Abstract
Acute aortic dissection (AAD) is a catastrophic emergency with high mortality and misdiagnosis rate. We aimed to determine whether circulating microRNAs allow to distinguish AAD from healthy controls and chest pain patients without AAD (CP). Plasma microRNAs expression were determined in 103 participants, including 37 AAD patients, 26 chronic aortic dissection patients, 17 healthy volunteers, 23 patients without AAD. We selected 16 microRNAs from microarray screening as candidates for further testing via qRT-PCR. The results showed that plasma miR-15a in patients with AAD (n = 37) had significantly higher expression levels than it from control group (n = 40; P = 0.008). By receiver operating characteristic curve analysis, the sensitivity was 75.7%; the specificity was 82.5%; and the AUC was 0.761 for detection of AAD. Furthermore, 37 patients with AAD had significantly higher plasma expression levels of let-7b, miR-15a, miR-23a and hcmv-miR-US33-5p compared with 14 CP patients of 40 controls (P = 0.000, 0.000, 0.026 and 0.011, respectively). The corresponding sensitivity were 79.4%, 75.7%, 91.9% and 73.5%, respectively; the specificity were 92.9%, 100%, 85.7% and 85.7%, respectively; and the AUCs of these microRNAs were 0.887, 0.855, 0.925 and 0.815, respectively. These data indicate that plasma miR-15a and miR-23a have promising clinical value in diagnosing AAD.
Similar content being viewed by others
Introduction
Acute aortic dissection (AAD) is one of the most catastrophic cardiovascular diseases with high mortality1,2, while diagnosis and treatment of AAD have been greatly improved in recent years, including noninvasive imaging diagnosis3 and endovascular treatment from descending aorta to ascending aorta4,5,6. An early and correct diagnosis may warrant immediate initiation of effective therapy to potentially reduce the mortality rate7. Current diagnosis of AAD requires definitive imaging such as computed tomography, magnetic resonance imaging, or transesophageal echocardiography, but the use of each investigation is based on an index of clinical suspicion, and each incurs a further logistical delay in patient management8. Diagnosis is therefore not immediate, and the varied presentation allows the diagnosis to be missed, misdiagnosed, or overlooked in up to 39% of cases9, sometimes only being established at post-mortem10.
A biomarker has great potential to improve the early diagnosis of AAD11. There are a number of candidate biomarkers, including smooth muscle myosin heavy chain, calponin, soluble elastin fragments and D-dimer which were shown to be useful within a short time-window or without thrombosis of the case11,12,13,14. Another biomarker with higher sensitivity and specificity for diagnosis of AAD that can extend the time-window to later stages would thus be expected.
MicroRNAs are a set of small non-coding RNAs, with the length of 18 to 22 nucleotide15,16, several of which have been reported to play an important role in cardiovascular development and diseases17,18,19. We have demonstrated that miR-30 might play important roles in the pathogenesis of AAD by regulation of the mitogen-activated protein kinase pathway20. Recent studies have shown that the levels of some circulating microRNAs were associated with the diagnosis and prognosis for diseases21. Our previous study demonstrated that plasma miR-208a might be a novel biomarker for acute myocardial infarction22. In this study, we investigated the possibility of plasma microRNAs as biomarkers for AAD diagnosis.
Methods
Population
63 aortic dissection (AD) patients (37 AAD and 26 subacute aortic dissection [SAD]), and 23 non-aortic dissection (non-AD) patients admitted to the Department of Vascular Surgery in Shanghai Changhai Hospital between February 2011 and July 2012 were recruited in the study. The diagnosis of AD was based on computed tomography angiography with contrast confirming a dissected aorta containing both a true and false lumen. The non-AD patients include 11 patients with acute myocardial infarction (AMI), 10 patients with aortic aneurysm (AA) and 2 patients with pulmonary embolism (PE). In addition, 17 healthy volunteers (without cardiovascular disease history, 15 [88.2%] males, [55.0 ± 5.3] years) were enrolled in this study. The control groups included all participants without aortic dissection (n = 40). Among them, 14 participants have symptoms of chest pain (CP), including 11 AMI patients, 2 PE patients and 1 AA patient.
The protocol of this study was carried out according to the principles of the Declaration of Helsinki and approved by the Medical Ethics Committee in Shanghai Changhai Hospital. Written informed consent was obtained from all the participants before enrollment.
Plasma Collection and Storage
Blood samples for microRNAs detection were collected from patients in the emergency department or the vascular surgery laboratory and were processed within 1 hour of collection by two-step centrifugation. The supernatant was transferred to RNase/DNase-free tubes and stored at −80 °C.
RNA Isolation
Total RNA were isolated from plasma specimens using miRNeasy Mini Kit (Qiagen, Hilden, Germany) following the instructions from the manufacturer. The Caenorhabditis elegans microRNA (cel-miR-39) was synthesized for the spiked-in control. 0.4 pmol cel-miR-39 was supplemented into each 200 ul sample, which had been mixed with 1.0 ml QIAzol lysis Reagent (Qiagen, Hilden, Germany). The quality of total RNA samples was detected by NanoDrop ND-2000 Spectrophotometer (Thermo Scientific Wilmington, DE, USA). Only the samples with A260/280 > 2.0 and A260/230 > 1.8 were used for further analyses.
Microarray and qRT-PCR
Human MicroRNA Microarrays (Release 16.0, 8 × 60k) (Agilent Technologies, Santa Clara, CA) were used to identify candidate microRNAs. The Complete Labeling and Hyb Kit (Agilent Technologies, Santa Clara, CA, US) was used to label the miRANs samples, and hybridized with Cy3-tagged RNA. Then the quantization of the image information was performed with Feature Extraction Software 10.7. Quantile algorithm (Gene Spring Software 12.6) was applied to normalize the raw data. The samples with intra-array coefficients of variation (CV) above 15% would be discarded. The prediction analysis of microarrays (PAM) was used for microarray analysis. All microarray data has been submitted to GEO (GSE 92427).
For testing of candidate microRNAs acquired on microarray, quantitative reverse transcriptase-polymerase chain reaction (qRT-PCR) was performed. The RNA sample with high quality was used for reverse transcribed reaction which was carried out by the TaqMan miRNA Reverse Transcription Kit (Applied BioSystems, Foster City, CA). Then qRT-PCR reaction was performed using Taqman microRNA assays (Applied Biosystems, Foster City, CA) in a ABI PRISM 7900HT Sequence Detection System (Applied Biosystems, Foster City, Calif., U.S.A.). The total volume of the reaction was 10 μL, containing 5 μL of Universal Master Mix (Applied Biosystems, Foster City, Calif., U.S.A.), 2 μL cDNA template, and 3 μL. water. The primers used in the study were designed according to the sequences of the candidate miRNAs. The amplification condition was as follows: 95 °C template denaturation for 10 min, followed by 40 cycles at 95 °C for 15 seconds and 60 °C for 1 minute. U6 served as an internal control. Each reaction was repeated in triplicate. The data were analyzed with an automatic setting for assigning baseline; the threshold cycle was defined as the fractional cycle number at which the fluorescence exceeds the given threshold. The microRNAs which showed CT values above 40 cycles in greater than 20% of the 103 was considered not pass the quality control. The plasma levels of microRNAs were detected and analyzed by two investigators who were blinded to the clinical data of patients. The data obtained by real-time PCR were translated in log2 (relative level).
Statistical Analysis
The prediction analysis of microarrays (PAM) was used for microarray analysis. The quantitative data were evaluated whether they followed the normal distribution by the Shapiro-Wilk test. The basis for declaring a certain parameter as normally distribution was P = 0.20. For the data that did not fit the normal distribution, the Kruskal-Wallis test was performed, while Levene’s test of homogeneity of variance was used for the analysis of data with normal distribution,. When the data fit the homogeneity of variance, one-way analysis of variance was applied, and for the data that did not fit the homogeneity of variance, the Kruskal-Wallis test was performed. The qualitative data were compared with Fisher’s exact test. The predicted probability of being diagnosed with AAD was used as a surrogate marker to construct receiver operating characteristic (ROC) curves. The area under the ROC curve (AUC) was used as an accuracy index for evaluating the diagnostic performance of the selected microRNAs. All P-values were two-sided, and any value less than 0.05 was considered statistically significant.
Results
Characteristics of Study Population
The characteristics of the study subjects were presented in Table 1. There were no statistically significant differences between AAD patients and controls (healthy volunteers and non-AD patients) for any of the considered variables (P > 0.05 for all).
MicroRNAs Screening and Training
A microarray containing probes for 1,205 human microRNAs and 142 viral microRNAs was initially used to screen the significant differential expression levels of microRNAs between the AAD (n = 8) and control groups (n = 16, 8 aortic aneurysm patients and 8 healthy volunteers). As shown in Fig. 1, the results illustrated the hierarchical clustering of the differentially expressed microRNAs in the pairwise comparison of the AAD and healthy groups, and AAD and aortic aneurysm, respectively. There were seven microRNAs, including let-7b, miR-15a, miR-16, miR-23a, miR-451, miR-518e* and miR-663, with significantly higher expression levels in the AAD group than in the healthy group. In contrast, miR-150, miR-192, miR-1182, miR-3131, miR-3162, miR-3196, miR-3682 and hcmv-miR-US33-5p exhibited significantly lower expression levels in the AAD group than that in the healthy group. When compared with patients with aortic aneurysm, patients with AAD showed up-regulated expression of miR-23a and miR-223. In summary, 16 abnormally expressed microRNAs were identified as candidates for further testing via qRT-PCR.
Process for Selection of Candidate MicroRNAs. Microarrays with 1,205 human microRNAs and 142 viral microRNAs were used for screening candidate diagnostic markers in the 3 categories of subjects from 24 plasma samples including AAD, healthy and AA subjects. There were two microRNAs overlapping in the 3 group comparisons. Finally, 16 candidate microRNAs discovered via microarrays were selected for the further validation. AAD, acute aortic dissection; AA, aortic aneurysm.The red flag indicates higher expression, and the green flag indicates lower expression.
The 16 candidate microRNAs were tested using the entire sample set (63 AD and 40 without AD) with qRT-PCR, and 8 of the 16 microRNAs passed quality control. Among the miRNAs, miR-15a had a significantly different pattern between the AAD and control groups (P = 0.008). There were four up-regulated microRNAs, including let-7b, miR-15a, miR-23a and US33-5p, in the AAD group compared with CP group (including 11 AMI, 2 PE and 1 AA patients; P = 0.000, 0.000, 0.026 and 0.011, respectively).
MicroRNA Levels in AAD Patients
The different expression levels of validated plasma microRNAs between AD and control groups were shown in Fig. 2. The elevated level of miR-15a was observed in patients with AAD compared to the control groups (P = 0.008, fold change = 2.795). Furthermore, plasma level of miR-15a in AD patients was significantly higher than that in control groups (P = 0.016, fold change = 3.207) (Table 2). ROC analysis yielded an optimal cutoff value for miR-15a with a sensitivity of 75.7% and a specificity of 82.5% for detection of AAD; and with a sensitivity of 66.7% and specificity of 82.5% for AD detection (including AAD and SAD). The diagnostic accuracy of miR-15a measured by AUC, was 0.761 and 0.724, respectively (Fig. 3A,B; Table 2).
Plasma MicroRNAs Different Expression Level Between AD and Control Groups. The different expression levels of validated plasma microRNAs (including let-7b (A), miR-15a (B), miR-23a (C), hcmv-miR-US33-5p (D) between AD and control groups. AAD, acute aortic dissection; SAD, subacute aortic dissection; AA, aortic aneurysm; AMI, acute myocardial infarction; PE, pulmonary embolism.
Receiver operating characteristic (ROC) curve analysis for acute aortic dissection diagnosis. Area under the curve (AUC) estimation for miR-15a in (A) acute aortic dissection (AAD) and control groups, (B) aortic dissection (including AAD and SAD) and control groups; D-dimer in (C) AAD and control groups. Calculation of optimal cutoff value between AAD and chest pain groups by ROC analysis with respect to let-7b (D), miR-15a (E), miR-23a (F), US33-5p (G) and D-dimer (H), respectively. Control groups (n = 40) includes 17 healthy participants, 11 AMI patients, 10 AA patients and 2 PE patients. Chest pain groups (n = 14) includes 11 AMI patients, 2 PE patients and 1 AA patients.
Based on clinical requirements, we further explored whether the plasma microRNA could be employed as biomarkers to distinguish between AAD and CP. When compared with CP groups, patients with AAD had significantly higher expression of let-7b, miR-15a, miR-23a and hcmv-miR-US33-5p (P = 0.000, 0.000, 0.026 and 0.011, respectively; fold change = 8.462, 8.938, 17.851 and 21.975, respectively; Table 2). ROC analysis for these four microRNAs exhibited sensitivity of 79.4%, 75.7%, 91.9% and 73.5%, respectively. And specificity of them were 92.9%, 100%, 85.7% and 85.7%, respectively. The corresponding AUCs were 0.887, 0.855, 0.925 and 0.815, respectively (Fig. 3D–G; Table 2). The ROC analysis exhibited microRNAs more specific than D-dimer for AAD detection in this study (Table 2; Fig. 3).
MicroRNA Expression Profile and the Type of AAD
Plasma let-7b, miR-15a, miR-23a and hcmv-miR-US33-5p levels in the enrolled patients were compared according to type of dissection (Stanford type A, n = 15 and type B, n = 48). Among the patients with AAD, the hcmv-miR-US33-5p level was significantly higher in patients with type A AAD than in those with type B (P = 0.048). The levels of miR-15a and miR-23a tended to be higher in patients with type A AAD than in those with type B (P = 0.142 and 0.071, respectively). The let-7b expression level did not exhibit any significant differences between the two types (P = 0.531) (Fig. 4).
MicroRNA Expression Profile and the Type of AAD. Among the patients with AAD (Stanford type A, n = 15 and type B, n = 48), the let-7b (A) expression level did not exhibit any significant differences between the two types (P = 0.531). The levels of miR-15a (B) and miR-23a (C) tended to be higher in patients with type A AAD than in those with type B (P = 0.142 and 0.071, respectively). The hcmv-miR-US33-5p (D) level was significantly higher in patients with type A AAD than in those with type B (P = 0.048).
MicroRNA Expression Profile and the Time from Onset
Further analysis of plasma let-7b, miR-15a, miR-23a and hcmv-miR-US33-5p levels in the enrolled patients according to time from onset (0~6 h, ~12 h, ~24 h, ~3d, ~7d, ~14d and subacute-phase) was shown in Fig. 5. Blood samples were collected from patients with AD on admission, and divided into 7 subgroups according to the time from onset of symptoms. By linear regression analysis, there was no correlation between plasma microRNA levels and the time from onset of symptoms (R 2 = 0.013, 0.005, 0.0007 and 0.003, respectively). This indicated that the detected microRNAs from patients with AAD were maintained at high levels accompanied by the persistence of a pathological state.
MicroRNA Expression Profile and Outcome
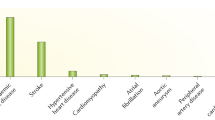
A total of four deaths (6.35%) occurred during the first 30 days. Major adverse events during postoperative course were documented in eleven patients (17.5%), including postoperative heart failure, severe bleeding complications, postoperative intervention and bacteremia. We compared the expression profiles of the microRNA in AAD patients according to their outcomes. Plasma miR-23a, miR-223 and hcmv-miR-US33-5p levels tended to be higher in patients with major adverse events than in those without major adverse events during the in-hospital period (P = 0.099, 0.114 and 0.192, respectively). However, the expression levels of let-7b and miR-15a did not show obvious association with major adverse events (P = 0.870 and 0.759, respectively) (Fig. 6).
Serial plasma miR-15a measurements during hospitalization of seven patients with AAD were presented in Fig. 7A. All of them underwent emergency endovascular repair. In the six patients without major adverse events, serial miR-15a measurements demonstrated a decline over time, while in one patient with occurrence of bacteremia a sharp increase was observed.
Serial Measurements of Plasma miR-15a. (A) Serial plasma miR-15a measurements during hospitalization of 7 AAD patients. In the 6 patients with no major adverse events, serial miR-15a measurements demonstrate a decline over time, while in one patient who occurrence of bacteremia (♂, 48 y, *), sharp increase was found. (B) Serial plasma miR-15a measurements during follow-up of 5 AAD patients. 2 of 5 patients with postoperative condition were stable. The plasma level of miR-15a was found to be lower than their respective plasma levels during the onset of AAD. **2 patients with recurrent AAD, their plasma miR-15a level were increased significantly. *One patient with occasional fever, his miR-15a level was sustained at a higher level and slightly higher than in-hospital.
A follow-up was performed to further determine whether plasma miR-15a in AAD patients presented any changes after endovascular repair (Fig. 7B). The blood was collected from five patients with AAD for serial miR-15a detection. Two of the five patients with postoperative conditions were stable. The plasma level of miR-15a was found to be lower than their respective plasma levels during the onset of AAD. Two patients with recurrent AAD experienced significantly increased levels of plasma miR-15a. One patient experienced occasional fevers, sustained a high level of plasma miR-15a which was slightly higher than in-hospital.
Characterization of Plasma microRNA Stability
The expression stability was an important prerequisite for plasma miRNAs serving as a biomarker. In the current, we investigated the stability of microRNAs in plasma. Incubating the plasma at room temperature for up to 24 hours (Fig. 8A,B) and subjecting it to up to four cycles of freeze-thawing (Fig. 8C,D) had minimal effect on miR-15a and miR-23a levels as measured by qRT-PCR.
We sought to determine if microRNA measurements were substantially different between plasma and serum, by measuring the miR-15a and miR-23a in matched samples of plasma and serum collected from a participant at the same blood draw. From the measurements obtained there was no discernible difference between either samples.
Discussion
At the present time, the diagnosis of AAD is based on clinical presentation and mainly relies on imaging techniques. However, the diagnostic performance of these modalities is unsatisfactory for the diagnosis of AAD. At this time, up to 39% of patients remain undiagnosed until necropsy9. A diagnostic biomarker for AAD will reduce delays in diagnosis and promote the early diagnosis rate, potentially improving the prognosis of patients. Until now, there are lack of a biomarker with high enough sensitivity and specificity, nor with a long enough time-window for positively identifying AAD. The recent discovery of aberrant expression of microRNAs in aortic tissue from AAD patients has paved the way for analyzing circulating microRNAs for the purpose of AAD diagnosis.
Blood-based biomarker test is convenient, low cost, fast and effective that may be a promising approach for routine screening and surveillance of AAD. In this study, we found that circulating microRNAs were significantly altered in AAD. In particular, our study revealed that patients with AAD had significantly higher expression levels of let-7b, miR-15a, miR-23a and hcmv-miR-US33-5p when compared with CP groups. Among them, miR-23a exhibited the highest diagnostic accuracy for AAD. When compared with control groups, patients with AAD had significantly higher expression levels of miR-15a, reveling its good diagnostic accuracy for detection of both AAD and SAD. Then, plasma miR-23a might have considerable clinical value for the initial diagnostic work-up of patients with CP and suspected AAD. Plasma miR-15a was important in monitoring the progression of and screening for AAD from the varied presentation, including asymptomatic patients.
Previous studies suggested a number of candidate biomarkers for AAD. Determination of smooth muscle myosin heavy chains and soluble elastin fragments showed high sensitivity and specificity in diagnosis of AAD, but these tests were not practical in a clinical emergency setting. Experience with calponin was limited, further investigation was required to improve sensitivity and specificity. D-dimer levels exhibited an excellent sensitivity for the detection of AAD, but had a lower specificity (46.6% to 68.6%)23,24,25,26. And subacute aortic dissection will be missed as a result of the short biological half-life of D-dimer26. Interestingly, circulating microRNAs showed higher diagnostic accuracy and a longer time-window for diagnosis of AAD in this study.
In previous studies, the control group forms of healthy volunteers and cases with similar symptoms of AAD, which was undoubtedly the primary factor to consider. However, there was a similar organizational structure in AA, along with similar risk factors and a series of hemodynamic changes, which might be applied as improved biomarkers in diagnosing aortic dissection. In this study, AA was used as a case control in the process of screening and validating differentially expressed circulating microRNAs in AAD. We believed that the results obtained in this way was more specific and reliable.
In earlier studies on cardiovascular diseases, microRNAs have also been proposed to possible be used for their diagnosis and prognosis as well. For example, miR-23a was found to be elevated in patients with coronary artery disease, which suggested that it might served as a biomarker for the disease development27; while miR-23 was reported to be enriched in the plasma of CAD cases, implying its potential function as a diagnostic tool28. All these evidences along with our findings supported the promising role of microRNAs in diagnosing cardiovascular diseases. In our study, when compared with CP groups (includes 11 AMI patients, 2 PE patients and 1 AA patients), patients with AAD had significantly higher expression of miR-23a (P = 0.026; fold change = 17.851). Discriminating between AAD and AMI provides important therapeutic implications. Recent reports have led us to identify a set of microRNAs that can be clinically practicable biomarkers for AMI diagnosis. Our present work also suggested that microRNAs in plasma might be sensitive and specific biomarkers for AAD. Circulating microRNAs may provide a helpful and reliable tool in the diagnosis of AAD or AMI.
Study limitations
The outcome of the patients in this study comes from in-hospital observations and short-term follow-ups, limiting our current ability for prognostic analysis. The data in our study from a single center study has lower dispersion. Additionally, the sample size is relatively small in the current study. Therefore, confirmation in multicenters studies with larger sample size is still necessary.
Conclusion
In summary, plasma miR-15a levels are significantly different between AAD and control groups (including healthy controls, AMI, AA and PE). And plasma miR-23a has a high sensitivity and specificity for differentiating AAD from CP groups. They have a long time-window and high degree of accuracy for detecting AAD. Our study demonstrates that plasma miR-23a exhibits considerable clinical value for the differential diagnosis of AAD. Plasma miR-15a can be used in monitoring and screening of AAD. The newly identified biomarkers may promote the early diagnosis rate of AAD, thus improving the prognosis of the patients. So that more patients with acute chest pain can benefit from the optimal therapy who would have otherwise missed the curative treatment window.
References
Hagan, P. G. et al. The International Registry of Acute Aortic Dissection (IRAD): new insights into an old disease. JAMA. 283, 897–903 (2000).
Pacini, D. et al. Acute aortic dissection: Epidemiology and outcomes. Int J Cardiol. 167, 2806–2812 (2013).
Nienaber, C. A. et al. The diagnosis of thoracic aortic dissection by noninvasive imaging procedures. N Engl J Med. 328, 1–9 (1993).
Nienaber, C. A. et al. Nonsurgical reconstruction of thoracic aortic dissection by stent-graft placement. N Engl J Med. 340, 1539–1545 (1999).
Dake, M. D. et al. Endovascular stent-graft placement for the treatment of acute aortic dissection. N Engl J Med. 340, 1546–1552 (1999).
Lu, Q. et al. Endovascular Repair of Ascending Aortic Dissection: A Novel Treatment Option for Patients Judged Unfit for Direct Surgical Repair. J Am Coll Cardiol. 61, 1917–1924 (2013).
Golledge, J. & Eagle, K. A. Acute aortic dissection. Lancet. 372, 55–66 (2008).
Ranasinghe, A. M. & Bonser, R. S. Biomarkers in Acute Aortic Dissection and Other Aortic Syndromes. J Am Coll Cardiol. 56, 1535–1541 (2010).
Olin, J. W. Acute Aortic Dissection: The Need for Rapid, Accurate, and Readily Available Diagnostic Strategies. Arterioscler Thromb Vasc Biol. 23, 1721–1723 (2003).
Khan, I. A. & Nair, C. K. Clinical, diagnostic, and management perspectives of aortic dissection. Chest. 122, 311–328 (2002).
Suzuki, T. et al. Preliminary experience with the smooth muscle troponin-like protein, calponin, as a novel biomarker for diagnosing acute aortic dissection. Eur Heart J. 29, 1439–1445 (2008).
Katoh, H. et al. Diagnosis of aortic dissection by immunoassay for circulating smooth muscle myosin. Lancet. 345, 191–192 (1995).
Suzuki, T. et al. Diagnostic implications of elevated levels of smooth-muscle myosin heavy-chain protein in acute aortic dissection. The smooth muscle myosin heavy chain study. Ann Intern Med. 133, 537–541 (2000).
Shinohara, T. et al. Soluble elastin fragments in serum are elevated in acute aortic dissection. Arterioscler Thromb Vasc Biol. 23, 1839–1844 (2003).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 116, 281–297 (2004).
Lee, R. C., Feinbaum, R. L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 75, 843–854 (1993).
Urbich, C., Kuehbacher, A. & Dimmeler, S. Role of microRNAs in vascular diseases, inflammation, and angiogenesis. Cardiovasc Res. 79, 581–588 (2008).
Zou, J. et al. Two functional microRNA-126s repress a novel target gene p21-activated kinase 1 to regulate vascular integrity in zebrafish. Circ Res. 108, 201–209 (2011).
Li, Q. et al. Attenuation of microRNA-1 derepresses the cytoskeleton regulatory protein twinfilin-1 to provoke cardiac hypertrophy. J Cell Sci. 123, 2444–2452 (2010).
Liao, M. et al. A microRNA profile comparison between thoracic aortic dissection and normal thoracic aorta indicates the potential role of microRNAs in contributing to thoracic aortic dissection pathogenesis. J Vasc Surg. 53, 1341–1349 e3 (2011).
Reid, G., Kirschner, M. B. & van Zandwijk, N. Circulating microRNAs: Association with disease and potential use as biomarkers. Crit Rev Oncol Hematol. 80, 193–208 (2011).
Wang, G. K. et al. Circulating microRNA: a novel potential biomarker for early diagnosis of acute myocardial infarction in humans. Eur Heart J. 31, 659–66 (2010).
Weber, T. D-dimer in Acute Aortic Dissection. Chest. 123, 1375–1378 (2003).
Eggebrecht, H. et al. Value of plasma fibrin D-dimers for detection of acute aortic dissection. J Am Coll Cardiol. 44, 804–809 (2004).
Suzuki, T. et al. Diagnosis of Acute Aortic Dissection by D-Dimer: The International Registry of Acute Aortic Dissection Substudy on Biomarkers (IRAD-Bio) Experience. Circulation. 119, 2702–2707 (2009).
Dong, J. et al. Diagnostic implication of fibrin degradation products and D-dimer in aortic dissection. Sci. Rep. 7, 43957, https://doi.org/10.1038/srep43957 (2017).
Han, H. et al. MiR-34a, miR-21 and miR-23a as potential biomarkers for coronary artery disease: a pilot microarray study and confirmation in a 32 patient cohort. Exp Mol Med. 47, e138 (2015).
Di, Y. et al. miR-23 regulate the pathogenesis of patients with coronary artery disease. Int J Cardiol. 8, 11759–11769 (2015).
Acknowledgements
We thank all members of the laboratory for their helpful discussions and comments during the study. We also thank our clinical colleagues, as well as Yufeng Zhuang, Jiannan Wang, Chenqi Xing, Min Wang and Rong Li for their technical assistance. This work was supported by the National Natural Science Foundation of China [81370441, 81700434].
Author information
Authors and Affiliations
Contributions
Conception and design: Jian Dong, Junmin Bao, Jian Zhou, Zaiping Jing; Analysis and interpretation: Jian Dong, Rui Feng, Jian Zhou, Zaiping Jing; Operation participation and Data collection: Jian Dong, Jian Zhou, Rui Feng, Qingsheng Lu, Guokun Wang, Zhiqing Zhao, Junmin Bao, Haiyan Li, Dingfeng Su, Qing Jing, Zaiping Jing; Writing the article: Jian Dong, Jian Zhou, Zaiping Jing, Rui Feng; Critical revision of the article: Jian Dong, Qingsheng Lu, Guokun Wang, Qing Jing, Zaiping Jing; Final approval of the article: Jian Dong, Jian Zhou, Junmin Bao, Qingsheng Lu, Guokun Wang, Zhiqing Zhao, Haiyan Li, Dingfeng Su, Qing Jing, Zaiping Jing; Statistical analysis: Jian Dong; Zhiqing Zhao; Rui Feng. Obtained funding: Jian Dong, Junmin Bao.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dong, J., Bao, J., Feng, R. et al. Circulating microRNAs: a novel potential biomarker for diagnosing acute aortic dissection. Sci Rep 7, 12784 (2017). https://doi.org/10.1038/s41598-017-13104-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-13104-w
This article is cited by
-
Diagnostic biomarkers and aortic dissection: a systematic review and meta-analysis
BMC Cardiovascular Disorders (2023)
-
HCMV-miR-US33-5p promotes apoptosis of aortic vascular smooth muscle cells by targeting EPAS1/SLC3A2 pathway
Cellular & Molecular Biology Letters (2022)
-
Altered human cytomegalovirus-encoded miRNAs in host circulation: novel disease biomarkers and potential aetiological agents
ExRNA (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.