Abstract
Medullary thymic epithelial cells (mTECs) ectopically express a diversity of peripheral tissue-restricted antigens (PTAs) and provide unique cues for the expansion, maturation and selection of a repertoire of functionally diverse T lymphocytes. Genetic deletion of all mature microRNAs in thymic epithelial cells (TECs) results in premature thymic involution, progressive disorganisation of the thymic epithelium, and alteration in thymic T cell lineage commitment, consequently eliciting autoimmune disorders. In the present study, we identified that microRNA-449a (miR-449a), a member of miR-449 cluster, regulated mTEC differentiation. Expression of miR-449a was induced by RANK ligand in mouse fetal thymus. In in vitro studies, overexpression of miR-449a induced thymic epithelial progenitor cells (TEPCs) differentiation into mature mTECs. Despite abundant expression of miR-449a in developing thymus, miR-449a-mutant mice exhibited normal thymic development. This might be partially due to in miR-449a-mutant thymus the up-regulation of miR-34a which shared similar seed sequence with miR-449a. However, thymic expression of miR-449/34 sponge which was able to neutralize the function of miR-449/34 family members significantly reduced the number of mature Ly51-MHCIIhi mTECs. Taken together, our data suggested that miR-449a modulated mTEC differentiation, and members of miR-34 cluster functioned redundantly to rescue miR-449a deficiency in thymus development.
Similar content being viewed by others
Introduction
The thymus containing thymic epithelial cells (TECs) that form a complex three-dimensional meshwork structure provides the microenvironment to drive the differentiation of bone marrow-derived hematopoietic precursors to mature T lymphocytes1. TECs consist of cortical thymic epithelial cells (cTECs) and medullary thymic epithelial cells (mTECs) which form discrete intrathymic microenvironments, thymic cortex and thymic medulla respectively. Each is specialized for mediating a particular aspect of thymocytes development2,3. The bone marrow-derived progenitors go through a consecutive process including release from bone marrow niches into the blood4,5 and exit from the circulation to settle in the thymus at the cortico-medullary junction (CMJ)6,7. On entering the thymus, these early thymic progenitors (ETP) undergo an ordered process of development from CD4/CD8 double negative stage (CD4−/CD8−) to double positive stage (CD4+/CD8+) during their migration from CMJ through the cortex to the outer subcapsular zone (SCZ) and then back to the cortex8,9,10. During this process, progenitors become committed to the αβ or γδT cell lineage11,12 at DN3 stage and undergo β-selection13; the resulting immature single positive (ISP) intermediate cells then differentiate into double positive cells and undergo positive selection by recognization of self-peptide/MHC complexes expressed on cTECs14,15. Positively selected thymocytes then enter the medulla, where T cells express T cell receptors with high affinity for self-antigens are clonally depleted by apoptosis16,17,18.
The crucial role of mTEC in establishing T cell central tolerance is attributed to the expression and presentation of a diversity of peripheral tissue-restricted antigens17,19,20,21. Recent studies have elucidated a battery of regulators underlying the differentiation and function of mTEC, among which the identification of autoimmune regulator (Aire) was a breakthrough in the study of mTEC biology19. Members of the tumor necrosis factor receptor (TNFR) family and their downstream canonical/alternative NF-κB pathways are involved in the differentiation and function of mTECs. Mice deficient with RANK22,23, CD4023, lymphotoxin β receptor24,25, NF-κB inducing kinase (NIK)26, IκB kinase (IKK) α27, RelB28, and Traf629 exhibited variable defects in mTEC.
MicroRNAs (miRNAs) are a class of small (19~22nt), noncoding RNAs that mediate sequence-dependent post-transcriptional gene repression by translational inhibition and/or mRNA destabilization. Approximately over 10,000 miRNAs have been identified to exert their effects in normal organism development and pathogenesis30,31,32. However, the molecular mechanisms of miRNAs underlying thymus development remain less understood. Adrian Liston’s group reported the function of thymic epithelial miRNA network in infection-associated thymic involution by Foxn1 mediated conditional knockout of Dicer in mouse model33. The deletion of Dicer and therefore all mature miRNAs in thymic epithelial cells result in premature thymic involution, progressive disorganisation of the thymic epithelium, alteration in thymic T cell lineage commitment, and consequently elicit autoimmune disorders33,34. In accordance with these discoveries, Foxn1 mediated conditional knockout of DGCR8, a gene whose product is responsible for canonical cleavage of miRNAs, results in severe loss of Aire+ mTECs and breaks down the negative selection in thymus35. A transcriptome analysis of murine thymic epithelial cells reveals miRNA expression profiling that is closely correlated with age-related thymic atrophy, indicating the function of miRNAs in control of thymic aging process36. Thus, miRNAs are reasonably playing crucial roles in thymic epithelial cell differentiation and function.
We identified miR-449a in 2-DG FTOC (2-DG FTOC, 2′-deoxyguanosine (2-DG) treated Fetal Thymus Organ Culture) treated with recombinant human RANK ligand. Expression of miR-449a was induced by RANK ligand in fetal thymus and overexpression of miR-449a could induce TEPC differentiation in vitro. Neutralization of miR-449a and other miR-449/34 family members reduced the number of mature MHCIIhi mTECs in thymus. Our results revealed a new function of miR-449a in regulation of mTEC differentiation.
Materials and Methods
Animals
The miR-449a mutant mice (miR-449aIns/Ins and miR-449aDel/Del) were generated by Shanghai Bioray Biotech Co., Ltd. All animals were housed and maintained in specific-pathogen-free conditions. All animal experiments were performed in compliance with the guide for the care and use of laboratory animals and were approved by the institutional biomedical research ethics committee of the Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences.
Primers used for genotyping:
F1: CACAATTCTATCTCTAGGCC
F2: GCTGGTTGAGTATGTGAG
R: GGGCAAATACACAAGGC
F1 and R primer pair for miR-449aIns/Ins genotyping, a 336 bp band indicates insert mutation, and further sequencing is needed to distinguish heterozygous and homozygous mutation. F2 and R primer pair for miR-449aDel/Del genotyping, a 314 bp band indicates wild type or heterozygous mutation, homozygous mutation results in no amplification.
Cell lines and cell culture
TSC (thymic epithelial progenitor cell line) cells were established in our lab previously37. TSC cells were treated with 100 ng/ml recombinant human RANKL (R&D, 390-TN-010) for 2 or 4 days.
To stably overexpress miR-449a in TSC cells, TSC cells were infected with empty control lentivirus or lentivirus expressing miR-449a and selected with 1ug/ml puromycin for 4 days. The stable cell lines were thereafter named as TSC Control or TSC miR-449a. Overexpression of miR-449a (>300 fold) was then confirmed by q-PCR.
Fetal Thymus Organ Culture, FTOC
Thymic lobes were isolated from E14.5 embryos and cultured for 4 days on nucleopore filters (Whatmann, NJ) placed in RPMI1640 (Invitrogen), supplemented with 10% fetal bovine serum (Invitrogen), 2 mM L-glutamine (Invitrogen), 50 µM 2-mercaptoethanol (Sigma-Aldrich) and 1.35 mM 2′-deoxyguanosine (2-DG, Sigma-Aldrich), as previously described23. Fetal thymic lobes (2-DG FTOC) were cultured in RPMI1640-10%FBS medium and infected with lentivirus carrying miR-449a or miR-449/34 sponge expressing vectors. For in vitro differentiation, fetal thymic lobes (2-DG FTOC) were cultured with 100ng/ml recombinant human RANKL (R&D, 390-TN-010) for 4 days.
Plasmids construction
MiR-449a was amplified from mouse genome using forward primer: 5'-TGA ATT CAC TTA GCC TCA GCC ACT C-3' and reverse primer: 5'-TGT CTA GAT AAT GTC AAG CTA GGA C-3' and cloned into plvx-IRES-EGFP (Clontech).
To clone miR-449/34 sponge, forward sequence: 5'-gatccACCAGCTAACTATCACTGCC ACGATACCAGCTAACTATCACTGCCAACGCGACCAGCTAACTATCACTGCCACGATACCAGCTAACTATCACTGCCAACG CGACCAGCTAACTATCACTGCCACGATACCAGCTAACTATCACTGCCAttttttg-3' and reverse sequence: 5'-aattcAAAAAATGGCAGTGATAGTTAGCTGGTATCGTGGCAGTGATAGTTAGCTGGTCGCGTTGGCA GTGATAGTTAGCTGGTATCGTGGCAGTGATAGTTAGCTGGTCGCGTTGGCAGTGATAGTTAGCTGGTATCGTGGCAGTGA TAGTTAGCTGGT g-3' were synthesized, annealed and cloned into plvx-shRNA2 (Clontech).
Western blot
The cells were harvested and washed with cold phosphate buffer solution (PBS) once. Add 100 µl lysis/loading buffer (0.25 M Tris-HCl, 10% SDS, 0.5% Bromophenol blue, 50% Glycerinum, 7.71% Dithiothreitol), boiling for 10 minutes.
For each sample, 10-20 μl of protein lysate was separated by SDS-PAGE, transferred electrophoretically to a PVDF membrane (Immobilon P, Millipore) and immunoblotted with primary and peroxidase-conjugated secondary antibodies in 5% non-fatty milk. Detection of the bound antibody was performed by SuperSignal west pico Chemiluminescent Substrate (Pierce). Antibodies to Aire (N-20), GAPDH (G-9), SATB2 (SATBA4B10), RelB (A-9) and NF-κB p52 (K-27) were purchased from Santa Cruz Biotech Inc. Monoclonal antibody to DNMT3a was generated in Dr. Guoliang Xu’s lab (Institute of Biochemistry and Cell Biology, SIBS, CAS).
Thymic in situ injection
Virus expressing Control-GFP or miR-449/34 sponge-GFP were packaged according to Lenti-X™ shRNA Expression Systems User Manual (Clontech). Control-GFP virus and miR-449/34 sponge-GFP virus were concentrated by ultracentrifugation and stored at −80 °C. The GFP protein acted as a reporter gene of miR-449/34 sponge expression.
3-week old C57BL/6 mice were narcotized, injected with about 30 μl of virus per lobe of thymus at the interstice between the second and third rib. Injection was conducted once every other day for 3 times. 2 days after the last injection, the thymus were harvested for analysis.
Flow cytometry
The thymi were harvested, washed with cold PBS twice, minced and digested with Liberase TH (Roche) and DNase I (Roche). Thymic cells were stained with antibodies: EpCAM (G8.8, Biolegend, 118204), Ly51 (Biolegend, 108312), MHCII (BD sciences, 562366), CD45 (eBioscience, 45-0451-82) for 30 min on ice. mTECs were gated as CD45−EpCAM+Ly51−MHCII+.
For Aire staining, cells were stained with Foxp3/Transcription Factor Staining Buffer Set (eBioscience) and stained with Anti-Aire antibody (eBioscience, 51-5934-82) for 30 min on ice.
Immunofluorescence
Frozen thymuses embedded in OCT compound were sliced into 8 µm-thick sections. Frozen sections were fixed with cold acetone for less than 5 minutes and stained with the following antibodies: rabbit polyclonal antibodies to Alexa-488 K5 (Covance), Alexa-647 K8 (Troma-1, Developmental Studies Hybridoma Bank), FITC-EpCAM (G8.8, Developmental Studies Hybridoma Bank), K14 (Covance) and Aire (M-300, Santa Cruz Biotech Inc.) followed by Alexa Fluor 568-conjugated anti-rabbit IgG antibody (Molecular Probes). Images were analyzed with TSC SP2 confocal laser-scanning microscope.
RNA Extraction, RT-PCR and Q-PCR
RNA was isolated from cell lines or thymus samples using TRIzol Reagent (Invitrogen) and reverse transcripted using Transcript First Strand Synthesis Supermix (TransGen Biotech) according to the manufacturer’s instructions. Reverse transcription polymerase chain reactions (RT-PCR) were conducted using 2X Taq PCR MasterMix (TianGen Biotech).
All quantitative PCR (q-PCR) were performed using a 7500 Fast Real-Time PCR System (Applied Biosystems) in SYBR Premix Ex Taq reaction system (TaKaRa). Each sample was analyzed in triple replication. Relative quantification (RQ) was derived from the difference in cycle threshold (Ct) between the target gene and tubulin (ΔCt) as compared to control cell lines using the equation RQ = 2−ΔΔCt. Error bars represent standard deviation (SD), and statistical significance was calculated using a one-tailed, unpaired t-test. Relative mRNA or miRNA expression was summarized using mean ± SEM. All these results were calculated using student t-tests. p < 0.05 was considered to be significant. All the primers are listed in Table S2.
Results
Expression of miR-449a was induced by RANK ligand
RANK and its downstream NF-κB signaling were previously demonstrated to regulate the differentiation and function of mTEC23. To see if downstream miRNAs of RANK signaling regulate thymus development, we treated fetal thymi with RANK ligand (RANKL) for 4 days in 2-DG FTOC system. By RANKL stimulation, miR-449a was dramatically induced in 2-DG FTOC (Fig. 1A). And consistent with previous report, the expression of Aire as well as Aire-dependent PTAs (Spt1 and Insulin) and Aire-independent PTAs (CRP) were also significantly up-regulated (Fig. 1A).
Expression of miR-449a was induced by RANKL. (A) q-PCR analyzed the expression of miR-449a, miR-34a, aire, spt1, Insulin and CRP in 2-DG FTOC treated with 100ng/ml recombinant human RANKL for 4 days. (B) q-PCR analyzed the expression of indicated genes in TSC cells treated with 100 ng/ml recombinant human RANKL for 2 days. (UD, undetectable). (C) q-PCR analysis of miR-449a expression in TSC cells overexpressing RelA/p50 or RelB/p52. (D) q-PCR analyzed the expression of indicated genes in RelB-deficient thymic epithelial cells. Bar graphs show means ± standard errors of at least three independent measurements. (*p < 0.05, **p < 0.01, n = 3).
We have previously established thymic epithelial progenitor cells (named as TSC) as an in vitro experimental system to study mTEC differentiation37. Stimulation with recombinant human RANKL induced the expression of Aire and Aire-dependent PTAs in TSC cells37. To see if expression of miR-449a could be induced by RANKL, TSC cells were treated with 100ng/ml RANKL for 2 days. Upon RANKL stimulation, expression of miR-449a was significantly increased while other members of miR-449 cluster were undetectable (Fig. 1B). MiR-449 cluster members (miR-449c/449b/449a) share similar seed sequence with miR-34 cluster members (miR-34a, miR-34b/34c) and constitute a conserved miRNA family38,39,40. Unlike miR-449a, expression of miR-34a, miR-34b and miR-34c remained unchanged or undetectable (Fig. 1B). In addition, overexpression of RelA/p50 or RelB/p52 in TSC cells induced miR-449a expression (Fig. 1C), indicating that RANKL and downstream canonical or non-canonical NF-κB activation could induce miR-449a expression. In contrast, RelB-deficient thymic epithelial cells showed decreased miR-449a expression as well as Aire and PTAs expression, which were consistent with previous report (Fig. 1D)28. These results demonstrated that RANKL and downstream canonical or non-canonical NF-κB activation were sufficient to induce miR-449a expression in TECs.
MiR-449a expression profiling during thymus development
We further analyzed the temporal expression profiling of these miRNAs in thymus at development stages from E14.5 to 5-week old mice. Expression of miR-449a was dramatically increased at E18.5 and peaked at postnatal 10-day (Fig. 2A). However, the other two members of miR-449 family, miR-449b and miR-449c, showed very low background expression and only miR-449c showed a slight increase in new-born thymi (Fig. 2A). Unlike miR-449 cluster, expression of miR-34 cluster was consistently decreased from the detecting point E14.5 (Fig. 2B). To delicately trace miR-449a expression, we analyzed its expression in thymic lobes at E13.5, E14.5, E15.5, E16.5, E17.5 and E18.5 days. Q-PCR revealed continuous increase of miR-449a expression and a sharp jump at E15.5 (Fig. 2C). Intriguingly, miR-449a and Aire showed very similar temporal expression profiling during thymus development (Fig. 2D), indicating that miR-449a may function to promote mTEC maturation.
miR-449a expression profiling during thymus development was positively correlated with that of Aire expression. (A) q-PCR analyzed the expression of miR-449a, miR-449b and miR-449c in thymic epithelial cells of E14.5, E18.5, New-born, 10-day old and 5-week old thymi as indicated. (B) q-PCR analyzed the expression of miR-34a, miR-34b and miR-34c in thymic epithelial cells at indicated stages. (C) q-PCR analysis of miR-449a expression in thymic epithelial cells at indicated stages. (D) Comparison of expression profiling of miR-449a with that of Aire (Gray line for miR-449a, Black line for Aire). Bar graphs show means ± standard errors of at least three independent measurements. (*p < 0.05, **p < 0.01, n = 3).
Overexpression of miR-449a induced TEPC differentiation into mature mTEC in vitro
Aire, a core transcription factor, interacts with a large set of proteins to regulate PTAs expression and is considered as a biomarker of functional mature medullary thymic epithelial cells19,41. To see if miR-449a could induce mTEC maturation and Aire expression, we further stably overexpressed miR-449a in TSC cells (labeled as TSC miR-449a cells). Immunofluonrescence staining (Fig. 3A, white arrow: Aire+; blue arrow: Aire−) and flow cytometry (Fig. 3C) identified about 50% of Aire-expressing TSCs 4 days after miR-449a overexpression. The TEPC population within the fetal thymus has been defined with K5+K8+ double positive phenotype42. Immunofluonrescence staining of TSCs revealed both K5 and K8 expression in control cells while loss of K8 expression in TSC miR-449a cells, thus resulting in K5+K8−cells reminiscent of K5+ mTEC in thymus (Fig. 3B). Counting the Aire+ TSC cells in 5 randomly selected view fields from immunofluonrescence images showed about 50% Aire+ TSCs in TSC miR-449a cultures while none in TSC control cultures (Fig. 3A and D). There were about 20% K5+K8−TSCs in TSC control cultures, and this proportion increased to about 55% in TSC miR-449a cultures (Fig. 3B and E).
Overexpression of miR-449a induced TEPC differentiation into mature mTEC in vitro. (A,B) Immunofluoresence staining of TSC miR-449a cells and control cells with antibodies: EpCAM (G8.8), Aire (M-300) (A), K5 and K8 (B). (upper panel: TSC Control, lower panel: TSC miR-449a, white arrow in (A): Aire+, blue arrow in (A): Aire−,white arrow in (B): K5+K8−). (C) Flow cytometry analysis of TSC miR-449a cells and TSC Control cells with anti-Aire-Alexa 647 (5H12) and isotype control antibody. (D) Percent of Aire+ TSCs in cultures of TSC miR-449a cells and TSC Control cells from 5 randomly selected view fields of immunofluoresence images in (A). (E) Percent of K5+K8−TSCs in cultures of TSC miR-449a cells and TSC Control cells from 5 randomly selected view fields of immunofluoresence images in (B). (F) Immunoblot analysis of Aire, RelB, p52, DNMT3a and SATB2 in protein extracts of TSC miR-449a cells and TSC Control cells. GAPDH, Tubulin act as loading control. (G) RT-PCR analysis of indicated genes in TSC miR-449a cells and TSC Control cells, Tubulin acts as a loading control. (n ≧ 3).
Western blot assay also detected Aire protein in TSC miR-449a cells (Fig. 3F). At the same time, RelB protein was also increased coupled with the processing of p52, indicating the activation of non-canonical NF-κB during TSC differentiation (Fig. 3F). Interestingly, DNMT3a and SATB2 (2 epigenetic regulators and SATB2 as a miR-449a target in our previous study43) were significantly decreased in TSC cells after miR-449a overexpression (Fig. 3F). Further RT-PCR analysis confirmed the thymic identity of TSC miR-449a cells (Fig. 3G).
Furthermore, ectopic overexpression of miR-449a in 2-DG FTOC via lentiviral transduction induced expression of Aire and Aire-dependent PTAs (Fig. 4). Introduction of miR-449/34 sponge that was used to silence miR-449a was sufficient to neutralize the function of miR-449a in 2-DG FTOC (Fig. 4). Taken together, these data demonstrated that in in vitro studies using TSC cells or 2-DG FTOC, miR-449a was able to induce mTEC differentiation.
Overexpression of miR-449a induced differentiation of mTEC in 2-DG FTOC. q-PCR analysis of mRNA expression for aire, spt1, Insulin and CRP in 2-DG FTOC (Control, 2-DG FTOC; miR-449a, 2-DG FTOC infected with lentivirus expressing miR-449a for 4 days; miR-449a/miR-449/34 sponge, 2-DG FTOC infected with lentivirus expressing miR-449a and miR-449/34 sponge for 4 days). Bar graphs show means ± standard errors of at least three independent measurements. (*p < 0.05, **p < 0.01, n = 3).
Mutation of miR-449a alone does not affect thymus development in mouse model
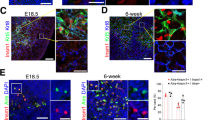
To investigate the function of miR-449a in vivo, we generated mouse mutants carrying an insert mutation or a deletion mutation in the miR-449a locus through CRISPR/Cas9 mediated gene editing (Fig. 5A)44. The insert mutant strain (miR-449aIns/Ins) bore a 20 bp insertion in miR-449a seed sequence following TGGCAGTGTATTGT while the deletion mutant strain (miR-449aDel/Del) bore a 30-bp deletion just after TGGCAGTGTATTGTTA (Fig. 5A). PCR analysis and further DNA sequencing demonstrated that both alleles of miR-449a were mutated in miR-449aIns/Ins and miR-449aDel/Del mice (Fig. S1A, B). Both mutant strains developed normally without gross defects in any organs and had normal reproductive ability.
Mutation of miR-449a alone did not affect thymus development in mouse model. (A) Sequence information of CRISPR/Cas9 mediated miR-449a mutants. (B) Immunofluoresence staining of thymus sections with antibodies to K5 (Green) and K8 (Blue). (C) Immunofluoresence staining of 12-week thymus sections with antibodies to K5 (Green) and Aire (Red). (n = 3).
We then analyzed the structure of mutant thymus. The mutant thymus displayed the same size as the heterozygous and wild type littermates (Fig. S2). Innunofluorescence staining of thymus sections showed normal thymic medulla and cortex distribution at 7-week, 9-week, and 12-week (Fig. 5B and S3). Aire expression in thymus sections had no significant difference (Fig. 5C). Transcripts for Aire and selected PTAs were quantified in thymic epithelial cells. Consistently, the expression of Aire and PTAs displayed no significant difference between miR-449a mutant mice and wild type littermates (Fig. S4A).
To further confirm if miR-449a was mutated in thymic epithelial cells, we analyzed the mature miR-449a expression in TECs. Q-PCR analysis revealed that mature miR-449a was nearly depleted in miR-449aIns/Ins (Fig. S4B) and miR-449aDel/Del mice (data not shown). However, the expression of miR-449b was slightly increased(Fig. S4B). The transcripts of miR-449a host gene CDC20b was unaffected indicating that the mutation of miR-449a had no side-effect on host gene expression (Fig. S4C). Collectively, these results suggested that the lack of miR-449a had no overall significant effects on thymus development.
Expression of miR-34a was increased in miR-449aIns/Ins thymus
The functional redundancy of miR-449/34 family has been previously reported38,39,40. We wondered if this functional redundancy was responsible for the lack of effects of miR-449a mutation on thymus development. We then analyzed the expression of miR-34 cluster members in TECs during thymus development at E14.5, E16.5, 3-week, 7-week and 12-week. To our surprise, miR-34a exhibited a marked increase in miR-449a deficient thymus (Fig. 6A). Mir-34b and miR-34c also showed an increase in E16.5 thymus but not in postnatal thymus (Fig. 6A).
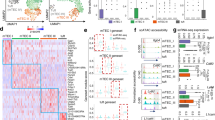
Expression of miR-34a was increased in miR-449aIns/Ins thymus. (A) q-PCR analysis of miR-34a, miR-34b and miR-34c expression in TECs of E14.5, E16.5, 3-week, 7-week, 12-week old miR-449aIns/Ins mice and their littermates. (B) In silico analysis of the miR-449 cluster and miR-34 cluster members. (C) 130 candidate targets of miR-34a and 128 candidate targets of miR-449a with target score >85 were predicted by miRDB, containing 124 overlapping candidates. Bar graphs show means ± standard errors of at least three independent measurements. (*p < 0.05, **p < 0.01, n = 3).
In silico analysis identified that members of the miR-449 cluster and miR-34 cluster possess similar mature sequences and seed regions (Fig. 6B). In addition, we analyzed the candidate targets of miR-34a and miR-449a by an online database miRDB45. Among the candidate targets with target score >85, most of the targets of miR-34a overlapped that of miR-449a (Fig. 6C and Table 1). Thus, these results supported the notion that the increased expression of miR-34a compensated for the absence of miR-449a in miR-449a-mutant mice.
Thymic in situ expression of miR-449/34 sponge reduced mature mTECs
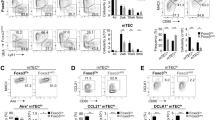
To further confirm the impact of miR-449a on mTEC development, we introduced miR-449/34 sponge-GFP in thymus through in situ injection of lentivirus in 3-week old mice. Expression of miR-449/34 sponge resulted in reduced GFP+ medulla (K14+GFP+) and augmented GFP+ cortex (K8+GFP+) in GFP-expression area (Fig. 7A 1st and 2nd panel). However, control-GFP expression could be detected in both medulla and cortex (Fig. 7A 3rd panel). Statistic analysis of the K14+ and K8+ zone in GFP-expressing area revealed that K14+ GFP+ mTEC was significantly reduced, while K8+ GFP+ cTEC was increased after expression of miR-449/34 sponge (Fig. 7B). Flow cytometry analysis of TEC subsets in infected thymus further confirmed reduction of mature MHCIIhi mTECs (Fig. 7C and D). Thus, these results indicated that expression of miR-449/34 sponge to interfere with miR-449a and other cluster members blocked normal maturation of mTECs.
Thymic in situ expression of miR-449/34 sponge reduced mature mTECs. (A) Immunofluoresence staining of thymus sections after in situ injection of control-GFP virus or miR-449/34 sponge-GFP virus with antibodies: K14 (Red) and K8 (Blue), Green indicated GFP expression. (B) Statistic analysis of the K14+GFP+ and K8+GFP+ zone in GFP-expressing area in 3 immunofluoresence images from (A). (C) Flow cytometry analysis of TECs after in situ injection of control-GFP virus or miR-449/34 sponge-GFP virus with antibodies: CD45-PEcy7, Ly51-Alexa 647, MHCII V500. (D) Frequencies of TEC populations (MHCIIhi mTEC: MHCIIhi Ly51−CD45−, MHClow mTEC: MHCIIlow Ly51−CD45−, cTEC: MHCII+ Ly51+CD45−) in (C). Data represent three independent experiments with at least three mice per group. *p < 0.05.
Discussion
The roles of miRNAs in adaptive immune system have been extensively investigated in T cells46,47, B cells48 and dendritic cells49 by conditional dysfunction of RNase III enzyme Dicer. Many miRNAs such as miR-14650, miR15551,52,53, miR-15054,55 have been reported to regulate T cell, B cell and dendritic cell development and function. The deletion of Dicer in TECs first revealed the global function of miRNAs in thymus33,34. However, the function of a single miRNA that regulates thymic epithelial cell development and function is rarely reported.
Here, analysis of miRNA expression in 2-DG FTOC identified that miR-449a was up-regulated by RANK ligand. Interestingly, by searching for the 3′UTR of mRNA sequence, miR-449a was predicted to target SATB2, a transcription factor that was highly expressed in embryonic stem cells56. Satb1 and the closely related Satb2 proteins regulate gene expression and higher-order chromatin structure of multigene clusters. The expression of Satb1 and Satb2 contributes to the plasticity of Nanog expression and SATB2 overexpression may functionally inhibit differentiation56. Consistently, expression of SATB2 was decreased in in vitro differentiation of TEPC into mature thymic epithelial cells by miR-449a from our data.
MiR-449a was highly expressed during thymus development and positively correlated with Aire expression. Furthermore, we demonstrated that miR-449a induced thymic epithelial progenitor cells differentiation into mature thymic epithelial cells in vitro. Despite the fact that mice with miR-449a mutation showed no discernible phenotype, the role of miR-449a in thymus development could not be entirely excluded because our data suggested that miR-34a compensated for the dysfunction of miR-449a. Although the expression levels of miR-449b, miR-449c, miR-34b and miR-34c were much lower than that of miR-449a, the expression abundance of miR-34a was comparable to that of miR-449a in developing thymus (data not shown). MiR-34a and miR-449a possessed similar seed sequence and had overlapped candidate targets as predicted. More importantly, expression of miR-34a was significantly up-regulated in miR-449a-mutant thymus, which in another way may indicate its compensation role.
To further confirm the impact of miR-449a on mTEC development, we introduced miR-449/34 sponge through thymic in situ injection of lentivirus in 3-week old mice. Injection of miR-449/34 sponge virus resulted in reduced GFP+ medulla and augmented GFP+ cortex, reflected on the extensive miR-449/34 sponge-GFP expression in cortex. Flow cytometry analysis of TEC subsets in infected thymus also showed reduction of mature MHCIIhi mTECs. Taken together, these results indicated that interference of miR-449a and miR-449/34 cluster blocked normal differentiation of mTEC and may also have impact on cTEC development. Clues from the expression profiling of miR-449/34 cluster during thymus development, miR-34 may function at early stage before E15.5 while miR-449 may regulate late differentiation of mTECs. The generation of miR-449/34 sponge transgenic mice or miR-449/34 knock out mice may help to reveal the molecular mechanisms of miR-449/34 in regulation of thymus development.
References
Klein, L., Hinterberger, M., Wirnsberger, G. & Kyewski, B. Antigen presentation in the thymus for positive selection and central tolerance induction. Nature reviews. Immunology 9, 833–844, https://doi.org/10.1038/nri2669 (2009).
Anderson, G. & Jenkinson, E. J. Lymphostromal interactions in thymic development and function. Nature reviews. Immunology 1, 31–40, https://doi.org/10.1038/35095500 (2001).
Blackburn, C. C. & Manley, N. R. Developing a new paradigm for thymus organogenesis. Nature reviews. Immunology 4, 278–289, https://doi.org/10.1038/nri1331 (2004).
Donskoy, E. & Goldschneider, I. Thymocytopoiesis is maintained by blood-borne precursors throughout postnatal life. A study in parabiotic mice. J Immunol 148, 1604–1612 (1992).
Schwarz, B. A. & Bhandoola, A. Circulating hematopoietic progenitors with T lineage potential. Nature immunology 5, 953–960, https://doi.org/10.1038/ni1101 (2004).
Mori, S., Shortman, K. & Wu, L. Characterization of thymus-seeding precursor cells from mouse bone marrow. Blood 98, 696–704 (2001).
Petrie, H. T. Cell migration and the control of post-natal T-cell lymphopoiesis in the thymus. Nature reviews. Immunology 3, 859–866, https://doi.org/10.1038/nri1223 (2003).
Lind, E. F., Prockop, S. E., Porritt, H. E. & Petrie, H. T. Mapping precursor movement through the postnatal thymus reveals specific microenvironments supporting defined stages of early lymphoid development. The Journal of experimental medicine 194, 127–134 (2001).
Ciofani, M. & Zuniga-Pflucker, J. C. The thymus as an inductive site for T lymphopoiesis. Annual review of cell and developmental biology 23, 463–493, https://doi.org/10.1146/annurev.cellbio.23.090506.123547 (2007).
Petrie, H. T. & Zuniga-Pflucker, J. C. Zoned out: functional mapping of stromal signaling microenvironments in the thymus. Annual review of immunology 25, 649–679, https://doi.org/10.1146/annurev.immunol.23.021704.115715 (2007).
Petrie, H. T., Scollay, R. & Shortman, K. Commitment to the T cell receptor-alpha beta or -gamma delta lineages can occur just prior to the onset of CD4 and CD8 expression among immature thymocytes. European journal of immunology 22, 2185–2188, https://doi.org/10.1002/eji.1830220836 (1992).
Ciofani, M., Knowles, G. C., Wiest, D. L., von Boehmer, H. & Zuniga-Pflucker, J. C. Stage-specific and differential notch dependency at the alphabeta and gammadelta T lineage bifurcation. Immunity 25, 105–116, https://doi.org/10.1016/j.immuni.2006.05.010 (2006).
Dudley, E. C., Petrie, H. T., Shah, L. M., Owen, M. J. & Hayday, A. C. T cell receptor beta chain gene rearrangement and selection during thymocyte development in adult mice. Immunity 1, 83–93 (1994).
Anderson, G., Jenkinson, E. J., Moore, N. C. & Owen, J. J. MHC class II-positive epithelium and mesenchyme cells are both required for T-cell development in the thymus. Nature 362, 70–73, https://doi.org/10.1038/362070a0 (1993).
Anderson, G., Owen, J. J., Moore, N. C. & Jenkinson, E. J. Thymic epithelial cells provide unique signals for positive selection of CD4+ CD8+ thymocytes in vitro. The Journal of experimental medicine 179, 2027–2031 (1994).
Derbinski, J., Schulte, A., Kyewski, B. & Klein, L. Promiscuous gene expression in medullary thymic epithelial cells mirrors the peripheral self. Nature immunology 2, 1032–1039, https://doi.org/10.1038/ni723 (2001).
Gallegos, A. M. & Bevan, M. J. Central tolerance to tissue-specific antigens mediated by direct and indirect antigen presentation. The Journal of experimental medicine 200, 1039–1049, https://doi.org/10.1084/jem.20041457 (2004).
Klein, L., Kyewski, B., Allen, P. M. & Hogquist, K. A. Positive and negative selection of the T cell repertoire: what thymocytes see (and don’t see). Nature reviews. Immunology 14, 377–391, https://doi.org/10.1038/nri3667 (2014).
Anderson, M. S. et al. Projection of an immunological self shadow within the thymus by the aire protein. Science 298, 1395–1401, https://doi.org/10.1126/science.1075958 (2002).
Kyewski, B. & Klein, L. A central role for central tolerance. Annual review of immunology 24, 571–606, https://doi.org/10.1146/annurev.immunol.23.021704.115601 (2006).
Oukka, M., Cohen-Tannoudji, M., Tanaka, Y., Babinet, C. & Kosmatopoulos, K. Medullary thymic epithelial cells induce tolerance to intracellular proteins. J Immunol 156, 968–975 (1996).
Rossi, S. W. et al. RANK signals from CD4(+)3(−) inducer cells regulate development of Aire-expressing epithelial cells in the thymic medulla. The Journal of experimental medicine 204, 1267–1272, https://doi.org/10.1084/jem.20062497 (2007).
Akiyama, T. et al. The tumor necrosis factor family receptors RANK and CD40 cooperatively establish the thymic medullary microenvironment and self-tolerance. Immunity 29, 423–437, https://doi.org/10.1016/j.immuni.2008.06.015 (2008).
Chin, R. K. et al. Lymphotoxin pathway directs thymic Aire expression. Nature immunology 4, 1121–1127, https://doi.org/10.1038/ni982 (2003).
Venanzi, E. S., Gray, D. H., Benoist, C. & Mathis, D. Lymphotoxin pathway and Aire influences on thymic medullary epithelial cells are unconnected. J Immunol 179, 5693–5700 (2007).
Kajiura, F. et al. NF-kappa B-inducing kinase establishes self-tolerance in a thymic stroma-dependent manner. J Immunol 172, 2067–2075 (2004).
Kinoshita, D. et al. Essential role of IkappaB kinase alpha in thymic organogenesis required for the establishment of self-tolerance. J Immunol 176, 3995–4002 (2006).
Burkly, L. et al. Expression of relB is required for the development of thymic medulla and dendritic cells. Nature 373, 531–536, https://doi.org/10.1038/373531a0 (1995).
Akiyama, T. et al. Dependence of self-tolerance on TRAF6-directed development of thymic stroma. Science 308, 248–251, https://doi.org/10.1126/science.1105677 (2005).
Alvarez-Garcia, I. & Miska, E. A. MicroRNA functions in animal development and human disease. Development 132, 4653–4662, https://doi.org/10.1242/dev.02073 (2005).
Sayed, D. & Abdellatif, M. MicroRNAs in development and disease. Physiological reviews 91, 827–887, https://doi.org/10.1152/physrev.00006.2010 (2011).
Ling, H., Fabbri, M. & Calin, G. A. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nature reviews. Drug discovery 12, 847–865, https://doi.org/10.1038/nrd4140 (2013).
Papadopoulou, A. S. et al. The thymic epithelial microRNA network elevates the threshold for infection-associated thymic involution via miR-29a mediated suppression of the IFN-alpha receptor. Nature immunology 13, 181–187, https://doi.org/10.1038/ni.2193 (2012).
Zuklys, S. et al. MicroRNAs control the maintenance of thymic epithelia and their competence for T lineage commitment and thymocyte selection. J Immunol 189, 3894–3904, https://doi.org/10.4049/jimmunol.1200783 (2012).
Khan, I. S., Taniguchi, R. T., Fasano, K. J., Anderson, M. S. & Jeker, L. T. Canonical microRNAs in thymic epithelial cells promote central tolerance. European journal of immunology 44, 1313–1319, https://doi.org/10.1002/eji.201344079 (2014).
Guo, Z. et al. Transcriptome analysis of murine thymic epithelial cells reveals ageassociated changes in microRNA expression. International journal of molecular medicine 32, 835–842, https://doi.org/10.3892/ijmm.2013.1471 (2013).
Chen, P. et al. Established thymic epithelial progenitor/stem cell-like cell lines differentiate into mature thymic epithelial cells and support T cell development. PloS one 8, e75222, https://doi.org/10.1371/journal.pone.0075222 (2013).
Bao, J. et al. MicroRNA-449 and microRNA-34b/c function redundantly in murine testes by targeting E2F transcription factor-retinoblastoma protein (E2F-pRb) pathway. The Journal of biological chemistry 287, 21686–21698, https://doi.org/10.1074/jbc.M111.328054 (2012).
Wu, J. et al. Two miRNA clusters, miR-34b/c and miR-449, are essential for normal brain development, motile ciliogenesis, and spermatogenesis. Proceedings of the National Academy of Sciences of the United States of America 111, E2851–2857, https://doi.org/10.1073/pnas.1407777111 (2014).
Song, R. et al. miR-34/449 miRNAs are required for motile ciliogenesis by repressing cp110. Nature 510, 115–120, https://doi.org/10.1038/nature13413 (2014).
Abramson, J., Giraud, M., Benoist, C. & Mathis, D. Aire’s partners in the molecular control of immunological tolerance. Cell 140, 123–135, https://doi.org/10.1016/j.cell.2009.12.030 (2010).
Bennett, A. R. et al. Identification and characterization of thymic epithelial progenitor cells. Immunity 16, 803–814 (2002).
Sun, X. et al. miR-449a inhibits colorectal cancer progression by targeting SATB2. Oncotarget. https://doi.org/10.18632/oncotarget.10900 (2016).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nature protocols 8, 2281–2308, https://doi.org/10.1038/nprot.2013.143 (2013).
Wang, X. & El Naqa, I. M. Prediction of both conserved and nonconserved microRNA targets in animals. Bioinformatics 24, 325–332, https://doi.org/10.1093/bioinformatics/btm595 (2008).
Muljo, S. A. et al. Aberrant T cell differentiation in the absence of Dicer. The Journal of experimental medicine 202, 261–269, https://doi.org/10.1084/jem.20050678 (2005).
Cobb, B. S. et al. T cell lineage choice and differentiation in the absence of the RNase III enzyme Dicer. The Journal of experimental medicine 201, 1367–1373, https://doi.org/10.1084/jem.20050572 (2005).
Koralov, S. B. et al. Dicer ablation affects antibody diversity and cell survival in the B lymphocyte lineage. Cell 132, 860–874, https://doi.org/10.1016/j.cell.2008.02.020 (2008).
Kuipers, H., Schnorfeil, F. M., Fehling, H. J., Bartels, H. & Brocker, T. Dicer-dependent microRNAs control maturation, function, and maintenance of Langerhans cells in vivo. J Immunol 185, 400–409, https://doi.org/10.4049/jimmunol.0903912 (2010).
Lu, L. F. et al. Function of miR-146a in controlling Treg cell-mediated regulation of Th1 responses. Cell 142, 914–929, https://doi.org/10.1016/j.cell.2010.08.012 (2010).
Thai, T. H. et al. Regulation of the germinal center response by microRNA-155. Science 316, 604–608, https://doi.org/10.1126/science.1141229 (2007).
Rodriguez, A. et al. Requirement of bic/microRNA-155 for normal immune function. Science 316, 608–611, https://doi.org/10.1126/science.1139253 (2007).
Lu, C. et al. miR-221 and miR-155 regulate human dendritic cell development, apoptosis, and IL-12 production through targeting of p27kip1, KPC1, and SOCS-1. Blood 117, 4293–4303, https://doi.org/10.1182/blood-2010-12-322503 (2011).
Xiao, C. et al. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell 131, 146–159, https://doi.org/10.1016/j.cell.2007.07.021 (2007).
Zhou, B., Wang, S., Mayr, C., Bartel, D. P. & Lodish, H. F. miR-150, a microRNA expressed in mature B and T cells, blocks early B cell development when expressed prematurely. Proceedings of the National Academy of Sciences of the United States of America 104, 7080–7085, https://doi.org/10.1073/pnas.0702409104 (2007).
Savarese, F. et al. Satb1 and Satb2 regulate embryonic stem cell differentiation and Nanog expression. Genes & development 23, 2625–2638, https://doi.org/10.1101/gad.1815709 (2009).
Acknowledgements
This work was supported by grants from National Natural Science Foundation of China (Grant No. 31401169).
Author information
Authors and Affiliations
Contributions
X.Z., P.C. and H.Z. designed all the experiments. P.C. wrote the initial draft of the manuscript. P.C. and H.Z. conducted the experiments and analyzed the data. Y.H., W.J., X.S., Z.L. and S.L. helped with the experiments. X.Z. supervised the study.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, P., Zhang, H., Sun, X. et al. microRNA-449a modulates medullary thymic epithelial cell differentiation. Sci Rep 7, 15915 (2017). https://doi.org/10.1038/s41598-017-16162-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-16162-2
This article is cited by
-
Epigenetic modifications in thymic epithelial cells: an evolutionary perspective for thymus atrophy
Clinical Epigenetics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.