Abstract
The abuse of antibiotics for disease treatment has led to the emergence of multidrug resistant bacteria. Antimicrobial peptides, found naturally in various organisms, have received increasing interest as alternatives to conventional antibiotics because of their broad spectrum antimicrobial activity and low cytotoxicity. In a previous report, Macropin, isolated from bee venom, exhibited antimicrobial activity against both gram-positive and negative bacteria. In the present study, Macropin was synthesized and its antibacterial and anti-biofilm activities were tested against bacterial strains, including gram-positive and negative bacteria, and drug resistant bacteria. Moreover, Macropin did not exhibit hemolytic activity and cytotoxicity to keratinocytes, whereas Melittin, as a positive control, showed very high toxicity. Circular dichroism assays showed that Macropin has an α-helical structure in membrane mimic environments. Macropin binds to peptidoglycan and lipopolysaccharide and kills the bacteria by disrupting their membranes. Moreover, the fractional inhibitory concentration index indicated that Macropin has additive and partially synergistic effects with conventional antibiotics against drug resistant bacteria. Thus, our study suggested that Macropin has potential for use of an antimicrobial agent for infectious bacteria, including drug resistant bacteria.
Similar content being viewed by others
Introduction
Since the discovery of penicillin, many antibiotics have been developed and used to treat infectious diseases caused by bacteria. Antibiotics kill bacteria by inhibiting cell wall synthesis, protein synthesis, or DNA synthesis. When an antibiotic is used in the clinic, consideration should be given to characteristics such as the antimicrobial range and mechanism of action. Nonetheless, the indiscriminate use of antibiotics has resulted in bacteria developing resistance to antibiotics. The number of resistant bacterial strains has increased, and some bacteria, referred to as superbugs, have emerged with resistance to most antibiotics, thus posing a serious health risk1. Therefore, there is a need to develop new drugs that are less likely to induce resistance to treat drug-resistant bacteria. Possible alternatives to conventional antibiotics are antimicrobial peptides (AMPs), which are part of the innate immune response2. AMPs have a broad spectrum of antimicrobial activity against bacteria, including multidrug resistant bacteria.
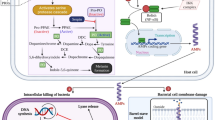
Generally, AMPs are found in nature and participate in host defense. AMPs are small proteins of between 12 and 50 amino acid residues. The secondary structures of AMPs include α-helices, β-sheets, extended structures, and loops3. In addition, AMPs tend to be amphipathic; i.e., they contain both hydrophilic and hydrophobic regions4,5. These characteristics promote the interaction of AMPs with the bacterial cell membrane, which comprises negatively charged lipids, such as phosphatidylglycerol (PG), phosphatidylethanolamine (PE), and cardiolipin (CL). Moreover, gram-negative bacteria contain lipopolysaccharide (LPS) in their outer membrane, and gram-positive bacteria have a peptidoglycan outside the bacterial plasma membrane, forming the cell wall. Different mechanisms of AMPs’ interaction with the membrane have been proposed and include the barrel stave model, the toroidal model, and the carpet model3. The barrel-stave model suggests that AMPs form pores by binding to the membrane surface. A hydrophilic region of the AMPs binds to the membrane’s lipid and hydrophobic regions, thus forming a channel6. The toroidal model is similar to the barrel-stave model; however, AMPs always contact the phospholipid head groups of the membrane. Peptides are then inserted into the membrane to form a pore7. In the carpet model, when a membrane reaches the threshold concentration of AMPs, it is disintegrated by the accumulation of peptides. AMPs then penetrate the membrane lipid layer to form a pore8,9. In addition, some AMPs inhibit DNA and protein synthesis10.
In many studies, treatment with two or more antibiotics revealed that this approach enhances the therapeutic effect, and it is now used extensively to treat diseases. For example, aminoglycoside and beta-lactam antibiotics have been tested in combination11 and with a β-lactamase inhibitor12,13. Combination therapy has a broader spectrum of bactericidal activity compared with that of a single antibiotic, reduces side effects, and decreases the risk of induced resistance. Recently, combinations of conventional antibiotics and AMPs were used to treat certain fungal and bacterial infections14,15,16.
In a previous study, a novel antimicrobial peptide named Macropin, with short α-helical structure, was isolated from venom of the solitary bee Macropis fulvipes (Hymenoptera: Melittidae) and was reported to have antimicrobial activity17. Bee venom, which comprises a complex mixture of active peptides, has been used for therapeutic purposes in various diseases and has immunological functions18.
In the present study, Macropin was tested for its antimicrobial activity against gram-negative and positive bacteria, including drug resistant bacteria. We also tested its cytotoxicity and hemolysis activities compared with those of melittin. Melittin is a well-known antimicrobial peptide from bee venom; however, it is highly cytotoxic19. The mechanism of action of Macropin was examined by n-phenyl-1-naphthylamine (NPN) uptake measurement and a 3,3′-dipropylthiadicarbocyanine iodide [DiSC3(5)] assay. Low vacuum scanning electron microscopy (SEM) was used to observe membrane destruction. Moreover, the synergistic effect of Macropin with conventional antibiotics was evaluated in a combination assay and by flow cytometry. Our results indicated that Macropin could potentially be used as an antimicrobial agent.
Results
Peptide synthesis
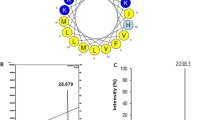
The Macropin sequence, theoretically calculated and observed molecular weights, retention time, hydrophobicity, hydrophobic moment, and net charge are summarized in Table 1. The observed molecular weight was consistent with the calculated molecular weight, indicating that the peptide was synthesized successfully. The wheel diagram represents the α-helical structure of Macropin (Fig. S1). The identity of the synthetic peptide was confirmed by reverse-phase high performance liquid chromatography (RP-HPLC) on a C18 column. The molecular weight of the peptide was verified by mass spectrometry (Fig. S2).
Minimum inhibitory concentration of macropin against microorganisms
The antimicrobial activities of AMP Macropin against gram-positive and negative bacteria are summarized in Table 2. Macropin showed antimicrobial activity against both gram-positive and negative bacteria in the concentration range of 3.13 μM to 25 μM. Macropin and nine antibiotics were then analyzed for their antimicrobial activity against drug-resistant bacteria. In the range of 2 to 32 μM, ciprofloxacin and levofloxacin showed activity against resistant S. aureus strains, whereas the other antibiotics showed no activity up to 128 μM. In the case of drug-resistant P. aeruginosa strains, tobramycin and gentamicin showed antimicrobial activity at concentrations of 32 μM and 64 μM (Table 3). Compared with the antibiotics, Macropin had stronger antimicrobial effects against drug-resistant bacterial strains.
Cytotoxicity and hemolysis
The cytotoxicity of Macropin was tested on HaCaT cells (keratinocytes) and Raw 264.7 cells (macrophages) using the MTT (3-(4,5-dimethylthiazol-2-Yl)-2,5-diphenyltetrazolium bromide) assay. At a concentration of 25 μM, Macropin produced cell survival rates of 82% and 41% in HaCaT and Raw cells, respectively (Fig. 1A and B). Melittin, a negative control, showed 100% toxicity at 12.5 μM in both cell lines. Hemolysis by Macropin was tested in an 8% suspension of red blood cells (RBCs) to test its toxicity to mammalian cells. The hemolysis rate of Macropin was approximately 5% at 25 μM. As negative control, Melittin was used because it is known to damage the membrane of RBCs. At 25 μM, melittin showed approximately 93% hemolytic activity (Fig. 1C). Thus, Macropin showed antimicrobial activity and low toxicity.
Cytotoxicity and Anti-biofilm Activity of Macropin. Cytotoxicity of peptide concentrations from 0 to 50 μM against (A) HaCaT cells, and (B) Raw 264.7 cells. After 24 h of incubation with the peptides, cytotoxicity was measured using an MTT (3-(4,5-dimethylthiazol-2-Yl)-2,5-diphenyltetrazolium bromide) assay. (C) Hemolytic activity of the peptide against red blood cells. The release of hemoglobin was measured using a microplate reader at 414 nm. (D) Biofilm formed by S. aureus ATCC 25923, P. aeruginosa ATCC 27853, and E. coli ATCC 25922. (E) Inhibition of biofilm formation by Macropin against the microorganisms. Each well contained 50 μL of the peptide and 50 μL of 5 × 105 CFU/mL suspension of bacteria. The biofilms were then stained with crystal violet and absorbance was measured at 595 nm.
Anti-biofilm activity
A biofilm is a social community of bacteria that produces multiple virulence factors. Bacteria in biofilms become more resistant to antibiotics, and biofilm-associated infections are difficult to treat. The degree of biofilm formation by the bacteria in the absence of the peptide was measured. Consequently, S. aureus ATCC 25923 and P. aeruginosa ATCC 27853 formed biofilms well (Fig. 1D). Next, we measured the inhibition of biofilm formation by Macropin. The maximal percentage of biofilm inhibition was 88%, 92%, and 84% against S. aureus, P. aeruginosa, and E. coli, respectively (Fig. 1E). We then evaluated the minimal biofilm inhibition concentration (MBIC) against drug-resistant strains of S. aureus and P. aeruginosa. In the concentration range of 12.5 μM to 50 μM, Macropin inhibited biofilm formation by S. aureus and P. aeruginosa (Table 4).
Structure of the peptide and circular dichroism spectroscopy
The secondary structure of Macropin in a membrane mimic environment was investigated using CD spectroscopy. In 10 mM sodium phosphate buffer (mimicking the aqueous environment), Macropin had a random coil structure. SDS and 2,2,2-trifluoroethanol (TFE) are widely used to mimic the cell membrane environment20. In 30 mM SDS (mimicking the microbial membrane environment) and 50% TFE (mimicking the hydrophobic environment of the microbial membrane), Macropin showed an α-helical conformation characterized by a positive band at 190 nm and negative bands at 208 and 222 nm (Fig. 2B). These results are consistent with the prediction that Macropin has an α-helical structure17, obtained using Mobyle@RPBS (Fig. 2A).
The structure analysis of the Macropin. (A) Three-dimensional structure simulations of Macropin. (B) Macropin formed an α-helical structure in 10 mM sodium phosphate buffer (pH 7.2), 30 mM SDS (mimicking microbial membrane environment), and 50% 2,2,2-trifluoroethanol (TFE) (mimicking the hydrophobic environment of the microbial membrane). The peptide concentration was fixed to 40 μM.
Large unilamellar vesicle (LUV) aggregation
Liposome aggregation indicates a peptide-lipid interaction, and liposome turbidity served as a measure of the degree of aggregation caused by Macropin to PE:PG, phosphatidyl choline (PC):cholesterol (CH), and PC:CH: sphingomyelin (SM) systems. Macropin induced LUV aggregation in PE:PG, which is similar to a bacterial membrane. PE:PG turbidity was increased up to an absorbance of 0.3 at a peptide/lipid (P/L) ratio of 0.2 (Fig. 3A). By contrast, Macropin did not induce LUV aggregation of the PC:CH and PC:CH:SM systems.
Aggregation of large unilamellar vesicles (LUVs) and Binding of the peptide to cell wall components. (A) A solution containing Macropin was added to 400 μM PC:CH (2:1, w/w), PE:PG (7:3, w/w), or PC:CH:SM (1:1:1, w/w). Liposome aggregation was monitored based on changes in the absorbance at 405 nm. PE, phosphatidylethanolamine; PG, phosphatidylglycerol; PC, phosphatidylcholine; CH; cholesterol; SM, sphingomyelin. (B) The peptide binds to the peptidoglycan of S. aureus. Lane 1: 5 μg of Macropin, lane 2: pellet of the mixture (5 μg of Macropin with 100 μg of peptidoglycan), lane 3: the supernatant of the mixture (5 μg of Macropin with 100 μg of peptidoglycan. (C) Binding affinity of the peptide for lipopolysaccharide (LPS), as measured using circular dichroism (CD) spectroscopy. Macropin (40 μM) was measured in the presence of 0.1% LPS. (D) Binding of Macropin to the peptidoglycan or LPS of the cell wall component was confirmed via antibacterial activity.
Binding of Macropin with cell wall components
To assess the ability of Macropin to bind to cell wall components, the peptidoglycan of S. aureus and the LPS of P. aeruginosa were used in binding assays. In sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) analysis, Macropin bound to the peptidoglycan was detected in the pellet of the mixture, but not in the supernatant (Fig. 3B). This result indicated that Macropin binds to the peptidoglycan, a cell wall component of gram-positive bacteria. In addition, we used CD spectra to study the conformational change caused by the interaction between the peptide and LPS. The CD spectra showed that Macropin has an α-helical conformation when interacting with LPS, a cell wall component of gram-negative bacteria (Fig. 3C). In previous studies, Macropin was found to bind to peptidoglycan and LPS. To confirm previous findings, Macropin’s antibacterial activity was measured by adding peptidoglycan into an S. aureus suspension and LPS into a P. aeruginosa suspension. As controls, only peptidoglycan or LPS was added to the culture medium of the bacteria. In the control, the bacteria grew constantly; however, in the experimental samples (peptide added), the absorbance of the mixture increased at 200 μg/mL of peptidoglycan or LPS (Fig. 3D). Blocking of the antibacterial activity of Macropin by peptidoglycan and LPS showed that peptide binds directly to a cell wall component.
Membrane permeability
An NPN uptake assay was performed to evaluate the ability of Macropin to permeabilize the outer membrane. The hydrophobic fluorescent probe NPN emits a weak fluorescent signal in an aqueous environment, but becomes strongly fluorescent in a hydrophobic environment when the cell membrane is disturbed21. Here, Macropin permeabilized the membrane in a dose-dependent manner in just a few minutes. After 5 min, NPN uptake increased to 60% in S. aureus and to 65% in P. aeruginosa at 4 × the MIC of Macropin. Thus, Macropin could permeabilize the outer membrane effectively (Fig. 4A and B).
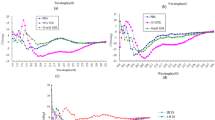
Effect of the peptide on the outer membrane and cytoplasmic membrane. Outer membrane permeabilization induced by Macropin as detected by n-phenyl-1-naphthylamine (NPN) uptake in (A) S. aureus ATCC 25923 and (B) P. aeruginosa ATCC 27853. The cytoplasmic membrane potential variation of (C) S. aureus and (D) P. aeruginosa treated by Macropin, and an assessment of the release of membrane potential sensitive dye DiSC3(5). The red arrows indicate the peptide-treated results, and control consisted of the buffer only (5 mM HEPES with 20 mM glucose and 100 mM KCl). (E) Flow cytometry analysis. S. aureus ATCC 25923 and P. aeruginosa ATCC 27853 were treated by 1 × the minimum inhibitory concentration (MIC) of Macropin for 1 h, and then stained with propidium iodide (PI). Macropin application resulted in increased fluorescence of 98.5% and 78.9% of S. aureus and P. aeruginosa cells, respectively.
The depolarization of the bacterial cytoplasmic membrane by Macropin was studied using the membrane potential-sensitive dye DiSC3(5). When the cytoplasmic membrane is depolarized and disrupted, DiSC3(5) is released into the medium and the fluorescence increases22. After stabilizing the bacteria with DiSC3(5) for 10 min, we added various concentrations of Macropin into each well of the plate. The fluorescence of DiSC3(5) increased compared with the control. As soon as we added the peptide, the fluorescence intensity increased by up to 70 arbitrary units (a.u.) in S. aureus and 100 a.u. in P. aeruginosa, whereas the fluorescence intensity of bacteria with DiSC3(5) without the peptide was 0 to 10 (Fig. 4C and D).
Flow cytometry
Propidium iodide (PI) is an intercalating agent that binds to nucleic acids and is used evaluate to cell viability. PI stains nucleic acid after cell membrane disruption, resulting in an increase in fluorescent signals. Thus, cell membrane damage was measured by PI staining together with flow cytometry. In the absence of Macropin, 93.1% of S. aureus and 99% of P. aeruginosa cells showed no staining with PI, indicating intact cell membranes (viable cells). After peptide treatment, 98.5% and 78.9% of the cells showed PI fluorescence in S. aureus and P. aeruginosa, respectively (Fig. 4E).
Low vacuum scanning electron microscopy (SEM)
This method was used to visualize the cell morphology after treatment of S. aureus and P. aeruginosa with Macropin. The control cells without Macropin showed a smooth surface. In contrast, treatment with 1 × the MIC of Macropin damaged the cell membrane. The membrane surface of S. aureus treated with Macropin showed blebs and damage. Part of the surface showed atrophy, destruction, and shrinking in P. aeruginosa treated with the peptide for 30 min (Fig. 5). These results suggested that Macropin induced the formation of membrane blebs on the surface of bacteria.
Low vacuum scanning electron microscopy (SEM) micrographs of peptide treated S. aureus and P. aeruginosa. The control bacteria without peptide were round and intact. S. aureus (ATCC 25923) and P. aeruginosa (ATCC 27853) were incubated with 1 × the minimum inhibitory concentration (MIC) of Macropin for 30 min. Macropin-induced changes in the bacterial membrane were observed.
Synergistic effect of Macropin and antibiotics
Combination therapy is used widely to treat diseases because of its better effect and lower costs. A combination of AMPs and antibiotics was found to have a better bactericidal effect than AMP or antibiotics alone. The S. aureus and P. aeruginosa strains used here are resistant to antibiotics; however, the addition of Macropin to the antibiotics improved the inhibition of bacterial growth. Combination therapy with Macropin and antibiotics showed antibacterial activity at a lower dose of the peptide or antibiotics compared with the peptide or antibiotics alone (Tables S1 and S2). The fractional inhibitory concentration (FIC) index indicated partial synergy and additive effects. When the S. aureus strains were treated with Macropin combined with gentamycin, tobramycin, ciprofloxacin, levofloxacin, piperacillin, or oxacillin, the FIC index revealed an additive effect. The combination of Macropin and oxacillin showed partial synergy (FIC index 0.52) against S. aureus 949987 (Fig. 6A). The P. aeruginosa strains treated with Macropin in combination with antibiotics showed the highest additive effect. In P. aeruginosa 3320, the combinations of Macropin with gentamycin, tobramycin, piperacillin, or oxacillin showed synergy (FIC index 0.52; Fig. 6B). Generally, the antibiotics and Macropin showed synergistic or additive effects.
The activity of the peptide combined with traditional antibiotics against S. aureus and P. aeruginosa strains. Macropin combined with ciprofloxacin, levofloxacin, tobramycin, gentamicin, oxacillin, or piperacillin showed partial synergy or additive effects against (A) S. aureus strains and (B) P. aeruginosa strains. Especially, Macropin combined with oxacillin (in S. aureus 949987) and Macropin with piperacillin (in P. aeruginosa 3320) showed a partial synergy effect (FIC index 0.52). Fractional inhibitory concentration (FIC) indices were interpreted as follows: FICi ≤ 0.5 indicates synergy, 0.5 < FICi ≤ 1.0 indicates additive, 1 < FICi ≤ 4 indicates indifference, and FICi > 4 indicates antagonism. (C) The bacteriostasis rate represents the inhibition activity. Oxacillin and Macropin were used individually or in combination against S. aureus 949987. (D) Piperacillin and Macropin were used individually or in combination against P. aeruginosa 3320.
Bactericidal activity of the synergistic combinations
The combinations of Oxacillin and Macropin (for S. aureus) and Piperacillin and Macropin (for P. aeruginosa) increased the bacteriostasis rate rapidly up to 60 min, and approximately 90% of S. aureus 949987 and P. aeruginosa 3320 were killed at 180 min. The bacteriostasis rate increased gradually for bacteria treated with only the antibiotic. In the combination group treated with Macropin and antibiotics at less than the active concentration, the bacteriostasis rate increased more rapidly compared with rate observed on treatment with only antibiotics (Fig. 6C and D). Thus, Macropin accelerated the bacteriostasis rate; i.e., the number of living bacteria decreased.
Inhibition of biofilm formation using the synergistic combinations
The anti-biofilm activity of Macropin, single antibiotics, and the combination groups were assessed using the MBIC. The MBIC values of Macropin against S. aureus and P. aeruginosa were 50 and 25 μM, respectively. All antibiotics used in experimental group showed no anti-biofilm activity at concentrations above 64 μg/mL. However, most combinations of the peptide and antibiotics improved the inhibition of biofilm formation. The MBIC value of the peptide and antibiotics decreased in combination groups. Macropin with oxacillin showed an additive effect (FIC index 0.63) against S. aureus. In the case of P. aeruginosa, the peptide with piperacillin, tobramycin, and gentamicin revealed strong synergistic effects, with FIC indices of 0.34, 0.26, and 0.63, respectively (Table 5).
Flow cytometry assessment of the synergistic groups
The synergistic effects of the peptide and antibiotics were evaluated by flow cytometry. In the control, the bacterial cells were not stained, but in the experimental group, peptide treatment increased the PI fluorescence to 96.7% and 89.25% in S. aureus 949987 and P. aeruginosa 3320, respectively. When antibiotic treatment had no antimicrobial effect on the bacteria, the fluorescence of PI was only present in 13.8% of S. aureus and 2.9% of P. aeruginosa cells. In contrast, when the bacterial cells were treated with the combination of the peptide and the antibiotics, the percentage of PI staining increased. The combination of Macropin with oxacillin in S. aureus yielded PI staining in 97.5% of the cells. In case of P. aeruginosa, combinations of Macropin with piperacillin, tobramycin, or gentamicin were examined. The results revealed 95.3%, 88.7%, and 86% of PI, respectively. These data indicated that Macropin killed bacteria by disrupting the cell membrane, and the combination of Macropin with certain antibiotics had synergistic effects (Fig. 7).
The combined effect as analyzed using flow cytometry analysis. (A) S. aureus 949987 and (B) P. aeruginosa 3320 were treated with the peptide, antibiotics, or both. (A) No peptide (5.1%). (A-1) oxacillin 64 μg/mL (13.8%), (A-2) Macropin 3.13 μM (96.7%), (A-3) Macropin 1.56 μM plus oxacillin 2 μg/mL (97.5%), (B) no peptide (7.4%), (B-1) piperacillin 64 μg/mL (2.9%), (B-2) Macropin 3.13 μM (89.2%), (B-3) Macropin 1.56 μM plus piperacillin 2 μM (95.3%).
In vivo bactericidal effect of Macropin on S. aureus-infected mice
We further tested whether Macopin has therapeutic activity in an air pouch model. Anesthetized BALB/c mice were infected with S. aureus in the presence or absence of Macropin, and tissues were removed and cultured on an agar plate. The mice in the group treated with Macropin (1 mg/kg) showed a decrease in the number of cultured colonies compared to the mouse group treated with S. aureus only (Fig. 8). Thus, Macropin has antibacterial activity in vivo.
Effect of Macropin on S. aureus ATCC 25923 infected in the air pouch model in vivo. (A) Bacteria were cultured on agar plate overnight on 1, 2, 3, 4, 7 day after injection. (B) Bacterial colony is indicated as percentage in BALB/c mouse skin caused by S. aureus infection in the presence or absence of Macropin.
Discussion
The prevalence of drug-resistant bacteria is increasing worldwide; therefore, there is a need to develop novel antimicrobial drugs. Additionally, diseases associated with biofilms, such as chronic infections and medical-appliance-related infections, are difficult to treat, which cause problems to public health that require novel solutions23. AMPs, a vital component of the innate immune system, represent novel therapeutic agents to treat microbial infections, because they are effective against pathogenic microorganisms and are less likely to induce drug resistance24. Although AMPs are attracting attention as a new drug, they have the disadvantage of high cost. Moreover, Melittin, a well-known peptide with strong antimicrobial activity from the bee venom, is cytotoxic at a concentration lower than the active concentration25. In a previous report, the AMP Macropin, from bee venom, was identified and reported to be composed of 13 amino acids17. Thus, Macropin is shorter than Melittin, which makes it more economical to synthesize. Therefore, in this study, we were interested in the possibility of Macropin being used drug.
Antimicrobial assays showed that Macropin possesses antimicrobial activity against both gram-positive and negative bacteria, including drug-resistant bacteria, especially S. aureus and P. aeruginosa. Furthermore, an assay of conventional antibiotics against resistant bacteria was conducted. Most of the traditional antibiotics used were ineffective against resistant bacteria. By contrast, Macropin showed antimicrobial activity against the resistant bacteria. Macropin’s hemolytic activity on RBCs and cytotoxicity toward keratinocytic (HaCaT) and macrophagic cells (Raw 264.7) were lower than those of melittin, which is a representative peptide derived from bees26. The active concentration of Macropin is from 3.13 μM to 6.25 μM, similar to that of Melittin, and the cytotoxicity effect of Macropin was reported at a concentration of 25 μM, which is about 8-times the active concentration. On the other hand, Melittin showed hemolysis and cytotoxicity at a concentration lower than the active concentration of 3.13 μM. Macropin displayed antimicrobial activity at a concentration similar to the active concentration of Melittin and was not cytotoxic up to about 10-times the active concentration. In addition, Macropin could inhibit biofilm formation in S. aureus and P. aeruginosa strains, including resistant bacteria, at concentrations ranging from 12.5 μM to 50 μM. Thus, we demonstrated that Macropin is a suitable antimicrobial agent and must be tested against relevant infections in humans.
Most AMPs have one of four types of secondary structures: an α-helix, a β-sheet, an extended structure or a loop type structure. CD spectra are used to determine the secondary structure of a peptide. CD assays performed in the present study showed that Macropin appeared as a random coil in aqueous solution, whereas in membrane-mimicking environments, such as SDS and TFE solutions, it formed an α-helix. LUVs are often used as a model of the bacterial membrane to study the interaction between a cationic peptide and the membrane. Prokaryotic and eukaryotic cells differ in the composition of their cellular membranes. For example, PC and PE have no net charge, whereas SM and cholesterol are neutral (not charged). PG and PS have a net negative charge27. The cell membrane of bacterial pathogens comprises PG and PS. In contrast, the membrane of human cells, such as erythrocytes, is enriched in PC, PE, and SM. Macropin induced aggregation of the PE:PG liposome (in the bacterial membrane) but not of the PC:CH (erythrocyte) and PC:CH:SM (mammalian cell) liposomes. These results correlated with the hemolytic and cytotoxicity properties of Macropin, suggesting that Macropin has cell selectivity for negative bacterial membranes compared to mammalian cell membranes.
AMPs are attracted to bacterial membranes via electrostatic bonding between the cationic peptide and the negatively charged membrane surface. LPS is the major component on the outer leaflet of the outer membrane of gram-negative bacteria28. The binding of LPS by AMPs, which plays an important role in killing bacteria, might proceed via an amphipathic sequence. AMPs that can bind to LPS and disrupt the membrane have attracted increased interest in terms of drug development29. Gram-positive bacteria possess structural components such as a thick peptidoglycan layer and lipoteichoic acid. These components perform permeability barrier functions, and the ability of a peptide to bind to such a cell wall component is a prerequisite for antibacterial activity. After a peptide binds to a cell wall component through electrostatic interaction, an AMP could be inserted into the bacterial membrane30. While binding to the cell wall, the peptide comes into contact with the cytoplasmic membrane. Macropin binds to the peptidoglycan of S. aureus and to the LPS of P. aeruginosa. To examine the mechanism of action of Macropin, outer membrane permeability and cytoplasmic membrane depolarization assays were performed. Macropin induced an increase in NPN and DiSC3(5)-related fluorescence. Our experiments confirmed that Macropin disturbs the outer membrane and depolarizes the cytoplasmic membrane. Furthermore, flow cytometry analysis indicated that Macropin can destroy a bacterial membrane, allowing PI to bind nucleic acids. Observation of morphological changes using low-vacuum SEM confirmed that the Macropin caused damage to the bacterial membrane. This agreed with a previous study that showed that the bacterial membrane is disrupted by AMPs such as LL3731 and melittin25, and that this action causes a loss of functional integrity of the membrane32. In particular, melittin is a peptide derived from bees and exerts a bactericidal effect by damaging the bacterial membrane. In this paper, the superiority of Macropin compared with melittin showed that the activity of Macropin is similar but its cytotoxicity is lower.
Traditional antibiotics exert their antimicrobial activity by inhibiting cell wall synthesis and the transcription and replication of DNA. However, mutations in the components of these mechanisms result in drug-resistant bacteria33,34. Compared with antibiotics, AMPs have several unique characteristics, such as a wide range of antibacterial activities against antibiotic-resistant pathogens and a relatively low rate of mutation induction in bacteria35. Thus, AMPs are promising antimicrobial agents that can be used to treat several infectious diseases, either alone or in combination with traditional antibiotics. Using killing tests and antimicrobial activity assessments of the combination group, we observed synergistic effects of the combination of Macropin with traditional antibiotics. The results of this study showed that the Macropin-Oxacillin and Macropin-Piperacillin combinations have an additive synergy for S. aureus 94998 and P. aeruginosa 3320, respectively. Furthermore, Macropin and antibiotic combinations inhibited biofilm formation of drug-resistant bacteria. The Macropin-Oxacillin and Macropin-Piperacillin combinations displayed synergistic activity against biofilm formation by S. aureus and P. aeruginosa, respectively. The decrease in MBIC concentration of the peptide combined with antibiotics indicated that it can inhibit biofilm formation successfully36. The synergistic action of the peptide and antibiotic offers an alternative strategy to treat biofilm-related infections. The mechanisms of the synergistic effect of the combination are still unclear. One of the proposed mechanisms is that the AMP increases the permeability of the cell membrane and facilitates the entry of antibiotics into the cytoplasm to access the intracellular target37. Thus, a combination of the peptide and traditional antibiotics can improve their antimicrobial activity, which suggested that combination therapy could be used against resistant bacteria.
In conclusion, Macropin binds to negatively charged components, such as peptidoglycan or LPS, of bacterial membranes and then kills the bacteria through membrane disruption and cell penetration. Macropin forms an α-helix structure in a bacteria-mimicking environment. In addition, Macropin can inhibit the growth of drug-resistant bacteria and biofilm formation, but has little or no hemolytic activity or cytotoxicity toward mammalian cells at microbiologically effective concentrations. To verify the effect of Macropin in vivo, we performed an air pouch model experiment in mice, which demonstrated that Macropin has antibacterial activity in vivo. Furthermore, combinations of the peptide and antibiotics have synergistic effects against certain drug-resistant bacteria. These effects can increase antimicrobial activity and help delay the emergence of drug resistance. Consequently, Macropin is a promising candidate for an antimicrobial drug to treat bacterial infection and as part of a combination therapy with antibiotics against multi-antibiotic-resistant bacteria.
Materials and Methods
Materials
Rink amide 4-mehylbenzhydrylamine resin, fluoren-9-ylmethoxycarbonyl (Fmoc), amino acids, and other reagents for peptide synthesis were purchased from Calbiochem-Novabiochem (La Jolla, CA, USA). L-α-phosphatidylethanolamine (PE), sphingomyelin (SM), cholesterol (CH), L-α-phosphatidylglycerol (PG), and egg yolk L-α-phosphatidylocholine (PC) were obtained from Avanti Polar Lipids (Alabaster, AL, USA). LPS (from P. aeruginosa), peptidoglycan (from S. aureus), 3,3′-Dipropylthiadicarbocyanine iodide [DISC3(5)], n-phenyl-1-naphtylamine (NPN), propidium iodide (PI), ciprofloxacin, levofloxacin, gentamicin, kanamycin, tobramycin, cefotaxime, oxacillin, erythromycin, ampicillin, piperacillin, and vancomycin were obtained from Sigma-Aldrich (St Louis, MO, USA). Staphylococcus aureus ATCC 25923, Staphylococcus aureus ATCC 29213, Escherichia coli ATCC 25922, Pseudomonas aeruginosa ATCC 27853, and Pseudomonas aeruginosa ATCC 15692 were obtained from the American Type Culture Collection (ATCC). Listeria monocytogenes KCTC 3710 was obtained from the Korea Collection for Type Cultures (KCTC). Staphylococcus aureus 95085, 691054, 254348, 254422, 547582, 771687, and 949987 were antibiotic resistant strains isolated from hospital patients with cholelithiasis. Pseudomonas aeruginosa 3320, 3318, 1034, 3241, 1162, and 3399 were antibiotic resistant strains isolated from patients with otitis media in Chonnam National University hospital. The patients provided informed consent for the use of their clinical isolate.
Peptide synthesis
The peptide was synthesized according to a solid-phase-9-fluorenylmethoxycarbonyl1 (Fmoc) method on a Rink amide 4-methylbenzhydrylamine resin using a Liberty microwave peptide synthesizer (CEM Co. Matthews). Then, 0.1 M N-hydroxy benzotriazole in piperidine and dimethylformamide (DMF), 0.45 M 2-(1H-benzotriazole-1-yil)-1,1,3,3-tetramethyluroniumhexafluorophosphate in DMF and 2 M N,N-diisopropyl ethylamine in N-methylprrolidone were used as linkage reagents. After washing with dichloromethane (DMF), cleavage was performed in a solution of trifluoroacetic acid, phenol, water, and trisopropylsaline for 2 h at room temperature. Finally, the dried peptide was purified by reverse-phase high-performance liquid chromatography (RP-HPLC) on a Jupiter C18 column (4.6 × 250 mm, 300 Å, 5 μm). The molecular mass of the peptide was confirmed using a matrix-assisted laser desorption/ionization time-of-flight mass spectrometer (Kratos Analytical Inc., Chestnut Ridge, NY, USA).
Antibacterial activity assay
The antibacterial activity of the peptide was determined using its minimal inhibitory concentration (MIC). Gram positive and gram negative bacteria, and drug resistant bacteria strains were cultured overnight at 37 °C in Mueller-Hinton broth (MHB)38. The bacteria were diluted to a final concentration 2 × 105 colony-forming units (CFU)/mL in fresh MHB. The peptides and commercial antibiotics were diluted in 10 mM sodium phosphate buffer (pH 7.2). The bacteria was mixed with serially diluted peptide, or antibiotics in a 96 well plate and incubated at 37 °C for 18 h. After that, growth was measured by absorbance at 600 nm. The MICs were determined as the lowest concentration that inhibited bacterial growth. The result is represented as an average of the MIC obtained from three independent experiments.
Cell culture and cytotoxicity
HaCaT and Raw 264.7 cells were seeded in a 96-well plate at 104 cells/well and incubation for 24 h. The peptide was added to wells, and then incubated for 24 h. Then, 0.5 mg/mL MTT was added to each well and incubated 4 h at 37 °C. The supernatants were aspirated, and DMSO was added to dissolve the formazan crystals. Cytotoxicity was measured using microplate reader at 570 nm. The test was reproduced at least three times using two replicates.
Hemolytic activity
Red blood cells (RBCs) at a final concentration of 8% were mixed with the peptide in a 96-well plate and incubated for 1 h at 37 °C. Thereafter, the plate was centrifuged at 1500 × g for 10 min. The supernatant was transferred to a new 96-well plate and its absorbance was monitored at 414 nm. In the controls, 100% hemolysis consisted of RBC in 0.1% Triton X-100, whereas zero hemolysis control consisted of RBCs in PBS (pH 7.4). Each measurement was performed at least three times using two replicates. Percent hemolysis was calculated according to the following equation39:
Biofilm inhibition assay
Bacteria were diluted to a final concentration 5 × 105 CFU/mL in fresh MHB containing 0.2% of glucose. The bacteria was mixed with peptide in a 96-well plate and incubated. Thereafter, the biofilm was fixed with 100% methanol for 15 min, then aspirated and the wells were dried. Wells were stained with 0.1% crystal violet for 2 h. The crystal violet was removed and the wells were rinsed with distilled water. After air-drying, 95% ethanol was added to dissolve the biofilm, and the absorbance was measured using a microplate reader at 595 nm. The minimal biofilm inhibitory concentration (MBIC) was determined as the lowest concentration that inhibited the biofilm formation. Each measurement was performed at least three times using two replicates.
Circular dichroism (CD) spectroscopy
The CD spectra were recorded for 40 μM peptide in a buffer consisting of 10 mM sodium phosphate, 30 mM sodium dodecyl sulfate (SDS), and 50% 2,2,2-trifluoroethanol (TFE). The CD spectra were acquired on a Jasco 810 spectropolarimeter (Jasco, Tokyo, Japan) equipped with a quartz cell with a 0.1-cm path length. The spectra were recorded at wavelengths from 190 nm to 250 nm.
Preparation of liposome and liposome aggregation
Large unilamellar vesicles (LUV) were prepared by the freeze-thaw method40. The lipid concentration was determined using a standard phosphate assay41. Liposome aggregation was monitored by measurement of absorbance. The peptide at 5, 10, 20, 40, and 80 μM was added to 400 μM LUVs of PC:CH (10:1, w/w), PE:PG (7:3. w/w), or PC:CH:SM (1:1:1, w/w). The increase absorbance was measured at 405 nm, using a microplate reader.
Peptide binding to cell wall components
The insoluble peptidoglycan was added to 10 mM Tris-HCl (pH 7.5), containing 5 μg of peptide, and was incubated for 1 h. The supernatant and pellet were then subjected separately to SDS-PAGE in a 15% gel. The gel was stained for 30 min with a staining solution and then destained with destaining solution. The peptide in the presence or absence of LPS (0.1%) was used in a binding assay examined by CD spectroscopy. Peptidoglycan or LPS, peptide, and peptide and bacteria were incubated overnight. Then, absorbance is measured at 600 nm.
Outer membrane permeabilization assay
Outer membrane permeability was determined based on the uptake of NPN, a fluorescent dye. The bacteria were re-suspended to an optical density at 600 nm of 0.2 in 5 mM HEPES (pH 7.2). The bacteria and peptide were mixed with NPN solution (final concentration 10 μM) in a black 96-well plate. 0.1% Triton X-100 was used as a negative control. Each measurement was performed at least three times using two replicates.
Cytoplasmic membrane depolarization assay
Membrane depolarization by the peptide was quantified using a membrane-sensitive fluorescent dye, 3,3′-Dipropylthiadicarbocyanine Iodide [DiSC3(5)]. The bacteria were cultured and washed with washing buffer (5 mM HEPES and 20 mM glucose) three times, and re-suspended to an OD600 of 0.05 in buffer (5 mM HEPES, 20 mM glucose, and 100 mM KCl). The bacteria was incubated with at final concentration 1 μM DiSC3(5) solution for 1 h. Then, a solution of the peptides was added to the mixture of bacteria and DiSC3(5). Each measurement was performed at least three times using two replicates.
Flow cytometry
Antibiotics, and a mixture of peptide and antibiotics was incubated with the bacteria (OD600 of 0.5) for 1 h. The cells were harvested by centrifugation, washed, and incubated with propidium iodide (PI) at a final concentration 10 μg/mL for 30 min at 4 °C. Thereafter, the unbound dye was removed by washing with PBS.
Low vacuum scanning electron microscopy (SEM)
The bacteria cells (OD600 of 0.2) were incubated with peptide for 30 min at 37 °C. After that, the cells were fixed 2.5% glutaraldehyde in PBS at 4 °C overnight. The fixed samples were washed with PBS and post-fixed with 1% osmium tetroxide (OsO4) for 1 h. Then, the cells were dehydrated in 50, 60, 70, 80, 90, and 100% ethanol for 15 min each. Each sample was placed on a cover glass and coated with platinum and then examined under a low vacuum scanning electron microscope (JEOL JSM-IT300, Japan).
Synergy activity
The synergistic effects of Macropin and traditional antibiotics were assessed according to the broth microdilution checkerboard method42. Macropin, gentamycin, tobramycin, ciprofloxacin, levofloxacin, oxacillin, and piperacillin were diluted in 10 mM sodium phosphate buffer (pH 7.2). Then, each of different concentration of antibiotics and peptide was mixed and added into bacteria in a 96-well plate. The result represents an average of the FIC index obtained from three independent experiments. The effects of this combination were assessed using the fractional inhibitory concentration index (FIC index):
The FIC index was represented as follows: FICi ≤ 0.5 indicated synergy, 0.5 < FICi ≤ 1 indicated additive, 1 < FICi ≤ 4 indicated indifference, FICi > 4 indicated antagonism43.
Time-dependent killing of the synergistic group
The peptide and antibiotics were mixed with bacteria. Every 10 min, the mixture was plated onto MHB agar. The bactericidal activity of the antimicrobial agent was evaluated by the bacteriostasis rate as follows44;
Effects of Macropin In vivo
Six- to seven-week-old female BALB/c mice (20 ± 0.5 g) were used in this study. The back skin of the mice was using a razor blade. The mice were anesthetized with isoflurane and air cavities were produced by infecting 3 mL of air in the back45. The mice were then infected with 1 × 108 CFU/mL of S. aureus ATCC 25923 in the presence or absence of 1 mg/kg Macropin. Control was injected with PBS. On days 1, 2, 3, 4, and 7, tissues surrounding the air pouch were harvested and homogenized separately. The homogenates were plated on MHB agar plates. The plates were incubated at 37 °C overnight and then the colonies were counted.
Ethics Statement
This study conformed to the ethical standards of the Institutional Ethics Committee of Chosun University, and the protocol was approved by the Institutional Ethics Committee of Chosun University. All experiments were carried out in strict accordance with the National Institutes of Health Guidelines for the Ethical Treatment of Animals and the guidelines of the Center for Experimental Animals of Chosun University for Medical Science (Permit number: CIAUCUC2017-S0025).
References
Aiello, A. E. & Larson, E. Antibacterial cleaning and hygiene products as an emerging risk factor for antibiotic resistance in the community. Lancet Infect Dis 3, 501–506 (2003).
Hancock, R. E. & Sahl, H. G. Antimicrobial and host-defense peptides as new anti-infective therapeutic strategies. Nat Biotechnol 24, 1551–1557, https://doi.org/10.1038/nbt1267 (2006).
Bahar, A. A. & Ren, D. Antimicrobial peptides. Pharmaceuticals (Basel) 6, 1543–1575, https://doi.org/10.3390/ph6121543 (2013).
Hancock, R. E. & Rozek, A. Role of membranes in the activities of antimicrobial cationic peptides. FEMS Microbiol Lett 206, 143–149 (2002).
Andersson, D. I., Hughes, D. & Kubicek-Sutherland, J. Z. Mechanisms and consequences of bacterial resistance to antimicrobial peptides. Drug Resist Updat 26, 43–57, https://doi.org/10.1016/j.drup.2016.04.002 (2016).
Brogden, K. A. Antimicrobial peptides: pore formers or metabolic inhibitors in bacteria? Nat Rev Microbiol 3, 238–250, https://doi.org/10.1038/nrmicro1098 (2005).
Wu, M., Maier, E., Benz, R. & Hancock, R. E. Mechanism of interaction of different classes of cationic antimicrobial peptides with planar bilayers and with the cytoplasmic membrane of Escherichia coli. Biochemistry 38, 7235–7242, https://doi.org/10.1021/bi9826299 (1999).
Rotem, S. & Mor, A. Antimicrobial peptide mimics for improved therapeutic properties. Biochim Biophys Acta 1788, 1582–1592, https://doi.org/10.1016/j.bbamem.2008.10.020 (2009).
Guilhelmelli, F. et al. Antibiotic development challenges: the various mechanisms of action of antimicrobial peptides and of bacterial resistance. Front Microbiol 4, 353, https://doi.org/10.3389/fmicb.2013.00353 (2013).
Nicolas, P. Multifunctional host defense peptides: intracellular-targeting antimicrobial peptides. FEBS J 276, 6483–6496, https://doi.org/10.1111/j.1742-4658.2009.07359.x (2009).
Bliziotis, I. A., Samonis, G., Vardakas, K. Z., Chrysanthopoulou, S. & Falagas, M. E. Effect of aminoglycoside and beta-lactam combination therapy versus beta-lactam monotherapy on the emergence of antimicrobial resistance: a meta-analysis of randomized, controlled trials. Clin Infect Dis 41, 149–158, https://doi.org/10.1086/430912 (2005).
Hirano, L. & Bayer, A. S. Beta-Lactam-beta-lactamase-inhibitor combinations are active in experimental endocarditis caused by beta-lactamase-producing oxacillin-resistant staphylococci. Antimicrob Agents Chemother 35, 685–690 (1991).
Hopefl, A. W. Overview of synergy with reference to double beta-lactam combinations. DICP 25, 972–977 (1991).
Jiang, Y. et al. Antimicrobial activities of recombinant mouse beta-defensin 3 and its synergy with antibiotics. J Mater Sci Mater Med 23, 1723–1728, https://doi.org/10.1007/s10856-012-4645-z (2012).
Cirioni, O. et al. Efficacy of colistin/rifampin combination in experimental rat models of sepsis due to a multiresistant Pseudomonas aeruginosa strain. Crit Care Med 35, 1717–1723, https://doi.org/10.1097/01.ccm.0000266685.25436.03 (2007).
Nuding, S., Frasch, T., Schaller, M., Stange, E. F. & Zabel, L. T. Synergistic effects of antimicrobial peptides and antibiotics against Clostridium difficile. Antimicrob Agents Chemother 58, 5719–5725, https://doi.org/10.1128/aac.02542-14 (2014).
Monincova, L. et al. Structure-activity study of macropin, a novel antimicrobial peptide from the venom of solitary bee Macropis fulvipes (Hymenoptera: Melittidae). J Pept Sci 20, 375–384, https://doi.org/10.1002/psc.2625 (2014).
Palm, N. W. et al. Bee venom phospholipase A2 induces a primary type 2 response that is dependent on the receptor ST2 and confers protective immunity. Immunity 39, 976–985, https://doi.org/10.1016/j.immuni.2013.10.006 (2013).
Steiner, H., Hultmark, D., Engstrom, A., Bennich, H. & Boman, H. G. Sequence and specificity of two antibacterial proteins involved in insect immunity. Nature 292, 246–248 (1981).
Ma, Z. et al. Characterization of cell selectivity, physiological stability and endotoxin neutralization capabilities of alpha-helix-based peptide amphiphiles. Biomaterials 52, 517–530, https://doi.org/10.1016/j.biomaterials.2015.02.063 (2015).
Loh, B., Grant, C. & Hancock, R. E. Use of the fluorescent probe 1-N-phenylnaphthylamine to study the interactions of aminoglycoside antibiotics with the outer membrane of Pseudomonas aeruginosa. Antimicrob Agents Chemother 26, 546–551 (1984).
Dong, N. et al. Antimicrobial potency and selectivity of simplified symmetric-end peptides. Biomaterials 35, 8028–8039, https://doi.org/10.1016/j.biomaterials.2014.06.005 (2014).
Donlan, R. M. Biofilms: microbial life on surfaces. Emerg Infect Dis 8, 881–890, https://doi.org/10.3201/eid0809.020063 (2002).
Sato, H. & Feix, J. B. Peptide-membrane interactions and mechanisms of membrane destruction by amphipathic alpha-helical antimicrobial peptides. Biochim Biophys Acta 1758, 1245–1256, https://doi.org/10.1016/j.bbamem.2006.02.021 (2006).
Dempsey, C. E. The actions of melittin on membranes. Biochim Biophys Acta 1031, 143–161 (1990).
Habermann, E. Bee and wasp venoms. Science 177, 314–322 (1972).
Yeaman, M. R. & Yount, N. Y. Mechanisms of antimicrobial peptide action and resistance. Pharmacol Rev 55, 27–55, https://doi.org/10.1124/pr.55.1.2 (2003).
Erridge, C., Bennett-Guerrero, E. & Poxton, I. R. Structure and function of lipopolysaccharides. Microbes Infect 4, 837–851 (2002).
Yu, L. et al. Interaction of an artificial antimicrobial peptide with lipid membranes. Biochim Biophys Acta 1788, 333–344, https://doi.org/10.1016/j.bbamem.2008.10.005 (2009).
Martinez de Tejada, G. et al. Bacterial cell wall compounds as promising targets of antimicrobial agents I. Antimicrobial peptides and lipopolyamines. Curr Drug Targets 13, 1121–1130 (2012).
Henzler Wildman, K. A., Lee, D. K. & Ramamoorthy, A. Mechanism of lipid bilayer disruption by the human antimicrobial peptide, LL-37. Biochemistry 42, 6545–6558, https://doi.org/10.1021/bi0273563 (2003).
Li, L., Shi, Y., Cheserek, M. J., Su, G. & Le, G. Antibacterial activity and dual mechanisms of peptide analog derived from cell-penetrating peptide against Salmonella typhimurium and Streptococcus pyogenes. Appl Microbiol Biotechnol 97, 1711–1723, https://doi.org/10.1007/s00253-012-4352-1 (2013).
Livermore, D. M. Bacterial resistance: origins, epidemiology, and impact. Clin Infect Dis 36, S11–23, https://doi.org/10.1086/344654 (2003).
Andersson, D. I. Persistence of antibiotic resistant bacteria. Curr Opin Microbiol 6, 452–456 (2003).
Feng, Q., Huang, Y., Chen, M., Li, G. & Chen, Y. Functional synergy of alpha-helical antimicrobial peptides and traditional antibiotics against Gram-negative and Gram-positive bacteria in vitro and in vivo. Eur J Clin Microbiol Infect Dis 34, 197–204, https://doi.org/10.1007/s10096-014-2219-3 (2015).
Dosler, S. & Karaaslan, E. Inhibition and destruction of Pseudomonas aeruginosa biofilms by antibiotics and antimicrobial peptides. Peptides 62, 32–37, https://doi.org/10.1016/j.peptides.2014.09.021 (2014).
Giacometti, A., Cirioni, O., Barchiesi, F. & Scalise, G. In-vitro activity and killing effect of polycationic peptides on methicillin-resistant Staphylococcus aureus and interactions with clinically used antibiotics. Diagn Microbiol Infect Dis 38, 115–118 (2000).
Park, Y. et al. A Leu-Lys-rich antimicrobial peptide: activity and mechanism. Biochim Biophys Acta 1645, 172–182 (2003).
Lee, J. K., Park, S. C., Hahm, K. S. & Park, Y. Antimicrobial HPA3NT3 peptide analogs: placement of aromatic rings and positive charges are key determinants for cell selectivity and mechanism of action. Biochim Biophys Acta 1828, 443–454, https://doi.org/10.1016/j.bbamem.2012.09.005 (2013).
Mayer, L. D., Hope, M. J. & Cullis, P. R. Vesicles of variable sizes produced by a rapid extrusion procedure. Biochim Biophys Acta 858, 161–168 (1986).
Stewart, J. C. Colorimetric determination of phospholipids with ammonium ferrothiocyanate. Anal Biochem 104, 10–14 (1980).
Orhan, G., Bayram, A., Zer, Y. & Balci, I. Synergy tests by E test and checkerboard methods of antimicrobial combinations against Brucella melitensis. J Clin Microbiol 43, 140–143, https://doi.org/10.1128/jcm.43.1.140-143.2005 (2005).
Zong, L. et al. Mechanism of action of a novel recombinant peptide, MP1102, against Clostridium perfringens type C. Appl Microbiol Biotechnol 100, 5045–5057, https://doi.org/10.1007/s00253-016-7387-x (2016).
Zhang, Y. et al. In vitro synergistic activities of antimicrobial peptide brevinin-2CE with five kinds of antibiotics against multidrug-resistant clinical isolates. Curr Microbiol 68, 685–692, https://doi.org/10.1007/s00284-014-0529-4 (2014).
Sin, Y. M., Sedgwick, A. D., Chea, E. P. & Willoughby, D. A. Mast cells in newly formed lining tissue during acute inflammation: a six day air pouch model in the mouse. Ann Rheum Dis 45, 873–877 (1986).
Acknowledgements
This work was supported by a National Research Foundation of Korea (NRF) grant funded by the Korean Government (No. 2016R1A2A1A05005440), Global Research Laboratory (GRL) Grant (No. NRF-2014K1A1A2064460) and Institute for Information & communications Technology Promotion (IITP) grant funded by the Korea government (MSIT) (No. 2017-0-01714, Development of Antimicrobial Peptide using Deep Learning).
Author information
Authors and Affiliations
Contributions
S.J. Ko, M.K. Kim and Y. Park designed the study. S.J. Ko, J.K. Bang, C.H. Seo, T. Luchian and Y. Park performed experiments, analyzed data, and provided intellectual input into all aspects of this study. S.J. Ko and Y. Park wrote the manuscript; all authors contributed to its editing.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ko, S.J., Kim, M.K., Bang, J.K. et al. Macropis fulvipes Venom component Macropin Exerts its Antibacterial and Anti-Biofilm Properties by Damaging the Plasma Membranes of Drug Resistant Bacteria. Sci Rep 7, 16580 (2017). https://doi.org/10.1038/s41598-017-16784-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-16784-6
This article is cited by
-
A novel antimicrobial peptide YS12 isolated from Bacillus velezensis CBSYS12 exerts anti-biofilm properties against drug-resistant bacteria
Bioprocess and Biosystems Engineering (2023)
-
Synthesis of nano-Fe3O4/ZnO composites with enhanced antibacterial properties and plant growth promotion via one-pot reaction
Environmental Science and Pollution Research (2023)
-
Bee venom-derived antimicrobial peptide melectin has broad-spectrum potency, cell selectivity, and salt-resistant properties
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.