & Reppert, S. M. Rhythmic histone acetylation underlies transcription in the mammalian circadian clock. Nature 421, 177–182, https://doi.org/10.1038/nature01314 (2003).
DiTacchio, L. et al. Histone lysine demethylase JARID1a activates CLOCK-BMAL1 and influences the circadian clock. Science (New York, N.Y.) 333, 1881–1885, https://doi.org/10.1126/science.1206022 (2011).
Koike, N. et al. Transcriptional architecture and chromatin landscape of the core circadian clock in mammals. Science (New York, N.Y.) 338, 349–354, https://doi.org/10.1126/science.1226339 (2012).
Katada, S. & Sassone-Corsi, P. The histone methyltransferase MLL1 permits the oscillation of circadian gene expression. Nature structural & molecular biology 17, 1414–1421, https://doi.org/10.1038/nsmb.1961 (2010).
Kwok, R. S., Lam, V. H. & Chiu, J. C. Understanding the role of chromatin remodeling in the regulation of circadian transcription in Drosophila. Fly (Austin) 9, 145–154, https://doi.org/10.1080/19336934.2016.1143993 (2015).
Kwok, R. S., Li, Y. H., Lei, A. J., Edery, I. & Chiu, J. C. The Catalytic and Non-catalytic Functions of the Brahma Chromatin-Remodeling Protein Collaborate to Fine-Tune Circadian Transcription in Drosophila. PLoS Genet 11, e1005307, https://doi.org/10.1371/journal.pgen.1005307 (2015).
Black, J. C., Van Rechem, C. & Whetstine, J. R. Histone lysine methylation dynamics: establishment, regulation, and biological impact. Mol Cell 48, 491–507, https://doi.org/10.1016/j.molcel.2012.11.006 (2012).
Mosammaparast, N. & Shi, Y. Reversal of histone methylation: biochemical and molecular mechanisms of histone demethylases. Annu Rev Biochem 79, 155–179, https://doi.org/10.1146/annurev.biochem.78.070907.103946 (2010).
Shalaby, N. A. et al. Systematic discovery of genetic modulation by Jumonji histone demethylases in Drosophila. Sci Rep 7, 5240, https://doi.org/10.1038/s41598-017-05004-w (2017).
Chan, C. C. et al. Systematic discovery of Rab GTPases with synaptic functions in Drosophila. Curr Biol 21, 1704–1715, https://doi.org/10.1016/j.cub.2011.08.058 (2011).
Picot, M., Cusumano, P., Klarsfeld, A., Ueda, R. & Rouyer, F. Light activates output from evening neurons and inhibits output from morning neurons in the Drosophila circadian clock. PLoS Biol 5, e315, https://doi.org/10.1371/journal.pbio.0050315 (2007).
Lee, G. & Park, J. H. Hemolymph sugar homeostasis and starvation-induced hyperactivity affected by genetic manipulations of the adipokinetic hormone-encoding gene in Drosophila melanogaster. Genetics 167, 311–323 (2004).
Keene, A. C. et al. Clock and cycle limit starvation-induced sleep loss in Drosophila. Curr Biol 20, 1209–1215, https://doi.org/10.1016/j.cub.2010.05.029 (2010).
Fabian-Fine, R. et al. Endophilin promotes a late step in endocytosis at glial invaginations in Drosophila photoreceptor terminals. J Neurosci 23, 10732–10744 (2003).
Johard, H. A. et al. Peptidergic clock neurons in Drosophila: ion transport peptide and short neuropeptide F in subsets of dorsal and ventral lateral neurons. J Comp Neurol 516, 59–73, https://doi.org/10.1002/cne.22099 (2009).
Foltenyi, K., Greenspan, R. J. & Newport, J. W. Activation of EGFR and ERK by rhomboid signaling regulates the consolidation and maintenance of sleep in Drosophila. Nat Neurosci 10, 1160–1167, https://doi.org/10.1038/nn1957 (2007).
Liu, Q., Liu, S., Kodama, L., Driscoll, M. R. & Wu, M. N. Two dopaminergic neurons signal to the dorsal fan-shaped body to promote wakefulness in Drosophila. Curr Biol 22, 2114–2123, https://doi.org/10.1016/j.cub.2012.09.008 (2012).
Noma, A. et al. Expanding role of the jumonji C domain as an RNA hydroxylase. J Biol Chem 285, 34503–34507, https://doi.org/10.1074/jbc.M110.156398 (2010).
Wang, H. et al. Structure of the JmjC-domain-containing protein JMJD5. Acta Crystallogr D Biol Crystallogr 69, 1911–1920, https://doi.org/10.1107/S0907444913016600 (2013).
Reischl, S. & Kramer, A. Fbxl11 Is a Novel Negative Element of the Mammalian Circadian Clock. Journal of biological rhythms 30, 291–301, https://doi.org/10.1177/0748730415587407 (2015).
Jones, M. A. et al. Jumonji domain protein JMJD5 functions in both the plant and human circadian systems. Proceedings of the National Academy of Sciences of the United States of America 107, 21623–21628, https://doi.org/10.1073/pnas.1014204108 (2010).
Lu, S. X. et al. The Jumonji C domain-containing protein JMJ30 regulates period length in the Arabidopsis circadian clock. Plant physiology 155, 906–915, https://doi.org/10.1104/pp.110.167015 (2011).
Klose, R. J., Kallin, E. M. & Zhang, Y. JmjC-domain-containing proteins and histone demethylation. Nat Rev Genet 7, 715–727, https://doi.org/10.1038/nrg1945 (2006).
Mistlberger, R. E. Neurobiology of food anticipatory circadian rhythms. Physiol Behav 104, 535–545, https://doi.org/10.1016/j.physbeh.2011.04.015 (2011).
Donlea, J. M., Pimentel, D. & Miesenbock, G. Neuronal machinery of sleep homeostasis in Drosophila. Neuron 81, 860–872, https://doi.org/10.1016/j.neuron.2013.12.013 (2014).
Pinzón, J. H. et al. Alcohol-Induced Behaviors Require a Subset of JmjC-Domain Histone Demethylases in the Nervous System. Alcoholism: Clinical and Experimental Research 41(12), 2015–2024 (2017).
Acknowledgements
N.A.S. was supported by the American Heart Association and NIH (Training grant DK 7745–17). This work was supported in various phases by NIAAA (R01AA019526 to A.R., R21AA022404 to A.R. & M.B.), NIDDK (K08DK091316 and R01DK110358 to ARR), The March of Dimes (#5FY09–10), NIHGMS (1R01GM086647) and Cancer Prevention Research Institute of Texas (RP100516) to M.B. We would like to thank the Bloomington Drosophila Stock Center and the Developmental Studies Hybridoma Bank for reagents, Robin Hiesinger, Helmut Kramer, Lauren Tyra, Zeynep Okray, Bassem Hassan and members of the Buszczak and Rothenfluh labs for comments and advice. The authors declare no competing financial interests.
Author information
Authors and Affiliations
Contributions
M.B. and A.R. conceived the project. N.S., J.P., A.N., E.J.J., M.R., R.D., H.W., A.R.R., M.B. and A.R. designed and conducted the experiments. N.S., A.R.R., M.B. and A.R. analyzed the data, wrote and edited the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shalaby, N.A., Pinzon, J.H., Narayanan, A.S. et al. JmjC domain proteins modulate circadian behaviors and sleep in Drosophila. Sci Rep 8, 815 (2018). https://doi.org/10.1038/s41598-017-18989-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-18989-1
This article is cited by
-
Identification of ecdysone receptor target genes in the worker honey bee brains during foraging behavior
Scientific Reports (2023)
-
JMJD5 inhibits lung cancer progression by facilitating EGFR proteasomal degradation
Cell Death & Disease (2023)
-
Structural analysis of the 2-oxoglutarate binding site of the circadian rhythm linked oxygenase JMJD5
Scientific Reports (2022)
-
Whole-genome association analyses of sleep-disordered breathing phenotypes in the NHLBI TOPMed program
Genome Medicine (2021)
-
Proteomic analysis of Drosophila CLOCK complexes identifies rhythmic interactions with SAGA and Tip60 complex component NIPPED-A
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.