Abstract
Chagas disease is caused by the protozoan parasite Trypanosoma cruzi and affects 5–8 million people in Latin America. Although the nitroheterocyclic compound benznidazole has been the front-line drug for several decades, treatment failures are common. Benznidazole is a pro-drug and is bio-activated within the parasite by the mitochondrial nitroreductase TcNTR-1, leading to the generation of reactive metabolites that have trypanocidal activity. To better assess drug action and resistance, we sequenced the genomes of T. cruzi Y strain (35.5 Mb) and three benznidazole-resistant clones derived from a single drug-selected population. This revealed the genome-wide accumulation of mutations in the resistant parasites, in addition to variations in DNA copy-number. We observed mutations in DNA repair genes, linked with increased susceptibility to DNA alkylating and inter-strand cross-linking agents. Stop-codon-generating mutations in TcNTR-1 were associated with cross-resistance to other nitroheterocyclic drugs. Unexpectedly, the clones were also highly resistant to the ergosterol biosynthesis inhibitor posaconazole, a drug proposed for use against T. cruzi infections, in combination with benznidazole. Our findings therefore identify the highly mutagenic activity of benznidazole metabolites in T. cruzi, demonstrate that this can result in multi-drug resistance, and indicate that vigilance will be required if benznidazole is used in combination therapy.
Similar content being viewed by others
Introduction
Chagas disease results from infection with the insect-transmitted protozoan parasite Trypanosoma cruzi. It is a life-long infection and a major cause of morbidity and mortality in many areas of Latin America1. Recently, the disease has also become more widespread, with large numbers of infected individuals being diagnosed within migrant populations, particularly in Europe and the USA2. In the initial ‘acute’ stage, which occurs 4–6 weeks post-infection, Chagas disease generally manifests as a transient low-level febrile condition. In children however, acute stage infections can lead to myocarditis or meningoencephalitis, and are sometimes fatal. Following suppression of the acute infection by a robust adaptive immune response3, the disease proceeds to a long-lasting asymptomatic chronic stage, which is characterised by an extremely low parasitemia. About one-third of those infected will eventually advance to a symptomatic stage, although this can take decades. Most of these individuals will develop cardiomyopathy4, or less commonly, digestive tract megasyndromes, outcomes for which there are few therapeutic options.
Although benznidazole has been used against chronic T. cruzi infections for almost 50 years, treatment failures are frequently reported, with non-curative outcomes varying between 6% and 50% in recent clinical trials5,6. Other problematic factors include the extended treatment length (often 60–90 days) and frequent toxic side-effects. Furthermore, there is a demonstrated risk for cross-resistance with the other front-line drug, nifurtimox. This arises because both of these nitroheterocyclic agents require to be activated by the T. cruzi flavin-dependent mitochondrial type 1 nitroreductase, TcNTR-17,8. Our understanding of how benznidazole kills the parasite and why treatment is often non-curative is limited. The extent to which this is due to acquired resistance or patient non-compliance remains to be determined. In addition, T. cruzi is an extremely diverse species and there is extensive natural variation in susceptibility to benznidazole9,10, both within and between lineages. Despite this, there is as yet no unequivocal evidence for a link between parasite taxonomic designation and treatment outcome in humans11,12, and no definitive mechanism demonstrated that accounts for natural variations in susceptibility. This is an area that requires further study.
Evidence suggests that trypanocidal activity could be mediated, at least in part, by DNA damage13,14 caused by reactive metabolites that are generated following reductive metabolism of the drug by TcNTR-115,16. Attempts to dissect the mechanisms underlying treatment failure have been compounded by the extremely low and focal nature of the parasite burden during chronic stage disease17,18 and technical difficulties in establishing parasitological cure/non-cure19. To gain insight into the mechanisms of resistance, we sequenced the genomes of three drug-resistant clones from a single parasite population that had undergone benznidazole-selection in vitro. Here, we show that long-term treatment with benznidazole can give rise to genome-wide mutagenesis in T. cruzi, and result in phenotypic changes that include the acquisition of multi-drug resistance.
Results
Genome-wide mutagenesis in benznidazole-resistant clones
Following the isolation of three drug-resistant clones from a single T. cruzi population exposed to benznidazole selection, we identified a stop-codon-generating mutation in the TcNTR-1 gene20. However, this nonsense mutation by itself was insufficient to account for the high level and inter-clonal diversity of benznidazole-resistance (9–26 fold), indicating that additional mechanisms must operate. To investigate this, we sequenced the genomes of the parental T. cruzi Y strain and the three benznidazole-resistant clones using an Illumina HiSeq. 2500 platform (x377 coverage). Genome size was 35.54 Mb (48.3% GC content), distributed over 150,022 contigs. Assembly, using the SPAdes program, and annotation performed with the Companion Sanger Pipeline (https://companion.sanger.ac.uk) on the 3,000 largest contigs, identified 8,289 putative protein coding genes, 5119 with predictable function. The T. cruzi CL Brener genome strain21 was used as reference. The parental Y strain genome sequence and those of the three drug-resistant clones were mapped using the BWA-mem alignment software, and single nucleotide polymorphisms (SNPs) were called using the SAMtools package.
We found evidence of the genome-wide accumulation of mutations in the drug-resistant clones. There was a total of 26,495 point mutations (Fig. 1a), with 0.42 SNPs/kb in clone 1, 0.49/kb in clone 2, and 0.45/kb in clone 3. 32% of SNPs were in coding regions, with an equivalent number of synonymous and non-synonymous changes (Fig. 1b). A total of 25 nonsense/nonstop mutations were identified (Supplementary Table S1), including the internal stop codon within TcNTR-1 20. Transitions (A ↔ G or C ↔ T) were significantly more frequent than transversions in clone 2, although in clones 1 and 3 the numbers were similar (Fig. 1c). 7,731 of the SNPs were present in all three clones and phylogenetic analysis suggested that the lineage giving rise to clones 2 and 3 diverged from a common ancestor of clone 1 during the drug-selection process (Fig. 1d). It can be inferred therefore that SNPs common to clones 1 and 2, and to clones 1 and 3 (Fig. 1a), arose from point mutations acquired by the ancestral drug-resistant line and that subsequent differential gene conversion led to their reversion in clones 3 and 2, respectively.
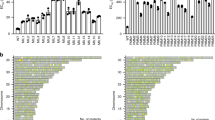
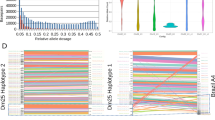
Analysis of SNPs identified in T. cruzi clones selected for benznidazole-resistance. (a) Venn diagram illustrating the distribution of 26,495 distinct SNPs identified by genome sequencing of three drug-resistant clones. (b) The total number of SNPs in coding and non-coding genomic regions are shown in purple and yellow. The smaller circle (right) illustrates SNP distribution within coding regions: missense (green), silent (grey) and nonsense/nonstop (orange). (c) Classification of missense SNPs as transitions (purine ↔ purine; blue) or transversions (purine ↔ pyrimidine; pink). (d) Maximum-likelihood phylogenetic tree based on nucleotide alignment (FastTree program), showing relationship between parental and drug-resistant T. cruzi clones. Scale bar represents the number of substitutions per site.
Copy number variation (CNV) and read coverage across the data set were also assessed. Trisomy was predicted for a large region of chromosome 13 in all three drug-resistant clones, on the basis of a 1.5-fold higher read-coverage compared to the genome-wide average (Fig. 2b), although the fragmented nature of the reference genome21 prevents us from making unequivocal inferences about the entire chromosome. We also found a similarly increased sequence coverage associated with regions (hundreds of kb) in chromosome 11 (clones 1 and 3), a finding supported by a control-freeC analysis. Genome plasticity has been widely observed in T. cruzi 22. With benznidazole, where resistance can arise by a “loss-of-function” mechanism, parasites missing an entire copy of the chromosome containing the TcNTR-1 gene have previously been isolated7. In this present study, where we observed that resistance was associated with copy number increases in chromosomes 11 and 13, the most parsimonious explanation is that enhanced expression of one or more of the linked genes contributes to the selected phenotype. In clone 2, the data suggest a reversion to disomy in chromosome 11 (Fig. 2b), following divergence from clone 3 (Fig. 1d), although the possibility that the increased copy number was acquired independently by clones 1 and 3 cannot be excluded.
Large-scale DNA copy-number variation in benznidazole-resistant clones. Histograms showing read-coverage for specific regions of chromosome 11 (a) and 13 (b) in parental and drug-resistant clones. Chromosome annotation was performed by ordering the contigs against the T. cruzi CL Brener reference genome21. High read peaks correspond to repetitive hypothetical protein genes, tandem repeats within kinesin-like genes (chromosome 11), linked histone H2A genes and short interspersed DNA sequences (chromosome 13). The red dotted line represents the average read-coverage across all chromosomes (single-copy genes). Assessment by the program control-freeC is consistent with an additional copy (trisomy) of the regions shown in chromosome 11 [from TCRUZ.110005800 (TcCLB.509349.30) to TCRUZ.110020000 (TcCLB.506443.80)] and chromosome 13 [from TCRUZ_130005000 (TcCLB.510165.60) to TCRUZ_130019000 (TcCLB.508325.90)]. Chromosome 11 in clone 2 did not display an increased read-coverage compared to the genome-wide average.
Deficiencies in DNA repair in drug-resistant parasites
Using gene ontology annotations, it was possible to link mutations to functional categories. The majority occurred in hypervariable regions, including the large and highly diverse surface antigen gene families, such as the mucins and trans-sialidases, which constitute much of the parasite genome23. In functional categories or metabolic pathways that have been tentatively associated with drug-resistance8,14,16,24,25,26,27, there were a range of SNP frequencies ranging from 0.47/kb in genes linked to electron transport to 0.11/kb in ABC transporter genes (Fig. 3a and b). Individually or collectively, it is implicit that changes to the sequence or expression of these, or other genes, have contributed to the enhanced levels of benznidazole-resistance. Given the extensive number of mutations in DNA repair enzymes, we investigated the susceptibility of the drug-resistant clones to DNA damaging agents. This revealed enhanced sensitivity to compounds that promote DNA alkylation (1-methyl-3-nitro-1-nitrosoguanidine) or inter-strand cross-linking (mechlorethamine), but not to methyl methanesulfonate, which causes double-strand breaks (Fig. 3c).
Genetic and phenotypic changes in benznidazole-resistant clones. (a) Total point mutations in genes linked to DNA repair, redox metabolism, electron transport, cell growth and proliferation, and ABC transporter activity, in each of the three clones. The colour scale indicates the number of mutations per gene: 1 (+), 2 (++), or 3 or more (+++). (b) Average SNPs/kb in genes from different functional categories in the 3 clones. Synonymous and non-synonymous SNPs are shown. (c) Sensitivity of parental and drug-resistant parasites to the DNA damaging agents methyl-3-nitro-1-nitrosoguanidine (MNNG), mechlorethamine (ME) and methyl methanesulfonate (MMS) (Methods). Statistical analysis was performed using one-way analysis of variance; p values, respectively: 0.02, 0.01, 0.02 (MNNG); 0.002, 0.003, 0.02 (ME). (d) Bioluminescence imaging of SCID mice infected i.p. with parental and drug-resistant clones (104 tissue culture trypomastigotes). Parasites were genetically transformed to express the luciferase reporter gene PpyRe9h 17,28 (Methods). Ventral images of representative mice (n = 3) are shown, with the number of days post-infection given above. Heat maps indicate the intensity of bioluminescence from low (blue) to high (red) (log10 scale); the minimum and maximum radiances for the pseudocolour scale are shown. The mice infected with the parental Y strain required euthanization on day 28, in accordance with animal welfare requirements (Methods).
To determine if mutagenesis had compromised the infectivity of the drug-resistant clones, we inoculated immunocompromised SCID mice with parasites transformed to express a red-shifted bioluminescence reporter17,28 (Methods). Whereas the non-resistant parental line resulted in a fulminant infection, which required the mice to be euthanised after 28 days, infections with drug-resistant clones were asymptomatic, and there was an extremely low parasite burden (Fig. 3d). This loss of fitness may reflect that infectivity was not a trait under selection during the continuous in vitro culturing used to generate benznidazole-resistant parasites (Methods). Studies had previously shown that there were no significant differences in the rate of intracellular amastigote replication or infectivity of cultured mammalian cells20.
Multi-drug resistance following benznidazole selection
As predicted for parasites with a disrupted TcNTR-1 gene, the three clones displayed cross-resistance to other nitroheterocyclic agents (Fig. 4a–d; Supplementary Table S2), including nifurtimox, a drug which is also used to treat T. cruzi infections, and fexinidazole, a drug which has undergone clinical trials against Chagas disease. The level of resistance (2–5 fold) was consistent with that previously observed for nitroheterocyclic compounds following disruption of one copy of TcNTR-1 7,29. It can therefore be inferred that the other, as yet unknown mechanisms that lead to the enhanced benznidazole-resistance, do not contribute significantly to additional levels of cross-resistance against other nitroheterocyclic drugs. Unexpectedly, we found that each of the clones were also highly resistant to the anti-fungal agent posaconazole (6–22 fold, Fig. 4e). Although posaconazole monotherapy recently failed in clinical trial against Chagas disease5,30, its use in combination with benznidazole has been discussed31,32,33 and subjected to preliminary trial in humans30. Posaconazole is an ergosterol biosynthesis inhibitor that targets lanosterol 14α-demethylase (CYP51)34,35, a mechanism of action that shares no known biochemical overlap with that of benznidazole. In the drug-resistant clones, there were no mutations detected in CYP51, and the gene is not located on either of the chromosomes (11 and 13) which displayed increased copy number. Mutations were noted in the 5’-upstream region of CYP51, and in the coding regions of other genes linked with ergosterol biosynthesis (Supplementary Figure S1). However, these did not affect the intracellular level of CYP51, as judged by binding between the enzyme and posaconazole conjugated with the fluorophore boron-dipyrromethene36,37 (Supplementary Figure 2). To exclude the possibility that TcNTR-1 depletion might perturb susceptibility to posaconazole, we tested its activity against two cloned T. cruzi cell lines (Methods) in which we had deleted one copy of the gene (Fig. 4f). There was no significant difference compared to the parental parasites.
Multi-drug resistance following benznidazole-selection. (a–e) EC50 values were obtained (Methods) for the parental Y strain (P) and drug-resistant clones 1–3 (epimastigotes) against benznidazole, nifurtimox, fexinidazole, fexinidazole sulfone and posaconazole. The fold-resistance is indicated above each bar. (f) Benznidazole and posaconazole susceptibility of parental parasites and two genetically modified clones in which one copy of the TcNTR-1 gene had been deleted. Each experiment was performed 3 times in triplicate.
Discussion
Investigating the underlying causes of treatment failure in humans has been challenging in the case of T. cruzi infections5,6. Reasons include the possible influences of host and parasite genetics, the effects of toxicity on compliance, difficulties in demonstrating parasitological cure, the unresolved contribution of drug-resistance and the possibility of re-infection/co-infection. With benznidazole, there are no data available from clinical trials that provide insight into the mechanisms of resistance or the precise mode of drug action in infected individuals. Evidence from in vitro studies suggest that the pivotal step in both drug activity and resistance involves bio-activation of the drug by the nitroreductase TcNTR-17,8,9,15,16. This gives rise to toxic metabolites that react with a range of biological molecules (for review, see ref.8,29). However, given the high level of resistance that is achievable in vitro (Fig. 4, as example), and the wide range of natural susceptibility to benznidazole9,10, it is clear that other mechanisms can also contribute. For example, several other parasite genes have been identified that could act through alternative bioactivation38, alleviation of oxidative stress39, or increased efflux26. In the context of DNA, electrophilic drug metabolites, increased oxidative stress resulting from drug adduct-thiol conjugation, and the generation of oxidised nucleotides such as 8-oxo-guanine, have been implicated as potential mediators of mutagenesis in benznidazole-treated T. cruzi 14,15,16,40,41.
Here, we show that benznidazole-induced mutagenic effects can be extensive and genome-wide in T. cruzi, with a range of phenotypic consequences, including reduced DNA repair capacity and multi-drug resistance. These types of outcome have many parallels with the situation in drug-resistant cancer, where selective pressure, coupled with the early acquisition of defects in the DNA damage repair machinery, can drive the development of multi-drug resistance through mechanisms linked with chromosome instability and an increased propensity to acquire mutations42,43. In a similar way, our results suggest that in T. cruzi under benznidazole-selection, the combined effects of electrophilic drug metabolites and deficiencies in DNA repair, which could arise early in the process, might also act in concert to create a hypermutagenic environment that can drive the development of enhanced resistance and other phenotypic changes (Fig. 3d). Reduced infectivity was one was one of the acquired traits common to the drug-resistant clones. This phenotype could have arisen as a result of sequence changes at a single locus, or could represent the collective effect of multiple mutations. It will be important to determine whether this loss of fitness is a frequent or inevitable consequence of benznidazole-resistance, or whether selective pressures encountered in an in vivo context would be sufficient to exclude this particular outcome.
The current benznidazole dosing regimen involves daily administration for periods of 60 days or more, with many patients failing to complete treatment because of toxic side-effects5,6,30. Partly to avoid this, combination therapy with posaconazole has been suggested as a treatment option that might allow a reduction in patient exposure to benznidazole30. On the basis that benznidazole and posaconazole activities are not thought to overlap, it has also been assumed that the generation of cross-resistance using this regimen would be unlikely. However, our results show that this need not be the case. Parasites selected on benznidazole alone were found to have developed high-level resistance to posaconazole (Fig. 4e). This outcome could not be attributed to changes in the CYP51 gene, and presumably results from the effects of other mutation(s), either individually or collectively. Whatever the precise mechanism, these findings have implications for future Chagas disease strategies, suggesting that combination therapy involving benznidazole will require careful monitoring, irrespective of the secondary drug used. Although caution must be displayed in extrapolating findings made in vitro, our data further highlight an urgent need for studies aimed at isolating parasites from patients following non-curative benznidazole therapy, to determine if induced mutagenesis and acquired drug-resistance have contributed to treatment failure. However, there will be significant technical and analytical issues associated with this. As mentioned previously44, these include complexity resulting from multi-clonal infections, possible stage-specific differences in parasite susceptibility, the tissue origin of recrudescence, interplay with the immune system, and finally, difficulties in isolating parasites from chronic chagasic patients, both before and after treatment.
Methods
Isolation of benznidazole-resistant clones
T. cruzi (Y strain) was cultured as epimastigotes in supplemented RPMI-1640 medium at 28 °C45. Parasites were selected for benznidazole-resistance by culturing in the continuous presence of 182 μM benznidazole for 10 passages. From the 11th until the 15th passage, the drug concentration was increased in 10 μM steps46. The total selection period was 4 months. Parasites were then cloned by limiting dilution20 and the extent of benznidazole-resistance assessed. The resistance phenotype was found to be unchanged after 6 months continuous culturing in the absence of benznidazole46.
Genome sequencing
Genomic DNA was prepared from three drug-resistant clones and the parental strain, and then submitted to paired-end sequencing (100 bp) using an Illumina HiSeq. 2500 platform. Assembly of high-quality reads was conducted using SPAdes assembly program, with genome annotation on the 3,000 largest contigs performed by the Companion Sanger Pipeline (https://companion.sanger.ac.uk), using the T. cruzi CL Brener strain21 as reference. Mapping of the parental population to the three drug-resistant clones and SNP analysis were performed using the BWA-mem alignment software and SAMtools package, respectively. Gene functional categories were classified based on GO annotations. Copy number variation was assessed using the program control-freeC (http://bioinfo-out.curie.fr/projects/freec/tutorial.html). Sequence data from this paper are available on the website PathogenSeq LSHTM (http://pathogenseq.lshtm.ac.uk/t_cruzi/) and at the European Nucleotide Archive (http://www.ebi.ac.uk/ena/) under the accession no. PRJEB14637; clone 1 (ERS1913021), clone 2 (ERS1913022), clone 3 (ERS1913023), parental Y strain (ERS1913024).
Genetic modification of T. cruzi and assessment of infectivity by in vivo imaging
The red-shifted luciferase gene PpyRE9h was integrated into rRNA loci in three drug-resistant clones and the parental Y strain using the construct pTRIX2-RE9h17 (Fig. 3d). Clones expressing comparable levels of bioluminescence were cultured under conditions that promote differentiation into metacyclic trypomastigotes17. The resulting parasites were used to infect rat myoblast L6 monolayers at a ratio of 10 metacyclics per host cell. Following incubation at 37 °C under 5% CO2 for 7 days, tissue culture trypomastigotes were collected from the supernatant and female CB17 SCID mice (aged 8–12 weeks and bred in-house) were infected i.p. with 104 trypomastigotes. Animals were maintained under specific pathogen-free conditions in individually ventilated cages, and experienced a 12 hour light/dark cycle, with access to food and water ad libitum. To monitor infections, mice were injected with 150 mg kg−1 d-luciferin every 7 days and imaged using an IVIS Lumina II system (Caliper Life Science), as described previously17. Mice injected with the parental line developed fulminant infections and were euthanized at or before humane end-points by exsanguination under terminal anaesthesia. All animal work was performed under UK Home Office licence 70/8207 and approved by the London School of Hygiene and Tropical Medicine Animal Welfare and Ethical Review Board.
To generate parasites in which one copy of the Tc-NTR-1 gene had been disrupted, bioluminescent T. cruzi CL Brener epimastigotes17,47 were transfected as described previously7. Briefly, parasites were electroporated with plasmid pKO-NTR-BLA which had been linearised by Kpn I/Sac II digestion, and blasticidin-resistant parasites were isolated, cloned and checked by PCR to confirm integration and disruption.
Parasite susceptibility to drugs or DNA damaging agents
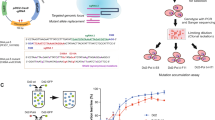
Epimastigotes were seeded at 2.5 × 105 ml−1 in 96-well plates in the presence of a range of concentrations of nitroheterocyclic compounds (benznidazole, nifurtimox, fexinidazole, or fexinidazole sulfone), posaconazole, or DNA damaging agents (MNNG, ME and MMS). Plates were incubated for 4 days at 28 °C, with resazurin added for the final 48 hours. The plates were then read in a Spectramax plate reader and the EC50 values determined using GraphPad Prism. Statistical analysis was performed using one-way analysis of variance, with significance for p values < 0.05 (*).
References
Bern, C. Chagas’ Disease. N. Engl. J. Med. 373, 456–466 (2015).
Pérez-Molina, J. A., Norman, F. & López-Vélez, R. Chagas disease in non-endemic countries: epidemiology, clinical presentation and treatment. Curr. Infect. Dis. Rep. 14, 263–274 (2012).
Tarleton, R. L. CD8 + T cells in Trypanosoma cruzi infection. Semin. Immunpath. 37, 233–238 (2015).
Cunha-Neto, E. & Chevillard, C. Chagas disease cardiomyopathy: immunopathology and genetics. Mediators of Inflammation 2014, 683230 (2014).
Molina, I. et al. Randomized trial of posaconazole and benznidazole for chronic Chagas disease. N. Engl. J. Med. 370, 1899–1908 (2014).
Morillo, C. A. et al. Randomized trial of benznidazole for chronic Chagas’ cardiomyopathy. N. Engl. J. Med. 373, 1295–1306 (2015).
Wilkinson, S. R., Taylor, M. C., Horn, D., Kelly, J. M. & Cheeseman, I. A mechanism for cross-resistance to nifurtimox and benznidazole in trypanosomes. Proc. Natl. Acad. Sci. USA 105, 5022–5027 (2008).
Wilkinson, S. R. & Kelly, J. M. Trypanocidal drugs: mechanisms, resistance and new targets. Expert Rev. Molec. Med . 11, e31 pp1-24 (2009).
Mejia, A. M. et al. Benznidazole-resistance in Trypanosoma cruzi is a readily acquired trait that can arise independently in a single population. J. Infect. Dis. 206, 220–228 (2012).
Villarreal, D., Barnabé, C., Sereno, D. & Tibayrenc, M. Lack of correlation between in vitro susceptibility to benznidazole and phylogenetic diversity of Trypanosoma cruzi, the agent of Chagas disease. Exp. Parasitol. 108, 24–31 (2004).
Teston, A. P. et al. In vivo susceptibility to benznidazole of Trypanosoma cruzi strains from the western Brazilian Amazon. Trop. Med. Internat. Health 18, 85–95 (2013).
Oliveira, M. T. et al. TcI, TcII and TcVI Trypanosoma cruzi samples from Chagas disease patients with distinct clinical forms and critical analysis of in vitro and in vivo behavior, response to treatment and infection evolution in murine model. Acta Tropica 167, 108–120 (2017).
La-Scalea, M. A., Serrano, S. H., Ferreira, E. I. & Brett, A. M. Voltammetric behavior of benznidazole at a DNA-electrochemical biosensor. J. Pharm. Biomed. Anal. 29, 561–568 (2002).
Rajão, M. A. et al. Unveiling benznidazole’s mechanism of action through overexpression of DNA repair proteins in Trypanosoma cruzi. Environ. Mol. Mutagen. 55, 309–321 (2014).
Hall, B. S. & Wilkinson, S. R. Activation of benznidazole by trypanosomal type I nitroreductases results in glyoxal formation. Antimicrob. Agents Chemother. 56, 115–123 (2012).
Trochine, A., Creek, D. J., Faral-Tello, P., Barrett, M. P. & Robello, C. Benznidazole biotransformation and multiple targets in Trypanosoma cruzi revealed by metabolomics. PLoS Negl. Trop. Dis. 22, e2844 (2014).
Lewis, M. D. et al. Bioluminescence imaging of chronic Trypanosoma cruzi infections reveals tissue-specific parasite dynamics and heart disease in the absence of locally persistent infection. Cell. Microbiol. 16, 1285–1300 (2014).
Lewis, M. D., Fortes Francisco, A., Taylor, M. C., Jayawardhana, S. & Kelly, J. M. Host and parasite genetics shape a link between Trypanosoma cruzi infection dynamics and chronic cardiomyopathy. Cell. Microbiol. 18, 1429–1443 (2016).
Fortes Francisco, A. et al. The limited ability of posaconazole to cure both acute and chronic Trypanosoma cruzi infections revealed by highly sensitive in vivo imaging. Antimicrob. Agents Chemother. 59, 4653–4661 (2015).
Campos, M. C., Leon, L. L., Taylor, M. C. & Kelly, J. M. Benznidazole-resistance in Trypanosoma cruzi: evidence that distinct mechanisms can act in concert. Mol. Biochem. Parasitol. 193, 17–19 (2014).
El-Sayed, N. M. et al. The genome sequence of Trypanosoma cruzi, etiologic agent of Chagas disease. Science 309, 409–415 (2005).
Obado, S. O., Taylor, M. C., Wilkinson, S. R., Bromley, E. V. & Kelly, J. M. Functional mapping of a trypanosome centromere by chromosome fragmentation identifies a 16 kb GC-rich transcriptional “strand-switch” domain as a major feature. Genome Res. 15, 36–43 (2005).
Weatherly, D. B., Peng, D. & Tarleton, R. L. Recombination-driven generation of the largest pathogen repository of antigen variants in the protozoan Trypanosoma cruzi. BMC Genomics 17, 729 (2016).
Portal, P. et al. Multiple NADPH-cytochrome P450 reductases from Trypanosoma cruzi: Suggested role in drug resistance. Mol. Biochem. Parasitol. 160, 42–51 (2008).
Alsford, S., Kelly, J. M., Baker, N. & Horn, D. Genetic dissection of drug resistance in trypanosomes. Parasitology 140, 1478–1491 (2013).
Franco, J., Ferreira, R. C., Ienne, S. & Zingales, B. ABCG-like transporter of Trypanosoma cruzi involved in benznidazole resistance: gene polymorphisms disclose inter-strain intragenic recombination in hybrid isolates. Infect. Genet. Evol. 31, 198–208 (2015).
Reigada, C. et al. Trypanosoma cruzi polyamine transporter: Its role on parasite growth and survival under stress conditions. J. Membr. Biol. 249, 475–481 (2016).
Branchini, B. R. et al. Red-emitting luciferases for bioluminescence reporter and imaging applications. Anal. Biochem. 396, 290–297 (2010).
Wyllie, S. et al. Nitroheterocyclic drug resistance mechanisms in Trypanosoma brucei. J. Antimicrob. Chemother. 71, 625–634 (2016).
Morillo, C. A. et al. Benznidazole and posaconazole in eliminating parasites in asymptomatic T. cruzi carriers: The STOP-CHAGAS trial. J. Am. Coll. Cardiol. 69, 939–947 (2017).
Cencig, S., Coltel, N., Truyens, C. & Carlier, Y. Evaluation of benznidazole treatment combined with nifurtimox, posaconazole or AmBisome® in mice infected with Trypanosoma cruzi strains. Int. J. Antimicrob. Agents 40, 527–532 (2012).
Diniz, L. & de, F. et al. Benznidazole and posaconazole in experimental Chagas disease: positive interaction in concomitant and sequential treatments. PLoS Negl. Trop. Dis. 7, e2367 (2013).
Bern, C. A new epoch in antitrypanosomal treatment for Chagas disease. J. Am. Coll. Cardiol. 69, 948–950 (2017).
Lepesheva, G. I., Villalta, F. & Waterman, M. R. Targeting Trypanosoma cruzi sterol 14α-demethylase (CYP51). Adv. Parasitol. 75, 65–87 (2012).
Buckner, F. S. & Urbina, J. A. Recent developments in sterol 14α-demethylase inhibitors for Chagas disease. Int. J. Parasitol. Drugs Drug Resist. 2, 236–242 (2012).
Pratt, A. et al. Evaluation of fungal-specific fluorescent labeled echinocandin probes as diagnostic adjuncts. Med. Mycol. 51, 103–107 (2013).
Campoli, P. et al. Pharmacokinetics of posaconazole within epithelial cells and fungi: Insights into potential mechanisms of action during treatment and prophylaxis. J. Infect. Dis. 208, 1717–1728 (2013).
Murta, S. M. et al. Deletion of copies of the gene encoding old yellow enzyme (TcOYE), a NAD(P)H flavin oxidoreductase, associates with in vitro-induced benznidazole resistance in Trypanosoma cruzi. Mol. Biochem. Parasitol. 146, 151–162 (2006).
Nogueira, F. B. et al. Increased expression of iron-containing superoxide dismutase-A (TcFeSOD-A) enzyme in Trypanosoma cruzi population with in vitro-induced resistance to benznidazole. Acta Trop. 100, 119–132 (2006).
Zahoor, A., Lafleur, M. V., Knight, R. C., Loman, H. & Edwards, D. I. DNA damage induced by reduced nitroimidazole drugs. Biochem. Pharmacol. 36, 3299–3304 (1987).
Diaz de Toranzo, E. G., Castro, J. A., Franke de Cazzulo, B. M. & Cazzulo, J. J. Interaction of benznidazole reactive metabolites with nuclear and kinetoplastic DNA, proteins and lipids from Trypanosoma cruzi. Experientia 44, 880–881 (1998).
Bouwman, P. & Jonkers, J. The effects of deregulated DNA damage signalling on cancer chemotherapy response and resistance. Nat. Rev. Cancer 12, 587–598 (2012).
Sakai, W. et al. Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers. Nature 451, 1116–1120 (2008).
Moreno, M. et al. Trypanosoma cruzi benznidazole susceptibility in vitro does not predict the therapeutic outcome of human Chagas disease. Mem. Inst. Oswaldo Cruz 105, 918–924 (2010).
Kendall, G., Wilderspin, A. F., Ashall, F., Miles, M. A. & Kelly, J. M. Trypanosoma cruzi glycosomal glyceraldehyde-3-phosphate dehydrogenase does not conform to the “hotspot” topogenic signal model. EMBO J. 9, 2751–2758 (1990).
Campos, M. C. et al. P-glycoprotein efflux pump plays an important role in Trypanosoma cruzi drug resistance. Parasitol. Res. 112, 2341–2351 (2013).
Lewis, M. D., Fortes Francisco, A., Taylor, M. C. & Kelly, J. M. A new experimental model for assessing drug efficacy against Trypanosoma cruzi infection based on highly sensitive in vivo imaging. J. Biomolecular Screening 20, 36–43 (2015).
Acknowledgements
M.C.C. was funded by the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Brazil. J.M.K. acknowledges financial support from the British Heart Foundation (Grant PG/13/88/30556) and the Drugs for Neglected Diseases initiative (DNDi). M.D.L. is supported by an EU Marie Curie Fellowship. J.P. is funded by a UK BBSRC LiDO PhD studentship. T.G.C. is funded by the UK MRC (Grant no. MR/K000551/1, MR/M01360X/1, MR/N010469/1). The authors would like to thank members of KAUST Bioscience Core Sequencing facility for sequencing the parasites. This part of the work was funded by Faculty baseline funding (BAS/1/1020–01–01) to A.P. We thank Arkady Mustaev (New Jersey Medical School –Rutgers) for kindly providing BODIPY conjugated posaconazole,
Author information
Authors and Affiliations
Contributions
M.C.C., J.M.K., J.P., A.P., and T.G.G. designed the experiments and analysed the data. M.C.C., A.F.F., M.C.T., M.D.L. and A.P. performed the experiments. J.M.K. and M.C.C. wrote the paper, with input from all co-authors.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Campos, M.C., Phelan, J., Francisco, A.F. et al. Genome-wide mutagenesis and multi-drug resistance in American trypanosomes induced by the front-line drug benznidazole. Sci Rep 7, 14407 (2017). https://doi.org/10.1038/s41598-017-14986-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-14986-6
This article is cited by
-
Nature-derived Peptides as Promising Antiparasitic Agents against Neglected Tropical Diseases
International Journal of Peptide Research and Therapeutics (2024)
-
Molecular Characterization of Four Mexican Isolates of Trypanosoma cruzi and Their Profile Susceptibility to Nifurtimox
Acta Parasitologica (2022)
-
Major changes in chromosomal somy, gene expression and gene dosage driven by SbIII in Leishmania braziliensis and Leishmania panamensis
Scientific Reports (2019)
-
Metabolomics, lipidomics and proteomics profiling of myoblasts infected with Trypanosoma cruzi after treatment with different drugs against Chagas disease
Metabolomics (2019)
-
Identification of di-substituted ureas that prevent growth of trypanosomes through inhibition of translation initiation
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.