Abstract
The main roles of equilibrative nucleoside transporters (ENTs) and concentrative nucleoside transporters (CNTs) are to transfer single nucleosides and analogues for the nucleic acid salvage pathway. Oligodeoxyribonucleotides (ODNs) can be transported into the cytoplasm or nucleus of cells under certain conditions. Among ODNs composed of a single type of nucleotide, the transport efficiency differs with the length and nucleotide composition of the ODNs and varies in different types of leukaemia cells; among the 5 tested random sequence ODNs and 3 aptamers with varying sequences, the data showed that some sequences were associated with significantly higher transport efficiency than others. The transport of ODNs was sodium, energy, and pH-independent, membrane protein-dependent, substrate nonspecific for ODNs and 4-nitrobenzylthioinosine (NBMPR)-insensitive, but it showed a low sensitivity to dipyridamole (IC50 = 35.44 µmol/L), distinguishing it from ENT1-4 and CNTs. The delivery efficiency of ODNs was superior to that of Lipofection and Nucleofection, demonstrating its potential applications in research or therapeutics. Moreover, this process was associated with p38 mitogen activated protein kinase (p38MAPK) instead of c-Jun N-terminal kinase (JNK) signalling pathways. We have denoted ODN transmembrane transport as equilibrative nucleic acid transport (ENAT). Overall, these findings indicate a new approach and mechanism for transmembrane transport of ODNs.
Similar content being viewed by others
Introduction
Despite the remarkable progress in the field of gene therapy for haematological diseases during the past several decades, limitations in gene delivery methods represent a barrier to its clinical application. Major limitations lie in low gene delivery efficiency due to the macromolecular and poly-anionic properties of the nucleic acids and the questionable safety of the delivery vectors1. Among various virus- or non-virus-based carrier systems, well-designed viral vectors (e.g., lentiviral vectors or adeno-associated virus [AAV] vectors) represent the gene delivery tools with the highest efficiency2. However, most of these vectors are associated with significant disadvantages. Lentiviral vectors can effectively integrate into the host cell genome, but genome integration may cause leukaemia if the insertion site is adjacent to a proto-oncogene3,4. AAV vectors can be applied to a broad range of hosts with very few insertion mutations, but they may induce a capsid-specific T cell immune response5,6 and integration-mediated tumourigenicity7. Non-virus (e.g., chemical vector) methods have shown a high delivery efficiency of Oligodeoxyribonucleotides (ODN) transport8,9,10, but there is a lack of long-term safety data1,11. Moreover, some physical methods may require expensive apparatuses or cause damage to samples11. Therefore, effective and safe gene delivery remains an area of improvement in gene therapy.
ODNs, short single-stranded DNA (ssDNA) molecules, have a broad range of applications in the field of targeted therapy. Single-stranded antisense ODNs can form DNA/RNA hybrids with specific mRNA strands to mediate mRNA degradation and can therefore be used to silence target gene expression12. Nucleic acid aptamers, oligonucleotide molecules that bind to a specific target molecule (protein or RNA) with high affinity, can be used to identify tumour cell biomarkers or mediate gene silencing for therapeutic purposes13,14,15,16,17. However, the delivery of ODNs into cells with high efficiency, especially without the aid of vectors, is very challenging.
Previous research has reported that nucleic acids, including ODNs, can be actively internalized into living cells18,19,20,21. The possible mechanism is receptor- mediated, temperature-dependent endocytosis. Following entry into the endocytic pathway, the majority of ODNs are degraded by lysosomal enzymes, and only a small fraction is released into the cytoplasm1. Thus, endocytosis is not suitable for ODN delivery. In this study, we observed that ODNs could be transported intact into the cytoplasm without any delivery tools in a temperature and energy-independent manner (Fig. 1C), although the exact mechanism was unclear. We hypothesized that the nucleoside transporter family, which is important for cellular nucleoside uptake, was involved in ODN transport. However, we found that ODN transport differed from concentrative nucleoside transporters (CNTs) or equilibrative nucleoside transporters (ENTs), and it involved the intracellular signalling pathway p38MAPK. Here, we describe the novel transport of ODNs in cells. To our knowledge, no other studies have examined ODN transmembrane transport and the underlying mechanism.
Characteristics of the transmembrane transport of ODNs. Treated clinical cells were incubated in various buffers to investigate the features of ODN transport. (A) In sodium or sodium-free buffer, the transmembrane transport of ODNs with different lengths and a single type of nucleotides were not significantly different (p > 0.05). (B) After trypsin digestion, the fluorescence intensity of T50-FAM clearly declined (p < 0.05), which indicated that ODN transport was membrane protein dependent. (C) To exhaust cellular ATP, cells were incubated in 10 mM sodium azide and 6 mM 2-deoxy-D-glucose DPBS buffer for 1 hr vs DPBS, and no significant difference in T50-FAM transport was observed in the groups (p > 0.05). (D) In gradient pH (4–8) phosphate buffer, there were no differences in T50-FAM transport (p > 0.05). All experiments were performed in triplicate or duplicate.
Results
Characteristics of ODN transmembrane transport
Because CNTs are sodium-dependent, we assessed ODN transmembrane transport in the presence or absence of sodium. Clinical leukaemia cells (AML, B-ALL, and B-CLL) were incubated with FAM-labelled A1, T1, C1, G1, A10, T10, C10, G10, A50, C50, T50 and T100 in either sodium or sodium-free buffer22. Figure 1A shows that the fluorescence intensity of the cells was not significantly different between sodium and sodium-free buffer (p > 0.05), indicating that ODN transmembrane transport was sodium-independent. As shown in Fig. 1B, pretreatment with trypsin for 2 minutes significantly reduced the fluorescence intensity of T50-FAM-treated cells, which suggested that ODN transmembrane transport was membrane protein-dependent. The fluorescence intensity did not decrease further with trypsin pretreatment longer than 2 minutes, possibly because of complete destruction of membrane proteins at this time point. As shown in Fig. 1C, there were no significant differences between cells pre-treated with sodium azide and 2-deoxy-D-glucose and the control group (p > 0.05). This result indicated that ODN transport was a passive and energy-independent mechanism. As shown in Fig. 1D, the T50-biotin-streptavidin-PE fluorescence of cells in different pH buffers was not significantly different, which suggested that ODN transport was insensitive to pH.
Transmembrane transport of ODNs in clinical leukaemia cells
Clinical leukaemia AML, B-ALL, B-CLL and T-ALL cells can transport various FAM-labelled ODNs into the cytoplasm. Figure 2A–D indicates that ODNs composed of a single type of nucleotides were transported in all 4 types of clinical leukaemia cells without delivery vectors, and the transport efficiency varied depending on the cell type and ODN sequence. Within each cell type, the order of decreasing ODN transport efficiency was as follows: for AML: T100 > T50 > T10 > C50 > G10 > C10 > A10 > A50; B-ALL: T100 > T50 > T10 > C50 > G10 > C10 > A50 > A10; B-CLL: T100 > T50 > C50 > G10 > T10 > C10 > A10 > A50; T-ALL: T100 > T50 > T10 > C50 > G10 > C10 > A10 > A50. When comparing transport efficiency across cell types, T-ALL cells generally had a lower fluorescence intensity than the other cell types for most ODNs. Figure 2E shows that the length of the ODNs affected the transport efficiency in clinical leukaemia cells (all 4 cell types mixed together). In general, the transport efficiency, as indicated by the fluorescence intensity, increased with increasing ODN length in the treated cells. Among the 10-nucleotide ODNs, T10 and G10 were transported with higher efficiency than A10 and C10 (p < 0.001). Among ODNs of 50–100 nucleotides, T50 and T100 demonstrated a higher transport efficiency than C50 and A50 (p < 0.001). Figure 2F shows that, in clinical leukaemia cells (all 4 cell types mixed together), the transport efficiency of A50 was increased after pre-incubation with T50 compared with the control. This finding seemed reasonable because T50 could bind and maintain a low concentration of A50 in the cytoplasm, thus maintaining the inward-directed A50 concentration gradient, which resulted in an influx of extracellular A50.
Flow cytometry quantification of the transmembrane transport efficiency of different ODNs. Clinical leukaemia cells were incubated with ODNs or DPBS for 30 minutes. Treated cells were incubated with solution A, and untreated cells were incubated with DPBS. (A–D) The fluorescence intensity of different types of clinical leukaemia cells, AML, B-ALL, B-CLL and T-ALL, incubated with various FAM-labelled ODNs composed of a single type of nucleotide vs control (incubated with DPBS). The transport efficiency of various ODNs, indicated by the intracellular fluorescence intensity, differed depending on the sequence and the cell type. (E) In the treated group, the fluorescence intensity of cells treated with FAM-labelled ODNs composed of a single type of nucleotides vs the length of the ODNs. The fluorescence intensity generally increased with increasing length of ODNs composed of the same type of nucleotide. (F) The fluorescence intensity of clinical leukaemia cells treated with FAM-labelled A50 with and without T50 pre-incubation vs control (incubated with DPBS). The cellular fluorescence intensity was significantly increased following T50 pre-incubation, indicating a higher transport efficiency of A50 in response to this procedure. (G–J) The fluorescence intensity of clinical AML, B-ALL, B-CLL, and T-ALL cells treated with various FAM-labelled random sequence ssDNAs (PMY6-10) and aptamers (sgc6, sga16, TD08) vs control (incubated with DPBS). While different types of cells showed varying fluorescence intensities when treated with the same ODNs, PMY7-FAM and TD08-FAM consistently displayed the highest fluorescence intensity among all random sequence ssDNAs and aptamers, respectively, in all cell types.
ODN transmembrane transport also occurred for random sequences of ssDNAs and aptamers. Figure 2G–J shows the fluorescence intensity in clinical AML, B-ALL, B-CLL and T-ALL cells treated with different random sequence ssDNAs (PMY6-10) and aptamers (sgc6, sga16, TD08). We found that B-CLL cells had a higher fluorescence intensity than other cell types incubated with PMY7 and TD08 (p < 0.05), but not other ODNs. Moreover, T-ALL cells showed the lowest fluorescence intensity among all cell types for most random sequence ssDNAs and aptamers. While the different types of leukaemia cells varied in fluorescence intensity when treated with the same ODNs, treatment with some ODNs consistently resulted in a higher fluorescence intensity than others across cell types. Among all random sequence ssDNAs, PMY7 treatment resulted in the highest fluorescence intensity. For aptamers, TD08 treatment was associated with higher fluorescence intensity than sgc6 and sga16. These results suggested that the transport efficiency for random sequence ssDNAs and aptamers was influenced by the DNA sequence.
Kinetics of ODN transmembrane transport
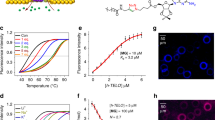
As shown in Fig. 3A, the fluorescence intensity of the various concentrations of T50-FAM-treated cells increased linearly and rapidly, within 1 minute. After 5 minutes, the increment of fluorescence slowed down significantly and gradually reached a plateau, indicating that the uptake of T50 had reached equilibrium. Therefore, we decided to estimate the initial rate of ODN transport at 30 s. The kinetics of ODN (C10, C50, T10, T50, T100) transport, which were determined in K562 cells at different concentrations of ODNs, are shown in Fig. 3B–F, and specific Km and Vmax values are listed in Table 1. It is noteworthy that the rate of T10 (Fig. 3D) uptake depended on its concentration in the buffer. At a low concentration (insert graph), T10 uptake conformed to the Michaelis-Menten equation; however, at a high concentration, it was similar to the pinocytosis of polyvinylpyrrolidone23.
Kinetic analysis of the transmembrane transport of ODNs. Time courses of the transmembrane transport of various concentrations of T50-FAM were studied in K562 cells. As shown in Fig. 3A, the fluorescence intensity of the T50-FAM in K562 cells increased rapidly within 1 minute, exhibiting a linear increase. After 5 minutes, the increment of fluorescence slowed down significantly and gradually reached a plateau, indicating that T50 uptake had reached equilibration. Thus, we decided to estimate the initial rate of ODN transport at 30 s. The kinetic analysis of ODN (C10, C50, T10, T50, T100) transport, which was determined at different concentrations in K562 cells, is shown in Fig. 3(B–F). It is noteworthy that the uptake of T10 (Fig. 3D) depended on its concentration in the buffer. At a low concentration (insert graph), the uptake of T10 conformed to the Michaelis-Menten equation. However, at a high concentration, the uptake of T10 was similar to the pinocytosis of polyvinylpyrrolidone. Km and Vmax values are listed in Table 1. The experiments were performed in triplicate.
Cellular localization of ODNs
To be qualified for gene therapy applications, a delivery method should locate the target DNA in the cytoplasm or the nucleus, where the exogenous genes retain their functions24. However, in previously reported ODN cellular internalization processes, the exogenous DNA was targeted to the endocytic pathway and degraded in the lysosome1. To assess the therapeutic potential of ODN transmembrane transport, we determined the cellular localization of the ODNs in mixed clinical leukaemia cells (AML, B-ALL, and T-ALL) using confocal microscopy. In T50-FAM-treated cells, strong fluorescent signals were observed on the cell membrane and in the cytoplasm (Fig. 4B); in C50-FAM-treated cells, fluorescent signals were detected in the cytoplasm (Fig. 4C). However, in G10-FAM (Fig. 4D) and T10-FAM (Fig. 4E)-treated cells, the fluorescent signals were observed in the nucleus (stained with DAPI), indicating a nuclear localization of these two ODNs. In addition, aptamer TD08-FAM was also localized in the nucleus, as demonstrated by the overlapping green fluorescence and DAPI signals (Fig. 4F). The differential cellular localization of the various ODNs may be dependent on the length and specific sequence of an ODN. The precise underlying mechanism requires further investigation.
Confocal microscopy analysis of the cellular localization of various ODNs in the treated cells. (A) Control image with no fluorescence. (B) T50-FAM-treated cells, in which strong green fluorescence was detected on the cell membrane and in the cytoplasm. (C) C50-FAM-treated cells, in which moderate green fluorescence was detected in the cytoplasm. (D) G10-FAM-treated cells, in which strong green fluorescence co-localized with DAPI staining of the nucleus (blue fluorescence). (E) T10-FAM-treated cells, in which strong green fluorescence co-localized with DAPI staining of the nucleus (blue fluorescence). (F) TD08-FAM-treated cells, in which green fluorescence co-localized with DAPI staining of the nucleus (blue fluorescence). A,B,C and F were taken at 20X magnification; D and E were taken at 1000X magnification to show the nucleus. Images of FAM-labelled A50, C10, A10, A1, C1, T1, and G1-treated cells are not shown because of the very weak fluorescence intensity.
Inhibition of ODN transport by dipyridamole but not by 4-nitrobenzylthioinosine (NBMPR)
Because ENTs have been reported to transport nucleosides into cells and NBMPR or coronary vasodilator dipyridamole can inhibit the transport function25,26, we speculated that transmembrane transport of ODNs might be blocked by ENT inhibitor. To test this hypothesis, we pre-incubated clinical leukaemia cells (AML [n = 1], B-ALL [n = 1], and B-CLL [n = 1]) with NBMPR before addition of T50-FAM and PMY6-10-FAM. NBMPR clearly did not impede ODN transport (Fig. 5A and C). We also incubated cells with FAM-labelled T50 or PMY6-10 before or after treatment with the other ENT inhibitor, dipyridamole. Flow cytometry analysis showed that pretreatment of the cells with dipyridamole decreased the fluorescence signal of FAM-labelled T50 (Fig. 5B) or PMY6-10 (Fig. 5D), which suggested that T50 and PMY6-10 uptake was inhibited by dipyridamole in concentration-dependent manner (p < 0.05). When the concentration of T50-FAM was 250 nmol/L, the IC50 of dipyridamole was 34.55 µM.
Inhibition of ODN transmembrane transport by dipyridamole but not by NBMPR. T50-FAM or PMY6-10-FAM uptake by cells could be blocked by pretreatment with dipyridamole (Fig. 5B,D) in a concentration-dependent manner (p < 0.05), but not by NBMPR (p > 0.05) (Fig. 5A,C). The uptake of T50 or PMY6-10 was largely inhibited by dipyridamole at a high concentration (1 mmol/L). When the concentration of T50-FAM was 250 nmol/L, the IC50 of dipyridamole was 34.55 μM. The experiment was repeated 3 times with similar results.
Based on these experiments, we conclude that ODN transmembrane transport can be entirely inhibited by a high concentration of dipyridamole and therefore may differ from ENT-dependent nucleoside transport.
Comparison of the efficiency of 3 kinds of transport methods and the transport of ssDNA (T50) and dsDNA (T = A50)
To further assess the therapeutic potential of ODN transmembrane transport in the clinical environment or research field, we compared the delivery efficiency between ODN transmembrane transport and transfection using Lipofection or Nucleofection in the K562 cell line. A leukaemia cell line was chosen for this experiment because it is easier to manipulate for transfection than clinical leukaemia cells. Equal numbers of cells were incubated with 250 nmol/L FAM-labelled T50 or transfected with the same amount of FAM-labelled T50 using Lipofection or Nucleofection. The delivery efficiency was quantified by flow cytometry. As shown in Fig. 6A, T50-FAM transport by vector-independent transmembrane transport showed a higher fluorescence intensity than those transfected by Lipofection or Nucleofection (p < 0.05). This result suggests that ODN transmembrane transport achieves a higher delivery efficiency compared with Lipofection or Nucleofection.
Comparison of the efficiency of 3 transport methods and the transport of ssDNA (T50) and dsDNA (T = A50). (A) The efficiency of FAM-labelled T50 delivery in K562 cells, as indicated by the fluorescence intensity, was significantly higher in the presence of ENAT (equilibrative nucleic acid transporter) compared with Lipofection or Nucleofection (p < 0.05). (B) The fluorescence intensity of clinical AML cells treated with FAM-labelled single-stranded T50 and double-stranded T = A50 vs control (incubated with DPBS). While strong fluorescent signals were detected in T50-treated cells, the signal intensity in T = A50 treated cells was very low (p < 0.05), indicating minimal dsDNA transmembrane transport. The experiments were performed in triplicate.
To examine whether double-stranded DNA (dsDNA) is also transported via vector-independent transmembrane transport, we tested FAM-labelled T = A50 in clinical AML cells. As shown in Fig. 6B, the transport efficiency of T = A50 was substantially decreased compared with single-stranded T50 (p < 0.05), which indicated that vector-independent transport of dsDNA was minimal. Compared with the ODN-treated cells, all control cells displayed minimal fluorescence intensity (p < 0.05).
Signal transduction pathway involved in ODN transmembrane transport
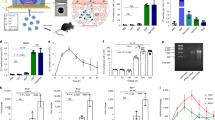
To further explore the signal transduction mechanism of ODN transport in cells, we investigated the influence on ODNs transport of several pathways inhibitors, such as the PKC inhibitor Gö6983 and the apoptosis inhibitor Z-VAD-FMK, which had no impact on T50 transport (p > 0.05) as shown in Fig. 7A. We also detected the phosphorylation of JNK and p38MAPK by western blotting (cropped blots were showed here and full-length and multiple exposures data are included in Supplementary file) and found that p38MAPK was lower but p-p38MAPK expression was clearly higher than the control group (Fig. 7B, p < 0.05), and the expression levels of both JNK & p-JNK were lower than in the control group (Fig. 7C, p < 0.05). Using the p38MAPK inhibitor SB203580, T50 uptake ability of the cells was largely decreased (Fig. 7E, p < 0.05). However, anisomycin, a potent activator of SPAKs/JNKs, did not increase the uptake of T50 (Fig. 7F, p > 0.05). Quantification values of the western blot bands normalized to internal and untreated control are shown in Fig. 7D. These results suggested that the p38MAPK rather than the JNK, PKC or apoptosis signalling pathway was involved in ODN transport.
Signalling pathway involved in ODN transport. (A) The PKC inhibitor Gö6983 and apoptosis inhibitor Z-VAD-FMK had no effect on T50 transport (p > 0.05). (B) Cropped western blot images showing the total p38, p-p38 and GAPDH protein abundance, among which p-p38 was elevated and p38 was reduced in the treated sample compared with the NC (normal control) (both p < 0.05). (C) Cropped western blot showing total JNK, p-JNK and Actin protein abundance, among which both p-JNK and JNK were lower in the treated sample than the NC (p < 0.05), and p-JNK was much lower in treated sample than in the NC. Full-length blots and multiple exposures are provided in Supplementary file online. Quantification of the western blot bands in 7B and 7 C normalized to the internal and untreated controls are shown in (D), in which “*” represents a significant difference compared with the control at p < 0.05. In the blots, the samples are derived from the same experiment, and the gels/blots were processed in parallel. (E) Using the p38MAPK inhibitor SB203580, T50 uptake by the cells was largely decreased (p < 0.05). (F) Anisomycin, a potent activator of SPAKs/JNKs, should increase but slightly decreased the uptake of T50 (p > 0.05). These results suggested that rather than JNK, PKC or apoptosis signalling pathways, p38MAPK is involved in ODN transport. The experiments were performed in triplicate.
Discussion
It is well known that nucleoside transporters are classified into two categories, Na+-independent ENTs and Na+-dependent CNTs27. ENTs are a family of transmembrane proteins that comprises four members: ENT1-4. They mediate the bidirectional, concentration gradient-driven transport of nucleosides and their analogues into the cytoplasm. CNTs are also transmembrane proteins and have three members, CNT1-328, which transport nucleosides and their analogues across the membrane in a unidirectional manner. Among all ENT members, ENT1 and ENT2 are sensitive to certain cardiovascular drugs, such as NBMPR, dipyridamole, and delazep28,29. They are also widely distributed in most tissue and cell types and have the ability to transport a broad range of purine and pyrimidine nucleosides into the cells30,31. In the present study, we showed that leukaemia cells could transport ODNs into the cytoplasm or nucleus in a sodium-independent (Fig. 1A) and energy-independent manner (Fig. 1C). Given that the activity of CNTs is cation-dependent and an active transport process28, we reasoned that CNTs were not involved in ODN transmembrane transport. The ODN transmembrane transport is insensitive to NBMPR, whereas it shows only weak sensitivity to dipyridamole with an IC50 of 35.44 µM for T50 (250 nmol/L, Fig. 5B), which is approximately 10–1000-fold higher than that of ENT1 or ENT2 for the relevant substrate32,33. In addition, the Vmax of ODN transport (Table 1) was 6 orders of magnitudes lower than the Vmax of substrate uptake for ENT1 and ENT233,34; therefore, it is distinct from ENT1/2. Furthermore, ODN transport was cell membrane protein-dependent (Fig. 1B), pH-independent (Fig. 1D) and broadly permissive to substrates for ODNs. In contrast, ENT3 is primarily located in intracellular membranes such as the membranes of lysosomes and mitochondria30,31. Additionally, ENT4 is pH-dependent and specific for adenosine35, and hence is distinguished from ENT3 and ENT4. Moreover, some intracellular membrane proteins such as ANT (adenine nucleotide translocase, SLC25A4) and Sidt2, among others, which are located in mitochondrial membrane or lysosomal organelles and involved in ADP/ATP energy metabolism or RNA uptake36,37, also differ in protein location and functional aspects of ODN transport across the plasma membrane. The IC50 values of known proteins inhibited by dipyridamole, such as p-glycoprotein (IC50 1.5 ± 1.5 μM)38, breast cancer resistance protein (BCRP) (IC50 6.4 ± 0.9 μM)39, and ATP binding cassette transporter 5 (ABCC5) (Ki 1.2 μM or 5.5 μM)40, are one order of magnitude lower than that of ODN transport (IC50 35.44 µM). Furthermore, in contrast to these active transporter proteins, ODN transport is energy-independent (Fig. 1C).
We also compared the transport efficiency of ODNs composed of a single type of oligonucleotide (A, T, C, or G) with varying lengths and in different types of leukaemia cells. We found that ODNs with different compositions differed in their transport efficiency. Overall, among the 10-nucleotide ODNs, T10 exhibited the highest transport efficiency, whereas A10 had the lowest efficiency. In contrast, T chain ODNs with a length of 50 or 100 nucleotides demonstrated a higher transport efficiency than A chain and C chains with the same length (G chain ODNs were not investigated because of technical limitations). While this trend was consistent across the 4 types of leukaemia cells tested in this study, the transport efficiency of each individual ODN may differ in different cell types. The transport efficiency of different ODNs decreased in the following order for AML: T100 > T50 > T10 > C50 > G10 > C10 > A10 > A50; for B-ALL: T100 > T50 > T10 > C50 > G10 > C10 > A50 > A10; B-CLL: T100 > T50 > C50 > G10 > T10 > C10 > A10 > A50; and for T-ALL: T100 > T50 > T10 > C50 > G10 > C10 > A10 > A50. We found that ODNs with different compositions showed the lowest fluorescence in T-ALL among all the tested clinical leukaemia cells, which is not consistent with the high mRNA expression of ENT1/2 in T-ALL41. This finding also confirmed that the high level of ENT1/2 mRNA expression was not correlated with the ODN transport activity in T-ALL. Additionally, the order of nucleotide preference of ODN transport differed from that of ENT1 and ENT2. Ward et al. has reported that the binding affinity of transporters for different nucleosides decreases in the following order for ENT1: adenosine > guanosine > isodine > uridine > thymidine >cytidine; and in the following order for ENT2: isodine > adenosine > uridine > thymidine > guanosine > cytidine33. Another study also reported that thymidine uptake is equally distributed between ENT1 and ENT242. One possible explanation for the discrepancy in nucleotide preference between ODN transport and nucleoside transport is that cells possess two distinctive transport functions under different conditions; one is to transport nucleosides, and the other is to transport ODNs. Together, these data support that ODN transport is mediated by a novel nucleic acid transport system that is distinct from ENTs and CNTs. The protein identity of ODN transport is presently not known. Despite the lack of protein identity and literature evidence, we have named the ODN transporter equilibrative nucleic acid transporter (ENAT).
In the present study, we were particularly concerned about the transmembrane transport of A50 because of possible interference from intracellular mRNA poly(A) tails. To eliminate such interference, we pre-treated the cells with T50 before the incubation with A50. Our results demonstrated that the cells were able to transport A50 into the cytoplasm, and pre-incubation with T50 increased the transport efficiency. Taken together, ENAT was able to transport ODNs with varying sequences, although some sequence-dependent differences in transport efficiency were observed. Furthermore, we confirmed that ENAT was able to transport random sequence ssDNA and aptamers into the cytoplasm or the nucleus. Overall, ENAT is broadly selective for pyrimidine and purine nucleotide strands and is a highly efficient transport system distinct from endocytosis, leading to nucleic acid degradation in the lysosome1. These findings suggest potential research or therapeutic applications for ENAT, particularly in the gene therapy field. Currently, gene therapy primarily relies on viral vectors to deliver various DNA sequences. However, viral vectors have potential side effects such as virus-mediated pathogenicity, oncogenicity, or immunogenicity, as well as the need to reserve longer DNA fragments to accommodate the viral genes. Therefore, ENAT would be very beneficial for clinical usage because it can deliver ODNs into the cytoplasm or the nucleus in the absence of vectors. Furthermore, ENAT demonstrated a higher delivery efficiency than transfection using Lipofection or Nucleofection. However, although it may not be suitable for the transport of double-stranded ODNs (Fig. 6B), ENAT provides a potentially safe and effective approach for ODN transport in gene therapy.
It is also important to understand the regulatory mechanism of ENAT to determine its potential clinical value. Recently, two studies indicated that ENT inhibition prevented p38MAPK and c-JNK activity and EC barrier dysfunction43,44. Huang M et al. found that nucleoside uptake of K562 cells was prevented by the inactive p38MAPK inhibitor SB202474, which suggested that nucleoside uptake inhibition by SB203580 was independent of p38MAPK inhibition but linked to the compound itself45. A study suggested that the phosphorylation of multiple pathways, such as PKC and PKA, was involved in ENT146. We speculate that the pathways responsible for modulating ENTs are related to ENAT. Thus, we investigated the influence of different signalling pathway inhibitors (SB203580 for p38MAPK, Gö6983 for PKC and Z-VAD-FMK for apoptosis) on ENAT. The results showed that the inhibitor of p38MAPK weakened the activity of ENAT, but inhibition of PKC and apoptosis had no effect (Fig. 7F). Western blotting results showing increased p-p38MAPK and decreased p38MAPK but decreased p-JNK and JNK, compared with the control sample, also support the involvement of p38MAPK phosphorylation in ENAT. These results facilitate our understanding of the mechanism by which ENAT is modulated under certain condition, i.e., solution A treatment (recipe pending patent application).
In conclusion, we demonstrated ODN transmembrane transport occurs via ENAT in clinical leukaemia cells; however, the exact underlying mechanisms require further investigation. Other areas of future research include determination of the transport function regulation of ENAT and whether oligoribonucleotides can be transported across the membrane in a vector-independent manner.
Materials and Methods
Synthesis of ODNs and random sequence ODNs from library amplification
The random sequence ssDNA library, PCR primers, fluorescein (FAM)-labelled aptamers (sgc 6, sga 16, TD 08), ODNs composed of a single type of nucleotides with varying lengths (A1, T1, C1, G1, A10, T10, C10, G10, A50, C50, T50, T100) labelled with 5′-FAM and ODNs (C10, C50, T10, T50, T100) labelled with 5′-FAM and phosphorothioate at 3 base of both ends were synthesized by Shanghai Biological Engineering Co., Ltd. (Shanghai, China). Phosphorothioate A10 and A50 were not synthesized because A10 and A50-FAM exhibited very dim fluorescence in cells. G50 and phosphorothioate G10 were not synthesized due to technical limitations.
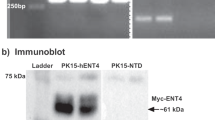
The random sequence ssDNA library contained the following sequences: 5′-GCAATGGTACGGTACTTCC (50 N) CAAAAGTGCACGCTACTTTGCTAA-3′, where “50 N” represented random sequences of 50 nucleotides. The ssDNA library was amplified by PCR using the following forward primer 5′-GCA ATG GTA CGG TAC TTC C-3′ and reverse primer 5′-TTA GCA AAG TAG CGT GCA CTT TTG-3′. Hot Master Taq DNA polymerase, 10 × PCR buffer, and dNTPs were purchased from TIANGEN BIOTECH Co., Ltd. (Beijing, China). The PCR products were purified by agarose gel electrophoresis and sequenced by Beijing Nohezhiyuan Co., Ltd. (Beijing, China). The 5 sequences with the highest abundance (PMY6-10) were further synthesized for FAM labelling in large quantities.
Cell lines, culture conditions, and transfection
The human chronic myeloid leukaemia cell line K562 was obtained from the Cell Research Center, Centre South University XiangYa School of Medicine. K562 cells were cultured in RPMI-1640 medium (GIBCO BRL, Burlington, MA, USA) containing 10% foetal bovine serum at 37 °C in humidified air supplied with 5% CO2.Cells in the logarithmic growth phase were harvested and seeded in 6-well plates at a concentration of 1 × 106/ml. Cells were transfected with T50-FAM (final concentration 250 nmol/L) using Lipofection or Nucleofection. Lipofection was performed using the RiboFECT™ CP Transfection Kit (Guangzhou RiboBio Co., Ltd, Guangzhou, China) according to the manufacturer’s instructions. Nucleofection was performed using the Amaxa™ Human Stem Cell Nucleofector™ kit 2 (Amaxa Biosystems, Gaithersburg, MD, USA) according to the manufacturer’s instructions. Cells were collected 24 hours after Lipofection and immediately after Nucleofection for flow cytometry (FACSCalibur or FACSCanto II, BD Bioscience, San Jose, CA, USA) to determine the transfection efficiency.
Cell isolation and treatment
Human clinical leukaemia cells were isolated from bone marrow or peripheral blood cell samples from 28 patients who had signed the informed consent. The patients ranged in age from 1 to 76 years, and 14 of them were male. Of the 28 patients, 16 had acute myeloid leukaemia (AML), 5 had B-cell acute lymphoblastic leukaemia (B-ALL), 4 had B-cell chronic lymphocytic leukaemia (B-CLL), and 3 had T-cell acute lymphoblastic leukaemia (T-ALL). The study protocol was approved by the institutional review boards at Xiangya Hospital, Central South University, and the research design and methods were performed in accordance with the regulatory requirements and procedures regarding human subject protection laws such as GCP and ICH-GCP. Mononuclear cells were separated using lymphocyte separation medium (Sigma-Aldrich, St Louis, MO, USA) following routine procedures. In brief, clinical samples were gently added to equal volumes of lymphocyte separation medium in centrifuge tubes and centrifuged at 1500 revolutions per minute (rpm) for 5 minutes. The middle layer containing the white blood cells (between the plasma layer on the top and the separation medium at the bottom) were collected by aspiration, washed once, and re-suspended in Dulbecco’s phosphate-buffered saline (DPBS; Shanghai Lifei biological technology Co., Ltd, Shanghai, China).
Clinical cells and K562 cells were divided into treatment and control groups. Treatment cells were incubated for 10 minutes in solution A (recipe pending patent application) at a 1:9 ratio and then washed 3 times and re-suspended in DPBS. Control group cells were also washed 3 times and suspended in DPBS. Both the treatment and control groups were adjusted to a concentration of 1 × 106/ml.
Two buffers (sodium and sodium-free solution), which have been described previously, i.e., 1 mM MgCl2, 5 mM KCl, 10 mM HEPES, 25 mM CaCl2, 137 mM choline chloride or NaCl22, were prepared for ODN transport to evaluate sodium dependence. Treatment group cells were washed 3 times and re-suspended in the two buffers described above. All the ODNs were also prepared in two buffers at a concentration of 500 nmol/L. The same volume of cells and ODNs were mixed in the dark at 4 °C for 30 minutes for flow cytometry analysis. As shown in Fig. 1A, ODN transport was sodium-independent, and thus, all subsequent experiments were conducted in DPBS or sodium buffer.
To determine whether ODN transport was membrane protein-dependent, trypsin was added to the treated cells at a concentration of 0.25%. After 2−5 minutes of incubation, trypsin was inactivated by addition of foetal bovine serum, and then the cells were washed 3 times and re-suspended in DPBS. Finally, the cells were incubated with T50-FAM in the dark at 4 °C for 30 minutes for flow cytometry analysis.
To examine whether ODN transport was energy dependent, a portion of the treatment group cells was pre-incubated at 4 °C with 10 mM sodium azide and 6 mM 2-deoxy-D-glucose for 1 hr to delete cellular ATP47. The other cells served as a control. The two cell groups were then incubated with T50-FAM in the dark at 4 °C for 30 minutes for flow cytometry analysis.
To investigate whether ODN transport was pH-dependent, the treatment group cells were washed 3 times and re-suspended in pH 5 to pH 8 phosphate buffer. The cells were then incubated with T50-biotin in the dark at 4 °C for 20 minutes. The cells were washed once with pH 5–8 phosphate buffer and incubated with streptavidin-PE (BD Bioscience, Bedford, MA, USA) in the dark at 4 °C for 20 minutes. After another wash, the cells were re-suspended in pH 5–8 phosphate buffer for flow cytometry analysis.
ODN Transmembrane transport
Various ODNs or dsDNA labelled with FAM at a concentration of 500 nmol/L were added to the treated and control cells, and both groups were incubated in the dark at 4 °C for 30 minutes. After incubation, the cells were washed and re-suspended in DPBS for flow cytometry or confocal microscopy analysis. For confocal microscopy, pre-DAPI staining (100 ng/ml for 20 minutes) was performed followed by re-suspension in DPBS. A FluoViewTM FV1000 laser confocal microscope (Olympus, Tokyo, Japan) was used.
To determine the linearity of ODN transport, the same volume of cells and various concentrations of T50 -FAM (0.1–3.2 μmol/L) were mixed in the dark at 4 °C, and then, T50 transport into cells was examined at different time points up to 30 minutes by flow cytometry analysis.
Transport kinetic assays were performed as described previously48. Flow cytometry was used to analyse the uptake of phosphorothioate ODNs, in which the oxygen atom of the phosphate group was replaced with a sulphur atom to resist nuclease degradation49. Briefly, the cells were suspended in PBS, and then, the same volume of cells and various concentrations of phosphorothioate ODNs were mixed and immediately analysed by flow cytometry. The relative fluorescence intensity of the cells at 30 s was converted into MEF by comparing the single cell relative units of the channel number with the fluorescence of SPHEROTM Easy Calibration Fluorescent Particles (Catalogue No. ECFP-FI-5K, Spherotech, Lake Forest, IL, USA) under the same conditions. Treated cells (total uptake) diminish the control cells (non-specific binding) are defined as mediated uptake.
Pre-incubation with dipyridamole and NBMPR
Treated cells were pre-incubated with various concentrations (from 1 μmol/L to 1 mmol/L) of dipyridamole (Dalian Mellon biological technology Co. Ltd, Dalian, China) and (from 1 nmol/L to 100 μmol/L) NBMPR (Sigma-Aldrich, St Louis, MO, USA), a hENT1 inhibitor, for 30 minutes before incubation with T50-FAM or PMY6-10-FAM for 30 minutes. The cells were then re-suspended for analysis by flow cytometry or confocal microscopy.
Protein extraction and Western Blotting
First, K562 cells were incubated for 10 minutes in solution A, which contained 1 × protease inhibitor and 1 × PhosStop cocktail (Roche, France), and then the treated cells were lysed in RIPA buffer (Thermo Fisher, Waltham, MA, USA). Control cells were lysed in RIPA buffer in the presence of 1 × protease inhibitor and 1 × PhosStop cocktail for 20 minutes. After sonication three times, the lysed cell product was centrifuged at 12000 rpm for 10 minutes at 4 °C. The protein concentration was measured using a Pierce® BCA protein assay kit (Thermo Fisher, Waltham, MA, USA). Western blotting analysis was performed as described previously50. Anti-human actin, GAPDH, p38MAPK, p-p38MAPK, JNK, and p-JNK were obtained from Santa Cruz (Santa Cruz, CA, USA).
Signalling pathway inhibition or activation
Before incubation with T50-FAM for 30 minutes, the treated cells were pre-incubated with the p38MAPK inhibitor SB203580 (0.5,1, 2 µmol/L) for 1 hr; a potent activator of SPAKs/JNKs, anisomycin (0.2, 0.5, 1, 2, 5 µmol/L) for 3 hr; the PKC inhibitor Gö6983 (100 nmol/L) for 1 hr; or the apoptosis inhibitor Z-VAD-FMK (50 nmol/L) for 1 hr. The cells were finally re-suspended for analysis by flow cytometry. (SB203580, Anisomycin, Gö6983 and Z-VAD-FMK were purchased from MedChem Express, Monmouth Junction, NJ, USA.)
Statistical analysis
All data are presented as the mean ± SD except where indicated. Paired or multiple group unpaired data were analysed with the Student’s paired t test or two-way ANOVA, respectively. Km, Vmax and IC50 values were calculated by fitting the data to a non-linear regression Michaelis-Menten equation or dose-response inhibition curve, respectively. All analyses, linear regression lines and curves were defined using GraphPad software 6.0 (GraphPad Software Incorporated, San Diego, CA, USA). *p < 0.05 was considered a statistically significant difference.
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
References
De Rosa, G., De Stefano, D. & Galeone, A. Oligonucleotide delivery in cancer therapy. Expert opinion on drug delivery 7, 1263–1278, https://doi.org/10.1517/17425247.2010.527942 (2010).
Ajith, T. A. Strategies used in the clinical trials of gene therapy for cancer. Journal of experimental therapeutics & oncology 11, 33–39 (2015).
Hacein-Bey-Abina, S. et al. Insertional oncogenesis in 4 patients after retrovirus-mediated gene therapy of SCID-X1. The Journal of clinical investigation 118, 3132–3142, https://doi.org/10.1172/JCI35700 (2008).
Hacein-Bey-Abina, S. et al. Efficacy of gene therapy for X-linked severe combined immunodeficiency. The New England journal of medicine 363, 355–364, https://doi.org/10.1056/NEJMoa1000164 (2010).
Nathwani, A. C. et al. Long-term safety and efficacy of factor IX gene therapy in hemophilia B. The New England journal of medicine 371, 1994–2004, https://doi.org/10.1056/NEJMoa1407309 (2014).
Ferreira, V. et al. Immune responses to intramuscular administration of alipogene tiparvovec (AAV1-LPL(S447X)) in a phase II clinical trial of lipoprotein lipase deficiency gene therapy. Human gene therapy 25, 180–188, https://doi.org/10.1089/hum.2013.169 (2014).
Kaeppel, C. et al. A largely random AAV integration profile after LPLD gene therapy. Nature medicine 19, 889–891, https://doi.org/10.1038/nm.3230 (2013).
Wang, S., Zhang, X., Yu, B., Lee, R. J. & Lee, L. J. Targeted nanoparticles enhanced flow electroporation of antisense oligonucleotides in leukemia cells. Biosensors & bioelectronics 26, 778–783, https://doi.org/10.1016/j.bios.2010.06.025 (2010).
Yu, B. et al. Targeted nanoparticle delivery overcomes off-target immunostimulatory effects of oligonucleotides and improves therapeutic efficacy in chronic lymphocytic leukemia. Blood 121, 136–147, https://doi.org/10.1182/blood-2012-01-407742 (2013).
Jin, Y. et al. Targeted delivery of antisense oligodeoxynucleotide by transferrin conjugated pH-sensitive lipopolyplex nanoparticles: a novel oligonucleotide-based therapeutic strategy in acute myeloid leukemia. Molecular pharmaceutics 7, 196–206, https://doi.org/10.1021/mp900205r (2010).
Geinguenaud, F., Guenin, E., Lalatonne, Y. & Motte, L. Vectorization of Nucleic Acids for Therapeutic Approach: Tutorial Review. ACS chemical biology 11, 1180–1191, https://doi.org/10.1021/acschembio.5b01053 (2016).
Falzarano, M. S., Passarelli, C. & Ferlini, A. Nanoparticle delivery of antisense oligonucleotides and their application in the exon skipping strategy for Duchenne muscular dystrophy. Nucleic acid therapeutics 24, 87–100, https://doi.org/10.1089/nat.2013.0450 (2014).
Yang, M. et al. Developing aptamer probes for acute myelogenous leukemia detection and surface protein biomarker discovery. Journal of hematology & oncology 7, 5, https://doi.org/10.1186/1756-8722-7-5 (2014).
Zhu, P. et al. [In vitro selection of single strand deoxyribonucleic acid aptamers binding to cells from patients with acute myeloblastic leukemia]. Zhong nan da xue xue bao. Yi xue ban = Journal of Central South University. Medical sciences 37, 771–776, https://doi.org/10.3969/j.issn.1672-7347.2012.08.003 (2012).
Sefah, K. et al. Molecular recognition of acute myeloid leukemia using aptamers. Leukemia 23, 235–244, https://doi.org/10.1038/leu.2008.335 (2009).
Pan, Y. et al. Selective collection and detection of leukemia cells on a magnet-quartz crystal microbalance system using aptamer-conjugated magnetic beads. Biosensors & bioelectronics 25, 1609–1614, https://doi.org/10.1016/j.bios.2009.11.022 (2010).
Deng, T. et al. A sensitive fluorescence anisotropy method for the direct detection of cancer cells in whole blood based on aptamer-conjugated near-infrared fluorescent nanoparticles. Biosensors & bioelectronics 25, 1587–1591, https://doi.org/10.1016/j.bios.2009.11.014 (2010).
Yan, A. & Levy, M. Cell internalization SELEX: in vitro selection for molecules that internalize into cells. Methods in molecular biology 1103, 241–265, https://doi.org/10.1007/978-1-62703-730-3_18 (2014).
Zhang, K. et al. A novel aptamer developed for breast cancer cell internalization. ChemMedChem 7, 79–84, https://doi.org/10.1002/cmdc.201100457 (2012).
Xiao, Z., Shangguan, D., Cao, Z., Fang, X. & Tan, W. Cell-specific internalization study of an aptamer from whole cell selection. Chemistry 14, 1769–1775, https://doi.org/10.1002/chem.200701330 (2008).
Soundararajan, S. et al. Plasma membrane nucleolin is a receptor for the anticancer aptamer AS1411 in MV4-11 leukemia cells. Molecular pharmacology 76, 984–991, https://doi.org/10.1124/mol.109.055947 (2009).
Fernandez Calotti, P. et al. Modulation of the human equilibrative nucleosidetransporter1 (hENT1) activity by IL-4 and PMA in B cells from chronic lymphocytic leukemia. Biochemical pharmacology 75, 857–865, https://doi.org/10.1016/j.bcp.2007.10.017 (2008).
Yakubov, L. A. et al. Mechanism of oligonucleotide uptake by cells: involvement of specific receptors? Proceedings of the National Academy of Sciences of the United States of America 86, 6454–6458 (1989).
Patil, S. D., Rhodes, D. G. & Burgess, D. J. DNA-based therapeutics and DNA delivery systems: a comprehensive review. The AAPS journal 7, E61–77, https://doi.org/10.1208/aapsj070109 (2005).
Deniskin, R., Frame, I. J., Sosa, Y. & Akabas, M. H. Targeting the Plasmodium vivax equilibrative nucleoside transporter 1 (PvENT1) for antimalarial drug development. International journal for parasitology. Drugs and drug resistance 6, 1–11, https://doi.org/10.1016/j.ijpddr.2015.11.003 (2016).
Endres, C. J., Sengupta, D. J. & Unadkat, J. D. Mutation of leucine-92 selectively reduces the apparent affinity of inosine, guanosine, NBMPR [S6-(4-nitrobenzyl)-mercaptopurine riboside] and dilazep for the human equilibrative nucleoside transporter, hENT1. The Biochemical journal 380, 131–137, https://doi.org/10.1042/bj20031880 (2004).
Podgorska, M., Kocbuch, K. & Pawelczyk, T. Recent advances in studies on biochemical and structural properties of equilibrative and concentrative nucleoside transporters. Acta biochimica Polonica 52, 749–758 (2005).
Young, J. D., Yao, S. Y., Baldwin, J. M., Cass, C. E. & Baldwin, S. A. The human concentrative and equilibrative nucleoside transporter families, SLC28 and SLC29. Molecular aspects of medicine 34, 529–547, https://doi.org/10.1016/j.mam.2012.05.007 (2013).
Molina-Arcas, M., Casado, F. J. & Pastor-Anglada, M. Nucleoside transporter proteins. Current vascular pharmacology 7, 426–434 (2009).
Baldwin, S. A. et al. Functional characterization of novel human and mouse equilibrative nucleoside transporters (hENT3 and mENT3) located in intracellular membranes. The Journal of biological chemistry 280, 15880–15887, https://doi.org/10.1074/jbc.M414337200 (2005).
Govindarajan, R. et al. Facilitated mitochondrial import of antiviral and anticancer nucleoside drugs by human equilibrative nucleoside transporter-3. American journal of physiology. Gastrointestinal and liver physiology 296, G910–922, https://doi.org/10.1152/ajpgi.90672.2008 (2009).
Visser, F. et al. Mutation of residue 33 of human equilibrative nucleoside transporters 1 and 2 alters sensitivity to inhibition of transport by dilazep and dipyridamole. The Journal of biological chemistry 277, 395–401, https://doi.org/10.1074/jbc.M105324200 (2002).
Ward, J. L., Sherali, A., Mo, Z. P. & Tse, C. M. Kinetic and pharmacological properties of cloned human equilibrative nucleoside transporters, ENT1 and ENT2, stably expressed in nucleoside transporter-deficient PK15 cells. Ent2 exhibits a low affinity for guanosine and cytidine but a high affinity for inosine. The Journal of biological chemistry 275, 8375–8381 (2000).
Hughes, S. J., Cravetchi, X., Vilas, G. & Hammond, J. R. Adenosine A1 receptor activation modulates human equilibrative nucleoside transporter 1 (hENT1) activity via PKC-mediated phosphorylation of serine-281. Cellular signalling 27, 1008–1018, https://doi.org/10.1016/j.cellsig.2015.02.023 (2015).
Zhou, M., Duan, H., Engel, K., Xia, L. & Wang, J. Adenosine transport by plasma membrane monoamine transporter: reinvestigation and comparison with organic cations. Drug metabolism and disposition: the biological fate of chemicals 38, 1798–1805, https://doi.org/10.1124/dmd.110.032987 (2010).
Maldonado, E. N. et al. ATP/ADP Turnover and Import of Glycolytic ATP into Mitochondria in Cancer Cells Is Independent of the Adenine Nucleotide Translocator. The Journal of biological chemistry 291, 19642–19650, https://doi.org/10.1074/jbc.M116.734814 (2016).
Aizawa, S. et al. Lysosomal putative RNA transporter SIDT2 mediates direct uptake of RNA by lysosomes. Autophagy 12, 565–578, https://doi.org/10.1080/15548627.2016.1145325 (2016).
Verstuyft, C. et al. Dipyridamole enhances digoxin bioavailability via P-glycoprotein inhibition. Clinical pharmacology and therapeutics 73, 51–60, https://doi.org/10.1067/mcp.2003.8 (2003).
Zhang, Y. et al. BCRP transports dipyridamole and is inhibited by calcium channel blockers. Pharmaceutical research 22, 2023–2034, https://doi.org/10.1007/s11095-005-8384-4 (2005).
Aronsen, L., Orvoll, E., Lysaa, R., Ravna, A. W. & Sager, G. Modulation of high affinity ATP-dependent cyclic nucleotide transporters by specific and non-specific cyclic nucleotide phosphodiesterase inhibitors. European journal of pharmacology 745, 249–253, https://doi.org/10.1016/j.ejphar.2014.10.051 (2014).
Homminga, I. et al. In vitro efficacy of forodesine and nelarabine (ara-G) in pediatric leukemia. Blood 118, 2184–2190, https://doi.org/10.1182/blood-2011-02-337840 (2011).
Roovers, K. I. & Meckling-Gill, K. A. Characterization of equilibrative and concentrative Na+ -dependent (cif) nucleoside transport in acute promyelocytic leukemia NB4 cells. Journal of cellular physiology 166, 593–600, doi:10.1002/(SICI)1097-4652(199603)166:3<593::AID-JCP14>3.0.CO;2-8 (1996).
Lu, Q. et al. Sustained adenosine exposure causes lung endothelial apoptosis: a possible contributor to cigarette smoke-induced endothelial apoptosis and lung injury. Am J Physiol-Lung C 304, L361–L370, https://doi.org/10.1152/ajplung.00161.2012 (2013).
Lu, Q. et al. Sustained adenosine exposure causes lung endothelial barrier dysfunction via nucleoside transporter-mediated signaling. American journal of respiratory cell and molecular biology 47, 604–613, https://doi.org/10.1165/rcmb.2012-0012OC (2012).
Huang, M., Wang, Y., Mitchell, B. S. & Graves, L. M. Regulation of equilibrative nucleoside uptake by protein kinase inhibitors. Nucleosides, nucleotides & nucleic acids 23, 1445–1450, https://doi.org/10.1081/NCN-200027667 (2004).
Reyes, G. et al. The Equilibrative Nucleoside Transporter (ENT1) can be phosphorylated at multiple sites by PKC and PKA. Molecular membrane biology 28, 412–426, https://doi.org/10.3109/09687688.2011.604861 (2011).
Rezgui, R., Blumer, K., Yeoh-Tan, G., Trexler, A. J. & Magzoub, M. Precise quantification of cellular uptake of cell-penetrating peptides using fluorescence-activated cell sorting and fluorescence correlation spectroscopy. Biochimica et biophysica acta 1858, 1499–1506, https://doi.org/10.1016/j.bbamem.2016.03.023 (2016).
Natarajan, A. & Srienc, F. Dynamics of glucose uptake by single Escherichia coli cells. Metabolic engineering 1, 320–333, https://doi.org/10.1006/mben.1999.0125 (1999).
Eckstein, F. P. essential components of therapeutic oligonucleotides. Nucleic acid therapeutics 24, 374–387, https://doi.org/10.1089/nat.2014.0506 (2014).
Carito, V. et al. Localization of nerve growth factor (NGF) receptors in the mitochondrial compartment: characterization and putative role. Biochimica et biophysica acta 1820, 96–103, https://doi.org/10.1016/j.bbagen.2011.10.015 (2012).
Acknowledgements
We thank professor Wei-hong Tan of state key laboratory of Chem/Biosensing and Chemometrics at Hunan University for support with the experimental techniques and equipment.
Author information
Authors and Affiliations
Contributions
Minyuan Peng participated in the study design, contributed to the data analysis, and wrote the manuscript. Yanming Li collected and analysed the data and participated in writing the manuscript. Jian Zhang, Xiaoyang Yang and Ye Lei participated in the flow cytometry and confocal microscopy procedures. Yong Wu carried out the Nucleofection experiment. Mao Ye provided technical guidance and data analysis of the confocal microscopy results. Xu Han and Yijin Kuang performed the western blotting and Lipofection. Jing Liu and Xielan Zhao participated in the study planning and provided experimental suggestions. Fangping Chen participated in the overall experimental design and helped to edit the manuscript. All authors provided final approval of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Peng, M., Li, Y., Zhang, J. et al. Vector-independent transmembrane transport of oligodeoxyribonucleotides involves p38 mitogen activated protein kinase phosphorylation. Sci Rep 7, 13571 (2017). https://doi.org/10.1038/s41598-017-14099-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-14099-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.