Abstract
Human activities can cause habitat degradation that may alter the types and quality of available food resources and thus influence the microbiomes of wild animal populations. Furthermore, seasonal shifts in food availability may cause adaptive responses in the gut microbiome to meet the need for different metabolic capabilities. Here, we demonstrate local-scale population structure in the gastrointestinal microbiotas of Chlorocebus monkeys, in southern Ethiopia, in response to varying degrees of human encroachment. We further provide evidence of adaptation to ecological conditions associated with the dry and wet seasons, and show seasonal effects to be more pronounced in areas with limited human activity. Finally, we report species-level microbiota differences between the endemic Ethiopian Bale monkey, an ecological specialist, and generalist Chlorocebus species from the same geographical region.
Similar content being viewed by others
Introduction
Much recent research has documented the important roles that the microbes inhabiting the gastrointestinal (GI) system play in host physiology1. Several studies have documented the role of diet in shaping the GI microbiota of humans2,3 and other mammals4,5. A recent study even demonstrated humanization of the GI microbiota in zoo monkeys, due to reduced intake of dietary fiber in the captive primates, indicating a high degree of plasticity in response to diet6.
The Chlorocebus is a genus of non-human primates widespread in sub-Saharan Africa that includes grivets (C. aethiops, Fig. 1a), vervets (C. pygerythrus, Fig. 1b) and Bale monkeys (C. djamdjamensis, Fig. 1c). Grivets are distributed across northern Ethiopia and surrounding areas, while vervets are found across large parts of southeast Africa. The habitats of vervets and grivets overlap in southern Ethiopia where interbreeding is believed to occur7 (Fig. 2). The Bale monkey is endemic to the Bale Mountains of Southern Ethiopia, and is thus parapatric with grivets and vervets8. Grivets and vervets are considered dietary generalists, feeding on a variety of fruits, seeds and pods9,10,11. Bale monkeys are considered dietary specialists who, along with the golden monkeys (Cercopithecus kandti) of Uganda12 and the lemurs of the genus Hapalemur (bamboo lemurs) of Madagaskar13, are the only primate species adapted to eating mostly bamboo14. Due to a large increase in human settlements, much of the original bamboo forest that forms the natural Bale monkey habitat has been cut down. This habitat fragmentation has resulted in populations becoming isolated in habitats with extensive human influence8. Two such areas are Afursa and Kokosa (Fig. 2). In Afursa (hereby referred to as NBF = no bamboo forest) bamboo growth is almost completely eradicated, while Kokosa (hereby referred to as DBF = degraded bamboo forest) consists of spread out patches of bamboo growth15. Bale monkeys living here have largely abandoned a bamboo-dominated diet. In contrast, Odobullu (hereby referred to as CBF = continuous bamboo forest; Fig. 2) covers a large area of predominantly dense bamboo growth, representing a natural habitat.
Map of the sampling area. The red outline shows the habitat range of the Bale monkeys. The map was generated using a 30 meter digital elevation model in ArcGIS 10.4 (www.arcgis.com).
In this study we use 16S rRNA gene amplicon sequencing to compare GI microbiotas in Bale monkeys from the three study sites, addressing three main hypotheses. First, ecological differences on a local spatial scale, driven by habitat degradation, may cause GI microbiota differences among Bale monkeys at different sampling sites. Secondly, seasonal differences in food availability may cause GI microbiota variation within Bale monkey sampling sites. Lastly, ecological specializations in Bale monkeys may cause their GI microbiota to differ from the more generalist Chlorocebus species. We also make an effort to interpret our microbiota data using chloroplast 16S rRNA gene sequence reads, produced as a bi-product of amplicon sequencing, as a potential source of dietary information.
Results
GI microbiota composition of the Bale monkeys in the three study sites
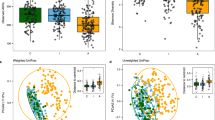
The three Bale monkey populations clustered according to study site (p < 0.001, ANOSIM). The main dimension of variation distinguished between NBF, on the one hand, and DBF and CBF on the other (Fig. 3a). The second dimension mostly distinguished between the DBF and CBF populations. On the phylum level, differences between the three populations were mainly related to the Firmicutes vs. Bacteroidetes balance (Fig. 3b). The population mean relative abundance of Firmicutes was highest in in NBF (64.9%), followed by CBF (58.2%), and DBF (54.7%) (p < 0.001 for all comparisons, Wilcoxon rank sum test). Conversely, Bacteroidetes relative abundances were lower in NBF (13.4%) than in CBF (23.0%) or DBF (23.7%) (p < 0.001 for both comparisons). We did not observe significant differences in either Firmicutes or Bacteroidetes abundances between the DBF and CBF populations. Other significant phylum-level differences between populations were elevated abundances of Spirochaetes in DBF relative to both of the other populations (p = 0.003 and 0.021 for comparison with NBF and CBF, respectively), higher levels of Euryarchaeota (methanogenic archaea) in CBF relative to NBF and DBF (p < 0.001 and p = 0.01, respectively), and higher levels of Tenericutes in NBF (p < 0.001 and p = 0.004 compared with DBF and CBF, respectively). A relatively rare phylum, Lentisphaera, was enriched almost nine fold in NBF (0.68%) compared with CBF (0.08%) (p < 0.001) and four fold relative to DBF (0.17%) (p < 0.001).
We also carried out exact tests for enrichments of specific OTUs among the three populations. Using a strict significance cut-off of both p-value and false discovery rate (FDR) ≤0.01 we found 83 OTUs with differential occurrence between NBF and DBF, 54 between NBF and CBF and 37 between DBF and CBF (Supplementary Tables 1–3). There was extensive sharing of enriched OTUs when comparing one of the populations with the two others. In particular, when comparing NBF with either DBF or CBF, the top two OTUs discriminating between the populations were enriched more than a hundredfold in both of the two latter populations. Both of these OTUs were classified with low confidence, one to the genus Parabacteroides (OTU56, phylum Bacteroidetes) and the other to the genus Asteroleplasma (OTU26, phylum Tenericutes). A total of 38 OTUs showed significant differential occurrence in NBF relative to both DBF and CBF, and out of these OTUs 36 co-varied in the sense that they were either enriched or depleted in both DBF and CBF. We saw a similar trend when looking at OTUs with significant differential occurrence in DBF relative to NBF and CBF, with 17 out of 22 shared OTUs co-varying. For OTUs with shared differential occurrence in CBF relative to NBF and DBF, 16 out of 17 co-varied. Thus, there was a trend that an OTU that distinguished a first population from a second also distinguished the first population from the third.
We did find some effects of sampling site on the observed OTU diversity (Supplementary Fig. 1), when considering all the bale monkey samples together regardless of season. DBF had consistently lower diversity as measured by Shannon entropy, OTU richness and the Chao1 diversity estimator. However, results from significance testing were somewhat inconsistent (Supplementary Table 4).
Seasonal plasticity
Fecal samples from Bale monkeys were collected between October and June, with October through February being a dry season and March through June a wet season. In order to look for seasonal effects in the three Bale monkey populations we compared the microbiota composition of samples collected during these two periods. In the NBF population we observed significant differences in the microbiotas dependent on season (Fig. 4a; p < 0.001, ANOSIM), with the distinction between the dry and wet season explaining 4.3% of inter-sample variation (PERMANOVA). For the DBF and CBF populations we saw stronger seasonal effects (Fig. 4b,c; p < 0.001 for both tests) with season explaining 9.1% and 10.6% of inter-sample variation, respectively.
NMDS and phylum level relative abundances in the Bale monkey samples with samples split into those collected during the dry and wet season. The panels show the populations in NBF (a and d), DBF (b and e) and CBF (c and f). The NMDS plots (a–c) shows the two main dimensions of variation, with dots colour coded according to season (black = dry, red = wet). Relative abundances (d–f) are shown for the top eight phyla.
At the phylum level, in NBF there was a marginally significant (p = 0.048, Wilcoxon rank sum test) decrease in Bacteroidetes abundances from the dry to the wet season (Fig. 4d), while Spirochaetes and Actinobacteria increased significantly (p = 0.010 and 0.021, respectively). In both DBF (Fig. 4e) and CBF (Fig. 4f), Firmicutes decreased in the wet relative to the dry season (p = 0.002 and 0.056 in DBF and CBF, respectively), with a concurrent increase in Bacteroidetes (p = 0.011 and 0.006). In DBF there were significant increases in Proteobacteria and Euryachaeota (p < 0.001 in both cases) and decreases in Actinobacteria and Fibrobacteres (p < 0.001 and p = 0.001), from the dry to the wet season. In CBF there were significant decreases in Spirochaetes and Actinobacteria (p = 0.029 and 0.026) as well as a marginally significant decrease in Verrucomicrobia (p = 0.057).
On the 97% OTU level, bacterial groups with significant enrichment in one season relative to the other were found in all three Bale monkey populations, with 18, 47 and 14 OTUs observed with a p-value and FDR of less than 0.01 in NBF, DBF and CBF, respectively (Supplementary Tables 5–7). No OTU with differential seasonal abundance was observed in all three populations, but five of the same OTUs were found to have significant seasonal enrichment in both NBF and DBF, one (tentatively classified to the Proteobacterial genus Vampirovibrio, a bacterial predator) during the wet season and four (all Firmicutes) during the dry season. Also, a single OTU (a Bacteroidetes tentatively classified as Barnsiella) was enriched in the wet season in both DBF and CBF. Thus, it appears that, in spite of a few communalities, the seasonal response on the 97% OTU level is particular to the area in which the monkeys live. We did not observe significant differences in OTU diversity between seasons in any of the Bale monkey populations, except for a marginally significant (p = 0.049, unpaired t-test) increase in the Chao1 diversity estimator in the wet relative to the dry season in CBF (Supplementary Fig. 2).
Comparison between specialist and generalist Chlorocebus species
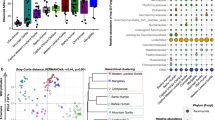
In order to look for differences in the GI microbiota between Ethiopian Chlorocebus species, we compared the three Bale monkey populations to the five populations of vervets and grivets. Sixty-four (4.9% of total) OTUs were found in all 219 samples from Bale, grivet and vervet monkeys. 334 OTUs (25.7% of total) were common to at least 90% of samples, suggesting a core microbiota in Chlorocebus monkeys. There was a highly significant separation between the Bale monkeys and the combined vervet and grivet populations (Fig. 5a; p < 0.001 ANOSIM), with the first NMDS dimension providing the main distinction. By comparing sample scores along the main NMDS dimension (Supplementary Fig. 3) the DBF population of Bale monkeys appeared more similar to vervets and grivets than Bale monkeys from NBF and CBF (mean score 0.05, p < 0.001 for both comparisons, Wilcoxon rank sum test), while the two latter populations did not differ significantly from one another (mean scores 0.15 and 0.18, respectively). We did not observe significant differences when comparing all three grivet populations to the one pure vervet population (Supplementary Fig. 4). However, we observed a strong geographical effect among the five vervet and grivet populations (p < 0.001, ANOSIM), with sampling site explaining 27.2% of inter-sample variation (PERMANOVA). At the southernmost site, Arba Minch, extensive hybridization is thought to occur between vervets and grivets7. Samples from this site clustered poorly (Supplementary Fig. 4), suggesting that interbreeding may affect the microbiota. Looking at all eight Chlorocebus populations, we did not find any evidence to suggest that geographical proximity causes GI microbiotas to be more similar (rho = −0.08, p = 0.70, Spearman’s rank correlation test).
NMDS and phylum level relative abundances for the three Bale monkey populations (BM) combined and the five vervet and grivet populations (V/G) combined. (a) The NMDS plot shows the two main dimensions of variation, with dots colour coded according to species (top right corner). (b) Relative abundances are shown for the top eight phyla.
On the phylum level there were extensive differences between Bale monkeys and vervets/grivets (Fig. 5b). Firmicutes, Tenericutes and Fibrobacteres were more abundant in the Bale monkeys (p = 0.001 for Firmicutes and p < 0.001 for the two others), while Proteobacteria, Verrucomicrobia and Actinobacteria were more abundant in vervets and grivets (p = 0.006 and 0.032 and p < 0.001, respectively). 94 of the most abundant OTUs were significantly (p-value and FDR ≤ 0.01) enriched in either Bale monkeys or vervets/grivets (Supplementary Table 8). The majority (66.0%) of enriched OTUs were classified as Firmicutes, but the three most highly enriched OTUs were all Bacteroidetes, one tentatively classified to the genus Barnsiella and two as Prevotella. These three OTUs were enriched more than a hundredfold in the vervet and grivet populations. The GI microbiota of the Bale monkey populations was more diverse than that of the vervets and grivets (Supplementary Fig. 5), both in terms of Shannon entropy (p < 0.001, unpaired t-test), OTU richness (p = 0.011) and the Chao1 diversity estimator, although the difference in Chao1 diversity was not significant.
Chloroplasts’ composition: implication for geographical and seasonal differentiation
The NBF population had the highest number of chloroplast reads (8.7% of total) followed by CBF (5.5%) and DBF (2.2%), while chloroplast sequences were relatively rare in the vervet and grivet populations (Supplementary Fig. 6). Thus, further analyses focuses only on the three Bale monkey populations. It should be noted that chloroplast 16S rRNA sequences do not provide good resolution as a taxonomic marker in plants, and this should be taken into account when interpreting the following results.
There was a clear geographical signature in the chloroplast sequences (Fig. 6a; p < 0.001, ANOSIM), with sampling site explaining 18% of between-sample variation (PERMANOVA). The chloroplast sequence data suggested that the NBF population had a much more diverse diet, as measured by Shannon entropy (Supplementary Fig. 7; p < 0.001 for both comparisons, unpaired t-test). Diversity in DBF was higher than in CBF with marginal significance (p = 0.042). We found 9 bacterial OTUs that had a significant (p ≤ 0.05, Spearman’s rank correlation tests) positive correlation with the relative abundance of sequences putatively classified as Poaceae (grass), and 7 of these OTUs were classified to the order Clostridia (Supplementary Table 9).
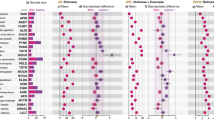
NMDS and relative abundances of OTUs classified as chloroplasts in 106 Bale monkey samples. (a) The NMDS plot shows the two main dimensions of variation, with plotted symbols colour coded according to sampling site (NBF = no bamboo forest, DBF = degraded bamboo forest, CBF = continuous bamboo forest) and shaped according to season. (b-d) Relative abundances of the seven top OTUs classified as chloroplasts. Samples from NBF (b), DBF (c) and CBF (d) are split into those collected during the dry and wet seasons. The colour key and classification shown on the right hand side give only the high level taxonomy (family and up).
While we did not observe significant seasonal effects of chloroplast sequence composition in the NBF population (Fig. 6b), we did see significant effects in DBF and CBF (Fig. 6c,d; p = 0.023 and 0.021, respectively; ANOSIM). We observed opposite trends at the two latter sites such that in DBF the chloroplast composition during the dry season was similar to that of NBF, with consumption of grassy plants (Poaceae) nearly halved relative to the wet season (44.4% vs. 82.8%; p < 0.001, Wilcoxon rank sum test). In CBF, on the other hand, grass consumption was greatly reduced in the wet relative to the dry season (66.2% vs. 91.5%, p < 0.001). Recovery of chloroplast sequences was higher in samples collected during the dry relative to the wet season in DBF and CBF (Supplementary Fig. 8).
Discussion
It is well established that the GI microbiota is instrumental to digestive function in animals, and that diet and microbiota composition are intimately linked. This is especially true for herbivorous creatures that rely on a high proportion of recalcitrant carbohydrates for nutrition, such as ruminants16,17 and many primate species18. There is also documentation that fragmentation and other kinds of habitat disturbance can cause dietary shifts in animal populations relative to those residing in pristine habitats19,20. Indeed, two previous studies of primates concluded that habitat degradation can significantly affect the GI microbiota21,22.
CBF represents a uniform and continuous environment with bamboo, a member of the family Poaceae, accounting for more than 86% of the total vegetation15. The Bale monkey population in this area relies on a single species of bamboo (Arundinaria alpina) for most of their dietary intake (Mekonnen et al., in review). Thus, these animals are considered ecological specialists, and until recently they were thought to exist only in the relatively undisturbed bamboo forests of the Bale Mountains. However, the discovery of populations living in small and isolated forest fragments in human-dominated areas suggests that they are more ecologically flexible than previously thought8.
In DBF bamboo has recently been found to account for ~40% of the vegetation, while the corresponding percentage in NBF was only 1.6%. Thus, although both forest fragments are subject to high anthropogenic influence, DBF is intermediate in terms of bamboo access (Mekonnen et al., in review). Access to bamboo is in rough correspondence with the number of chloroplast reads putatively classified as Poaceae recovered from the three sampling sites (Fig. 6, Supplementary Fig. 7). In CBF, averaged over both seasons, Poaceae constituted about 80% of the total chloroplast sequences. This corresponds very well with the empirically observed rates at which monkeys there feed on bamboo (Mekonnen et al., in review). In DBF, monkeys spend ~30% of their foraging time feeding on bamboo, while sequences matching Poaceae were ~60% of total chloroplasts. This population is known to spend large amounts of time in human agricultural areas15, feeding on grasses and herbs, as well as fruit and other crops, i.e. plant parts without chloroplasts. This may help explain the relatively low number of chloroplast reads recovered from these monkeys, as well as the discrepancy between observed feeding rates and recovered grass sequences. In NBF the amount of Poaceae sequences was greatly reduced relative to the two other sites. The main grass species there is the so called smelly grass (Bothriochloa radicans), which takes up ~15% of the Bale monkeys’ foraging time, and they have adopted a range of other flowering plants as major components of their green diet, including shrubs, herbs and trees.
We observed clear differentiation of the Bale monkey GI microbiota among the three habitat types (Fig. 3a). We found that monkeys from the two areas with better access to bamboo had more similar GI microbiotas compared to the low bamboo site, both having lower levels of Firmicutes and much higher levels of Bacteroidetes (Fig. 3b). We also found that that several OTUs were highly enriched in the high and intermediate bamboo sites relative to the low bamboo site (Supplementary Tables 1–3). Two OTUs, putatively classified to the genera Parabacteroides and Asteroleplasma, were particularly highly enriched in monkeys in CBF and DBF. Parabacteriodes is a typical component of mammalian GI microbiotas, including those of humans and chimpanzees23. Asteroleplasma is a poorly characterized genus within the Tenericutes, but two recent studies have identified these bacteria as part of the rumen microbiota24,25, suggesting that they may play a role in cellulose degradation. The taxonomic assignment of these OTUs should be interpreted with caution as both were classified with low confidence. The DBF population had consistently lower GI microbiota diversity than the two other populations, although the significance of differences between populations was inconsistent. We do not have a good explanation for this result. It is possible that it is related to a larger component of fruit, which is more easily digested than green plant parts, in the diet of the DBF population. It is known that herbivores have more diverse GI microbiotas than omnivores4, and this interpretation is supported by the number of recovered chloroplast sequences from the three Bale monkey populations (Supplementary Fig. 6).
Looking to the human GI microbiome, the most studied microbiome of any primate, it has been found that, although it is highly variable between individuals26, it is remarkably stable over time27. However, seasonal population level variation in the GI microbiota has been reported both in humans28 and non-human primates29,30, as well as wild wood mice31 and giant pandas32. In Bale monkeys we observed seasonal plasticity of the GI microbiotas of all three populations (Fig. 4a–c), with elevated levels of Firmicutes and lower levels of Bacteroidetes during the dry season in each of the populations (Fig. 4d–f). These effects are presumably related to the diversity of ingested food items and the relative amounts of bamboo leaves and other green grasses in their diet. A study of lowland and mountain gorillas similarly found that seasonal variation of the GI microbiota corresponded with fruit availability33. It is noteworthy that the seasonal signal was weakest in the NBF population (Fig. 4a and d), suggesting that these monkeys are becoming adapted to a food supply that is less dependent on seasonal availability, an observation that is supported by the chloroplast data (Fig. 6b). The seasonal effect was strongest in CBF (Fig. 4c and f), in accordance with a recent study of howler monkeys living in habitats with different degrees of disturbance which also found more pronounced seasonal effects on the GI microbiota of animals living in habitats with little human influence34. In the chloroplast data, we saw opposing seasonal trends in DBF and CBF, with the relative amount of Poaceae sequences increasing in the wet season in DBF while decreasing in CBF (Fig. 6c and d). These patterns may be explained by differences in the ecology at the two sites. In CBF, during the wet season, the Bale monkeys supplement their bamboo leaf intake with fresh shoots. The shoots do not contain chloroplasts, and thus are not observable in our data. In DBF the wet season brings a bloom in the abundance of many grass and herb species that are favoured by the monkeys, thus shifting their diets away from the flowering plants consumed during the dry season. Surprisingly, we observed only marginal effects of season on GI microbiota diversity within sampling sites. While it is fully possible that the Bale monkey GI microbiota does not respond strongly to seasonal ecological changes in terms of OTU diversity, it is also possible that sample sizes become too small to detect significant differences when splitting the data from each site into samples collected during the dry and wet season. This may be particularly important in the CBF population, where the sample size is smallest and we would expect the strongest seasonal effect.
In addition to feeding ecology, phylogeny has been found to be an important determinant of GI microbiome structure4, and differences have been found even among closely related non-human primates35. In our data we saw clear signs of differentiation between Bale monkeys and vervets/grivets (Fig. 5a). The vervet and grivet populations studied here were all from habitats with a high degree of human influence, and both of these species are well known ecological generalists. The Bale monkey populations of NBF and DBF have adapted to more generalist lifestyles than their conspecifics in CBF, and in particular the GI microbiota of the DBF population was more similar to those of vervets and grivets. However, the main contrast was between Bale monkeys on the one hand and vervets and grivets on the other, suggesting that in this case the phylogenetic signal overpowers the signal caused by the degree of dietary generalism. It is noteworthy that, although rare in all populations, the phylum Fibrobacteres was significantly enriched in Bale monkeys, by a factor of 3.4, relative to vervets and grivets. This phylum is mainly associated with cellulose digestion in ruminants36, and enrichment of these bacteria in Bale monkeys may signify adaptation to a high-fiber diet relative to the other Chlorocebus species. We also found that OTU diversity was elevated in Bale moneys relative to vervets and grivets (Supplementary Fig. 5). A previous study found that, in mammals, herbivores harbour more diverse GI microbiotas than omnivores4, while a study of two human populations found higher diversity in the one with a higher fiber diet2. Thus, our result may reflect an adaptation in Bale monkeys to a diet richer in cellulose relative to other Chlorocebus species. Our results also support the notion that, in general, ecological conditions and phylogeny rather than geographical proximity cause differentiation of the GI microbiota, with no detectable correlation between the mean pairwise distances between the GI microbiotas of the various Chlorocebus populations and the physical distance between their habitats. Within the grivet and vervet populations our results indicate that ecology was more important than phylogeny, with samples generally structuring according to population rather than species (Supplementary Fig. 4). Notably, the GI microbiota profiles of the population at the southernmost sampling site, Arba Minch, did not form one consistent cluster, and this site is thought to be a hybrid zone7. Effects of hybridization on the GI microbiome have rarely been documented37,38, and in our case we cannot say for sure whether the lack of clustering was due to hybrids, or that we were sampling a mixed population.
In 16S rRNA gene amplicon-based microbiota studies chloroplast sequences are normally treated as contaminants and steps are routinely taken to eliminate them39,40. 16S rRNA gene sequences are highly conserved, and the chloroplast version of this gene is not commonly used as a marker in phylogenetic studies due to poor resolution on lower taxonomic levels, although it has been used for deep branching lineages41. Thus, results from this part of the study should be interpreted with some caution. Relating the relative abundances of specific OTUs with the relative abundance of sequence reads classified as Poaceae, in individual Bale monkeys, only identified 9 relatively rare OTUs (Supplementary Table 9) with a significant correlation. Most likely the number of chloroplast reads recovered from a sample from a specific individual only reflects feeding behaviour on a very short time scale. Within each of the three habitats under study here, Individual Bale monkeys have very similar diets, and the observed composition of the GI microbiota in a given sample may not be an accurate reflection of short term feeding behaviour. On the population level, however, the trends observed in our chloroplast data make good sense in terms of feeding behaviour both across sampling sites and seasons (Fig. 6 and Supplementary Fig. 7).
In this study we have conducted the first analysis of the microbiota of the Ethiopian Bale monkey, one of very few primate species of bamboo specialists. We have shown that anthropogenic degradation of their habitats has the potential to cause an ecologically plastic response in their GI microbiota on the local scale. Whether this response is beneficial to the host is uncertain. We have further shown that the Bale monkey GI microbiota can have a strong seasonal signal, but that this signal may be partially disrupted by human encroachment. We also provide evidence that in spite of moving towards a more generalist lifestyle, the Bale monkey GI microbiota retains a distinct species signature relative to other Chlorocebus monkeys. Finally, we found that chloroplast sequence reads, produced as a by-product of GI microbiota 16S rRNA gene amplicon sequencing, can potentially provide useful information about the diversity of green plants in animal diets.
Methods
Study sites
The three Bale monkey populations in this study live in either the continuous Odobullu Forest (CBF = continuous bamoboo forest) or one of the two fragmented forests Afursa (NBF = no bamboo forest) and Kokosa (DBF = degraded bamboo forest) in the southern Ethiopian Highlands (Fig. 2, Table 1). CBF is dominated by bamboo, but also has tree-dominated sections, shrubland and occasional stretches of grassland42. DBF and NBF lie ~160 km west of CBF. DBF consists of degraded bamboo forest with large trees set amidst a matrix of human settlements, cultivated land, shrubland and grazing land. NBF is set upon a hilltop and is a mix of secondary forest, shrubland and Eucalyptus plantation with grass cover underneath, and it is surrounded by an anthropogenic matrix including cultivated lands, pastures, and human settlements15. Bamboo here has been nearly extirpated. Both DBF and NBF were dominated by bamboo forest only three decades ago, and although they are only 9 km apart they have been separated by human settlements and agriculture for many decades8.
Samples from vervet and grivet monkeys were collected from 5 sites in the Oromia and Southern Nations, Nationalities, and Peoples’ (SNNP) regions surrounding the current range of the Bale monkeys (Fig. 2, Table 1). All of these five sites represent habitats with a high degree of human contact.
Sampling and DNA extraction
Fecal samples from Bale monkeys were collected from October 2013 to June 2014 covering both the wet and dry seasons. In addition, we collected feces from vervet and grivet monkeys once at each of the five sites in 2016. We collected fresh feces immediately after defecation to avoid contamination. To minimize the risk of repeated fecal sampling from a single individual, we collected samples within a short period of time, for up to five consecutive days per month. For each sample approximately 2 grams of fecal material was transferred to a 15 ml sterile plastic vial with 8–10 ml 97% ethanol. Sample tubes were then stored at Addis Ababa University pending transport to the University of Oslo. DNA extraction from the fecal samples was carried out with the PowerSoil 96 well DNA isolation kit (MO BIO Laboratories Inc., Carlsbad, CA, USA). Permission to conduct this research was granted by the Ethiopian Wildlife Conservation Authority in compliance with the Convention on International Trade in Endangered Species of Wild Fauna and Flora (CITES). Fecal samples were collected non-invasively without harming or disturbing the animals. This study meets all animal care policies and adheres to the legal requirements of Ethiopia and Norway. It also complied with the ethical and legal requirements of the American Society of Primatologists Principles for the Ethical Treatment of Nonhuman Primates.
Illumina sequencing
Library preparation for Illumina sequencing of the V4 region of the 16S rRNA gene was carried out according to de Muinck et al.43, using the primers 515 f (GTGYCAGCMGCCGCGGTAA) and 806r (GGACTACNVGGGTWTCTAAT). Sequencing was carried out on an Illumina HiSeq. 2500 apparatus (Illumina, San Diego, CA, USA) using the 2 × 250PE rapid run mode. An overview of the sequencing libraries can be found in Supplementary Table 10.
The total raw number of 16S rRNA gene sequence reads after quality trimming and paired read merging was 30,092,962, and the mean per sample read number was 137,411 (±36,180 s.d.). Importantly, 1,349,220 (4.5%) of the sequence reads were classified as eukaryotic chloroplast DNA. These reads were removed before analysis of the bacterial/archaeal communities and analyzed separately.
Data processing and analyses
Low quality reads were trimmed and Illumina adapters were removed using Trimmomatic v0.3644 with default settings. Reads mapping to the PhiX genome (NCBI id: NC_001422.1) were removed using BBMap v36.0245. De-multiplexing of data based on the dual index sequences was carried out using custom scripts43. Internal barcodes and spacers were removed using cutadapt v1.4.146 and paired reads were merged using FLASH v1.2.1147 with default settings.
Further processing of sequence data was carried out using a combination of vsearch v2.0.348 and usearch v8.1.186149. Specifically, dereplication was performed with the ‘derep_fulllength’ function in vsearch with the minimum unique group size set to 2. Operational taxonomic unit (OTU) clustering, chimera removal, taxonomic assignment and OTU table building were carried out using the uparse pipeline50. Taxonomic assignment to the genus level was done against the RDP-15 training set. Classification was generally poor, with only 9.3% classified to the genus level with a confidence of 0.9 or higher and 63.3% classified to the phylum level with that degree of confidence. Between-sample differences in sequencing library size were normalized by common scaling51. After library scaling the data were filtered to retain only OTUs with at least 0.1% relative abundance (corresponding to 48 reads in the common scaled data) in at least one sample. These filtering criteria are used throughout unless other criteria are specifically stated. The total number of 97% bacterial/archaeal OTUs, in the raw data was 4289. After filtering, in order to eliminate OTU artefacts, the number dropped to 1298.
1,349,220 (4.5%) of the 16S rRNA gene sequences were classified as eukaryote chloroplasts. These sequences were further classified to 141 OTUs. For analysis of reads classified as chloroplast sequence, OTUs with fewer than a total of 1000 reads were removed from the data, as well as samples where chloroplast reads made up less than 1% of the total, leaving 106 (n = 55 for NBF, n = 32 for DBF, and n = 19 for CBF) samples and 20 OTUs that made up more than 99% of total chloroplast sequences. Reads in each sample were normalized to the original library size. Further classification of sequences was done manually by BLAST search52 against the GenBank nucleotide collection. We assigned the chloroplast OTUs to taxonomic groups based on the top hits from different publication events. The taxonomic level of these groups relied on the degree of consistency between the hits for each OTU. Accessions for uncultured organism clones were ignored.
All statistical analyses were done in R53. Permutational Multivariate Analysis of Variance Using Distance Matrices (PERMANOVA) and Analysis of Similarities (ANOSIM) were carried out using the’adonis’ and ‘anosim’ functions in the ‘vegan’ package, respectively, with Bray-Curtis dissimilarities and 10,000 permutations. Non-metric multidimensional scaling (NMDS) of Bray Curtis distance matrices was carried out using the ‘isoMDS’ function in the ‘MASS’ package. We used Spearman correlations, as implemented in the function ‘cor.test’ to test for associations between chloroplast abundances and specific OTUs, applying the Bonferroni correction for multiple hypothesis testing. To test whether populations at sampling sites that are closer together have more similar GI microbiotas than populations at sites further apart we computed mean Bray-Curtis distances between populations in all possible pairwise combinations of the eight sites. The resulting vector of distances was then tested for significant Spearman correlation with the geographical distances (km) between the corresponding pairs of sites. Exact tests for differences in means between two groups of negative binomially distributed counts were carried out using the edgeR package51,54. For these tests we did not use common scaled data, as library size differences are accounted for as part of the statistical algorithm. Specific filtering criteria were used for exact tests in order to focus on abundant OTUs present in many individuals, i.e. an OTU had to be observed at least 0.1% in at least one third of the samples in the data set being analyzed.
Data availability
All sequence data produced in this study are available at the NCBI Sequence Read Archive under BioProject ID PRJNA407723.
References
Sommer, F. & Backhed, F. The gut microbiota - masters of host development and physiology. Nat Rev Microbiol 11, 227–238 (2013).
De Filippo, C. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and ruralAfrica. P Natl Acad Sci USA 107, 14691–14696 (2010).
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559-+ (2014).
Ley, R. E. et al. Evolution of mammals and their gut microbes. Science 320, 1647–1651 (2008).
Muegge, B. D. et al. Diet Drives Convergence in Gut Microbiome Functions Across Mammalian Phylogeny and Within Humans. Science 332, 970–974 (2011).
Clayton, J. B. et al. Captivity humanizes the primate microbiome. P Natl Acad Sci USA 113, 10376–10381 (2016).
Haus, T. et al. Mitochondrial Diversity and Distribution of African Green Monkeys (Chlorocebus Gray, 1870). Am J Primatol 75, 350–360 (2013).
Mekonnen, A. et al. Newly Discovered Bale Monkey Populations in Forest Fragments in Southern Ethiopia: Evidence of Crop Raiding, Hybridization With Grivets, and Other Conservation Threats. Am J Primatol 74, 423–432 (2012).
Isbell, L. A., Pruetz, J. D. & Young, T. P. Movements of vervets (Cercopithecus aethiops) and patas monkeys (Erythrocebus patas) as estimators of food resource size, density, and distribution. Behav Ecol Sociobiol 42, 123–133, https://doi.org/10.1007/s002650050420 (1998).
Enstam, K. L. & Isbell, L. A. The guenons: Polyspecific associations in socioecological perspective. In Primates in perspective (eds Campbell, C. J. et al.) 277–300 (Oxford University Press, 2007).
Barrett, A. S., Barrett, L., Henzi, P. & Brown, L. R. Resource selection on woody plant species by vervet monkeys (Chlorocebus pygerythrus) in mixed-broad leaf savanna. Afr J Wildl Res 46, 14–21, https://doi.org/10.3957/056.046.0014 (2016).
Twinomugisha, D., Chapman, C. A., Lawes, M. J., Worman, C. O. D. & Danish, L. M. How does the golden monkey of the Virungas cope in a fruit-scarce environment? In Primates of Western Uganda (eds Newton-Fisher, N. E., Notman, H., Paterson, J. D. & Reynolds, V.) 45–60 (Springer, New York, 2006).
Grassi, C. Variability in habitat, diet, and social structure of Hapalemur griseus in Ranomafana National Park, Madagascar. Am J Phys Anthropol 131, 50–63, https://doi.org/10.1002/ajpa.20423 (2006).
Mekonnen, A., Bekele, A., Fashing, P. J., Hemson, G. & Atickem, A. Diet, Activity Patterns, and Ranging Ecology of the Bale Monkey (Chlorocebus djamdjamensis) in Odobullu Forest, Ethiopia. Int J Primatol 31, 339–362 (2010).
Mekonnen, A. et al. Impacts of habitat loss and fragmentation on the activity budget, ranging ecology and habitat use of Bale monkeys (Chlorocebus djamdjamensis) in the southern Ethiopian Highlands. Am J Primatol, https://doi.org/10.1002/ajp.22644 (2017).
Hess, M. et al. Metagenomic discovery of biomass-degrading genes and genomes from cow rumen. Science 331, 463–467, https://doi.org/10.1126/science.1200387 (2011).
Henderson, G. et al. Rumen microbial community composition varies with diet and host, but a core microbiome is found across a wide geographical range. Sci Rep-Uk 5 (2015).
Lambert, J. E. & Fellner, V. In Vitro Fermentation of Dietary Carbohydrates Consumed by African Apes and Monkeys: Preliminary Results for Interpreting Microbial and Digestive Strategy. Int J Primatol 33, 263–281 (2012).
Abbas, F. et al. Landscape fragmentation generates spatial variation of diet composition and quality in a generalist herbivore. Oecologia 167, 401–411, https://doi.org/10.1007/s00442-011-1994-0 (2011).
Chaves, O. M., Stoner, K. E. & Arroyo-Rodriguez, V. Differences in Diet Between Spider Monkey Groups Living in Forest Fragments and Continuous Forest in Mexico. Biotropica 44, 105–113 (2012).
Amato, K. R. et al. Habitat degradation impacts black howler monkey (Alouatta pigra) gastrointestinal microbiomes. ISME J 7, 1344–1353 (2013).
Barelli, C. et al. Habitat fragmentation is associated to gut microbiota diversity of an endangered primate: implications for conservation. Sci Rep-Uk 5 (2015).
Moeller, A. H. et al. Chimpanzees and humans harbour compositionally similar gut enterotypes. Nat Commun 3, 1179 (2012).
Huang, Q. et al. Fecal microbiota of lambs fed purple prairie clover (Dalea purpurea Vent.) and alfalfa (Medicago sativa). Arch Microbiol (2017).
Zhan, J. et al. Effects of alfalfa flavonoids extract on the microbial flora of dairy cow rumen. Asian-Australas J Anim Sci 30, 1261–1269 (2017).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227, https://doi.org/10.1038/nature11053 (2012).
Faith, J. J. et al. The long-term stability of the human gut microbiota. Science 341, 1237439, https://doi.org/10.1126/science.1237439 (2013).
Davenport, E. R. et al. Seasonal variation in human gut microbiome composition. PLoS One 9, e90731, https://doi.org/10.1371/journal.pone.0090731 (2014).
Amato, K. R. et al. The gut microbiota appears to compensate for seasonal diet variation in the wild black howler monkey (Alouatta pigra). Microb Ecol 69, 434–443, https://doi.org/10.1007/s00248-014-0554-7 (2015).
Sun, B. et al. Marked variation between winter and spring gut microbiota in free-ranging Tibetan Macaques (Macaca thibetana). Sci Rep 6, 26035, https://doi.org/10.1038/srep26035 (2016).
Maurice, C. F. et al. Marked seasonal variation in the wild mouse gut microbiota. Isme J 9, 2423–2434, https://doi.org/10.1038/ismej.2015.53 (2015).
Xue, Z. et al. The bamboo-eating giant panda harbors a carnivore-like gut microbiota, with excessive seasonal variations. MBio 6, e00022–00015, https://doi.org/10.1128/mBio.00022-15 (2015).
Gomez, A. et al. Temporal variation selects for diet-microbe co-metabolic traits in the gut of Gorilla spp. Isme J 10, 514–526 (2016).
Amato, K. R. et al. Phylogenetic and ecological factors impact the gut microbiota of two Neotropical primate species. Oecologia 180, 717–733 (2016).
Yildirim, S. et al. Characterization of the Fecal Microbiome from Non-Human Wild Primates Reveals Species Specific Microbial Communities. PLoS One 5 (2010).
Ransom-Jones, E., Jones, D. L., McCarthy, A. J. & McDonald, J. E. The Fibrobacteres: an important phylum of cellulose-degrading bacteria. Microb Ecol 63, 267–281 (2012).
Brucker, R. M. & Bordenstein, S. R. The hologenomic basis of speciation: gut bacteria cause hybrid lethality in the genus Nasonia. Science 341, 667–669, https://doi.org/10.1126/science.1240659 (2013).
Wang, J. et al. Analysis of intestinal microbiota in hybrid house mice reveals evolutionary divergence in a vertebrate hologenome. Nat Commun 6, 6440, https://doi.org/10.1038/ncomms7440 (2015).
Russell, J. A. et al. Uncovering symbiont-driven genetic diversity across North American pea aphids. Mol Ecol 22, 2045–2059, https://doi.org/10.1111/mec.12211 (2013).
Hanshew, A. S., Mason, C. J., Raffa, K. F. & Currie, C. R. Minimization of chloroplast contamination in 16S rRNA gene pyrosequencing of insect herbivore bacterial communities. J Microbiol Methods 95, 149–155, https://doi.org/10.1016/j.mimet.2013.08.007 (2013).
Manhart, J. R. Chloroplast 16S rDNA sequences and phylogenetic relationships of fern allies and ferns. Am Fern J 85, 182–192 (1995).
Mekonnen, A., Bekele, A., Hemson, G., Teshome, E. & Atickem, A. Population size and habitat preference of the Vulnerable Bale monkey Chlorocebus djamdjamensis in Odobullu Forest and its distribution across the Bale Mountains, Ethiopia. Oryx 44, 558–563, https://doi.org/10.1017/S0030605310000748 (2010).
de Muinck, E. J., Trosvik, P., Gilfillan, G. D., Hov, J. R. & Sundaram, A. Y. M. A novel ultra high-throughput 16S rRNA gene amplicon sequencing library preparation method for the Illumina HiSeq platform. Microbiome 5, 68, https://doi.org/10.1186/s40168-017-0279-1 (2017).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120, https://doi.org/10.1093/bioinformatics/btu170 (2014).
Bushnell, B. BBMap short read aligner. https://sourceforge.net/projects/bbmap/ (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17 (2011).
Magoc, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963, https://doi.org/10.1093/bioinformatics/btr507 (2011).
Rognes, T. F. T., Nichols, B., Quince, C., Mahé, F. VSEARCH: a versatile open source tool for metagenomics. PeerJ Preprints 4 (2016).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461, https://doi.org/10.1093/bioinformatics/btq461 (2010).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10, 996–998, https://doi.org/10.1038/nmeth.2604 (2013).
McMurdie, P. J. & Holmes, S. Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput Biol 10, e1003531, https://doi.org/10.1371/journal.pcbi.1003531 (2014).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J Mol Biol 215, 403–410, https://doi.org/10.1016/S0022-2836(05)80360-2 (1990).
R Core Team. R: A language and environment for statistical computing (R Foundation for Statistical Computing Vienna, Austria, 2016).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140, https://doi.org/10.1093/bioinformatics/btp616 (2010).
Acknowledgements
This study was funded by Research Council of Norway grant 230796/F20, Rufford Small Grants 11727–1 and the People’s Trust for Endangered Species. We would like to thank the Department of Zoological Sciences of Addis Ababa University for logistical support. We are grateful to the Ethiopian Wildlife Conservation Authority for granting permission to conduct this study. We thank Mengistu Birhan and Mamar Dilnesa for their valuable assistance with the field research. We would also like to thank the local guides Firdie Sultan, Jemal Kedir, Mudie Kedir, Matiyos Yakob, Omer Hajeleye and Hassen Wolle.
Author information
Authors and Affiliations
Contributions
P.T., E.K.R., E.D.M. and A.M. developed the study concept and design. A.M. and A.M. carried out field work. P.T. and E.D.M. carried out lab work. P.T., E.K.R., E.D.M. and A.M. carried out data analysis and wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Trosvik, P., Rueness, E.K., de Muinck, E.J. et al. Ecological plasticity in the gastrointestinal microbiomes of Ethiopian Chlorocebus monkeys. Sci Rep 8, 20 (2018). https://doi.org/10.1038/s41598-017-18435-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-18435-2
This article is cited by
-
Comparative analysis of gut microbiota between common (Macaca fascicularis fascicularis) and Burmese (M. f. aurea) long-tailed macaques in different habitats
Scientific Reports (2023)
-
Gut microbiota of ring-tailed lemurs (Lemur catta) vary across natural and captive populations and correlate with environmental microbiota
Animal Microbiome (2022)
-
Microbial rewilding in the gut microbiomes of captive ring-tailed lemurs (Lemur catta) in Madagascar
Scientific Reports (2022)
-
The gut microbiome as an indicator of habitat disturbance in a Critically Endangered lemur
BMC Ecology and Evolution (2021)
-
Identifying molecular pathways and candidate genes associated with knob traits by transcriptome analysis in the goose (Anser cygnoides)
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.