Abstract
The multifunctional, transmembrane glycoprotein human CD38 catalyses the synthesis of three key Ca2+-mobilising messengers, including cyclic adenosine 5′-diphosphate ribose (cADPR), and CD38 knockout studies have revealed the relevance of the related signalling pathways to disease. To generate inhibitors of CD38 by total synthesis, analogues based on the cyclic inosine 5′-diphosphate ribose (cIDPR) template were synthesised. In the first example of a sugar hybrid cIDPR analogue, “L-cIDPR”, the natural “northern” N1-linked D-ribose of cADPR was replaced by L-ribose. L-cIDPR is surprisingly still hydrolysed by CD38, whereas 8-Br-L-cIDPR is not cleaved, even at high enzyme concentrations. Thus, the inhibitory activity of L-cIDPR analogues appears to depend upon substitution of the base at C-8; 8-Br-L-cIDPR and 8-NH2-L-cIDPR inhibit CD38-mediated cADPR hydrolysis (IC50 7 μM and 21 µM respectively) with 8-Br-L-cIDPR over 20-fold more potent than 8-Br-cIDPR. In contrast, L-cIDPR displays a comparative 75-fold reduction in activity, but is only ca 2-fold less potent than cIDPR itself. Molecular modelling was used to explore the interaction of the CD38 catalytic residue Glu-226 with the “northern” ribose. We propose that Glu226 still acts as the catalytic residue even for an L-sugar substrate. 8-Br-L-cIDPR potentially binds non-productively in an upside-down fashion. Results highlight the key role of the “northern” ribose in the interaction of cADPR with CD38.
Similar content being viewed by others
Introduction
The calcium-releasing second messengers, cyclic adenosine 5′-diphosphate ribose (cADPR, 1, Fig. 1)1 and adenosine 5′-diphosphate ribose (ADPR)2 are synthesised in humans by CD38 from nicotinamide adenine dinucleotide (NAD+). Under acidic conditions, CD38 can also generate the most potent Ca2+-releasing second messenger known to date, nicotinic acid adenine dinucleotide 2′-phosphate (NAADP)3, from NADP.
The transmembrane glycoprotein CD38 functions both as a surface receptor in the immune system and a multifunctional ADP-ribosyl cyclase (ADPRC) ectoenzyme. Its catalytic domain may be either extracellular (type II) or intracellular (type III)4. We recently confirmed the presence of both CD38 activities in Jurkat T-cells using the non-membrane permeant CD38 inhibitor araF-NAD5. CD38 is a marker in AIDS progression6 and a negative prognostic marker of chronic lymphocytic leukaemia7. The CD38-cADPR pathway is implicated in the pathogenesis of asthma8 and Alzheimer’s disease9. It acts to regulate intracellular levels of NAD+ and therefore is intricately linked to energy homeostasis, signal transduction and aging10,11,12,13. CD38 is a clinical target for antibody therapy in treating multiple myeloma with encouraging efficacy in patients14. Its emerging role in disease states is thus stimulating the search for new CD38 modulators and particularly small molecule inhibitors to provide structural clues for drug design and as potential therapeutic candidates. To date, the reported inhibitors of CD38 are either mechanism-based covalent inhibitors15, or reversible, competitive, non-covalent inhibitors. Competitive inhibitors are diverse in structure, including NAD+ analogues16, flavonoids17 and those developed from library hits18,19.
cADPR Acts as a principal second messenger, mobilising intracellular calcium20,21,22,23. We are interested in exploiting the common intermediate in cADPR formation and hydrolysis by CD3824,25 using product-like inhibitors. cADPR Analogues have been accessed by either a chemo-enzymatic route, modelled on its biosynthesis from NAD+, or by total chemical synthesis. Chemo-enzymatic routes rely on Aplysia californica cyclase recognising an NAD+ analogue as a substrate and cyclising at the desired N1 position. This limits their use, particularly for analogues that are modified at the locus of the forming N1-glycosidic bond [although see24]. However, cADPR itself is unattractive for inhibitor design since it is readily hydrolysed at the unstable N1 link to give the linear adenosine 5′-diphosphoribose (ADPR), itself a calcium-mobilising second messenger26,27,28. We previously reported a chemo-enzymatic route to cyclic inosine 5′-diphosphate ribose via its 8-bromo derivative [N1-cIDPR (abbreviated as cIDPR), 2 and 8-Br-cIDPR, 3, Fig. 1]29. Chemically and biologically stable (but vide infra), cIDPR and 8-Br-cIDPR both inhibit cADPR hydrolysis by the catalytic domain of CD38 (shCD38; IC50 of 276 and 158 µM respectively).
Many total synthetic routes to cADPR analogues required considerable modification of the “northern” ribose in order to generate a stable N1-link30,31,32,33,34. The difficulty of selective N1-ribosylation meant that this tended to be the site at which structural modifications were carried out to overcome issues with synthetic tractability. We developed modified Vorbrüggen conditions that effect completely stereo- and regiospecific introduction of an acetylated ribose at the N1-position of a protected inosine35,36,37. Unlike the chemo-enzymatic or other synthetic routes, this permits further exploration of the structure-activity relationship at the locus of CD38 catalytic activity using the stable cIDPR template.
Crystallography of shCD38 has identified the mechanism by which NAD+ is cyclised to cADPR and ADPR38. Glu146 is critical in regulating the multi-functionality of CD38-mediated NADase, ADP-ribosyl cyclase and cADPR hydrolysis activities and Glu226 is the catalytic residue, since its mutation essentially eliminates catalytic activity39. Crystal structures obtained with shCD38 and cADPR analogues40,41 suggest that the “northern” ribose monophosphate region is highly conserved. In the catalytic site, cADPR forms two hydrogen bonds through N6 and N7 to Glu146 and interacts with Glu226 through the 2″- and 3″-OH of the “northern” ribose. Taken together, this suggests a critical role for the base and “northern” ribose in the binding of cADPR analogues to CD38, as might be predicted for the locus of both cADPR formation and degradation. We have previously shown that small fragments consisting of only these elements (N1-inosine 5′-monophosphates, N1-IMP), or analogues of cIDPR with the “southern” ribose replaced by a butyl linker, inhibit cADPR hydrolysis by shCD38 with IC50 values in the low µM region36,41. Crystallography of a hydrolysed cADPR analogue revealed that interactions with Glu146 and Glu226 were maintained even after hydrolysis of the N1-glycosidic bond36.
cADPR possesses two ribose sugar motifs and both of these are of the D-configuration. Analogues where the ribose is substituted for an alternative sugar have not previously been explored. Therefore, we designed and synthesised the hybrid cIDPR analogue “L-cIDPR”, where the unnatural sugar enantiomer L-ribose is employed to generate an analogue that presents the “northern” ribose hydroxyl groups in a different spatial arrangement. We hoped that an L-configuration in an analogue at this site would probe the interaction between the hydroxyl groups of the “northern” ribose and the catalytic residue Glu226 of shCD38, and predicted that modification of this critical interaction might affect binding affinity.
We report here the synthesis of L-cIDPR and its evaluation as an inhibitor of cADPR hydrolysis by shCD38. L-cIDPR is surprisingly able to access the catalytic machinery of shCD38, being hydrolysed at high enzyme concentrations. We also report the synthesis of further base-modified analogues of L-cIDPR and their activity as inhibitors of cADPR hydrolysis by shCD38 and HPLC studies to examine the stability of the novel analogues towards hydrolysis by shCD38.
Results and Discussion
Synthesis of L-ribose cIDPR analogues
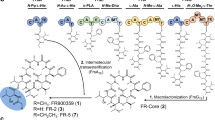
Starting from 5′-O-(tert-butyldiphenylsilyl)-2′,3′-O-isopropylidene-8-bromoinosine35 (8, Fig. 2) we introduced 2,3,5-O-triacetyl-β-L-ribofuranose by N1-ribosylation. The optimised reaction conditions for stereo- and regio-selective N1-coupling gave the desired product 9 in 85% yield, with no trace of the unwanted O-6 regioisomer. The 5″-OH was then revealed by sequential deprotection of the triol using methanolic ammonia and reprotection of the 2″,3″-diol was accomplished as its isopropylidene ketal. Treatment of 11 with N,N-diisopropyldibenzylphosphoramidite and 5-phenyl-1-H-tetrazole generated the 5″-O-phosphite, which was oxidised under basic conditions using H2O2 and triethylamine to generate 12. Deprotection of the 5′-O-TBDPS ether was carried out under neutral conditions using tetrabutylammonium fluoride (TBAF) and acetic acid to prepare 13. The second protected phosphate was introduced at the 5′-OH using S,S-diphenylphosphorodithioate (PSS), 2,4,6-triisopropylbenzenesulfonyl chloride (TPS-Cl) and 5-phenyl-1H-tetrazole to generate 14. Sequential deprotection of the phosphates was carried out using 50% trifluoroacetic acid (TFA) to remove the tert-butyl phosphate esters from the “northern” ribose and the two isopropylidene groups, then 0.1 M NaOH in dioxane to selectively deprotect one thiophenyl ester from the “southern” ribose. Cyclisation of 16 was carried out using iodine and 3 Å molecular sieves in pyridine under dilute conditions to prepare 8-Br-L-cIDPR 6 (Note: The notation ‘L’ refers to the stereochemistry of the “northern” ribose only and is not systematic.). The 8-bromo substituent was removed using Pd/C catalysis under an atmosphere of H2 to prepare the L-cIDPR analogue 5. Treatment of 6 with trimethylsilyl azide (TMSN3) in DMF was monitored by RP-HPLC (λ = 255 nm → 277 nm) and the crude material treated with dithiothreitol to generate 8-amino L-ribose cIDPR, 7.
Synthesis of L-ribose cIDPR analogues. Reagents i) (a) DBU, DCM (b) Tetraacetyl-L-ribose, TMSOTf, 85%; ii) NH3, MeOH, 100%; iii) pTsOH, H3CC(OMe)2CH3, Acetone, 100%; iv) (a) (tBuO)2PN(iPr)2, 5-Ph-1H-tetrazole, DCM (b) H2O2, Et3N, 60%; v) TBAF.3H2O, AcOH, 96%; vi) PSS, TPS-Cl, 5-Ph-1H-tetrazole, pyridine, 78%; vii) 50% TFA (aq.), 70%; viii) 0.1 M NaOH-dioxane; ix) I2, 3 Å MS, pyridine, 18% over 2 steps; x) H2, Pd/C, NaHCO3, EtOH-H2O, 46%; xi) (a) TMSN3, DMF, (b) Dithiothreitol, 0.05 M TEAB, 23%.
Inhibition of cADPR hydrolysis by CD38
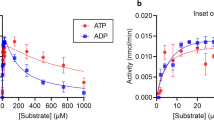
The ability of compounds 5-7 to inhibit hydrolysis of cADPR by human shCD38 was evaluated by a fluorimetric cycling assay42. The analogues inhibit hydrolysis of cADPR in a concentration-dependent manner (Fig. 3). 8-Br-L-cIDPR 6 and 8-NH2-L-cIDPR 7 are good inhibitors, with IC50 values of 7 μM and 21 μM respectively. In contrast, the parent compound, L-cIDPR 5, is a poor inhibitor with IC50 of 526 μM, Table 1.
The contrasting inhibitory activity upon substitution of the “northern” D-ribose by L-ribose was unexpected. L-cIDPR 5 is a roughly twofold poorer inhibitor of hydrolysis than cIDPR – which could be ascribed to changes within the “northern” ribose hydroxyl stereochemistry, meaning that critical interactions with the binding pocket are not fully realised. This might be anticipated prima facie, but is nevertheless somewhat surprising given the apparent stereochemical insult afforded by the L-sugar to the CD38 enzymatic machinery. Astonishingly, by comparison, 8-Br-L-cIDPR 6 is highly active with an IC50 of 7 μM. We had already seen that substitution of cIDPR to generate 8-Br-cIDPR improves the inhibitory activity of this analogue from 276 μM to 158 μM41; however, addition of the 8-bromo atom when the “northern” ribose is in the L-configuration generates a 75-fold increase in inhibitory activity (from 526 μM to 7 μM). In our previous studies, the introduction of an 8-NH2 substituent on the adenine ring resulted in a further improvement of inhibitory activity (approximately 5-fold increase compared to the parent molecule and 3-fold increase compared to the 8-Br substituted analogue). This increase in binding affinity was shown, by modelling and crystallography, to result from an additional interaction with Asp-15541. A comparable trend was not observed for the L-ribose cIDPR analogues prepared here, and this may suggest that the binding of these analogues has changed to adapt to the L-ribose substitution.
Conformational analysis
In solution, the ribose ring of nucleosides and nucleotides exists in conformational equilibrium between the C2′-endo and C3′-endo forms. As illustrated in Fig. 4A, the particular conformation adopted affects the spatial presentation of the hydroxyl groups and consequently would be expected to affect the interaction of a ligand with the binding pocket. Indeed, the conformation adopted by the southern ribose in cADPR analogues was shown to underpin their activity at the sea urchin cADPR receptor43. Using the method established by Altona and Sundaralingham44, the ratio of C2′-endo/C3′-endo forms may be mathematically calculated from the observed coupling constants in the 1H-NMR spectrum.
(A) Schematic representation of the ribofuranose ring in both C2-endo and C3-endo conformations; (B) From 1H-NMR data, cIDPR (2) in solution is predicted to display a C3-endo configuration in the “northern” ribose and 61% C2-endo configuration in the “southern” ribose; (C) L-cIDPR (5) is predicted to display a 59% C3-endo and 77% C2-endo configuration, respectively.
We used the 1H-NMR spectra of analogues 5-7 to determine the syn/anti-conformation of the purine base45 relative to the N9-ribose and to examine the ribose ring pucker of both the N9 and N1 sugars, Table 2.
As would be expected for conformationally restricted, cyclised analogues with two intact riboses, all three analogues adopt a syn conformation. Analysis of the N9-ribose conformation predicted that 5-7 would adopt a primarily C2-endo ring pucker of the “southern” ribose in free solution – matching that of cIDPR. For the “northern” N1-ribose, the three L-ribose analogues are predicted to have a predominantly C3-endo conformation, calculated using the coupling constant between H-1″ and H-2″ whereas cIDPR displays only a singlet for H-1″, suggesting a dihedral angle of 90° and a C3-endo conformation. The effect of the predominant conformation on 2″- and 3″-hydroxyl group orientation is illustrated for cIDPR (Fig. 4B) and L-cIDPR (Fig. 4C).
The “northern” ribose anomeric proton of L-cIDPR is shifted downfield by 0.3 ppm compared to cIDPR, suggesting it is more deshielded and the ring protons H-2″-4″ shifted upfield by 0.2 ppm. These changes reflect the different environments occupied by the ring protons and hydroxyl groups as a result of the change in sugar stereochemistry (Fig. 5).
HPLC studies
We recently discovered36 that high concentrations of shCD38 were in fact able to hydrolyse the cIDPR scaffold, when crystallography of 8-NH2-N9-butyl-cIDPR revealed an N1-hydrolysed product in the active site. This was somewhat surprising, as initial studies at lower concentrations had led to the cIDPR template being described as non-hydrolysable46. Hydrolysis would however not be predicted to occur at the concentrations found in enzyme assays, or those likely under physiological conditions (Supplementary Fig. S1, top two panels), so the cIDPR template can still be assumed to be biologically stable. Study of cIDPR analogues under these conditions can thus be used to evaluate the ability of the enzyme to turn over an analogue, and of an analogue to access the CD38 catalytic machinery. Interestingly, cIDPR is also enzymatically hydrolysable by the related Aplysia cyclase under extreme conditions, as revealed by X-ray crystallographic studies. Incubation of 30 mM cIDPR with wild-type Aplysia cyclase showed that cIDPR was clearly hydrolysed to IDPR at the N1 bond. In this instance, both linear IDPR and the cyclic N1-cIDPR are present and close to each other, with cIDPR in an upside-down orientation compared to cIDPR in CD38 (Q Liu, C Moreau, Q Hao, B V L Potter unpublished data, Supplementary Figure S2). Such an upside down binding mode for cIDPR in the Aplysia enzyme had also been observed earlier47. However, cIDPR bound in this way is clearly not an active conformation for cleavage in relation to the known catalytic residue and therefore this mode would be likely to lead to enzyme inhibition.
Incubation of 5 (1 mM final concentration) with 4 mg/mL shCD38 [enzyme in heavy excess] was monitored using RP-HPLC. The peak corresponding to 5 (RT = 12.5 min) reduced in intensity over time, alongside the appearance of a new peak (RT = 15.6 min), that was characteristic of an ADPR analogue (Fig. 6, Supplementary Fig. S1 third panel). No change in the original peak was observed in a parallel control experiment containing no shCD38. The rate of hydrolysis was similar, or slightly faster than that observed for cIDPR (2) under the same conditions. Surprisingly, treatment of analogue 6 under the same conditions, resulted in no hydrolysis, even after 22 hours (Fig. 6 and Supplementary Fig. S1, lower panel).
The observed hydrolysis of L-cIDPR at high concentrations of shCD38 demonstrates the flexibility of the CD38 catalytic machinery. The inability of CD38 to hydrolyse 8-Br-L-cIDPR perhaps suggests that this ligand adopts a different binding pose, presumably due to its additional 8-Br substituent combined with the change in hydroxyl stereochemistry resulting from introduction of the L-ribose at N1. Such a change in binding mode would help rationalise the observed difference in inhibition of CD38-mediated cADPR hydrolysis. It would seem feasible that 8-Br-L-cIDPR could be an excellent inhibitor of CD38 and not a substrate by virtue of binding in an upside-down fashion. Such a mode for a hybrid “northern” L-ribose analogue would put the “southern” D-ribose in a pseudo-northern ribose position, and the L-ribose in a pseudo-southern ribose position. We know from previous studies that the southern ribose is not essential for CD38 activity and can be replaced36. Unfortunately, attempts to crystallise 8-Br-L-cIDPR with CD38 to determine the exact binding pose were not successful.
Molecular Modelling
Docking of the “northern” ribose modified analogues using the modelling software GOLD generated several different poses in the binding site. No pose, by either visual inspection or by comparison of the docking scores, stood out as being obviously better than any of the others. As an alternative approach, the modified ligands were prepared by amending the stereochemistry of the natural “northern” ribose of cIDPR in the published 2PGJ and 3U4H crystal structures. The 8-substituent was also altered accordingly to 8-H, 8-Br, or 8-NH2, to generate a set of ligands in both crystal structures. The protein-ligand complexes were minimised and the binding interactions compared. It is important to note that the latter method is limited, in that it assumes that the L-ribose compounds bind to CD38 with the same basic pose as the natural D-ribose compounds and we would not be able to identify any potential completely different binding poses adopted by the L-cIDPR ligands. However, the approach is justified by the fact that L-ribose cIDPR must be able to access the catalytic residue of CD38 to be a substrate.
All analogues had similar poses to cIDPR after minimisation, and showed good overlap of the southern ribose sugar and pyrophosphate backbone. The L-configuration “northern” ribose is rotated around its N1-C1 and C4-C5 bonds, placing the furanose ring oxygen in the centre of the macrocycle and directing the two ring hydroxyl groups towards the binding pocket. The 2″- and 3″-hydroxyl group orientation is significantly changed due to the altered “northern” ribose stereochemistry, and without this rotation, they would point towards the centre of the binding pocket and not be able to contribute to binding. Therefore, the rotation of the ribose maximises the possibility of H-bonds to the catalytic residue Glu226 (Fig. 7).
(A) cIDPR (pink) and minimised L-cIDPR (yellow) in the 2PGJ crystal structure. Hypoxanthine ring tilt and good overlay of the southern ribose and pyrophosphate regions; (B) cIDPR (yellow) crystal structure indicates H-bonds from 2″- and 3″-OH to Glu226, stacking of hypoxanthine ring with Trp189, H-bond from 6 = O to Glu146; (C) L-cIDPR (yellow) minimised structure suggests disruption of π-stacking with Trp189 and significant rotation of the L-ribose to maintain at least one H-bond with Glu226 and potentially form a second H-bond between the 2″-OH and Ser-193. H-bonds are shown as yellow dashed lines with distances labelled in Å, interacting residues as shown as cyan sticks.
In all the L-ribose ligand complexes we generated, the L-ribose linked hypoxanthine ring is tilted, suggesting that these ligands may have disrupted π-stacking with Trp189 (Fig. 7C). The potential interaction of the L-ribose itself with the catalytic residue Glu226 is very interesting; the altered stereochemistry of the L-configuration ribose appears to prevent binding of both 2″-OH and 3″-OH to this residue meaning that only one H-bond is formed with Glu226 and the 3″-OH. However, the 2″-OH in the L-ribose conformation is correctly aligned to make a new H-bond with Ser193 (Fig. 7C). This suggests that the catalytically active conformation of L-cIDPR that allows scission of the N1-bond in the active site may differ from cIDPR (Fig. 8). Nevertheless, it is anticipated that only relatively minimal movement of the critical residue Glu226 should be required to accommodate an interaction with the L-ribose hydroxyls and facilitation of attack at the glycosidic bond, in comparison to the natural D-isomer. It is worth noting the immense value of L-nucleosides and derivatives such as Lamivudine, approved against HIV and HBV, that have been shown to have profound antiviral activity with many analogues in clinical trials and abundant evidence for their inhibition of and processing by host and viral enzymes48.
Schematic representing (A) cIDPR in the active site (from 2JPG) and active site residues proposed to lead to hydrolysis36; (B) L-cIDPR in the active site – twisted base and non-natural ribose sugar may alter catalysis in the active site. Possible additional interaction with Ser193 in the binding site is predicted by modelling.
We have previously predicted and confirmed by crystallography that the introduction of an 8-NH2-substituent can introduce an additional H-bond to Asp-155 in the shCD38 binding pocket41. For the 8-substituted L-cIDPR analogues our modelling suggested that the tilt of the hypoxanthine ring to accommodate the rotation of the L-configuration N1-ribose does not interfere with the H-bond to Asp-155 (Supplementary Figure S3). These observations give some rationale to the improved affinity of both 8-Br-L-cIDPR and 8-NH2-cIDPR compared to L-cIDPR but seem unlikely to be solely responsible for the extent of the difference in inhibitory activity that is observed.
Conclusion
Synthesis of the proposed ligands was facilitated by selective N1-glycosylation, and three hybrid L-ribose cIDPR-based analogues, substituted in the 8-position of adenine were prepared. Such analogues are unlikely to be accessible via a chemo-enzymatic route and illustrate the utility of our recently developed N1-ribosylation strategy to explore modifications to this sensitive region where both cyclisation and hydrolysis takes place. 8-Br-L-cIDPR is a potent (7 μM) inhibitor of cADPR hydrolysis by CD38. 8-NH2-L-cIDPR is similar in potency (21 μM), not showing the enhanced activity relative to its parent that has been previously observed with an extra 8-amino group for another analogue series41. Most surprising was the comparable lack of inhibitory activity of the parent compound, L-cIDPR, and its hydrolysis by CD38, albeit at high enzyme concentrations, showing that this analogue can still access the catalytic machinery. In contrast, 8-bromo-L-cIDPR was unaffected by high concentrations of CD38. In our past studies with cADPR analogues docking to CD38 we have been able to employ GOLD to reproduce very accurately the observed crystallographic binding modes, both with a compound from the inosine-based template, 8-amino-cIDPR, and also with another based upon cADPR itself, the carbocylic analogue cADPcR41. Docking studies with the L-ribose-modified ligands based on published crystal structures, however, struggled to predict clear preferred poses for such compounds with the catalytic site, highlighting potentially the critical nature of the stereochemical insult afforded by the L-ribose sugar. Adapting the natural ligand in the active site suggested that interactions could be maintained with the catalytic residue Glu226 by twisting the hypoxanthine ring in the binding pocket and this is supported by the substrate activity of L-cIDPR. It seems feasible, however, that the resistance to hydrolysis of the 8-bromo-L-cIDPR could be ascribed to a non-productive binding mode, possibly one in an upside-down configuration. Taken together, the analogues prepared in this study demonstrate that the “northern” ribose is an area that is very sensitive to modification, presents key binding partners for CD38 and offers some surprising insights.
Experimental Section
General
All reagents and solvents were of commercial quality and were used without further purification, unless described otherwise. The human CD38 catalytic domain (shCD38) was expressed and purified as described previously36,39. Unless otherwise stated, all reactions were carried out under an inert atmosphere of argon. 1H, 13C, and 31P NMR spectra were collected on a Jeol Delta 270 MHz, Varian Mercury 400 MHz or Bruker Avance III 500 MHz Spectrometer. All 1H- and 13C-NMR assignments are based on gCOSY, gHMBC, gHSQC, and DEPT-135 experiments. Abbreviations for splitting patterns are as follows: b, broad; s, singlet; d, doublet; t, triplet; m, multiplet etc. Coupling constants are given in Hertz (Hz). High resolution time-of-flight mass spectra were obtained on a Bruker Daltonics micrOTOF mass spectrometer using electrospray ionisation (ESI). Analytical HPLC analyses were carried out on a Waters 2695 Alliance module equipped with a Waters 2996 Photodiode Array Detector (210–350 nm). The chromatographic system consisted of a Hichrom Guard Column for HPLC and a Phenomenex Synergi 4μ MAX-RP 80 A column (150 × 4.60 mm), eluted at 1 mL/min with the following ion-pair buffer: 0.17% (m/v) cetrimide and 45% (v/v) phosphate buffer (pH 6.4) in MeOH. Synthetic phosphates were assayed and quantified by Ames phosphate test49.
N1-(2′′,3′′,5′′-Tri-O-acetyl-β-L-ribofuranosyl)-5′-O-(tert-butyldiphenylsilyl)-2′,3′-O-isopropylidene-8-bromoinosine (9)
5′-O-TBDPS-2′,3′-O-isopropylidene-8-bromoinosine 8 35 (980 mg, 1.57 mmol) was taken up in DCM (10.0 mL) and DBU (703 µL, 4.70 mmol) added. After 30 minutes, 1,2,3,5-tetra-O-acetyl-β-L-ribofuranose (548 mg, 1.72 mmol) was added and the solution cooled to -78 °C. Trimethylsilyl trifluoromethanesulfonate (1.13 mL, 6.26 mmol) was added dropwise and the solution stirred for a further 45 min before warming to rt. After 2.5 H, NaHCO3 (sat. aq.) was added and the crude material extracted into DCM (×3). The combined organic fractions were dried (Na2SO4), and solvent evaporated under reduced pressure. The residue was purified by column chromatography on silica gel eluting with PE:EtOAc (1:0 → 0:1 v/v) to afford the title compound (1.18 g, 85%) as a colourless glass; R f = 0.69 (EtOAc:PE 3:1 v/v); 1H-NMR (400 MHz, CDCl3) δ 7.96 (s, 1H, H-2), 7.57 (dd, 2H, J = 8.0, 1.4), 7.53 (dd, 2H, J = 8.0, 1.4), 7.37-7.28 (m, 4H), 7.23 (t, 2H, J = 8.0), 6.33 (d, 1H, J = 4.2, H-1′′), 6.14 (d, 1H, J = 2.2, H-1′), 5.45 (dd, 1H, J = 6.4, 2.2, H-2′), 5.42-5.37 (m, 2H, H-2′′ and H-3′′), 5.00 (dd, 1H, J = 6.4, 4.2, H-3′), 4.41 (dd, 1H, J = 4.9, 3.8, H-4′′), 4.37-4.36 (m, 2H, 2 ×H-5′′), 4.31 (dd, 1H, J = 4.2, 1.4, H-4′), 3.83 (dd, 1H, J = 11.0, 5.2, H-5′a), 3.74 (dd, 1H, J = 11.0, 6.6, H-5′b), 2.11, 2.08, 2.07 (each s, 3H, 3 × OAc), 1.61 (s, 3H, CH3), 1.37 (s, 3H, CH3), 1.00 (s, 9H, tBu) ppm; 13C-NMR (100 MHz, CDCl3) 169.9, 169.5, 169.4, 154.5, 147.6, 143.8, 135.43 (2C), 135.42 (2C), 133.3, 132.8, 129.6 (2C), 127.5 (2C), 127.4 (2C), 126.3, 124.7, 114.5, 90.9, 87.7, 87.1, 83.3, 81.3, 80.0, 74.1, 69.9, 63.9, 62.8, 27.2, 26.7 (3C), 25.4, 20.6, 20.4, 20.3, 19.1 ppm; HRMS (ESI+) found m/z [M + Na]+ 905.2013, 907.1994; C40H47N4O12 79BrSiNa requires 905.2035, C40H47N4O12 81BrSiNa requires 907.2015.
N1-(β-L-ribofuranosyl)-5′-O-(tert-butyldiphenylsilyl)-2′,3′-O-isopropylidene-8-bromoinosine (10)
Intermediate 9 (3.56 g, 4.03 mmol) was taken up in MeOH (20 mL) in a pressure tube. The solution was saturated with NH3 (g) at 0 °C, then stirred at rt for 12H. The solvent was evaporated and the residue purified by column chromatography on silica gel eluting with DCM:Acetone (1:0 → 0:1 v/v) to afford the title compound (3.05 g, 100%) as an off-white amorphous solid; R f = 0.24 (PE:EtOAc 1:3 v/v); 1H-NMR (400 MHz, CDCl3) δ 8.13 (s, 1H, H-2), 7.58 (dd, 2H, J = 8.0, 1.3), 7.52 (dd, 2H, J = 8.0, 1.3), 7.40-7.31 (m, 4H), 7.22 (t, 2H, J = 7.1), 6.12 (d, 1H, J = 2.2, H-1′), 5.87 (d, 1H, J = 4.1, H-1′′), 5.49 (dd, 1H, J = 6.3, 2.2, H-2′), 5.03 (dd, 1H, J = 6.3, 3.8, H-3′), 4.78 (d, 1H, J = 2.9, 2′′-OH), 4.49-4.46 (m, 1H, H-2′′), 4.42-4.39 (m, 1H, H-3′′), 4.36-4.32 (m, 1H, H-4′), 4.31-4.28 (m, 1H, H-4′′), 3.99-3.93 (m, 1H, H-5′′a), 3.93-3.75 (m, 2H, H-5′a and H-5′′b), 3.73-3.70 (m, 1H, H-5′b), 3.65 (d, 1H, J = 4.0, 3′′-OH), 2.62 (bs, 1H, 5′′-OH), 1.60 (s, 3H, CH3), 1.37 (s, 3H, CH3), 1.00 (s, 9H, tBu) ppm; 13C-NMR (100 MHz, CDCl3) δ 156.0, 148.4, 144.6, 135.5 (2C), 135.4 (2C), 133.3, 132.9, 129.7, 129.6, 127.6 (2C), 127.4 (2C), 126.8, 124.6, 114.4, 93.9, 91.3, 87.8, 86.4, 83.1, 81.6, 75.6, 70.8, 63.9, 61.9, 27.2, 26.7 (3C), 25.5, 19.1 ppm; HRMS (ESI+) found m/z [M + Na]+ 779.1701 and 781.1700, C34H41N4NaO9 79BrSi requires 779.1718, C34H41N4NaO9 81BrSi requires 781.1698.
N1-(2′′,3′′-O-isopropylidene-β-L-ribofuranosyl)-5′-O-(tert-butyldiphenylsilyl)-2′,3′-O-isopropylidene-8-bromoinosine (11)
p-TsOH (116 mg, 0.61 mmol) was added to 10 (460 mg, 0.61 mmol) in acetone-2,2-dimethoxypropane (10 mL, 4:1 v/v). After 45 min, DCM and NaHCO3 (sat. aq.) were added, and the organic layer dried (Na2SO4) and evaporated. The residue was taken up in MeOH (5 mL) and DOWEX 50WX8H+ resin (15 mg) added to convert any unwanted 5″-O-hemi-acetal side product into product. After 30 min the resin was removed by filtration under gravity and the solvent evaporated to obtain the title compound (484 mg, 100%) as a white foam; R f = 0.74 (DCM:Acetone 1:1 v/v); 1H-NMR (400 MHz, CDCl3) δ 7.62 (dd, 2H, J = 7.0, 1.4), 7.53 (s, 1H, H-2), 7.52 (dd, 2H, J = 7.0, 1.4), 7.42-7.32 (m, 4H), 7.22 (t, 2H, J = 5.8), 6.12 (d, 1H, J = 2.2, H-1′), 5.55 (d, 1H, J = 2.6, H-1′′), 5.44 (dd, 1H, J = 6.4, 2.2, H-2′), 5.18 (dd, 1H, J = 6.5, 2.6, H-2′′), 5.11 (dd, 1H, J = 6.5, 3.6, H-3′′), 5.04 (dd, 1H, J = 6.4, 3.8, H-3′), 4.36-4.32 (m, 2H, H-4′ and H-4′′), 3.89-3.80 (m, 2 H, H-5′′a and H-5′′b), 3.83 (dd, 1H, J = 10.8, 5.7, H-5′a), 3.77 (dd, 1H, J = 10.8, 6.2, H-5′b), 3.08 (bd, 1H, J = 4.2, 5′′-OH), 1.61 (s, 6 H, 2 × CH3), 1.37 (s, 6H, 2 × CH3), 1.01 (s, 9H, tBu) ppm; 13C-NMR (100 MHz, CDCl3) δ 155.0, 148.0, 146.2, 135.5 (2 C), 135.4 (2C), 133.5, 132.9, 129.7 (2C), 127.6 (2C), 127.5 (2C), 126.7, 125.5, 114.4, 114.2, 96.9, 91.3, 88.2, 87.7, 83.6, 83.2, 81.5, 80.6, 63.8, 62.8, 27.3, 27.2, 26.7 (3C), 25.5, 25.2, 19.2 ppm; HRMS (ESI+) found m/z [M + Na]+ 819.2005 and 821.1993; C37H45N4O9 79BrSiNa requires 819.2031, C37H45N4O9 81BrSiNa requires 821.2011.
N1-[2′′,3′′-O-isopropylidene-5′′-O-(di-tert-butyl)-phosphoryl-β-L-ribofuranosyl]-5′-O-(tert-butyldiphenylsilyl)-2′,3′-O-isopropylidene-8-bromoinosine (12)
5-Phenyl-1H-tetrazole (425 mg, 2.46 mmol) and N,N-diisopropyldibutylphosphoramidite (584 µL, 1.85 mmol) were added to a solution of 11 (985 mg, 1.23 mmol) in DCM (10 mL). After 2H, the solution was cooled to 0 °C and Et3N (1.03 µL, 7.38 mmol) and H2O2 (35%, 269 µL, 3.08 mmol) added. The solution was allowed to warm to rt and stirred for a further 2H, after which DCM (100 mL) and H2O were added. The organic layer was washed with NaHCO3 (sat. aq.), then brine, dried (Na2SO4) and evaporated to dryness. The residue was purified by column chromatography on silica gel eluting with PE:EtOAc (1:0 → 0:1 v/v), where both solvents contained 0.5% v/v pyridine, to afford the title compound (736 mg, 60%) as a colourless glass; R f = 0.67 (PE:EtOAc 1:3 v/v); 1H-NMR (400 MHz, CDCl3) δ 7.59 (dd, 2H, J = 8.0, 1.4), 7.57 (s, 1H, H-2), 7.51 (dd, 2H, J = 8.0, 1.3), 7.41-7.31 (m, 4H), 7.22 (t, 2H, J = 7.7), 6.13 (d, 1H, J = 2.1, H-1′), 5.75 (d, 1H, J = 1.6, H-1′′), 5.46 (dd, 1H, J = 6.4, 2.1, H-2′), 5.03 (dd, 1H, J = 6.4, 3.9, H-3′), 4.99 (dd, 1H, J = 6.2, 1.6, H-2′′), 4.96 (dd, 1H, J = 6.2, 3.8, H-3′′), 4.43 (ddd, 1H, J = 6.4, 5.8, 3.8, H-4′′), 4.34 (ddd, 1H, J = 6.0, 5.8, 3.9, H-4′), 4.24-4.11 (m, 2H, H-5″a and H-5″b), 3.80 (dd, 1H, J = 10.8, 5.6, H-5′a), 3.74 (dd, 1H, J = 10.8, 6.4, H-5′b), 1.61 (s, 3H, CH3), 1.60 (s, 3H, CH3), 1.428 (s, 9H, tBu), 1.427 (s, 9H, tBu), 1.37 (s, 3H, CH3), 1.36 (s, 3H, CH3), 1.00 (s, 9H, tBu) ppm; 13C-NMR (100 MHz, CDCl3) δ 154.5, 147.9, 145.6, 135.5 (2C), 135.4 (2C), 133.5, 132.8, 129.7, 129.6, 127.6 (2C), 127.4 (2C), 126.4, 125.3, 114.4, 114.2, 95.0, 91.2, 87.9, 87.2 (d, J = 7.8), 85.0, 83.3, 82.6 (d, 2C, J = 7.2), 81.6 81.5, 66.4 (d, J = 6.2), 63.9, 29.8 (3C), 29.7 (3C), 27.2, 27.1, 26.7 (3C), 25.5, 25.2, 19.2 ppm; 31P-NMR (162 MHz, 1H decoupled, CDCl3) δ -10.2 ppm; HRMS (ESI+) found m/z [M + Na]+ 1011.2911 and 1013.2900; C45H62N4O12 79BrSiNaP requires 1011.2947, C45H62N4O12 81BrSiNaP requires 1013.2926.
N1-[2′′,3′′-O-isopropylidene-5′′-O-(di-tert-butyl)-phosphoryl-β-L-ribofuranosyl]-2′,3′-O-isopropylidene-8-bromoinosine (13)
Acetic acid (133 µL, 2.32 mmol) and TBAF.3H2O (698 mg, 2.21 mmol) were stirred in DMF (2 mL) for 30 min, after which the solution was cooled to 0 °C and 12 (730 mg, 0.74 mmol) in DMF (5 mL) added. The resulting solution was allowed to warm to rt and stirred for a further 4 h. The solution was diluted with ether, and NaHCO3 (sat. aq.) and NH4Cl (sat. aq.) added. The organic layer was separated, and the aqueous layer extracted with ether (×3). The combined organic layers were dried (Na2SO4), evaporated to dryness and the residue purified by column chromatography on silica gel eluting with DCM:Acetone (1:0 → 0:1 v/v), where both solvents contained 0.5% v/v pyridine, to afford the title compound (530 mg, 96%) as a colourless glass; R f = 0.70 (DCM:Acetone 1:1 v/v); 1H-NMR (400 MHz, CDCl3) δ 8.10 (s, 1H, H-2), 6.06 (d, 1H, J = 4.7, H-1′), 5.96 (d, 1H, J = 1.8, H-1′′), 5.20 (dd, 1H, J = 5.8, 4.7, H-2′), 5.06-5.02 (m, 2H, H-2′′ and H-3′), 4.96 (dd, 1H, J = 6.4, 3.8, H-3′′), 4.44-4.41 (m, 2H, H-4′ and H-4′′), 4.24-4.12 (m, 2H, H-5′′a and H-5′′b), 3.87 (dd, 1H, J = 12.5, 2.5, H-5′a), 3.73 (dd, 1H, J = 12.5, 2.5, H-5′b), 1.64 (s, 3H, CH3), 1.56 (s, 3H, CH3), 1.45 (s, 9H, tBu), 1.43 (s, 9H, tBu), 1.36 (s, 3H, CH3), 1.33 (s, 3H, CH3) ppm; 13C-NMR (100 MHz, CDCl3) δ 154.3, 147.6, 146.1, 126.1, 125.8, 114.40, 114.38, 94.5, 93.0, 86.9 (d, J = 7.9), 85.8, 85.0, 82.9, 82.8 (d, J = 7.2), 82.7 (d, J = 7.2), 81.3, 81.2, 66.3 (d, J = 6.2), 62.8, 29.74 (d, 3C, J = 4.2), 29.72 (d, 3C, J = 4.2), 27.5, 27.1, 25.3, 25.2 ppm; 31P-NMR (162 MHz,1H decoupled, CDCl3) δ -10.4 ppm; HRMS (ESI+) found m/z [M + Na]+ 773.1744 and 775.1711; C29H44N4O12 79BrNaP requires 773.1769, C29H44N4O12 81BrNaP requires 775.1748.
N1-[2′′,3′′-O-isopropylidene-5′′-O-(di-tert-butyl)phosphoryl-β-L-ribofuranosyl]-5′-O-[(diphenylthio)phosphoryl]- 2′,3′-O-isopropylidene-8-bromoinosine (14)
Intermediate 13 (250 mg, 0.33 mmol) was evaporated from pyridine (3 × 2 mL) and taken up in pyridine (2.5 mL). This solution was added to PSS (381 mg, 1.00 mmol) which had also been evaporated from pyridine (3 × 2 mL). 5-Phenyl-1H-tetrazole (146 mg, 1.00 mmol) and TPS-Cl (201 mg, 0.67 mmol) were added and the solution stirred at rt for 5H. DCM and H2O were added, the organic layer separated and the aqueous layer washed with DCM (×2). The combined organic layer was washed with brine, dried (Na2SO4) and evaporated to dryness. The residue was purified by column chromatography on silica gel eluting with PE:EtOAc (1:0 → 0:1 v/v), where both solvents contained 0.5% v/v pyridine, to afford the title compound (263 mg, 78%) as a white foam; R f = 0.76 (DCM:Acetone 1:1 v/v); 1H-NMR (400 MHz, CDCl3) δ 7.97 (s, 1H, H-2), 7.47-7.25 (m, 10H, Ar-H), 6.17 (d, 1H, J = 2.0, H-1′), 5.79 (d, 1H, J = 1.5, H-1′′), 5.51 (dd, 1H, J = 6.2, 2.0, H-2′), 5.08 (dd, 1H, J = 6.2, 3.6, H-3′), 5.04 (dd, 1H, J = 6.4, 1.5, H-2′′), 4.97 (dd, 1H, J = 6.4, 3.9, H-3′′), 4.44-4.31 (m, 4H, H-4′, H-4′′, both H-5′), 4.24-4.21 (m, 2H, both H-5′′), 1.62 (s, 3H, CH3), 1.55 (s, 3H, CH3), 1.47 (s, 9H, tBu), 1.45 (s, 9H, tBu), 1.39 (s, 3H, CH3), 1.30 (s, 3H, CH3) ppm; 13C-NMR (100 MHz, CDCl3) δ 154.5, 148.0, 146.4, 135.3 (d, 2C, J = 5.5), 135.1 (d, 2C, J = 5.5), 129.6-129.5 (m, 2C), 129.4-129.3 (m, 4C), 126.3, 125.6-125.5 (m, 2C), 125.4, 114.8, 114.2, 95.5, 91.2, 87.5 (d, J = 7.8), 85.5 (d, J = 8.4), 84.8, 83.5, 83.1 (d, J = 7.4), 83.0 (d, J = 7.4), 81.7, 81.1, 66.8 (d, J = 6.0), 66.2 (d, J = 8.2), 29.8 (d, 3C, J = 4), 29.7 (d, 3C, J = 4), 27.2, 27.0, 25.4, 25.2 ppm; 31P-NMR (162 MHz,1H decoupled, CDCl3) δ 50.7, −10.8 ppm; HRMS (ESI+) found m/z [M + Na]+ 1037.1639 and 1039.1569; C41H53N4O13 79BrNaP2S2 requires 1037.1601, C41H53N4O13 81BrNaP2S2 requires 1039.1581.
N1-[5′′-O-phosphoryl-β-L-ribofuranosyl]-5′-O-[(diphenylthio)phosphoryl]-8-bromoinosine (15)
Intermediate 14 (250 mg, 0.25 mmol) was stirred in 50% aq. TFA (4 mL) at 0 °C for 4H. All solvents were evaporated and the residue co-evaporated with MeOH (×4). The residue was purified by column chromatography on silica gel eluting with EtOAc:MeOH:H2O (1:0:0 → 4:2:0 → 7:2:1 v/v/v) to afford the title compound (147 mg, 70%) as a colourless glass; R f = 0.14 (EtOAc:MeOH:H2O 7:2:1 v/v/v); 1H-NMR (500 MHz, d4-MeOD) δ 8.78 (s, 1H, H-2), 7.44-7.37 (m, 6H), 7.32-7.28 (m, 4H) (10 × Ar-H), 6.15 (d, 1H, J = 3.9, H-1′), 6.01 (d, 1H, J = 3.3, H-1′′), 5.22 (dd, 1H, J = 5.4, 3.3, H-2′′), 4.86 (t, 1H, J = 5.9, H-3′′), 4.50-4.43 (m, 2H, both H-5′′), 4.32 (dd, 1H, J = 4.7, 3.9, H-3′), 4.28 (dd, 1H, J = 4.7, 3.9, H-2'), 4.21-4.10 (m, 4H, H-4′, H-4′′ and both H-5′) ppm; 13C-NMR (125 MHz, d4-MeOD) δ 157.2, 150.0, 146.9, 136.6 (d, 2C, J = 5.3), 136.4 (d, 2C, J = 5.4), 131.0 (d, J = 3.1), 130.9 (d, J = 3.1), 130.5 (4C), 128.7, 126.8 (d, J = 6.4), 126.7 (d, J = 6.4), 125.4, 93.3, 90.6, 85.5 (d, J = 8.6), 83.9 (d, J = 8.0), 77.3, 73.7, 71.1, 70.9, 68.3 (d, J = 8.3), 64.9 (d, J = 5.0) ppm; 31P-NMR (202 MHz, 1H decoupled, d4-MeOD) δ 52.1, 0.7 ppm; HRMS (ESI−) found m/z [M − H]− 820.9762 and 822.9735; C27H28N4O13 79BrP2S2 requires 820.9758, C27H28N4O13 81BrP2S2 requires 822.9738.
N1-(5′′-O-phosphoryl-β-L-ribofuranosyl)-5′-O-(phenylthio)phosphoryl-8-bromoinosine (16)
Intermediate 15 (24 mg, 0.029 mmol) was taken up in dioxane:H2O (1 mL, 1:1 v/v). NaOH (100 µL, 1 M) was added and the solution stirred for 30 min at rt before addition of HCl (100 µL, 1 M). The solution was diluted with H2O and washed with hexane (×3) before evaporation of all solvents to give a colourless glass which was converted to the TEA salt as described below. 1H-NMR (500 MHz, D2O) δ 8.40 (s, 1H, H-2), 7.01 (d, 2H, J = 7.9), 6.92 (t, 1H, J = 7.5), 6.80 (t, 2H, J = 7.7) (5 × Ar-H), 5.95 (d, 1H, J = 6.0, H-1′), 5.78 (d, 1H, J = 2.8, H-1″), 5.54 (t, 1H, J = 6.0, H-2′), 4.43 (dd, 1H, J = 6.0, 2.8, H-3′), 4.29-4.24 (m, 2H), 4.21-4.17 (m, 4H), 4.08-4.01 (m, 2H) ppm; 13C-NMR (125 MHz, D2O) δ 156.0, 148.8, 145.0, 131.5 (d, 2C, J = 5.5), 129.7 (d, J = 5.6), 128.8, 128.4 (2C), 127.2, 123.7, 90.4, 89.9, 84.5 (d, J = 10.6), 82.8 (d, J = 8.8), 75.0, 70.8, 70.3, 68.8, 65.8 (d, J = 5.4), 63.9 (d, J = 4.8) ppm; 31P-NMR (202 MHz, D2O, 1H-decoupled) δ 17.3, 0.0 ppm; HRMS (ESI−) calcd for C21H23N4O14P2S79BrNa 750.9493 [(M + Na − 2H)−], found 750.9493; and calcd for C21H23N4O14P2S81BrNa 752.9473 [(M + Na − 2H)−], found 752.9458. Conversion to TEA salt: The Na+ salt was passed through pre-washed DOWEX H+ resin. Acidic fractions were neutralised with TEAB (2 mL, 1 M). All solvents were evaporated and the residue co-evaporated with H2O to remove excess buffer. The colorless glass obtained was used directly for cyclisation.
Cyclic-8-bromoinosine 5′-diphosphate-L-ribose (8-Br-L-cIDPR, 6)
Intermediate 16 (0.029 mmol) was evaporated from pyridine (2 mL, ×3). The residue was taken up in pyridine (20 mL) and added over 15H to a solution of iodine (140 mg, 0.591 mmol) and 3 Å molecular sieves (1.0 g) in pyridine (40 mL), in the dark. The solution was filtered through celite, washed with H2O. After addition of TEAB (2 mL) all solvents were evaporated, and the residue partitioned between H2O and CHCl3. The aqueous layer was washed with CHCl3 and evaporated to dryness. The residue was purified by semi-preparative reverse phase HPLC eluted at 5 mL/min with acetonitrile:0.1 M TEAB (1:19 → 13:7 v/v) over 25 min. Fractions were analysed by analytical RP-HPLC and appropriate fractions collected and evaporated under vacuum to give the title compound (3.2 mg, 18% over 2 steps); UV (H2O, pH 7), λmax 255 nm (ε 12,400); 1H-NMR (500 MHz, D2O) δ 8.80 (s, 1H, H-2), 6.34 (d, 1H, J = 3.8, H-1″), 6.05 (d, 1H, J = 6.8, H-1′), 5.15 (dd, 1H, J = 6.8, 5.0, H-2′), 4.61 (dt, 1H, J = 10.6, 1.9, H-5″a), 4.54 (t, 1H, J = 5.0, H-3″), 4.51 (dd, 1H, J = 5.0, 3.8, H-2″), 4.47 (dd, 1H, J = 5.0, 1.8, H-3′), 4.35 (dd, 1H, J = 6.4, 1.8, H-4′), 4.27-4.22 (m, 2H, H-4″ and H-5′a), 4.09-4.02 (m, 2H, H-5′b and H-5″b) ppm; 13C-NMR (125 MHz, D2O) δ 156.8, 148.8, 145.3, 128.1, 123.7, 91.0, 87.9, 85.2 (d, J = 10.6), 83.6 (d, J = 9.0), 75.1, 73.3, 70.9, 69.5, 64.9 (d, J = 4.8), 64.3 (d, J = 5.0) ppm; 31P-NMR (202 MHz, D2O, 1H-decoupled) δ -10.39 (d, J = 14.1), −10.71 (d, J = 14.1) ppm; HRMS (ESI−) calcd for C15H18N4O14P2 79Br 618.9484 [(M − H)−], found 618.9493; and calcd for C15H18N4O14P2 81Br 620.9463 [(M−H)−], found 620.9464.
Cyclic-inosine 5′-diphosphate-L-ribose (L-cIDPR, 5)
Cyclic-8-bromoinosine 5′-diphosphate-L-ribose (6, 10 mg, 0.016 mmol) was taken up in MilliQ H2O (4 mL) and NaHCO3 (13 mg, 0.161 mmol) added. EtOH (2 mL) and Pd/C (5 mg) were added and the flask placed under an atmosphere of H2 for 16H. The solution was filtered and all solvents were evaporated. The residue was purified by semi-preparative reverse phase HPLC eluted at 5 mL/min with acetonitrile:0.1 M TEAB (1:19 → 13:7 v/v) over 25 min. Fractions were analysed by analytical RP-HPLC and appropriate fractions collected and evaporated under vacuum to give the title compound (4.0 mg, 46% over 2 steps); UV (H2O, pH 7), λmax 251 nm (ε 9,900); 1H-NMR (500 MHz, D2O) δ 8.81 (s, 1H, 2H-2), 8.11 (s, 1H, H-8), 6.35 (d, 1H, J = 3.5, H-1″), 5.93 (d, 1H, J = 6.5, H-1′), 5.09 (dd, 1H, J = 6.5, 4.9, H-2′), 4.59-4.52 (m, 3H, H-5″a, H-3″, H-2″), 4.49 (dd, 1H, J = 4.9, 1.9, H-3′), 4.35-4.33 (m, 1H, H-4′), 4.29-4.27 (m, 1H, H-4″), 4.23 (dd, 1H, J = 11.7, 3.5, H-5′a), 4.09-4.04 (m, 2H, H-5′b and H-5″b) ppm; 13C-NMR (125 MHz, D2O) δ 158.0, 147.6, 145.1, 142.4, 123.7, 90.5, 88.1, 84.9 (d, J = 11.1), 83.7 (d, J = 9.3), 75.1, 73.4, 71.0, 69.6, 65.0 (d, J = 5.5), 64.2 (d, J = 4.6) ppm; 31P-NMR (202 MHz, D2O, 1H-decoupled) δ −10.40 (d, J = 10.1), −10.75 (d, J = 10.1) ppm; HRMS (ESI−) calcd for C15H19N4O14P2 541.0378 [(M − H)−], found 541.0390.
Cyclic-8-aminoinosine 5′-diphosphate-L-ribose (8-NH2-L-cIDPR, 7)
Cyclic-8-bromoinosine 5′-diphosphate-L-ribose (6, 3.6 mg, 5.8 μmol) was converted to the H+ form by stirring with DOWEX 50WX8H+ resin in MilliQ (2 mL) for 15 mins. The resin was removed by filtration and all solvent evaporated. The residue was co-evaporated with DMF (4 × 1 mL) and taken up in DMF (1 mL). TMSN3 (50 μL) was added and the solution stirred at 70 °C in the dark for 16H at which point the HPLC showed 95% conversion to the 8-azido product. MilliQ (5 mL) was added and all solvents evaporated. The residue was taken up in TEAB (0.05 M, 5 mL) and dithiothreitol (10 mg) added. The resulting solution was stirred for 12H, when HPLC analysis showed complete consumption of the 8-azido intermediate. The solution was purified by RP-HPLC eluted at 5 mL/min with acetonitrile:0.1 M TEAB (1:19 → 13:7 v/v) over 25 min. Fractions were analysed by analytical RP-and appropriate fractions collected and evaporated under vacuum to give the title compound (1.35 μmol, 23% over 2 steps); UV (H2O, pH 7), λmax 261 nm (ε 11,700); 1H-NMR (500 MHz, D2O) δ 8.60 (s, 1H, H-2), 6.37 (d, 1H, J = 3.5, H-1″), 5.80 (d, 1H, J = 6.7, H-1′), 5.20 (dd, 1H, J = 6.7, 4.9, H-2′), 4.56-4.53 (m, 3H, H-5″a, H-3″, H-2″), 4.48 (dd, 1H, J = 4.9, 1.8, H-3′), 4.30 (dd, 1H, J = 6.2, 1.8, H-4′), 4.27-4.25 (m, 1H, H-4″), 4.20-4.18 (m, 1H, H-5′a), 4.06 (d, 1H, H-5′b), 4.02 (dt, 1H, J = 10.8, 4.0, H-5″b) ppm; 13C-NMR (125 MHz, D2O) δ 152.6, 149.9, 145.9, 145.1, 110.5, 88.6, 88.3, 85.1 (d, J = 10.8), 83.4 (d, J = 8.6), 74.8, 72.9, 70.5, 69.2, 64.9 (d, J = 3.8), 64.2 (d, J = 4.3) ppm; 31P-NMR (202 MHz, D2O, 1H-decoupled) δ −10.27 (d, J = 16.2), −10.51 (d, J = 16.2) ppm; HRMS (ESI−) calcd for C15H20N5O14P2 556.0487 [(M − H)−], found 556.0471.
Enzymatic Assay for cADPR Hydrolysis
The inhibition of cADPR hydrolysis by various concentrations of analogue (0–1 mM) was determined by incubating 1 μM cADPR with 1 μg/ml of CD38 for 10 min at 20–24 °C in 25 mM sodium acetate, pH 4.5. The reaction was stopped by the addition of 150 mM HCl. The precipitated protein was filtered, and the pH was neutralised with Tris base. After diluting the mixture 20-fold, the concentration of the unhydrolysed cADPR present in the diluted reaction mixture was assayed by the fluorimetric cycling assay as described previously42.
Modelling of the L-ribose compounds: Method A
Three cIDPR analogues with L-ribose in the “northern” position but differing in the substituent at the 8-position (hydrogen, bromine or amino) were built using the Schrödinger software and docked into the 2PGJ structure using GOLD50. The binding site was defined as a sphere of 5 Å radius centred on the centroid of the cIDPR: the centroid of the docked ligand has to lie within this sphere. Each ligand was docked twenty-five times. The best ranked poses of each the ligand were merged with the protein structure and the resulting complexes passed through a minimisation procedure using the Schrödinger software.
Method B
The ligand in the prepared 2PGJ and 3U4H crystal structures was modified so that the “northern” ribose was of the L-configuration. An 8-amino or 8-bromo group was added to the 8-position of the 2PGJ ligand, and the 8-amino group of the 3U4H ligand was either amended to an 8-bromo substituent or a hydrogen. The resulting complexes were put through 500 rounds of minimisation using the Schrödinger software or minimised using the Discovery Studio software from Accelrys. All figures were prepared using PyMOL.
HPLC studies
The solution containing the CD38 catalytic domain was adjusted to the desired concentration using Tris-HCl buffer (20 mM, pH 8) and 50 μL therefrom was added to the inhibitor (0.05 μmole) in an Eppendorf tube at room temperature. At a given time point, a sample of 5 μL was removed and diluted with 95 μL MilliQ water. 10 μL Of this sample was injected directly into the analytical HPLC system (see General Experimental), eluting at 1 mL/min with an isocratic ion-pair buffer: 0.17% (m/v) cetrimide and 45% (v/v) phosphate buffer (pH 6.4) in MeOH.
Data availability
The crystallographic data for 2PGJ and 3U4H are available in the PDB repository (www.pcsb.org). All other data generated or analysed during this study are included in this published article and Supplementary Information file.
References
Howard, M. et al. Formation and hydrolysis of cyclic ADP-ribose catalyzed by lymphocyte antigen CD38. Science 262, 1056–1059 (1993).
Partida-Sanchez, S. et al. Chemotaxis of mouse bone marrow neutrophils and dendritic cells is controlled by adp-ribose, the major product generated by the CD38 enzyme reaction. J. Immunol. 179, 7827–7839 (2007).
Cosker, F. et al. The ecto-enzyme CD38 is a nicotinic acid adenine dinucleotide phosphate (NAADP) synthase that couples receptor activation to Ca2+ mobilization from lysosomes in pancreatic acinar cells. J. Biol. Chem. 285, 38251–38259 (2010).
Zhao, Y. J., Lam, C. M. C. & Lee, H. C. The Membrane-Bound Enzyme CD38 Exists in Two Opposing Orientations. Sci. Signal. 5, ra67 (2012).
Fliegert, R. et al. 2′-Deoxyadenosine 5′-diphosphoribose is an endogenous TRPM2 agonist. Nat. Chem. Biol., 13, 1036-1044 (2017).
Savarino, A. et al. Human CD38 interferes with HIV-1 fusion through a sequence homologous to the V3 loop of the viral envelope glycoprotein gp120. FASEB J. 17, 461-463 (2003).
Malavasi, F. et al. CD38 and chronic lymphocytic leukemia: a decade later. Blood 118, 3470–3478 (2011).
Deshpande, D. A. et al. CD38 in the pathogenesis of allergic airway disease: Potential therapeutic targets. Pharmacol. Ther. 172, 116–126 (2017).
Blacher, E. et al. Alzheimer’s disease pathology is attenuated in a CD38-deficient mouse model. Ann. Neurol. 78, 88–103 (2015).
Aksoy, P., White, T. A., Thompson, M. & Chini, E. N. Regulation of intracellular levels of NAD: A novel role for CD38. Biochem. Biophys. Res. Commun. 345, 1386–1392 (2006).
Young, G. S., Choleris, E., Lund, F. E. & Kirkland, J. B. Decreased cADPR and increased NAD+ in the Cd38−/− mouse. Biochem. Biophys. Res. Commun. 346, 188–192 (2006).
Chini, E. N. CD38 as a Regulator of Cellular NAD: A Novel Potential Pharmacological Target for Metabolic Conditions. Curr. Pharm. Design 15, 57–63 (2009).
Mouchiroud, L., Houtkooper, R. H. & Auwerx, J. NAD+ metabolism: A therapeutic target for age-related metabolic disease. Crit. Rev. Biochem. Mol. Biol. 48, 397–408 (2013).
Lokhorst, H. M. et al. Targeting CD38 with Daratumumab Monotherapy in Multiple Myeloma. New England Journal of Medicine 373, 1207–1219 (2015).
Kwong, A. K. Y. et al. Catalysis-Based Inhibitors of the Calcium Signaling Function of CD38. Biochemistry 51, 555–564 (2012).
Wall, K. A. et al. Inhibition of the intrinsic NAD+ glycohydrolase activity of CD38 by carbocyclic NAD analogues. Biochem. J. 335, 631–636 (1998).
Kellenberger, E., Kuhn, I., Schuber, F. & Muller-Steffner, H. Flavonoids as inhibitors of human CD38. Bioorg. Med. Chem. Lett. 21, 3939–3942 (2011).
Zhou, Y. et al. Design, Synthesis and Biological Evaluation of Noncovalent Inhibitors of Human CD38 NADase. Chem Med Chem 7, 223–228 (2012).
Haffner, C. D. et al. Discovery, Synthesis, and Biological Evaluation of Thiazoloquin(az)olin(on)es as Potent CD38 Inhibitors. J. Med. Chem. 58, 3548–3571 (2015).
Lee, H. C. Multiplicity of Ca2+ messengers and Ca2+ stores: A perspective from cyclic ADP-ribose and NAADP. Curr. Mol. Med. 4, 227–237 (2004).
Guse, A. H. Biochemistry, biology, and pharmacology of cyclic adenosine diphosphoribose (cADPR). Curr. Med. Chem. 11, 847–855 (2004).
Potter, B. V. L. & Walseth, T. F. Medicinal chemistry and pharmacology of cyclic ADP-ribose. Curr. Mol. Med. 4, 303–311 (2004).
Guse, A. H. Second messenger function and the structure-activity relationship of cyclic adenosine diphosphoribose (cADPR). FEBS J. 272, 4590–4597 (2005).
Sauve, A. A., Deng, H. T., Angeletti, R. H. & Schramm, V. L. A covalent intermediate in CD38 in responsible for ADP-ribosylation and cyclisation reactions. J. Am. Chem. Soc. 122, 7855–7859 (2000).
Kim, H., Jacobson, E. & Jacobson, M. Synthesis and degradation of cyclic ADP-ribose by NAD glycohydrolases. Science 261, 1330–1333 (1993).
Ashamu, G. A., Sethi, J. K., Galione, A. & Potter, B. V. L. Roles for adenosine ribose hydroxyl groups in cyclic adenosine 5′-diphosphate ribose-mediated Ca2+-release. Biochemistry 36, 9509–9517 (1997).
Guse, A. H. et al. Novel hydrolysis-resistant analogues of cyclic ADP-ribose: Modification of the “northern” ribose and calcium release activity. Biochemistry 41, 6744–6751 (2002).
Bailey, V. C., Fortt, S. M., Summerhill, R. J., Galione, A. & Potter, B. V. L. Cyclic aristeromycin diphosphate ribose: A potent and poorly hydrolysable Ca2+-mobilising mimic of cyclic adenosine diphosphate ribose. FEBS Lett. 379, 227–230 (1996).
Wagner, G. K., Guse, A. H. & Potter, B. V. L. Rapid synthetic route toward structurally modified derivatives of cyclic adenosine 5′-diphosphate ribose. J. Org. Chem. 70, 4810–4819 (2005).
Shuto, S. et al. Total synthesis of cyclic ADP-carbocyclic ribose, a stable mimic of Ca2+-mobilizing second messenger cyclic ADP-ribose. J. Am. Chem. Soc. 123, 8750–8759 (2001).
Gu, X. et al. Synthesis and biological evaluation of novel membrane-permeant cyclic ADP-ribose mimics: N1-[(5″-O-phosphorylethoxy)methyl]-5′-O-phosphorylinosine 5′,5″-cyclic pyrophosphate (cIDPRE) and 8-substituted derivatives. J. Med. Chem. 47, 5674–5682 (2004).
Guse, A. H. et al. A minimal structural analogue of cyclic ADP-ribose - Synthesis and calcium release activity in mammalian cells. J. Biol. Chem. 280, 15952–15959 (2005).
Tsuzuki, T. et al. Design, Synthesis, and Chemical and Biological Properties of Cyclic ADP-4-Thioribose as a Stable Equivalent of Cyclic ADP-Ribose. Messenger 3, 35–51 (2014).
Mahal, A. et al. Synthesis of cyclic N1-pentylinosine phosphate, a new structurally reduced cADPR analogue with calcium-mobilizing activity on PC12 cells. Beilstein Journal of Organic Chemistry 11, 2689–2695 (2015).
Swarbrick, J. M. & Potter, B. V. L. Total Synthesis of a Cyclic Adenosine 5′-Diphosphate Ribose Receptor Agonist. J. Org. Chem. 77, 4191–4197 (2012).
Swarbrick, J. M. et al. Cyclic Adenosine 5′-Diphosphate Ribose Analogs without a “Southern” Ribose Inhibit ADP-ribosyl Cyclase–Hydrolase CD38. J. Med. Chem. 57, 8517–8529 (2014).
Swarbrick, J. M. et al. ‘Click cyclic ADP-ribose’: a neutral second messenger mimic. Chem. Commun. 50, 2458–2461 (2014).
Graeff, R. et al. Mechanism of cyclizing NAD to cyclic ADP-ribose by ADP-ribosyl cyclase and CD38. J. Biol. Chem. 284, 27629–27636 (2009).
Munshi, C. et al. Identification of the enzymatic active site of CD38 by site-directed mutagenesis. J. Biol. Chem. 275, 21566–21571 (2000).
Liu, Q. et al. Catalysis associated conformational changes revealed by human CD38 complexed with a non-hydrolysable substrate analog. J. Biol. Chem. 282, 24825–24832 (2007).
Moreau, C. et al. CD38 Structure-Based Inhibitor Design Using the N1-Cyclic Inosine 5′-Diphosphate Ribose Template. PLOS ONE 8, e66247 (2013).
Graeff, R. M. & Lee, H. C. A novel cycling assay for cellular cADP-ribose with nanomolar sensitivity. Biochem. J. 361, 379–384 (2002).
Moreau, C. et al. Synthesis of cyclic adenosine 5′-diphosphate ribose analogues: a C2′ endo/syn “southern” ribose conformation underlies activity at the sea urchin cADPR receptor. Org. Biomol. Chem. 9, 278–290 (2011).
Altona, C. & Sundaralingam, M. Conformational analysis of sugar ring in nucleosides and nucleotides - Improved method for interpretation of proton magnetic resonance coupling constants. J. Am. Chem. Soc. 95, 2333–2344 (1973).
Rosemeyer, H. et al. Syn-anti conformational analysis of regular and modified nucleosides by 1D H-1 NOE difference spectroscopy - A simple graphical-method based on conformationally rigid molecules. J. Org. Chem. 55, 5784–5790 (1990).
Kirchberger, T. et al. Cellular effects and metabolic stability of N1-cyclic inosine diphosphoribose and its derivatives. Brit. J. Pharmacol. 149, 337–344 (2006).
Liu, Q. et al. Structural basis for enzymatic evolution from a dedicated ADP-ribosyl cyclase to a multifunctional NAD hydrolase. J. Biol. Chem. 284, 27637–27645 (2009).
Mathé, C. & Gosselin, G. L-nucleoside enantiomers as antivirals drugs: a mini-review. Antiviral Research 71, 276–281 (2006).
Ames, B. N. & Dubin, D. T. The Role of Polyamines in the Neutralization of Bacteriophage Deoxyribonucleic Acid. J. Biol. Chem. 235, 769–775 (1960).
Jones, G., Willett, P., Glen, R. C., Leach, A. R. & Taylor, R. Development and validation of a genetic algorithm for flexible docking. J. Mol. Biol. 267, 727–748 (1997).
Acknowledgements
BVLP is a Wellcome Trust Senior Investigator (Grant 101010). We also acknowledge RCGAS 201105159001 (to R.G.).
Author information
Authors and Affiliations
Contributions
J.M.W. and B.V.L.P. designed the study. J.M.W. synthesised the cADPR analogues, carried out the HPLC studies and the conformational analysis. R.G. carried out the enzyme assays. M.P.T. carried out the modelling studies. J.M.W. and B.V.L.P. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Watt, J.M., Graeff, R., Thomas, M.P. et al. Second messenger analogues highlight unexpected substrate sensitivity of CD38: total synthesis of the hybrid “L-cyclic inosine 5′-diphosphate ribose”. Sci Rep 7, 16100 (2017). https://doi.org/10.1038/s41598-017-16388-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-16388-0
This article is cited by
-
Synthetic cADPR analogues may form only one of two possible conformational diastereoisomers
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.