Abstract
The Crumbs (Crb) complex is a key epithelial determinant. To understand its role in morphogenesis, we examined its function in the Drosophila pupal wing, an epithelium undergoing hexagonal packing and formation of planar-oriented hairs. Crb distribution is dynamic, being stabilized to the subapical region just before hair formation. Lack of crb or stardust, but not DPatj, affects hexagonal packing and delays hair formation, without impairing epithelial polarities but with increased fluctuations in cell junctions and perimeter length, fragmentation of adherens junctions and the actomyosin cytoskeleton. Crb interacts with Moesin and Yurt, FERM proteins regulating the actomyosin network. We found that Moesin and Yurt distribution at the subapical region depends on Crb. In contrast to previous reports, yurt, but not moesin, mutants phenocopy crb junctional defects. Moreover, while unaffected in crb mutants, cell perimeter increases in yurt mutant cells and decreases in the absence of moesin function. Our data suggest that Crb coordinates proper hexagonal packing and hair formation, by modulating junction integrity via Yurt and stabilizing cell perimeter via both Yurt and Moesin. The Drosophila pupal wing thus appears as a useful system to investigate the functional diversification of the Crb complex during morphogenesis, independently of its role in polarity.
Similar content being viewed by others
Introduction
The type I transmembrane protein Crumbs (Crb) is a key regulator of epithelial cell integrity, which has been strongly conserved across evolution1. In most fly epithelia, Crb localizes to a subapical region (SAR), a membrane region positioned just above adherens junctions (AJs) [refs2,3,4 and Fig. 1a], where it forms a complex with the intracellular adaptor Stardust [Sdt] (Pals1 in Vertebrates) and DPatj5,6. Crb has been initially identified in flies for its role in maintaining epithelial organization7 and then in the expansion of the apical membrane upon overexpression8. These results demonstrate the key role of Crb in the organization of the apical domain, as further supported by studies in vertebrates [reviewed in refs9,10,11]. During later Drosophila development, Crb is involved in the positioning and stability of adherens junctions12,13. Crb is also connected to the actin cytoskeleton by its intracellular FERM-binding domain that interacts with three actin-binding proteins: Moesin (moe)14, βH-spectrin14 and Yurt15. Moe and Yurt negatively regulate Crb association to the membrane in some epithelia15,16. Recent evidence shows that Crb regulates actomyosin dynamics specifically via Moe, during dorsal closure in the embryo17 and for the morphogenesis of the adult follicular epithelium16. Therefore, Crb sits at a key position at physical/functional intersection of the apical membrane domain, adherens junctions and actin cytoskeleton. Because crb mutant embryos usually present strong apical-basal (AP/BL) polarity defects, whether and how Crb could regulate apical organization during morphogenesis yet remains poorly understood.
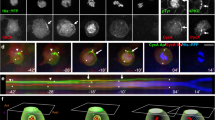
Crb displays a dynamic redistribution during pupal wing development. (a) Schematic drawing of a Drosophila epithelial cell, showing the position of the subapical region (SAR, in green) and of the adherens junctions (AJ, in red). (b–d and i,k) Crb (green) and Fmi (red) distribution in pupal wings at 25 °C at 16 h (b–d) or 30 h (i,k) APF; Red arrowheads in panel J show the Fmi zig-zag pattern oriented orthogonally to the PD axis. (e–h and l–o) Orthogonal sections of pupal wings at 16 h (e–h) or 30 h APF (l–o) stained for Crb (green), F-actin (red) and Dlg (blue). (p–r) Pupal wing at 32–34 h APF stained for Crb (blue) and F-actin (red). Red arrowheads in panel Q show Crb accumulation at the bottom of emerging hair. On the right of panels B–D and I–K drawn orthogonal views of a wing epithelial cell where the focal plane positions of the confocal image projections in the left panels are indicated (black line). All images are maximal projections of 2 up to 6 optical sections (every 0.2 μm). Distal is right, proximal left. Scale bar: 10 μm.
The Drosophila pupal wing represents a useful model to address the role of Crb in epithelia morphogenesis. Crb is not essential for AP/BL polarity in the third instar imaginal disc, the larval epithelium that develops into the pupal wing18,19. In the absence of intense cell proliferation, the pupal wing epithelium undergoes dramatic cell rearrangements, leading to a characteristic hexagonal cell packing. Hexagonal packing requires reorganization of the actin cytoskeleton and AJs, as well as polarized localization of proteins involved in Planar Cell Polarity (PCP)20,21,22. This eventually results in a monolayered epithelium, differentiating a single F-actin-rich prehair (trichome) at the distal vertex of each cell, with a defined proximal-distal (P/D) orientation. Mutations in genes that control wing morphogenesis lead to hair defects, as easily seen in the adult23,24,25. For instance, the loss-of-function of key cytoskeleton regulators such as Zipper (Myosin II heavy chain) leads to cells forming multiple hairs26,27,28,29,30,31,32. Thus, the apico-basal polarity, junction organization and apical cytoskeleton remodeling are intimately interconnected during wing differentiation33,34.
In this study, we investigated the role of Crb, Sdt and DPatj during pupal wing development. We found that both Crb and Sdt (but not DPatj) play a role in epithelial morphogenesis that is independent of the apico-basal or PCP pathways. Our data further indicate that Crb is necessary for the integrity and stability of E-cadherin (E-cad) and actomyosin at the adherens junctions at the end of hexagonal packing, a function likely mediated by Yurt. In addition, our results suggest a role of Crb in modulating opposed Moesin- and Yurt-dependent mechanisms for the regulation of the cell perimeter.
Results
Crb redistributes to the subapical region during pupal wing development
Although the putative function of Crb has never been examined in the development of adult wings that occurs during pupal stages, previous studies have noticed that Crb accumulates at the SAR of epithelial cells in the larval wing imaginal discs35,36,37, suggesting that Crb regulates epithelium morphogenesis at later stages of development.
As a first step, we investigated whether and where Crb accumulates in developing pupal wing cells. Pupal wing development comprises three major morphogenetic events: 1) cell packing [10–28 h after puparium formation (APF)], resulting from E-cad-dependent remodeling of cell contacts that allows irregularly shaped cells turning into honeycomb-packed hexagons; 2) establishment of the P/D axis (28–30 h APF), as visualized by typical pattern of the PCP protein Flamingo (Fmi) at the SAR; 3) a spectacular rearrangement of the actin cytoskeleton (32–34 h APF) for the formation of prehairs [reviewed in22,38,39].
We found that Crb is expressed throughout all morphogenetic stages at the apical region of pupal wing cells and displays highly dynamic redistribution during their differentiation (Fig. 1). During hexagonal packing (at 16 h APF), Crb exhibits a punctuated pattern both intracellularly and at the SAR and localizes just above a basolateral membrane marker, Discs large 1 tumor suppressor (Dlg)40 (Fig. 1b–h). Then, when the P/D axis is established, Crb distribution is less punctuated and more associated to the SAR (Fig. 1i–o), as in wing imaginal discs35,36,37. While Fmi becomes restricted to the P/D boundaries of the SAR (Fig. 1j and k, arrowheads), Crb has a uniform redistribution along the membrane (Fig. 1i and k). We found that Crb localization at the SAR is dependent of Sdt, since the genetic nullification of Sdt leads to the loss of Crb staining (Fig. S1a–c), as also reported in other epithelia41. In contrast, whereas the absence of DPatj leads to a decrease in Crb levels, it is not sufficient to prevent Crb accumulation at the SAR (Fig. S1g–i). Finally, when prehair formation takes place, Crb continues to associate to SAR and occasionally accumulates at the distal vertex of the cell (Fig. 1p–r, arrowheads).
Hence, Crb is expressed and apically localized within developing pupal wing cells, with a specific redistribution to the SAR coinciding with the end of hexagonal packing. These data therefore suggested that the Crb complex contributes to wing cell morphogenesis, a hypothesis we next evaluated using genetic analysis.
Crb is not required for apical/basal and planar polarities of the pupal wing epithelium, but participates in prehair formation and hexagonal packing during morphogenesis
To investigate the putative role of Crb in pupal wing morphogenesis, we examined prehair formation in tissues manipulated to inactivate the individual function of crb, sdt and dPatj using targeted expression of RNAi, as well as mosaic clones of null alleles (Figs 2 and S1). Compared to neighboring wild type (wt) cells (Figs 1p and 2a), cells homozygous for crb 11A22 (marked by loss of GFP, Fig. 2a–c) or expressing crbRNAi (Fig. 2d–f), exhibit a disorganized pattern of prehairs. Indeed, while ~90% of prehairs in wt tissue points to the P/D axis (with an angle between the prehair and the P/D axis of 0 to 15 degrees), prehairs alignment presents a higher variability in crb mutants cells, with an angle ranging from 0 to 90 degrees (Fig. 2g). Moreover, multiple prehairs per cell are often observed in crb mutant cells (Fig. 2a and d, arrowheads). We quantified 26.58% and 27.94% of cells showing multiple hairs in crbRNAi and crb 11A22 cells, respectively, compared to controls where multiple prehair cells are lower than 5% [nwings = 5, ncells = 100] (Fig. 2h). These defects therefore reinforce the hypothesis that Crb is required for the proper morphogenesis of pupal wing cells.
Depletion of crb expression affects prehair morphogenesis. (a–f) crb 11A22 clones (a–c) and crbRNAi cells (d–f) in pupal wings at 32–34 h APF, indicated by the absence of GFP or Crb (blue), respectively, stained for F-actin (red). Red arrowheads in panels A and D show double or triple mis-oriented prehair in crb mutant cells. Scale bar: 10 μm. (g) Histogram of angles (degrees) between prehair and the L3 vein in wt, crbRNAi, crb 11A22 cells at 32–34 h APF. (h) Tables showing the percentage of single and multiples prehair in wt, crbRNAi, crb 11A22 cells at 32–34 h APF.
Consistently, wing cells homozygous for a null allele of sdt (sdt k85) show a similar phenotype to that observed in the absence of Crb (Fig. S1d–f). In contrast, no defects in prehair formation are observed in DPatj 53 mutant cells (Fig. S1j–l). Surprisingly, wings containing crb 11A22 clones or crbRNAi knockdown produce normal-looking adult hairs (data not shown). It has been proposed that hair morphogenesis starts with the formation of multiple bundles of actin near the distal vertex, which merge over time to form a single hair20,26. The formation of multiple prehairs in crb mutants that eventually resolve into adult hairs of normal morphology, suggested a delay rather than an impairment of adult hair formation upon Crb depletion. Remarkably, mutations in mwh, a negative regulator of the actin cytoskeleton, induce a similar delay in hair development42. Therefore, our results argue for a cell-autonomous role of Crb in the regulation of hair development.
Crb is essential for apical domain organization and AP/BL polarity in most epithelia analyzed8,43. Therefore, we addressed whether prehair defects were associated with an alteration of cell polarity. Analysis of the basolateral membrane marker Dlg40 in crbRNAi and crb 11A22 cells does not show any changes compared to wt tissues (Fig. S2a–l and data not shown). Orthogonal sections show that the localization of AJs, visualized by E-cad staining, is indistinguishable in crbRNAi cells from wt cells (Fig. S2g–l). Also, no differences in cell height are detected between wt and crbRNAi cells (Fig. S2g–l). These observations argue that AP/BL polarity in crb mutants is not affected, as opposed to other Drosophila tissues8,43.
Although a function of Crb in PCP has not been previously addressed, the crb mutant phenotype in prehair formation could be linked to defects in the establishment and/or maintenance of the P/D axis, a hypothesis we next assayed through examining the distribution of Fmi. As in wt cells, the asymmetric localization of Fmi at the proximal-distal cell boundaries (giving rise to a characteristic zig-zag pattern of Fmi) is unchanged (red dots) in crbRNAi cells (Fig. S2m–r, red dots) and in crb 11A22 clones (not shown), showing that PCP polarity is not disrupted upon crb depletion.
Taken together, our data therefore indicate that Crb is not involved in the maintenance of AP/BL polarity or the establishment of PCP in pupal wing cells, but Crb is instead required for proper prehair formation through an independent pathway.
The proper formation of prehairs in pupal wing cells requires hexagonal packing22,38,39. During hexagonal packing cells change their shape and, concomitantly, increasingly point to the P/D axis in response to tissue stretching44,45. At the end of tissue remodeling (28–30 h APF), all cells are turned into a hexagon, displaying regular vertex to vertex distances and highly similar cell perimeters (Fig. 3a–c). This results in an ordered honeycomb-like pattern of cell junctions, the asymmetric distribution of PCP components along the P/D cell sides defining the distal-most apical vertex where prehairs start growing and become aligned on each other.
Crb is required for the integrity of adherens junctions and of the F-actin cytoskeleton belt and for the P/D orientation of epithelial cells during hexagonal packing (a–l) wt (a–c), crbRNAi (d–f) and crb 11A22 cells (g–l) in pupal wings at 28–30 h APF, stained for F-actin (red) and E-cad (green, a–i) or Dlg (green, j–l). crb 11A22 clones are indicated by the absence of GFP (blue). Red arrowheads in d, e, g and h show cortical gaps devoid of F-actin and E-cad. Red arrows in d show the intracellular accumulation of F-actin. On the right of panels a-l, drawn orthogonal views of a wing epithelial cell where the focal plane positions of the confocal image projections in the left panels are indicated (black line). All images are maximal projections of 2 up to 6 optical sections (every 0.2 mm). Distal is right, proximal left. (m) Histogram of vertex number in wt, crbRNAi, crb 11A22 cells at 28–30 h APF. Note that the percentage of cells with 6 vertices (hexagons) is similar between wt and crb mutant tissues. Bars indicate mean values ± SEM. nwings =9, ncells =180 crbRNAi and wt crbRNAi were analyzed. (n) wt (on the top) and crb 11A22 (on the bottom) cells at 30–34 h APF marked with E-cad. Cell orientation angle corresponding to the absolute angle between the longest axis of the cell and the vein L3 was drawn with bars in each cell. Red bars correspond to an angle <25° whereas green bars correspond to an angle >25°. (o) Histogram of the distribution of cell orientation angle in crbRNAi and its wt control, crb 11A22 and its wt twin clone cells at 30–34 h APF. Note that the percentage of cells with an orientation angle under 15° is lower in crb mutants than in wt cells. Bars indicate mean values of cells percentage ± SEM and statistical significance was analyzed by Student’s t-test {crb 11A22[0,15[=23.0% ± 10.65, versus wt[0,15[=78.6% ± 6.11; crb 11A22[15,30[=25.7% ± 6.80, versus wt[15,30[=17.1% ± 3.55; crb 11A22[30,90[=51.3% ± 17.45, versus wt[30,90[=4.6% ± 3.17; nwings=3, ncells=60, P < 0.05; crbRNAi[0,15[=61.2% ± 3.80, versus wt[0,15[=80.9% ± 3.74; crbRNAi[15,30[=25.9% ± 1.68, versus wt[15,30[=18.9% ± 4.20, crbRNAi[30,90[=13.0% ± 2.19, versus wt[30,90[=2.8% ± 1.31; nwings=9, ncells=180; P < 0.05]. Scale bar: 10 mm.
We observed that the inactivation of crb strongly disrupts tissue rearrangement, with a loss of the honeycomb-like pattern (Figs 3e and S2p). Quantification of the apical cell perimeter does not detect significant differences between wt and crb mutant cells (Fig. S3a, see Figure legend), and most crb mutant cells retain six vertices (Fig. 3m), without significant modification of the average vertex to vertex distance (Fig. S3b, see Figure legend). Next, we addressed whether hexagons point to the P/D axis by measuring the orientation of hexagonal cells, which we defined by the angle between the longest axis of a fitted ellipse and the vein 3 that is parallel to the P/D axis (See Materials and methods and Fig. 3n). Remarkably, we found that the alignment of hexagonal cells was altered in crb mutants. Indeed, while >80% of control cells displays an average angle ≤15 degrees, crb mutant cells exhibit a higher variability in their orientation and pointing to the P/D axis (Fig. 3n). These results thus suggest a role of Crb in regulating hexagonal packing through the P/D alignment of cells.
Crb is required during wing morphogenesis for the integrity and stability of adherens junctions and circumferential actomyosin belt
The defects in prehair orientation and hexagonal alignment of Crb mutant cells are also associated to strong alterations of cells junctions and actin cytoskeleton. Cell junctions are often wiggly (Fig. 4a) and show higher length fluctuations amplitude over time when compared to wt cells [Amplitude of junction length fluctuation over time in wt cells = 0.10 ± 0.01 A.U. versus crbRNAi = 0.17 ± 0.01 A.U. (nwings = 5, ncells = 40), P < 0.0001] (Fig. 4b), suggesting altered cortical tension. Consistently, the apical surfaces of crb mutant cells constrict and expand with higher amplitude than in wt cells [mean of cell perimeter length fluctuation over time in wt cells = 0.30 ± 0.02 A.U. versus crbRNAi = 0.78 ± 0.07 A.U., nwings = 5, ncells = 10, P < 0.0001] (Fig. S3c and d). We also observed a fragmentation of E-cad staining in both crbRNAi (Figs 3e and 4c and d, red arrowheads and movie 1) and crb 11A22 cells (Fig. 3g–i, red arrowheads). Live imaging of crbRNAi cells revealed that these gaps are transient (Fig. 4c and d, arrowheads, and movie 1), quickly forming when junctions expand and disappearing when junctions retract.
Crb is required for the stability of vertex-vertex length fluctuations. (a–d) wt and crbRNAi pupal wing cells expressing E-cad::GFP at 28–30 h APF were imaged in vivo to follow the evolution of vertex-vertex length (a,b) and the evolution of gaps devoid of E-cad (c,d). (a) Example of variation of vertex-vertex distance length. Vertex-vertex distance is color-coded (heat map from red to yellow) based on the percentage of vertex-vertex distance decrease calculated with respect to the vertex-vertex distance captured during imaging. (b) Graph of the evolution of vertex-vertex length variation amplitude expressed in A.U. in 10 wt (red) and 10 crbRNAi (black) cells. Each distance is normalized by its average length over time and expressed in arbitrary units (A.U.). (c) Example of the evolution of gaps devoid of E-cad (red arrowheads) in wt and crbRNAi pupal wing cells. (d) Higher magnification of a cell-cell contact imaged over the indicated time and its associated kymograph. Scale bar: 0.5 μm.
The defects observed in crb mutant cells supported an alteration in membrane tension, which results from the interaction between the E-cadherin adhesion system and the contractile actomyosin cytoskeleton [reviewed in ref.46]. The existence of E-cad and actomyosin clusters involved respectively in junctional stability and contractility has previously been described in mammalian cells47,48. Consistently, we observed that the junctional F-actin was abnormally fragmented in crb mutant cells, and regions lacking junctional F-actin are the ones also devoid of E-cad (Fig. 3d–j, arrowheads). Further aberration of the circumferential actomyosin belt was also visualized at the SAR, where F-actin is delocalized intracellularly in the subapical cytoplasm (Fig. 3d, arrows, and Fig. S2q). We did not, however, detect obvious alteration of basolateral F-actin (Fig. S2b and e), consistent with a specific role of Crb at the SAR. Similar actin defects are also observed in crb 11A22 null clones (Fig. 3j–l).
Taken together, these data show that the absence of Crb causes a correlated fragmentation of the cortical actin cytoskeleton and AJs, and a larger fluctuation in both junction length and apical cell perimeter. These results thus suggest that Crb stabilizes E-cad and the actin cytoskeleton at the adherens junctions to regulate proper hexagonal packing.
Loss of the FERM protein Moesin alters cell perimeter and F-actin accumulation but not the circumferential actomyosin belt and adherens junctions
In order to elucidate how Crb regulates the circumferential actomyosin belt and adherens junctions, we analyzed the contribution of Moe, a FERM protein that interacts with Crb and regulates actin-based cell shape14,49,50,51. Interestingly, Moe mostly localizes at the SAR at the end of hexagonal packing (Fig. 5a and c) and its localization is perturbed upon Crb depletion (Fig. 5b–d). To determine whether moe had a role in pupal wing morphogenesis, we first analyzed an available strong allele, moe PL106, resulting from a P-element insertion49,50. None of the defects observed for crb mutant cells were detected in clones of moe PL106 cells. However, immuno-staining revealed remnants of Moe levels in moe PL106 cells (Fig. S4e), precluding a definitive conclusion. We therefore generated a true moe loss-of-function allele, Δmoe, by inducing a targeted genomic deletion that removes most moe coding sequences (Fig. S4g and see Methods). Although Moe is clearly absent in mutant clones (Fig. 5f and i), cells lacking moe do not display the clear mis-organization of prehairs observed for crb mutants (Fig. 5k). However, we detected a strong apical constriction phenotype in Δmoe cells (Fig. 5e,h and Movie 2). Quantification of the cell perimeter of Δmoe cells localized in the middle of mutant clones indicated a strong decrease, compared to wt cells of the corresponding twin clones (Δmoe = 8.70 μm ± 0.10 versus wt = 11.30 μm ± 0.10, ncells = 100, P < 0.0001; Fig. 5l). Interestingly, the decrease in cell perimeter is less pronounced for Δmoe cells positioned along the clone border (Δmoe edge cells = 9.50 μm ± 0.10, versus wt edge cells = 10.30μm ± 0.10, ncells = 100, P < 0.001). The reduction in cell perimeter is also associated to a decrease in the average junction length (Δmoe = 1.60 μm ± 0.05, versus wt = 2.00 μm ± 0.05, ncells = 100, P < 0.001; Fig. 5l). Crb levels at the SAR appeared increased in Δmoe cells (Fig. 5m,n). Nonetheless, this increase in Crb accumulation can be explained by the associated decrease in the perimeter of Δmoe cells. Indeed, Crb intensity per length unit (pixel intensity average) remains mainly unchanged in Δmoe cells (Δmoe = 7.18 ± 0.08 × 106 A.U., versus wt = 6.96 ± 0.08 × 106 A.U., nwings = 5, ncells = 70; P > 0.05, Fig. 5o). Moreover, an accumulation at the membrane is also observed for other proteins, such as E-cad and Sdt (Fig. S5a–f), as well as Fmi, despite the absence of a significate increase in Fmi intensity per length unit (Δmoe = 6.29 ± 0.08 × 106 A.U. versus wt = 6.06 ± 0.08 × 106 A.U., nwings = 5, ncells = 100; P > 0.05, Fig. S5g).
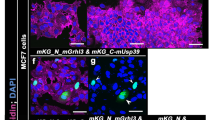
Loss of Moesin induces apical cell perimeter constriction, but does not affect adherens junctions. (a–d) Staining of Moe in wt (a), crbRNAi (b) or in crb 11A22 (c,d) pupal wing cells at 28–30 h APF. In crb 11a22 the wt twin clone is labelled by GFP expression (blue), Moe staining is in green (c,d). (e–k) Pupal wings containing Δmoe clones at 28–30 h (e–j) or 32–34 h (k) APF stained for E-cad (green), F-actin (red) or Moe (blue). E-cad and F-actin staining revealed defects in cell perimeter (e,h) or apical F-actin redistribution (h). On the right of panels a-j, drawn orthogonal views of a wing epithelial cell where the focal plane positions of the confocal image projections in the left panels are indicated (black line). All images are maximal projections of 2 up to 6 optical sections (every 0.2 μm). Distal is right, proximal left. Scale bar: 10 μm. (l) Quantification of cell perimeter length (Left) and vertex-vertex distances (Right) (μm) in wt and Δmoe clones (in the center of the clone and along the border of the clone) in wings at 28–30 h APF. Bars indicate mean values ± SEM and statistical significance was analyzed by Student’s t-test [Δmoe clones versus wt cells, ncells = 200, P < 0.0001, see the main text]. (m,n) Pupal wings containing Δmoe clones at 28–30 h APF stained for Crb (green). Moe depletion is revealed by the absence of GFP (blue). Distal is right, proximal left. Scale bar: 10 μm. (o) Quantification of Crb staining at the SAR in wt and Δmoe clones (in the center of the clone and along the border of the clone) in pupal wings at 28–30 h APF. Crb intensity per length unit (pixel intensity average) at the apico-lateral cortex was calculated and expressed in A.U. Bars indicate mean values of intensity ± SEM and statistical significance was analyzed by Student’s t-test [Δmoe = 7.18 ± 0.08 × 106 A.U., versus wt = 6.96 ± 0.08 × 106 A.U., nwings = 5, ncells = 70; P > 0.05].
Consistently with the role of Moe as a regulator of apical actin, F-actin accumulates at the apical cytoplasm in Δmoe cells, while some signal could still be detected at the AJs (Fig. 5h). However, the distribution of both E-cad (Fig. 5e) and F-actin (Fig. 5h) is not fragmented and we observed no gaps in the adherens junctions in Δmoe pupal wing cells, as opposed to the defects featuring observed for crb mutant cells (see Movie 2).
We thus concluded that Crb controls Moe distribution at the SAR, suggesting a functional connection between these proteins. Our data yet indicate that Moe is not directly involved in the stabilization of adherens junctions, but instead it mainly acts to regulate the apical cell perimeter during pupal wing morphogenesis.
The FERM protein Yurt is essential for the proper organization of the circumferential actomyosin belt and adherens junctions
Crb directly binds via its cytoplasmic FERM-binding region to Yurt52,53, which is known to negatively control Crb association to tight junctions in mammals15. To decipher how E-cad and F-actin gaps at the AJs are produced independently of Moe, we focused on the possible involvement of Yurt in this process. Interestingly, we observed that Yurt associates to the SAR at the end of hexagonal packing (as revealed by Yurt staining and comparing wt and yurtRNAi cells, Fig. 6a,b and d). Yurt localization at the SAR is dependent on Crb because this staining is lost upon crb inactivation (Fig. 6b and c). In contrast to other tissues or developmental stages, yurt knockdown by two distinct RNAi does not affect Crb accumulation at the SAR, even if its distribution is slightly more discontinuous (Fig. 6e,f, and not shown). In contrast to the lack of Moe, Yurt depletion leads to a strong disorganization of prehairs (Fig. 6g) and to prominent junctional defects similar to those observed in crb mutants, i.e., a correlated disruption of E-cad and F-actin at adherens junctions that could explain the observed discontinuities for Crb distribution (Fig. 6i–k). A phenotype not seen in the absence of Crb, however, is an increase in the perimeter of yurtRNAi cells (13.90 μm ± 0.20 versus wt 12.30 μm ± 0.08; nwings = 5, ncells = 200, P < 0.001), as also supported by an increased mean distance between vertices (yurtRNAi = 2.50 μm ± 0.07 versus wt = 1.90 μm ± 0.05; nwings = 5, ncells = 200, P < 0.001, Fig. 6h).
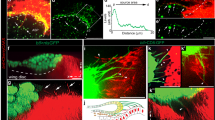
Loss of yurt induces cell perimeter expansion and affects the circumferential actomyosin belt and adherens junctions. (a–d) Staining of Yurt in wt (A, at the top of the wing), crbRNAi (a, at the bottom of the wing and c) and yurtRNAi (d) pupal wings (at 28–30 h APF). Higher magnifications of wt (b) and crbRNAi (c) tissues are shown. Note that while the cortical staining of Yurt disappears in yurtRNAi cells, the intracellular staining remained mainly unchanged suggesting that it is not specific (d). (e,f) Staining of Crb in wt (e) and yurtRNAi (f) pupal wings (28–30 h APF). Note that in yurtRNAi cells, although fragmented, Crb still associates to the SAR. (g) F-actin staining in yurtRNAi pupal wings at 32 h APF show defects in prehair organization. (h) Quantification of cell perimeter length and vertex–vertex distance (μm) in wt and yurtRNAi cells at 28–30 h APF. Bars indicate mean values ± SEM and significance was analyzed by Student’s t-test [yurtRNAi versus wt cells, ncells = 200, P < 0.0001, see main text]. (i–k) Staining of E-cad (green) and F-actin (red) in wt and yurtRNAi pupal wings (28–30 h APF) reveals defects in junctional integrity in yurtRNAi. Red arrowheads show gaps devoid of F-actin and E-cad staining at the adherens junctions. On the left of panels A–C and on the right of panels d–f and i–k, drawn orthogonal views of a wing epithelial cell where the focal plane positions of the confocal image projections in the left panels are indicated (black line). All images are maximal projections of 2 up to 6 optical sections (every 0.2 μm). Distal is right, proximal left. Scale bar: 10 μm.
Myosin II (Myo) has been implicated in the regulation of tension during epithelial morphogenesis and Myosin apical accumulation preceded the constriction and intercalation of embryonic cells48,54,55. Inactivation of myosin II (zipper) in wing cells leads to multiple hairs26,27,28,29,30,32. To better understand the cytoskeletal modifications observed in crb and yurt mutants, we next examined Myo distribution in these contexts (Fig. 7). In wt wing cells at the end of hexagonal packing Myo tightly associates to AJs and vertices (Fig. 7a–f), as previously shown in embryos54,56. In crbRNAi cells we found that Myo is missing from the gaps depleted in E-cad and F-actin (Fig. 7g), while strongly accumulating in rings around these gaps (Fig. 7g–l, arrowheads). In yurtRNAi cells, Myo was diffuse in the cytoplasm, being completely lost from the SAR (Fig. 7m–o) and correlating with an increase in the average vertex-vertex distance. Noticeably, a decrease in Myo has been previously associated to a loss of contractility and to an increase in junction length in pupal wing cells56.
Myo2 distribution is affected in crbRNAi and yurtRNAi cells. wt (a–f), crbRNAi (g–l) and yurtRNAi (m–o) pupal wing cells at 28–30 h APF stained for E-cad (green) and Myo2 (red). Red arrowheads in g and h show gaps at the cortex devoid of E-cad staining. Red arrows in j and l show a gap at the cortex devoid of E-cad and surrounded by Myo2 accumulation. All images are maximal projections of 2 up to 6 optical sections (every 0.2 μm). Distal is right, proximal left. Scale bar: 10 μm.
Taken together, our results show that Crb is required for the proper localization of Yurt at the SAR at the end of hexagonal packing. The similar defects following crb or yurt knockdown on the distribution of both E-cad and actomyosin cytoskeleton at the AJs further suggests a common function in the stabilization of junctions during pupal wing morphogenesis.
Discussion
Our study aimed at unveiling the function of the Crumbs complex in epithelial morphogenesis. Although Crb was discovered several decades ago in Drosophila 7, the severe apico-basal polarity defects associated to crb inactivation in embryos have hampered the full exploration of its function during epithelia development. Our results indicate that Crb also acts during pupal wing morphogenesis, where the absence of crb function does not impair AP/BL polarity and does not lead to the dramatic tissue alterations often seen in other tissues. The pupal wing thus represents an attractive model system, well suited to dissect additional functions of the Crb complex during epithelial morphogenesis, independently of its role in polarity.
The redistribution of Crb at the SAR at the end of hexagonal packing, as well as the defects in cells orientation observed in crb mutants suggest that Crb is required to stabilize the actin cytoskeleton and E-cadherin at the adherens junctions at the end of tissue rearrangement. Alterations in F-actin and Myo distribution in crb mutant cells strikingly mimic those observed in embryos mutant for the actin-binding protein Canoe/Afadin, which links the actomyosin network to AJs54. Canoe loss diminishes this coupling leading to reduced cell shape anisometry and defects in germ band elongation. As for crb, canoe mutant cells still retain some ability to change their shape and germ band elongation is delayed and not completely impaired. The defects observed in crb mutant cells support the hypothesis that Crb is a crucial regulator of the interconnection between the actomyosin cytoskeleton and AJs.
The fragmentation of AJs upon Crb depletion has been already described, for example in embryo2,17 or during follicular morphogenesis16. However, in these two systems the function of Crb has been related to the role of Moe in the regulation of the actomyosin cytoskeleton, while the role of Yurt has never been addressed or has been excluded. Our data support that in pupal wing cells the role of Crb in the stability of the AJs is likely established via Yurt. We show that Crb modulates Yurt localization at the SAR at the end of hexagonal packing and yurt mutant cells phenocopy crb mutant cortical defects. Nonetheless, previous studies in cultured cells have established that Yurt participates in epithelial polarity and organization of apical membranes by negatively regulating the activity of the Crb complex15,57. On the contrary, we show that, whereas Crb modulates Yurt distribution at the SAR at the end of hexagonal packing of wing cells, Yurt depletion does not impact Crb association to the SAR, with the exception of the E-cad- and F-actin-devoid gaps. Yurt and Crb similarly act on actomyosin and E-cad organization at the cell-cell junctions suggesting that the coordinated function of these two proteins is regulated by different mechanisms in different tissues. On the other hand, moe depletion does not specifically modify Crb distribution at the SAR, a finding coherent with the evidence that Moe is not implicated in stability of AJs in this tissue, as opposed to other models16.
Studies based on in vivo mechanical measurements or mathematical/physical modeling have proposed that epithelial cell packing results from a balance between intrinsic cell tension and extrinsic tissue-wide forces to establish a correct and robust order in the tissue44,46,58,59. Hence, the tension generated by the actomyosin cortex and the pressure transmitted through adherens junctions are the two main self-organizing forces driving tissue morphogenesis. Tension shortens cell-cell contacts and pressure of individual cells counteracts tension to maintain cell size44,48,60,61. Our data indicate that Crb recruits at SAR Moe and Yurt, which show opposite effects on pupal wing morphogenesis. While Moe promotes cell expansion, Yurt controls cell constriction and the stability of the AJs and of the actomyosin network. In crb mutant cells, the absence of variation in the cell perimeter might be explained by the simultaneous loss of positive and negative regulators. Therefore, Crb acts as a coordinator of the two self-organizing mechanisms implicated in morphogenesis. Additionally, the dynamic redistribution of Crb at the SAR at the end of hexagonal packing, together with the disruption of cell orientation in crb mutants, is consistent with the hypothesis that Crb is required to stabilize cell shape and pattern in order to properly progress throughout tissue development.
In conclusion, these functional analyses during pupal wing morphogenesis allowed us unraveling Crb-dependent mechanisms that are integrated to produce shape changes during development independently of epithelial polarity. Furthermore, our results show that the interplay between Crb and FERM proteins is tissue-regulated and that their epistatic interactions differ in a spatio-temporal manner.
Methods
Drosophila stocks and crosses
Control and driver strains (Bloomington Drosophila Stock Center, Indiana University); Transgenic lines used were UAS-crb RNAi (line 39177), UAS-Yurt RNAi (VDRC 107016 and 26674), (Vienna Drosophila RNAi Center, Vienna, Austria). These transgenic lines were crossed to ptc-GAL4 at 25 °C.
Mutant strains were crb 11A22 7; Sdt k85 62; DPatj 53, 63, Moe PL106 51. Mutant clones were generated using the FLP/FRT technique64. Crosses were grown at 25 °C and clones were recovered from pupae of the following genotypes:
hs > flp;; FRT82B, crumbs 11A22 /FRT82B, ubi > GFP;
Δmoe FRT19A/ubi > mRFP, hs > flp FRT19A
Sdt k85 FRT19A/ubi > mRFP; hs > flp FRT19A
hs > flp; DPatj 53 FRT2A/ubi > GFP, FRT2A
Mutant clones were generated by heat-shocking L2 larvae for 1 h at 37 °C. Pupae were dissected at 16–18, 28–30 or 32 h APF. Staging of pupal wing development at 25 °C were performed as described39,65.
CG12075 coding for Moesin extends from 8767045 to 8792365 bp on the X chromosome. A null allele was generated by targeted deletion of the Moesin coding region using site specific recombination between two Piggy Bac elements [e02421] and [e04400] (Exelixis) as described in66. The resulting deficiencies, carrying the recombinant (hybrid element), were characterized molecularly by PCR using transposon or genomic specific primers according to67. The recombinant hybrid element was subsequently eliminated by precise excision and the resulting null allele for Moesin, selected on the basis of white eyes (loss of mW+), was characterized molecularly by PCR and sequencing.
Immunofluorescence and antibodies
The head and the bottom of the pupae was dissected in PBS and quickly transferred in PFA 4% at room temperature for one hour. Dissected pupae were transferred into PBS-TN (PBS-0.3% Triton-20% NGS), the pupal case was removed and finally the wing was extracted. Washes were done in PBS-TN. Primary antibodies were incubated in PBS-TN, overnight at 4 °C. Secondary antibodies were incubated in PBS-TN for 1 hour at room temperature.
For Crb localization at SAR antibody staining in pupal wings were performed as previously described39,68 by using PBS 0.01% Triton X-100. Primary antibodies were: mouse anti-Fmi [1:20], mouse 4F3 anti-Dlg and rat anti-DE-Cad2 [1:100] from Developmental Studies Hybridoma Bank (DSHB, University of Iowa, USA); rat anti-Crumbs2-87, rabbit anti-DPatj69, rat anti-Sdt62, rabbit anti-Moesin70, rat anti-Yurt15; anti-Myosin II71. Secondary antibodies and Rhodamine-conjugated phalloidin were from Molecular Probes and Jackson ImmunoResearch Laboratories. Confocal images were acquired at 40x, 63x and 100x magnification on a LSM 510 Zeiss Confocal Microscope. Confocal sections were spaced 0.5 μm apart.
Quantifications
All quantifications were done on single optical sections corresponding to the AJ plane. For image analysis (cell perimeter, vertex-vertex distance, vertex number per cells, Crb mean intensity at SAR) we used the software “packing analyzer v2.0”45. Measurements of vertex-vertex length over time were done manually using ImageJ, and were normalized by its average vertex-vertex length over time.
For quantification of the orientation angle of hexagonal cells, cells were segmented and then analyzed with a custom written Matlab code based on the regionsprops function. Briefly, based on the segmented images, an ellipse was fitted on each cell (see below) and different parameters were extracted, such as the length of the Major and Minor axis allowing computing both the eccentricity and the orientation of the ellipse.
Cell eccentricity was computed using this equation (1):
If cells are totally round with a LengthMajorAxis = LengthMinor Axis, eccentricity value is 0. In this study we used 0.5 as threshold for an elongated (>0.5, Cell 1 blue, below) and anisotropic cell (<0.5, Cell 2 green, Table 1, below).
The ellipse orientation (in degrees ranging from 0° to 90°) is defined as the angle between the vein 3, which define the P/D axis, and the major axis of the ellipse, allowing to discriminate between a random oriented cell (angle >25°) and cell oriented in the plane of the wing elongation (angle <25°). We considered for quantification: for wt and crbRNAi 20 cells per wing, nwings = 9 with total ncells analyzed = 180; and for crb 11A22 and its wt twin clone 20 cells per wing, nwings = 3 with total ncells analyzed = 60.
References
Le Bivic, A. Evolution and cell physiology. 4. Why invent yet another protein complex to build junctions in epithelial cells? American journal of physiology. Cell physiology 305, C1193–1201, https://doi.org/10.1152/ajpcell.00272.2013 (2013).
Tepass, U. Crumbs, a component of the apical membrane, is required for zonula adherens formation in primary epithelia of Drosophila. Developmental biology 177, 217–225, https://doi.org/10.1006/dbio.1996.0157 (1996).
Bachmann, A., Schneider, M., Theilenberg, E., Grawe, F. & Knust, E. Drosophila Stardust is a partner of Crumbs in the control of epithelial cell polarity. Nature 414, 638–643, https://doi.org/10.1038/414638a (2001).
Harris, T. J. & Peifer, M. The positioning and segregation of apical cues during epithelial polarity establishment in Drosophila. The Journal of cell biology 170, 813–823, https://doi.org/10.1083/jcb.200505127 (2005).
Bulgakova, N. A. & Knust, E. The Crumbs complex: from epithelial-cell polarity to retinal degeneration. Journal of cell science 122, 2587–2596, https://doi.org/10.1242/jcs.023648 (2009).
Bazellieres, E., Assemat, E., Arsanto, J. P., Le Bivic, A. & Massey-Harroche, D. Crumbs proteins in epithelial morphogenesis. Frontiers in bioscience 14, 2149–2169 (2009).
Tepass, U., Theres, C. & Knust, E. crumbs encodes an EGF-like protein expressed on apical membranes of Drosophila epithelial cells and required for organization of epithelia. Cell 61, 787–799 (1990).
Wodarz, A., Hinz, U., Engelbert, M. & Knust, E. Expression of crumbs confers apical character on plasma membrane domains of ectodermal epithelia of Drosophila. Cell 82, 67–76 (1995).
Assemat, E., Bazellieres, E., Pallesi-Pocachard, E., Le Bivic, A. & Massey-Harroche, D. Polarity complex proteins. Biochimica et biophysica acta 1778, 614–630, https://doi.org/10.1016/j.bbamem.2007.08.029 (2008).
Pocha, S. M. & Knust, E. Complexities of Crumbs function and regulation in tissue morphogenesis. Current biology: CB 23, R289–293, https://doi.org/10.1016/j.cub.2013.03.001 (2013).
Campbell, K., Knust, E. & Skaer, H. Crumbs stabilises epithelial polarity during tissue remodelling. Journal of cell science 122, 2604–2612, https://doi.org/10.1242/jcs.047183 (2009).
Xu, N., Keung, B. & Myat, M. M. Rho GTPase controls invagination and cohesive migration of the Drosophila salivary gland through Crumbs and Rho-kinase. Developmental biology 321, 88–100, https://doi.org/10.1016/j.ydbio.2008.06.007 (2008).
Fan, S. S., Chen, M. S., Lin, J. F., Chao, W. T. & Yang, V. C. Use of gain-of-function study to delineate the roles of crumbs in Drosophila eye development. Journal of biomedical science 10, 766–773 (2003).
Medina, E. et al. Crumbs interacts with moesin and beta(Heavy)-spectrin in the apical membrane skeleton of Drosophila. The Journal of cell biology 158, 941–951, https://doi.org/10.1083/jcb.200203080 (2002).
Laprise, P. et al. The FERM protein Yurt is a negative regulatory component of the Crumbs complex that controls epithelial polarity and apical membrane size. Developmental cell 11, 363–374, https://doi.org/10.1016/j.devcel.2006.06.001 (2006).
Sherrard, K. M. & Fehon, R. G. The transmembrane protein Crumbs displays complex dynamics during follicular morphogenesis and is regulated competitively by Moesin and aPKC. Development 142, 1869–1878, https://doi.org/10.1242/dev.115329 (2015).
Flores-Benitez, D. & Knust, E. Crumbs is an essential regulator of cytoskeletal dynamics and cell-cell adhesion during dorsal closure in Drosophila. eLife 4, https://doi.org/10.7554/eLife.07398 (2015).
Pellikka, M. et al. Crumbs, the Drosophila homologue of human CRB1/RP12, is essential for photoreceptor morphogenesis. Nature 416, 143–149, https://doi.org/10.1038/nature721 (2002).
Izaddoost, S., Nam, S. C., Bhat, M. A., Bellen, H. J. & Choi, K. W. Drosophila Crumbs is a positional cue in photoreceptor adherens junctions and rhabdomeres. Nature 416, 178–183, https://doi.org/10.1038/nature720 (2002).
Lu, Q., Schafer, D. A. & Adler, P. N. The Drosophila planar polarity gene multiple wing hairs directly regulates the actin cytoskeleton. Development 142, 2478–2486, https://doi.org/10.1242/dev.122119 (2015).
Eaton, S. & Julicher, F. Cell flow and tissue polarity patterns. Current opinion in genetics & development 21, 747–752, https://doi.org/10.1016/j.gde.2011.08.010 (2011).
Adler, P. N. The frizzled/stan pathway and planar cell polarity in the Drosophila wing. Current topics in developmental biology 101, 1–31, https://doi.org/10.1016/B978-0-12-394592-1.00001-6 (2012).
Bryan, J., Edwards, R., Matsudaira, P., Otto, J. & Wulfkuhle, J. Fascin, an echinoid actin-bundling protein, is a homolog of the Drosophila singed gene product. Proceedings of the National Academy of Sciences of the United States of America 90, 9115–9119 (1993).
Cant, K., Knowles, B. A., Mooseker, M. S. & Cooley, L. Drosophila singed, a fascin homolog, is required for actin bundle formation during oogenesis and bristle extension. The Journal of cell biology 125, 369–380 (1994).
Mitchell, H. K., Roach, J. & Petersen, N. S. The morphogenesis of cell hairs on Drosophila wings. Developmental biology 95, 387–398 (1983).
Guild, G. M., Connelly, P. S., Ruggiero, L., Vranich, K. A. & Tilney, L. G. Actin filament bundles in Drosophila wing hairs: hairs and bristles use different strategies for assembly. Molecular biology of the cell 16, 3620–3631, https://doi.org/10.1091/mbc.E05-03-0185 (2005).
Turner, C. M. & Adler, P. N. Distinct roles for the actin and microtubule cytoskeletons in the morphogenesis of epidermal hairs during wing development in Drosophila. Mechanisms of development 70, 181–192 (1998).
Eaton, S., Wepf, R. & Simons, K. Roles for Rac1 and Cdc42 in planar polarization and hair outgrowth in the wing of Drosophila. The Journal of cell biology 135, 1277–1289 (1996).
Franke, J. D., Montague, R. A. & Kiehart, D. P. Nonmuscle myosin II is required for cell proliferation, cell sheet adhesion and wing hair morphology during wing morphogenesis. Developmental biology 345, 117–132, https://doi.org/10.1016/j.ydbio.2010.06.028 (2010).
Kiehart, D. P. et al. Drosophila crinkled, mutations of which disrupt morphogenesis and cause lethality, encodes fly myosin VIIA. Genetics 168, 1337–1352, https://doi.org/10.1534/genetics.104.026369 (2004).
Winter, C. G. et al. Drosophila Rho-associated kinase (Drok) links Frizzled-mediated planar cell polarity signaling to the actin cytoskeleton. Cell 105, 81–91 (2001).
Yan, J., Lu, Q., Fang, X. & Adler, P. N. Rho1 has multiple functions in Drosophila wing planar polarity. Developmental biology 333, 186–199, https://doi.org/10.1016/j.ydbio.2009.06.027 (2009).
Collinet, C. & Lecuit, T. Stability and dynamics of cell-cell junctions. Progress in molecular biology and translational science 116, 25–47, https://doi.org/10.1016/B978-0-12-394311-8.00002-9 (2013).
Tepass, U. The apical polarity protein network in Drosophila epithelial cells: regulation of polarity, junctions, morphogenesis, cell growth, and survival. Annual review of cell and developmental biology 28, 655–685, https://doi.org/10.1146/annurev-cellbio-092910-154033 (2012).
Hafezi, Y., Bosch, J. A. & Hariharan, I. K. Differences in levels of the transmembrane protein Crumbs can influence cell survival at clonal boundaries. Developmental biology 368, 358–369, https://doi.org/10.1016/j.ydbio.2012.06.001 (2012).
Hamaratoglu, F. et al. The Hippo tumor-suppressor pathway regulates apical-domain size in parallel to tissue growth. Journal of cell science 122, 2351–2359, https://doi.org/10.1242/jcs.046482 (2009).
Genevet, A. et al. The Hippo pathway regulates apical-domain size independently of its growth-control function. Journal of cell science 122, 2360–2370, https://doi.org/10.1242/jcs.041806 (2009).
Fristrom, D., Wilcox, M. & Fristrom, J. The distribution of PS integrins, laminin A and F-actin during key stages in Drosophila wing development. Development 117, 509–523 (1993).
Classen, A. K., Anderson, K. I., Marois, E. & Eaton, S. Hexagonal packing of Drosophila wing epithelial cells by the planar cell polarity pathway. Developmental cell 9, 805–817, https://doi.org/10.1016/j.devcel.2005.10.016 (2005).
Woods, D. F., Hough, C., Peel, D., Callaini, G. & Bryant, P. J. Dlg protein is required for junction structure, cell polarity, and proliferation control in Drosophila epithelia. The Journal of cell biology 134, 1469–1482 (1996).
Lin, Y. H. et al. AP-2-complex-mediated endocytosis of Drosophila Crumbs regulates polarity by antagonizing Stardust. Journal of cell science 128, 4538–4549, https://doi.org/10.1242/jcs.174573 (2015).
Nakajima, H. & Tanoue, T. Lulu2 regulates the circumferential actomyosin tensile system in epithelial cells through p114RhoGEF. The Journal of cell biology 195, 245–261, https://doi.org/10.1083/jcb.201104118 (2011).
Yan, J. et al. The multiple-wing-hairs gene encodes a novel GBD-FH3 domain-containing protein that functions both prior to and after wing hair initiation. Genetics 180, 219–228, https://doi.org/10.1534/genetics.108.091314 (2008).
Tepass, U. & Knust, E. Crumbs and stardust act in a genetic pathway that controls the organization of epithelia in Drosophila melanogaster. Developmental biology 159, 311–326, https://doi.org/10.1006/dbio.1993.1243 (1993).
Sugimura, K. & Ishihara, S. The mechanical anisotropy in a tissue promotes ordering in hexagonal cell packing. Development 140, 4091–4101, https://doi.org/10.1242/dev.094060 (2013).
Aigouy, B. et al. Cell flow reorients the axis of planar polarity in the wing epithelium of Drosophila. Cell 142, 773–786, https://doi.org/10.1016/j.cell.2010.07.042 (2010).
Heisenberg, C. P. & Bellaiche, Y. Forces in tissue morphogenesis and patterning. Cell 153, 948–962, https://doi.org/10.1016/j.cell.2013.05.008 (2013).
Wu, S. K. & Yap, A. S. Patterns in space: coordinating adhesion and actomyosin contractility at E-cadherin junctions. Cell communication & adhesion 20, 201–212, https://doi.org/10.3109/15419061.2013.856889 (2013).
Wu, S. K. et al. Cortical F-actin stabilization generates apical-lateral patterns of junctional contractility that integrate cells into epithelia. Nature cell biology 16, 167–178, https://doi.org/10.1038/ncb2900 (2014).
Polesello, C., Delon, I., Valenti, P., Ferrer, P. & Payre, F. Dmoesin controls actin-based cell shape and polarity during Drosophila melanogaster oogenesis. Nature cell biology 4, 782–789, https://doi.org/10.1038/ncb856 (2002).
Molnar, C. & de Celis, J. F. Independent roles of Drosophila Moesin in imaginal disc morphogenesis and hedgehog signalling. Mechanisms of development 123, 337–351, https://doi.org/10.1016/j.mod.2006.02.001 (2006).
Polesello, C. & Payre, F. Small is beautiful: what flies tell us about ERM protein function in development. Trends in cell biology 14, 294–302, https://doi.org/10.1016/j.tcb.2004.04.003 (2004).
Laprise, P. et al. Yurt, Coracle, Neurexin IV and the Na(+), K(+)-ATPase form a novel group of epithelial polarity proteins. Nature 459, 1141–1145, https://doi.org/10.1038/nature08067 (2009).
Sawyer, J. K. et al. A contractile actomyosin network linked to adherens junctions by Canoe/afadin helps drive convergent extension. Molecular biology of the cell 22, 2491–2508, https://doi.org/10.1091/mbc.E11-05-0411 (2011).
Royou, A., Field, C., Sisson, J. C., Sullivan, W. & Karess, R. Reassessing the role and dynamics of nonmuscle myosin II during furrow formation in early Drosophila embryos. Molecular biology of the cell 15, 838–850, https://doi.org/10.1091/mbc.E03-06-0440 (2004).
Bardet, P. L. et al. PTEN controls junction lengthening and stability during cell rearrangement in epithelial tissue. Developmental cell 25, 534–546, https://doi.org/10.1016/j.devcel.2013.04.020 (2013).
Gamblin, C. L., Hardy, E. J., Chartier, F. J., Bisson, N. & Laprise, P. A bidirectional antagonism between aPKC and Yurt regulates epithelial cell polarity. The Journal of cell biology 204, 487–495, https://doi.org/10.1083/jcb.201308032 (2014).
Cavey, M. & Lecuit, T. Molecular bases of cell-cell junctions stability and dynamics. Cold Spring Harbor perspectives in biology 1, a002998, https://doi.org/10.1101/cshperspect.a002998 (2009).
Rauzi, M., Lenne, P. F. & Lecuit, T. Planar polarized actomyosin contractile flows control epithelial junction remodelling. Nature 468, 1110–1114, https://doi.org/10.1038/nature09566 (2010).
Farhadifar, R., Roper, J. C., Aigouy, B., Eaton, S. & Julicher, F. The influence of cell mechanics, cell-cell interactions, and proliferation on epithelial packing. Current biology: CB 17, 2095–2104, https://doi.org/10.1016/j.cub.2007.11.049 (2007).
Ishihara, S. & Sugimura, K. Bayesian inference of force dynamics during morphogenesis. Journal of theoretical biology 313, 201–211, https://doi.org/10.1016/j.jtbi.2012.08.017 (2012).
Penalva, C. & Mirouse, V. Tissue-specific function of Patj in regulating the Crumbs complex and epithelial polarity. Development 139, 4549–4554, https://doi.org/10.1242/dev.085449 (2012).
Xu, T. & Rubin, G. M. Analysis of genetic mosaics in developing and adult Drosophila tissues. Development 117, 1223–1237 (1993).
Classen, A. K., Aigouy, B., Giangrande, A. & Eaton, S. Imaging Drosophila pupal wing morphogenesis. Methods in molecular biology 420, 265–275, https://doi.org/10.1007/978-1-59745-583-1_16 (2008).
Parks, A. L. et al. Systematic generation of high-resolution deletion coverage of the Drosophila melanogaster genome. Nature genetics 36, 288–292, https://doi.org/10.1038/ng1312 (2004).
Thibault, S. T. et al. A complementary transposon tool kit for Drosophila melanogaster using P and piggyBac. Nature genetics 36, 283–287, https://doi.org/10.1038/ng1314 (2004).
Mottola, G., Classen, A. K., Gonzalez-Gaitan, M., Eaton, S. & Zerial, M. A novel function for the Rab5 effector Rabenosyn-5 in planar cell polarity. Development 137, 2353–2364, https://doi.org/10.1242/dev.048413 (2010).
Bhat, M. A. et al. Discs Lost, a novel multi-PDZ domain protein, establishes and maintains epithelial polarity. Cell 96, 833–845 (1999).
Berger, S., Bulgakova, N. A., Grawe, F., Johnson, K. & Knust, E. Unraveling the genetic complexity of Drosophila stardust during photoreceptor morphogenesis and prevention of light-induced degeneration. Genetics 176, 2189–2200, https://doi.org/10.1534/genetics.107.071449 (2007).
Edwards, K. A., Demsky, M., Montague, R. A., Weymouth, N. & Kiehart, D. P. GFP-moesin illuminates actin cytoskeleton dynamics in living tissue and demonstrates cell shape changes during morphogenesis in Drosophila. Developmental biology 191, 103–117, https://doi.org/10.1006/dbio.1997.8707 (1997).
Levayer, R., Pelissier-Monier, A. & Lecuit, T. Spatial regulation of Dia and Myosin-II by RhoGEF2 controls initiation of E-cadherin endocytosis during epithelial morphogenesis. Nature cell biology 13, 529–540, https://doi.org/10.1038/ncb2224 (2011).
Acknowledgements
We thank B. Aigouy, M. Mavrakis and C. Toret for critical reading of this manuscript. This project was supported by CNRS and Aix-Marseille University, the labex INFORM (grant ANR-11-LABX-0054), ANR Ghearact (14-CE13-0013) and ANR Chrononet (14-CE10-0010). We thank the IBDM imaging facility for imaging support and acknowledge France-BioImaging infrastructure supported by the Agence Nationale de la Recherche (10-INSB-04-01, call “Grand Emprunt”). The Le Bivic group is an “Equipe labellisée 2008 de La Ligue Nationale contre le Cancer”.
Author information
Authors and Affiliations
Contributions
P.S. and G.M. conceived and conducted all the experiments; F.P. and P.V. generated moesin mutant; E.B. performed cell orientation quantifications. P.S., A.L.B. and G.M. analyzed the results and wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Salis, P., Payre, F., Valenti, P. et al. Crumbs, Moesin and Yurt regulate junctional stability and dynamics for a proper morphogenesis of the Drosophila pupal wing epithelium. Sci Rep 7, 16778 (2017). https://doi.org/10.1038/s41598-017-15272-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-15272-1
This article is cited by
-
Bi-allelic variations in CRB2, encoding the crumbs cell polarity complex component 2, lead to non-communicating hydrocephalus due to atresia of the aqueduct of sylvius and central canal of the medulla
Acta Neuropathologica Communications (2023)
-
Apical–basal polarity and the control of epithelial form and function
Nature Reviews Molecular Cell Biology (2022)
-
Multiple functions of the scaffold protein Discs large 5 in the control of growth, cell polarity and cell adhesion in Drosophila melanogaster
BMC Developmental Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.