Abstract
Next Generation Sequencing (NGS) strategies, like RNA-Seq, have revealed the transcription of a wide variety of long non-coding RNAs (lncRNAs) in the genomes of several organisms. In the present work we assessed the lncRNAs complement of Schistosoma mansoni, the blood fluke that causes schistosomiasis, ranked among the most prevalent parasitic diseases worldwide. We focused on the long intergenic/intervening ncRNAs (lincRNAs), hidden within the large amount of information obtained through RNA-Seq in S. mansoni (88 libraries). Our computational pipeline identified 7029 canonically-spliced putative lincRNA genes on 2596 genomic loci (at an average 2.7 isoforms per lincRNA locus), as well as 402 spliced lncRNAs that are antisense to protein-coding (PC) genes. Hundreds of lincRNAs showed traits for being functional, such as the presence of epigenetic marks at their transcription start sites, evolutionary conservation among other schistosome species and differential expression across five different life-cycle stages of the parasite. Real-time qPCR has confirmed the differential life-cycle stage expression of a set of selected lincRNAs. We have built PC gene and lincRNA co-expression networks, unraveling key biological processes where lincRNAs might be involved during parasite development. This is the first report of a large-scale identification and structural annotation of lncRNAs in the S. mansoni genome.
Similar content being viewed by others
Introduction
Over the past ten years, the advent of Next Generation Sequencing (NGS) strategies like RNA-Seq have revealed the transcription of a wide variety of non-protein-coding RNAs (ncRNAs) in the genomes of eukaryote organisms. Early on this age, it had become quite evident that life in the eukaryotic cells is orchestrated by complex regulatory networks in which ncRNA molecules appear to be the most numerous and main signaling effectors on many vital reactions1. Within this large class of ncRNA genes, long non-coding RNAs (lncRNAs) have been conventionally defined as being over 200 nt-long transcripts that have no protein-coding (PC) potential2. LncRNAs seem to display a number of regulatory functions and many of them now have a well-characterized role as epigenetic modulators3,4,5.
There are several subtypes of lncRNAs, which are classified by their architecture and/or location onto the genome rather than by their function. The following are the terminologies established for the already known lncRNAs’ subtypes: (i) antisense lncRNAs, which overlap the exons of PC genes on the opposite strand; (ii) sense intronic lncRNAs, which reside exclusively into PC gene introns; (iii) large intergenic/intervening non-coding RNAs (lincRNAs), which do not overlap PC genes and may act either in cis, like enhancer RNAs (eRNAs), or in trans, by binding to chromatin modifier protein complexes2, 3, 6, 7.
Despite sharing similar general traits with their PC gene counterparts such as being transcribed by RNA polymerase II, following the same chromatin state rules, often 5′-capped, spliced and polyadenylated2, it is known in vertebrates that lncRNAs exhibit several peculiar and distinct features when compared with mRNAs, such as: (i) rapid evolution across species from the same clade8, 9; (ii) both low and transient expression, being most of the times specimen-, tissue-, and/or cell-specific6, 7, 10,11,12; (iii) reduced splicing efficiency10; (iv) high propensity to cis-regulatory action immediately following or during their own transcription2, 13; (v) participation in trans-acting mechanisms when associated to polycomb repressive complex 2 (PRC2), for instance3, 14, 15; (vi) formation of RNA-DNA triplex structures that may mediate both cis- and trans-actions14, 16,17,18; (vii) more enriched in the nucleus than in the cytoplasm6.
Schistosoma mansoni was chosen by us as a target organism to unravel the class of lincRNA genes because of both the lack of lncRNAs annotation on its genome and its global health importance as a neglected tropical disease-causing agent. It is a flatworm endemic in both Africa and South America continents that causes an infectious and parasitic disease known as schistosomiasis that affects, together with other schistosome species, over 250 million people worldwide19. Besides its public health impact, this parasite is also an interesting model for studies on the Molecular Parasitology field because of its complex life cycle and, consequently, the drastic gene expression changes it faces during development. From the nearly 12,000 PC genes already mapped onto Schistosoma spp. genomes20,21,22, there are about 2,000 stage-specific differentially expressed (DE) ones among S. mansoni cercariae (the human infective larval stage), schistosomula (the first stage in a mammal host) and mature mixed adults residing in the mesenteric veins21, as well as between separate male and female adult forms of S. haematobium and their respective eggs22.
Up to date only one single study has been devoted to a large-scale mapping of ncRNA genes in schistosomes23. However, the authors have only adopted homology-based annotation through the use of bioinformatics tools in order to screen for structured RNAs on a comparison between S. mansoni and S. japonicum genomes23. No lncRNAs were reported on that investigation. On the other hand, our group has already pointed to the existence of lncRNAs in the S. mansoni transcriptome, by using both microarray and low-coverage RNA-Seq approaches24, 25.
Recently, several new bioinformatics tools have been developed with the aim of capturing novel lncRNAs mapped to the genomes26,27,28,29. However, they are mostly geared towards applying the machine learning techniques to the well-curated human and/or mouse transcriptome mapping annotations. For invertebrates, Caenorhabditis elegans (a nematode free-living worm) and Plasmodium falciparum (a protozoan parasite) have already had their transcriptomes screened for lncRNAs by independent groups who established their own ad-hoc pipelines for analyzing RNA-Seq data30, 31. They have unraveled ~1,000 transcripts among antisense lncRNAs and lincRNAs for each of the above two organisms.
In the present work, we thoroughly screened the S. mansoni genome and transcriptome and we present a novel repertoire of spliced lncRNA genes, with a main focus on the lincRNAs subtype. We first describe our computational pipeline aimed at retrieving lncRNAs from dozens of raw large-scale RNA-Seq samples (either public data available at the NCBI-SRA repository or data generated by us). Subsequently, we report their location and architecture onto the genome, the evidence for being transcribed, and their evolutionary conservation among other species. Finally, we analyze the lincRNAs putative functionality by building co-expression networks that associate them to their PC gene counterparts that are co-expressed across five different parasite developmental stages. This is the first study that provides a holistic systems overview that merges S. mansoni lincRNAs content with what is encoded into proteins and raises hypotheses about the lincRNAs functionality through the construction of co-expression networks.
Results
A computational pipeline is able to retrieve thousands of lncRNAs from S. mansoni RNA-Seq samples
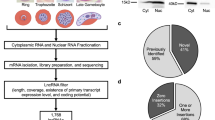
Our ad-hoc computational pipeline (Fig. 1) was designed based on two distinct transcriptome assembly approaches: (i) a de novo method using Trinity32 (Fig. 1, path A), and (ii) a reference genome/annotation-based method, using Tophat2 (ref. 33), Cufflinks and Cuffmerge34 (the Tuxedo suite) (Fig. 1, path B). As suggested by Ulitsky9, the use of different assembly algorithms increases the chance of detecting the lncRNAs complement of a cell or organism (more details on the Methods section). Table 1 shows the number of transcripts obtained with each assembly algorithm.
Computational workflow for rescuing and identifying novel putative lncRNAs from S. mansoni high-throughput transcriptomic data. Path A depicts the use of a de novo assembly method, whereas path B comprises reference-guided assembly. The red section is built in one single PERL script for the automation of tasks. On the bedtools intersect step (first red rectangle), one can choose whether the pipeline retrieves antisense lncRNAs or lincRNAs.
We have established a series of more stringent filtering steps in our pipeline (red border rectangles on Fig. 1) compared with the previously described ones for other invertebrates31, and we used a greater number of RNA-Seq samples as input (n = 88, see Methods). The first filtering step was the removal of PC-overlapping transcripts when rescuing lincRNAs, or the removal of both intervening and sense PC-overlapping transcripts when rescuing the antisense lncRNAs. As an example, Table 1 shows the number of lincRNA transcripts that remain at this step and at each of the further ones in our pipeline. The subsequent steps included the removal of repeats (low complexity and/or transposable elements), rRNAs, monoexonic transcripts, and spliced transcripts presenting all introns with less than 30 nt each and/or not showing canonical splice donor/acceptor (GT/AG) sites. Further, we removed transcripts presenting ORFs greater than 25 amino acids (aa) and covering 25% or more of the transcript lengths, as well as transcripts assigned as having coding potential probability by either CPC35 or TransDecoder36 algorithms, and, finally, transcripts containing Pfam and/or PANTHER domains as detected by InterProScan37 (see Methods).
At the end of the filtering pipeline we obtained two datasets: one with 7029 lincRNA isoforms (Table 1) located on 2596 genomic loci (at an average of 2.7 lincRNA isoforms per each lincRNA genomic locus), and another dataset with 402 lncRNAs that are antisense to 268 PC genes. The transcript sequences are on the Supplementary File S1. On Supplementary Table S1 we provide the genomic location and architecture of each lncRNA for the two datasets. We decided to focus our analyses on the lincRNAs gene set due to the greater amount retrieved, and also because the lincRNAs arise from PC-independent novel gene loci. For a better visualization of the lincRNA loci and architecture, we have created a track for them on the S. mansoni UCSC-like genome browser (http://schistosoma.usp.br), which we recently implemented and reported24. On Fig. 2 we selected three lincRNA loci to be viewed as genome browser pictures, where one can see different tracks on each of three different loci, showing evidence for transcription and evolutionary conservation of these novel lncRNA genes. Such evidence will be described further on this section.
Novel S. mansoni spliced lncRNA gene loci. All lncRNA isoforms depicted on the figure display both evolutionary conservation (phastCons score) among other schistosome species (S. haematobium and S. japonicum) and epigenetic marks (H3K4me3) that correspond to transcription initiation. (A) LncRNA antisense to a 78 kDa centrosomal protein gene (Smp_140230). The PC genes’ “tail to tail” orientation indicates that the H3K4me3 marks are exclusive to the lncRNA gene. (B) LincRNA locus upstream of a G protein coupled receptor gene (Smp_194740) of which only the last exon is shown. (C) LincRNA loci upstream of a cathepsin L proteinase gene (Smp_187140). (D) LincRNA loci downstream a hypothetical protein gene (Smp_194040). Both protein-coding and lncRNA gene IDs are searchable on the S. mansoni UCSC-like genome browser which our group has deployed (http://schistosoma.usp.br) and previously reported24.
Because there is no gold standard in silico protocol for detecting lincRNAs in ancient parasite genomes/transcriptomes, we ran a sensitivity test of our pipeline, namely trying to reproduce the list of previously identified lincRNAs from Plasmodium falciparum 30, and we successfully recovered 53% of the P. falciparum lincRNAs. We believe the 53% sensitivity rate reflects the more stringent filtering steps that we have deployed on our pipeline compared with the ones from Broadbent et al.30. We expect that low levels of false positive lncRNAs detection may be achieved in our S. mansoni data analysis due to our stringency choices.
A great variety of lincRNA transcript isoforms was detected
In order to check whether different assembly tools and datasets used as input in our pipeline (see Table 1) yielded similar transcripts, and to calculate the degree of redundant transcripts obtained, we performed a blastn38 alignment of the total 7029 lincRNA sequences against themselves. This all against all comparison has revealed only 15 identical pairwise alignments (100% identity and 100% coverage of both query and subject sequences) encompassing 21 lincRNAs (0.3% redundancy) (Supplementary Table S2). Excluding the requirement of complete identity and coverage, and computing the pairs in which one transcript is completely contained within a longer transcript isoform (i.e. lincRNA queries completely covered by the subject lincRNA with 100% identity and with partial subject coverage) the number of pairs was increased to 281 (4%). This reflects the presence of quite similar isoforms in our assembled transcripts dataset.
Further agreeing with the existence of multiple isoforms of each lincRNA per locus, we found that, keeping the 100% identity threshold but decreasing the requirement of the extent of query coverage (qcov), the all against all comparison has identified approximately similar numbers of lincRNA queries being aligned to the subject at the different qcov ranges: 1422 (90% ≤ qcov < 100%), 1310 (80% ≤ qcov < 90%), 1304 (70% ≤ qcov < 80%), 1327 (60% ≤ qcov < 70%), 1221 (50% ≤ qcov < 60%) (Supplementary Table S2).
In summary, sorting all these lincRNA query lists and eliminating the redundancy, we obtained 4212 (60%) lincRNAs that share 100% identity on at least half of their length with other lincRNAs in the dataset. This result indicates that different transcriptome assembly approaches plus different samples used as input lead to the detection of distinct lincRNAs transcript isoforms. These isoforms mapped to 2596 unique genomic loci, as seen above. Finding different lncRNA sets when using different RNA-Seq datasets and assembly tools was already reported when analyzing sets of different individuals, tissues and even at the single-cell level8,9,10, 12. We therefore decided to keep in our downstream analyses all the 7029 S. mansoni putative lincRNAs (hereafter called SmLINCs) as bona fide representatives of the complement of expressed S. mansoni lincRNAs.
We analyzed the density of lincRNAs mapped to the long, assembled chromosomes and found that the average density was 1.86 lincRNAs per 100 kb of genomic sequence (range 1.31 to 2.30) (Table S3), with no higher density in any particular chromosome. A linear correlation between chromosome size and number of lincRNAs per chromosome was obtained (coefficient of determination R2 = 0.97) (Table S3). A higher density of lncRNAs in certain chromosomes was found in vertebrates such as in the mouse, where there is a higher density of lncRNAs in chromosomes 2, 4 and 11 (ref. 39); this pattern was not observed in S. mansoni.
S. mansoni putative lincRNAs (SmLINCs) display evidence of being functional genes
We have selected four features to scrutinize our S. mansoni putative lincRNAs dataset in the search for evidence of their possible functional role: (i) presence of histone H3 lysine 4 trimethylation (H3K4me3) on the transcription start site (TSS) as an epigenetic mark for transcriptional activation; (ii) evolutionary conservation among two other schistosomes; (iii) differential expression across five parasite developmental stages; and (iv) positive or negative expression correlation to their PC gene neighbors.
H3K4me3 is a very well studied epigenetic mark present on the promoter region of actively transcribed genes40. Moreover, it is already known that such mark is not exclusive to PC genes, being also present on active lncRNA loci2. Therefore, we screened the lincRNAs’ TSS surrounding regions for the presence of significant H3K4me3 peaks in the ChIP-Seq data for both somula and adult forms of the parasite (publicly available data on SRA-NCBI41, 42). Forty percent of SmLINC (2878/7029) showed the H3K4me3 mark on their TSS surrounding regions. In addition, the pattern of distance between the lincRNAs TSS and the H3K4me3 closest peak (Fig. 3A) was quite similar to the pattern observed for PC genes (Fig. 3B).
S. mansoni lincRNAs display four different traits that may characterize them as functional genes. (A,B) transcriptional activation epigenetic mark (H3K4me3) surrounding the TSS on both lincRNAs (red) and PC genes (blue). (C) An empirical cumulative distribution function (ECDF) showed that lincRNAs’ (red) evolutionary conservation scores (phastCons) among three schistosome species are significantly different from genomic deserts (green) (arrow: one-sided KS-test p-value = 4.25e-06) and from PC genes (blue). (D) As a control, lincRNAs from human chromosome 1 (red) were compared with genomics deserts (green) and with PC genes (blue) from the same chromosome, and they display a phastCons score (comparison among 20 mammals provided by UCSC Table Browser for human genome) ECDF pattern similar to the one observed for S. mansoni (arrow: one-sided KS-test p-value = 2.67e-07). (E,F) Heatmap of 2450 PC genes (E) and 916 lincRNA transcripts (F) that were detected as differentially expressed with one-way ANOVA-like analysis comparing RNA-seq samples from somula 3 h (3S), 24 h (24S) and adults (male and female) against cercariae (the average expression of biological triplicates for each life cycle stage was used, and an adjusted p-value threshold of 0.01 was employed). For each gene (lines), the expression log-ratio between the indicated life cycle stage and cercariae was obtained (columns), and it was colored according to the scale indicated at the bottom of the panel F. (G–I) LincRNAs tend to be co-expressed with their PC gene neighbors (A curves, red) rather than with a set of 1000 randomly picked PC genes (R curves, cyan) during parasite development (same five life cycle stages assessed on E and F panels). Pearson correlation values’ ECDF for 1,572 expressed lincRNAs (TPM ≥ 1) that showed either r > 0.5 or r < −0.5 with their actual PC gene neighbors (G), for 893 differentially expressed lincRNAs (TPM ≥ 1) correlated with their actual PC gene neighbors (H) and for 402 DE lincRNAs (TPM ≥ 1) that also have H3K4me3 marks at their TSSs and are correlated with their actual PC gene neighbors (I) (arrows: KS-test p-value < 1e-12 for each of the three distributions).
Regarding the evolutionary conservation, we computed the phastCons scores43 assigned at the single-base level, after whole genomes multiple alignments had been obtained with the MULTIZ-TBA package of tools44. As already shown for the human genome45, phastCons scores are able to distinguish between features of high selective constraint (PC genes) and the ones of weak conservation (lncRNAs, ancestral repeats and desert regions), and even within the latter group it is still possible to differentiate genomic elements by their phastCons profile. After aligning three schistosomes’ genomes (S. mansoni, S. haematobium and S. japonicum), we obtained phastCons scores for about 56.9 Mb of genome sequence (see Methods), on which 3453 SmLINC were mapped and presented scored/conserved bases on at least one of their exons.
Table 2 presents an overview of three different S. mansoni genomic features (PC genes, lincRNAs and desert regions with neither annotation nor transcript evidence) and their phastCons evolutionary conservation scores’ content. As one can see by this rough summary, lincRNAs are more similar to desert regions than to PC genes, corroborating what was already seen in both human and mouse studies6, 46. By performing an empirical cumulative distribution function (ECDF) analysis of normalized phastCons scores per block together with a Kolmogorov-Smirnov (KS) test, we were able to distinguish a significantly different evolutionary conservation between SmLINC and desert regions (800 bp) in the S. mansoni genome, being the former higher than the latter (KS one-sided p-value = 4.2e-06, Fig. 3C). As a comparison, we calculated the phastCons scores for human chr1 annotated lincRNAs compared with desert regions (400 bp) on the same chromosome, using the 20 mammals phastCons scored bases provided by the UCSC genome browser (https://genome.ucsc.edu) for the human hg38 genome sequence assembly. Similar ECDF profiles were obtained (Fig. 3D, KS one-sided p-value = 2.6e-07). These results are indicative that almost half of the SmLINC reported herein (3453/7029) are under some sort of selective evolutionary pressure in the Schistosoma genus.
Differential expression along parasite development is another important trait that may characterize lincRNAs as transcripts that are required during the life cycle. We performed a one-way ANOVA-like analysis comparing somula (both 3 h and 24 h) and adult worms (male and female) against cercariae (see Methods for details). First, we identified 2450 differentially expressed (DE) PC genes (Smps) (Fig. 3E), a number close to what was identified by others comparing several developmental stages on both S. mansoni 21 and S. haematobium 22. Next, 916 SmLINCs were identified as DE on at least one developmental stage (p-value < 0.01) (Fig. 3F). The complete list of DE genes and their respective log2(fold-change), p-value and adjusted p-value are provided as a spreadsheet on Supplementary Table S4. We observed a lower frequency of DE lincRNAs (916/7029 = 13%) compared with DE PC genes (2450/11, 844 = 20.7%). We believe that we might have missed some lowly expressed lincRNAs on our final DE list (having TPM < 1 on all analyzed life cycle stages), since lncRNAs have lower expression levels compared with PCs (Supplementary Fig. S1) and because we have established for the ANOVA-like DE analysis a cutoff TPM ≥ 1 on at least one life cycle stage (three replicates). Similar to what can be seen for DE PC genes (Fig. 3E), DE lincRNAs also display stage-specific up-regulation patterns (Fig. 3F): adult-specific up-regulated genes can be seen on the heatmap’s top half, some somula-specific up-regulated lincRNAs are depicted at the middle portion of the heatmap, and cercariae up-regulated lincRNAs (i.e., somula and adults down-regulated lincRNAs) are observed at the bottom half of the heatmap (Fig. 3F). This result suggests that the lincRNA genes described herein might be important for the parasite development.
The fourth and last evidence sought by us was whether there is an expression correlation between SmLINCs and their PC gene neighbors in the genome, in comparison with control pairs of the SmLINCs and randomly-selected PC genes. First, we picked four PC gene neighbors (the first two upstream and two downstream) for each SmLINC (when it was possible, see below) and computed the pairwise (SmLINC-PC gene) Pearson expression correlation coefficient (r) (using TPM expression values for each gene from the same five developmental stages described above for the DE one-way ANOVA-like analysis, Fig. 3E,F). Next, we compared the expression correlation coefficients (r) against the ones from a negative control consisting of 1,000 times bootstrapped random PC genes paired to the same SmLINC set.
We started the analysis with 2039 expressed lincRNAs that had TPM ≥ 1 (on all three replicates of at least one life cycle stage), yielding 7269 SmLINC-neighbor_PC gene pairs (some lincRNAs may not have four PC gene neighbors because they can either be located on a scaffold bearing quite few (or no) PC genes or be at the extremities of assembled chromosomes/scaffolds). After comparing the ECDF profiles of all r-values computed for the actual 7269 SmLINC-neighbor_PC gene pairs against a negative control consisting of r-values for 7,269,000 pairs of SmLINC-randomly_selected_Smps, we observed a significant difference on the two distribution curves (KS one-sided p-value = 5.6e-25, Fig. 3G). It is noteworthy that 1572/2039 SmLINC-neighbor_PC gene pairs (77%) showed significant expression correlation (r) p-values < 0.05, whereas in the negative control only 2,547,966/7,269,000 SmLINC-randomly_selected_Smp pairs (35%) had r p-values < 0.05.
Next, we reduced the set of lincRNAs in the analysis by keeping only the 893 lincRNAs that are significantly differentially expressed (DE) across the stages within the set of 2039 lincRNAs with TPM ≥ 1. For these 893 DE lincRNAs there were 3167 SmLINC-neighbor_ PC gene pairs, which were compared with 3,167,000 SmLINC-randomly_selected_Smp bootstrapped pairs. The difference between their ECDF curves was also significant (KS one-sided p-value = 6.8e-20, Fig. 3H) and notoriously greater than the difference in the previous ECDF plot (Fig. 3G). Seven hundred thirty two out of the 893 SmLINC-neighbor_PC gene pairs (81.9%) showed significant expression correlation (r) p-values (<0.05), whereas 1,206,153/3,167,000 SmLINC-randomly_selected_Smp pairs (38%) had the same correlation significance.
Finally, filtering the 2039 SmLINCs by both being DE and having the H3K4me3 mark at their TSS-surrounding genomic region, we obtained 402 lincRNAs and 1406 SmLINC-neighbor_PC gene pairs. Their ECDF plot showed the greatest difference between the curves (KS one-sided p-value = 1.5e-13, Fig. 3I); in this set, a total of 325/402 SmLINC-neighbor_PC gene pairs (80.8%) had expression correlation (r) p-values < 0.05, while this same feature was seen for only 532,925/1,406,000 random pairs (37.3%) in the negative control.
The above results indicate that it is possible that SmLINCs may play a cis-acting role regulating their PC gene neighbors, as already known for vertebrates2, 5. In addition, as we decreased the lincRNAs’ set by keeping the ones displaying more evidence for being active, we observed an increase in the difference between the correlation of actual neighbors versus random pairs (Fig. 3GHI), suggesting that a set of lincRNAs is more prone to show a co-expression pattern with their PC gene neighbors.
An intersection diagram displaying the number of S. mansoni lincRNAs having from one to four of the analyzed traits that may characterize them as functional genes was obtained for the 7029 lincRNAs (Fig. 4), showing that only 1739 lincRNAs (25%) could not be associated with these traits. We decided to focus our downstream analyses on the main intersection group of 181 lincRNA genes (Fig. 4). The genes that were displayed as genome browser images on Fig. 2 are part of that main intersection set. The complete list of the 181 robust SmLINCs is provided as supplementary file (Supplementary File S2). Each of them can be accessed through our online genome browser (http://schistosoma.usp.br).
Hundreds of SmLINCs share two or more traits that may characterize them as functional genes. The UpSet intersection diagram shows the number of lincRNAs that were detected as being differentially expressed across five life cycle stages (DE), as having their expression correlated with the expression of protein-coding gene neighbors (Neighbor), as having histone active transcription marks at their TSSs (H3K4me3) and as being conserved among Schistosoma spp. (PhastCons). The 181 lincRNAs that share all traits were selected for further investigation using systems approaches.
RT-qPCR assays confirm the SmLINCs’ differential expression across distinct developmental stages
Sixteen out of the 181 robust SmLINCs’ set from the UpSet intersection diagram on Fig. 4 were selected for individual assessment of their expression levels on cercariae (C), somula 3 h (3S) and somula 24 h (24S) after mechanical transformation, male and female stages/forms. First, in order to confirm that the RNA samples herein used for RT-qPCR contained previously known stage-specific/enriched transcripts, we measured the expression levels of six protein-coding genes as positive controls (Supplementary Fig. S2). In the RT-qPCR assays (Supplementary Fig. S2), all six selected control PC transcripts corroborated the up-regulation pattern at the life cycle stage that had already been reported in the literature: Smp_044250 (Metalloprotease), shown as up-regulated in cercariae compared with somula47; Smp_033040 (Lactate dehydrogenase), up-regulated in somula compared with cercariae47; Smp_126730-5HTR and Smp_145140-WNT5, assessed as up-regulated in adult male compared with female24, Smp_000390-Trematode Eggshell and Smp_000430-EggShell, up-regulated in adult female compared with male24.
We then performed RT-qPCR to assess the expression levels of sixteen selected lincRNAs across the five S. mansoni stages studied. The primers used are listed on Supplementary Table S5. It was possible to detect differential expression of lincRNAs on all the four life stages: two more highly expressed in cercariae (SmLINC02630-IBu and SmLINC05716-IBu), ten in somula 3 h (SmLINC00001-IBu, SmLINC00133-IBu, SmLINC00282-IBu, SmLINC01084-IBu, SmLINC01122-IBu, SmLINC03930-IBu, SmLINC04271-IBu, SmLINC04521-IBu, SmLINC05284-IBu and SmLINC05720-IBu), two more highly expressed in somula 24 h (SmLINC01431-IBu and SmLINC06024-IBu) and two in male adults (SmLINC02394-IBu and SmLINC06535-IBu) (Fig. 5). Half of the genes showed an expression pattern concordant to those measured by RNA-Seq analysis (SmLINC02630-IBu, SmLINC00133-IBu, SmLINC00282-IBu, SmLINC03930-IBu, SmLINC04271-IBu, SmLINC05284-IBu, SmLINC05720-IBu and SmLINC02394-IBu). It is noteworthy that some SmLINCs from the other half, for which the qPCR did not corroborate the RNA-Seq stage of higher expression, had very low RNA-Seq read counts on all stages (SmLINC01084-IBu, SmLINC01122-IBu and SmLINC01431), which in turn may diminish the detection of differential expression of some lincRNAs by the high-throughput approach.
Confirmation by RT-qPCR of the differential expression of selected SmLincRNAs across the parasite life cycle stages. Sixteen SmLincRNAs were selected for validation by RT-qPCR at the parasite stages, namely cercariae (C), schistosomula after 3 hours of mechanical transformation (3S), schistosomula after 24 hours of mechanical transformation (24S), adult male and female (x axis). All sixteen lincRNA loci are among the 181 lincRNAs loci shown at the main intersection in Fig. 4. For each lincRNA plot, the individual sample with the lowest normalized expression value across all stages was chosen and arbitrarily set to 1. The expression values of the same lincRNA for all the other samples are represented as the relative expression compared with the lowest one (y axis). Bars represent standard deviation of the mean from four biological replicates for each stage. Three technical replicates were performed for each of the four biological replicates per stage. The Smp_092920 was used for internal normalization as the reference gene among the parasite stages (see Methods). The ANOVA Tukey test was used to calculate the statistical significance of the expression differences among the parasite forms (*p-value ≤ 0.05; **p-value ≤ 0.01; ***p-value ≤ 0.001; ****p-value ≤ 0.0001). For clarity purposes, it is shown only the highest p-value representation for the stage in which the lincRNA was detected with the highest expression.
Co-expression networks (lincRNAs-PC gene) raise hypotheses about lincRNAs’ functionality
A very often-adopted approach to hypothesize about lncRNAs’ function is the investigation of their co-expression patterns along with PC genes12, 45, 48, 49. We analyzed expression data for five different developmental stages of S. mansoni (15 RNA-Seq samples) in order to identify DE genes (previous topic), and we have selected a robust list of 181 SmLINCs that have strong evidence for being regulated genes. We therefore decided to perform an expression correlation analysis of these 181 lincRNAs against all DE PC genes (from the one way ANOVA-like heatmap on Fig. 3E) using read-counting values for each gene from the 15 RNA-Seq samples (biological triplicates for each life cycle stage). Among the correlated genes, we kept only gene pairs (lincRNA-PC gene) presenting r > 0.8 (classified as positively correlated) or r < −0.8 (negatively correlated). A co-expression network was built relying on that correlation information (Fig. 6A). It contains 181 lincRNAs (red nodes), 2359 PC genes (blue nodes), and 68,625 correlated pairs (edges), among which 92% (63,156/68,625) are for positive correlations (cyan edges) and 8% (5469/68,625) for negative ones (gray edges). As a negative control, a random network was generated with the same number of edges (68,625), 181 and 2359 randomly-picked lincRNAs and PC genes, respectively, in order to give support for the actual network significance. In this negative control dataset, we obtained 4788 edges that had either r ≥ 0.8 or r ≤ −0.8 out of 68,625 total edges (6.9%). Therefore, we could estimate a false discovery rate of 6.9% (4788/68,625), or a precision of ~ 93% for the correlations displayed on the actual network (see details on the Methods section).
Co-expression gene network (lincRNA-PC gene) pinpoints the processes in which lincRNAs may act. (A) The 181 robust lincRNAs (red nodes; lincRNAs from the main intersection in the UpSet Intersection Diagram on Fig. 4) were used as bait for catching 2359 PC genes (blue nodes) whose expression levels across five developmental stages are either positively (r ≥ 0.8, cyan edges) or negatively (r ≤ −0.8, gray edges) correlated with the expression levels of the lincRNAs. The network contains 68,625 edges among which 63,156 (92%) are positive correlations and 5469 (8%) negative ones. (B) A GO gene-enrichment analysis on the PC genes’ list revealed 16 significantly enriched GO terms (−log (p-value) on the x-axis), and among them there were membrane-related terms, as well as retrotransposons activity (orange dashed circles) and GPCR-associated pathways (purple dashed circles). The PC genes from the dash-circled GO categories were manually searched and clustered on the peripheral region from the network in (A), allowing one to see that both positive and negative correlations between lincRNA and PC genes are present on those processes.
A gene enrichment analysis for those 2359 PC genes was performed relying on the gene ontology (GO) terms assigned to them (see Methods). Sixteen GO categories comprise enriched genes with a significant hypergeometric test adjusted p-value < 0.01 (Fig. 6B). They are distributed into 10 cellular components (CC), 3 biological processes (BP) and 3 molecular functions (MF). Two terms from both BP and MF are highlighted: RNA-dependent DNA replication/RNA-directed DNA polymerase activity and G-protein coupled receptor (GPCR) signaling pathway/GPCR activity. The former represents genes involved in the transposition of retroelements, the major class of transposable elements (TEs) in S. mansoni genome, which is about 47% comprised by repeats50. TEs are also an important source for the birth of new lncRNAs within a genome9. The latter GO category is related to signal transduction via GPCR, which was already cited as a potential target in the context of drug discovery in S. mansoni 51. In addition, a recent study that built lncRNA-mRNA co-expression networks from venous congestion-subject human endothelial cells has identified GPCR as a potential pathway dynamically regulated by lincRNAs48.
Filtering the network by topological proximity between SmLINC-PC gene pairs
Since it is known that lincRNAs may act by regulating their flanking-PC gene neighbors (as already mentioned herein), and that they may form triplex structures (dsDNA-RNA) anchoring on different genomic loci and recruiting chromatin remodelers such as PRC2 (refs 13, 14), we idealized a two-step filtering method in order to build a co-expression network where the lincRNA-PC gene expression correlations would be linked to the topological proximity of the pair on the chromatin. The first step was to filter each gene pair (lincRNA-PC gene) by a pre-defined correlation coefficient (r) threshold. Starting with the same input used for building the network on Fig. 6A, we have now decreased the cutoff to r ≥ 0.5 or r ≤ −0.5. The second step, which we will hereafter call topological filtering step (TFS), was aimed at keeping only the SmLINC-PC gene neighbors and/or the pairs presenting a triplex structure predicted by the triplexator algorithm52 (see Methods for details). After those two rounds of gene pairs filtering, we obtained a network containing 326 nodes (89 lincRNAs and 237 PC genes) that comprised 62.5% positively correlated pairs (204/326 edges) and 37.5% negatively correlated ones (122/326 edges) (Supplementary Fig. S3).
We have also calculated the betweenness centrality (BC) score in order to identify important nodes within the network53. Roughly, BC measurement reflects the ratio of the number of shortest paths between two nodes (x, z) that pass through a node y by the number of total shortest paths between x and z, inferring, therefore, the propensity node y has to be a hub. The network layout that we show displays the node size proportional to the BC in each subnetwork (Fig. 7A and Supplementary Fig. S3). As examples, three subnetworks were selected and the expression profile across the five developmental stages of each target SmLINC and the correlated PC genes comprising each subnetwork have been displayed (Fig. 7B).
Topological filtering step (TFS) approach decreases the number of nodes and improves the visual inspection of lincRNAs’ subclusters. (A) Eighty nine out of the 181 robust lincRNAs from Fig. 4 passed at least one of the two TFS rules: (i) have a PC gene neighbor (Smp) as one of its correlated mates (putative cis-acting mechanism), and (ii) have an in silico computed probability to form a triplex structure (RNA-DNA) on its PC gene mate locus (putative trans-acting mechanism). The “Network Analyzer” function from cytoscape was used in order to obtain betweenness centrality score for each node, meaning the bigger the node, the higher propensity to be a hub. The 22 SmLINCs correlated with multiple PC genes are shown here. The entire network is shown on Supplementary Fig. S3 and it contains 326 nodes (89 SmLINCs and 237 Smps) with 62.5% positively correlated pairs (204/326, r ≥ 0.5, cyan edges) and 37.5% negatively correlated ones (122/326, r ≤ −0.5, gray edges). Dashed edges represent neighbor genes. (B) Gene expression patterns on three subnetworks that were manually selected by highlighting three SmLINC hubs, marked by arrows in (A) and named at the top of each panel in (B). Co-expression kinetics of the SmLINC (red line), positively correlated Smps (dark gray lines) and negatively correlated Smps (light gray lines) expressed across the five life cycle stages: cercariae (cerc), somula 3 h (s3h), somula 24 h (s24h), male and female.
We performed a k-means clustering analysis on all the 326 genes present in the entire network, being able to detect that 70% of the genes (230/326) grouped into clusters that had an expression pattern similar to what was observed for the majority of the genes composing the SmLINC-hubs’ subclusters depicted on Fig. 7B, that is, an up-regulation profile from cercariae to adult forms (Supplementary Fig. S4). It appeared to be concurrent to what was recently reported about cercariae transcriptional poised state42.
Another co-expression network building approach highlights PC gene hubs and their correlated lincRNAs
A second network building method, where we captured the most DE PC genes-correlated lincRNAs, was idealized. First we selected 307 PC genes (Smps) that were highly differentially expressed among the five life cycle stages on focus (log2FC ≥ 10 on at least one stage), and then we rescued all the other genes (Smps + SmLINCs) that presented a very stringent expression correlation (coefficient r ≥ 0.9 or r ≤ −0.9) with the 307 pre-chosen PC genes. A network with 2965 nodes (750 lincRNAs + 2215 PCs) and 52,913 edges (51,807 positive r and 1106 negative r) was generated by this approach (Supplementary Fig. S5 panel I). Due to their highest BC scores, three PC genes were detected as the most important nodes in the network: Smp_055340.2, Smp_038870.2 and Smp_036270.4. The first one (Smp_055340.2), with the highest BC value (~0.15, Supplementary Fig. S5 panel II), encodes the protein Lin-9, a nuclear tumor-suppressing agent in mammal cells54 that acts inhibiting DNA synthesis (G1/S transition). The second (Smp_038870.2) and third (Smp_036270.4) ones encode a NADH ubiquinone oxidoreductase and a splicing factor arginine/serine rich splicing factor 4, respectively. The expression patterns of SmLINCs that are either positively or negatively correlated with those three PC gene hubs along parasite differentiation are depicted on Supplementary Fig. S5 panel III.
Similarly to what was seen on the gene enrichment analysis for the 2359 Smps correlated with the 181 robust SmLINCs (Fig. 6A, B), the same assay performed on the different set of 2215 PC genes from this second network has also revealed “RNA-dependent DNA replication” and “DNA integration” as significantly enriched GO biological processes, as well as “Monovalent inorganic cation transmembrane transporter activity” as significantly enriched GO molecular functions (Supplementary Fig. S5 panel IV). Membrane and extracellular components’ terms were also significant in the gene enrichment.
Again, as a negative control we built a random network containing 2215 and 750 randomly-selected PC genes and lincRNAs, respectively, and the same number of edges present on the actual network. In this negative control, we retrieved 1550 out of 52,913 edges with r ≥ 0.9 or r ≤ −0.9, estimating a false discovery rate on the original network of 2.9%, or a precision of ~97%.
These results are indicative that SmLINC-Smp co-expression network construction may be considered as a reliable initial approach to hypothesize and direct further investigations on S. mansoni lincRNAs’ regulatory pathways.
Discussion
Long non-coding RNAs are striking molecules acting on a variety of biological processes within the cell, mostly related to regulation of gene expression2,3,4, 7, 8, 12, 14, 45. Due to this broad landscape of content and functionality and despite the increasing efforts to unravel novel lncRNAs (mainly in higher eukaryotes), as well as their respective roles, the hitherto understanding of these molecules action on the diverse intracellular regulatory pathways still corresponds to a small tip of a huge iceberg. Regarding lncRNAs from more ancient important organisms, such as parasites, this tip is even smaller and few studies are devoted to unravel such genes.
The S. mansoni genome, which had its first version published in 2009 (ref. 51), and improved with the use of a NGS strategy in 2012 (ref. 21), has a size of about 380 Mb and nearly 12,000 PC genes (Smps) already mapped/annotated. These static data pose the even greater challenge of understanding the molecular dynamics responsible for the peculiar features of the parasite biology. Post-transcriptional control of gene expression events, such as modulation of mRNA alternative splicing and silencing by RNAi, as well as epigenetic events such as chromatin remodeling by histone modifications and the consequent epigenetic mechanisms to activate transcription, are present in schistosomes41, 42, 55,56,57. As in other higher eukaryotes, it is believed that a variety of lncRNAs may mediate such reactions in the parasites.
Regulatory RNAs appear to occur in all forms of life on the earth and it has been described that the protein-coding fraction of genomes decreases according to the complexity of organisms, ranging from 80–95% in prokaryotes to only a minor proportion in mammals (~1.22% in humans) (refs 58 and 59). On a brief screening on the latest version of S. mansoni genome and its current predicted proteome21, we detected a protein-coding DNA content of about 4.5%. Thus, it is apparent that S. mansoni has a considerable “free space” on the non-protein-coding portion of the genome which can be a source of hundreds to thousands lncRNAs that may act as regulatory and adaptive elements on the strenuous environmental changes this parasite experiences in order to complete its life cycle.
Due to the weak evolutionary constraint regarding lncRNAs’ sequence conservation8, 9, conventional local alignment search tools, commonly used for identifying homology evidence between PC genes from quite divergent organisms, are not the ideal ones for any kind of non-coding RNA sequence analyses60. With the advent of RNA-Seq, it is now possible to screen a whole transcriptome, on a high-throughput manner, in order to catch such transcripts. Both GENCODE61 and FANTOM62 consortia made use of the NGS strategy providing a great volume of input data to their computational pipelines aimed to assemble, distinguish/classify and annotate the maximum number of transcripts in humans and mice, respectively.
In the present study, we relied on tens of RNA-Seq samples (n = 88) from different developmental stages (Supplementary Table S6) and we built a transcriptomics/bioinformatics computational pipeline (Fig. 1) that successfully retrieved over 7,000S. mansoni multiexonic lncRNA transcripts (402 antisense to PC genes and 7029 lincRNAs) (Supplementary File S1 and Supplementary Table S1). It is important to note that our datasets do not represent the entire lncRNAs’ complement from the S. mansoni transcriptome, since we ruled out monoexonic transcripts in order to reduce the chance of artifacts9.
The conservative RNA-Seq assembly and filtering approach used here has shown 53% sensitivity when applied to the P. falciparum lincRNAs dataset30, as described in the Results. Therefore, it indicates that we are probably missing some S. mansoni lincRNA loci (most probably monoexonic ones), however we are certainly decreasing our false-positive rate by the stringent selection criteria established in our pipeline, which gives us a lower-boundary for the lincRNAs complement in our target S. mansoni organism.
The search for evidence of being active genes revealed only few hundreds lincRNAs (181) that have a combination of the four investigated traits: presence of H3K4me3 marks on their TSS-surrounding genomic regions, Schistosoma spp. phastCons conservation score assigned to at least one exon, differential expression across five developmental stages/forms (cercariae, somula 3 h, somula 24 h, male and female) and a significant either positive or negative expression correlation with their PC gene neighbors (Fig. 3). Sixteen out of those 181 lincRNAs had their expression individually quantified by us across the five parasite life stages/forms through RT-qPCR. It is noteworthy that, while some lincRNAs were found to be upregulated in adults (such as SmLINC02394-IBu and SmLINC06535-IBu, Fig. 5), most of the lincRNAs showed upregulation in schistosomula. This might indicate an importance of lincRNAs on regulating some processes involved in the rapid adaptation of schistosomula after transition from the free-living larvae to the early mammal parasitic stage, such as the worm body remodeling and defense against the host immune system.
We were guided by studies on mice and humans that make use of co-expression approaches on the attempt of hypothesizing lincRNAs’ function12, 45, 49. After clustering lincRNA-PC genes by either positive or negative expression correlation (roughly mimicking either activation/stabilization or inhibition/destabilization, respectively) across a set of different samples, co-expression networks were built and/or gene enrichment analyses were performed in order to acquire a holistic systems overview of potential lincRNAs’ roles. Two approaches were used for the networks construction that basically differ only on their initial dataset (list of genes) used for rescuing the additional positively or negatively correlated genes along parasite development. The first approach started with a list of 181 robust lincRNA candidates used for capturing their correlated Smp protein-coding genes (Figs 6 and 7), whereas the second one began with a list of 307 highly DE PC genes that were used for capturing their correlated genes in general (both SmLINCs and other Smps) (Supplementary Fig. S5). One interesting feature displayed by these two different network construction approaches was the presence of common GO categories of enriched PC genes: “RNA-dependent DNA replication”, “DNA integration”, “Monovalent inorganic cation transmembrane transporter activity” (Fig. 6B and Supplementary Fig. S5 panel IV), indicating that SmLINCs might be regulating such processes.
Both topological filtering step (TFS) and betweenness centrality (BC) measurement allowed us to highlight important SmLINCs (Fig. 7 and Supplementary Figs S3 and S5). For instance, SmLINC01637-IBu and SmLINC04271-IBu (orange and pink arrows on Fig. 7) appeared to be interesting candidates for functional studies. The former (SmLINC01637-IBu), while positively correlated to only one PC gene (ataxin 2 - Smp_122830), is negatively correlated to several mRNAs coding for membrane and/or secreted proteins, such as: fibrillin 2 (Smp_001100), solute carrier family 1 (Smp_016600), sodium dependent glucose transporter 1 (Smp_139150), transmembrane protein 26 (Smp_026670), transmembrane protein 231 (Smp_081720), fras1 related extracellular matrix protein (Smp_149390), protocadherin 9 (Smp_151620), surface membrane antigen (Smp_195180) and saposin B domain containing protein (Smp_016490.1 and 0.2), which has already been investigated as a vaccine candidate63. Whereas the latter (SmLINC04271-IBu) displays a mix of positively and negatively correlated mRNAs: more than half (8/15) on the first set (r ≥ 0.5) code for positively correlated hypothetical proteins (and noteworthy, the set includes a gene coding for the transcription factor forkhead box protein P1 - Smp_212350); and on the second set (r ≤ −0.5) there are 35% (15/43) and 39.5% (17/43) negatively correlated mRNAs coding for hypothetical proteins and membrane/secreted proteins, respectively, of which several are shared with SmLINC01637-IBu as negatively correlated (Smp_001100, Smp_016490.1, Smp_016490.2, Smp_016600, Smp_031430, Smp_081720, Smp_139150, Smp_141180, Smp_149390 and Smp_195180). Other interesting mRNAs are also listed on the r ≤ −0.5 set from SmLINC04271-IBu, such as SPARC protein (Smp_171780), PDZ domain-containing protein GIPC3 (Smp_170870), fasciclin domain-containing protein (Smp_141680), transmembrane protein 145 (Smp_125130), MEG-5 (Smp_152580) and a couple of transcription factors (HNF 4 - Smp_174700 and engrailed 2 C - Smp_145200).
This is the first time an approach for rescuing lncRNAs plus another for proposing their possible functionality through co-expression networks’ construction are reported in a single study on a parasite’s genome/transcriptome landscape. Adding to what was already seen for P. falciparum regarding lncRNAs30, 64, the data presented herein reinforce that those transcripts are expressed and may play a role on the biology of neglected tropical disease-causing agents. Thus, parasites’ lncRNAs and their pathways of action should start being considered as possible new therapeutic targets on future investigations.
Conclusions
The established computational pipeline appears to be a robust tool for the identification of thousands of multiexonic S. mansoni lncRNAs and might be applicable to any other organism. Hundreds of the putative lincRNAs display evolutionary conservation within the Schistosoma genus, transcriptional activation epigenetic mark (H3K4me3) at their TSSs, differential expression across five developmental stages and expression correlation with their protein-coding gene neighbors. In addition, RT-qPCR assays for 16 SmLINCs have confirmed that they undergo differential expression along the parasite development.
The construction of co-expression networks allows a holistic systems overview that helps us to decipher the role of these intriguing molecules on S. mansoni biology. We believe that the networks built and disclosed herein have now paved the way, as an important initial source, for investigations on S. mansoni regulatory pathways involving lincRNAs and their correlated PC genes. Functional assays are necessary to characterize individual SmLINCs as regulatory elements of their PC gene counterparts within the same co-expression subcluster.
Methods
Full methods are available on-line in the Supplementary Materials section.
Ethics statement
All protocols involving animals were conducted in accordance with the Ethical Principles in Animal Research adopted by the Brazilian College of Animal Experimentation (COBEA), and the protocol/experiments have been approved by the Ethics Committee for Animal Experimentation of Instituto Butantan (CEUAIB Protocol number 1777050816).
Genomic and transcriptomic data
The Schistosoma genomes analyzed herein were downloaded from the Wellcome Trust Sanger Institute ftp site for S. mansoni (http://ftp.sanger.ac.uk/pub/pathogens/Schistosoma/mansoni/Latest_assembly_annotation_others/Schistosoma_mansoni_v5.2.fa) and from wormbase ftp for both S. haematobium (http://ftp.ebi.ac.uk/pub/databases/wormbase/parasite/releases/WBPS8/species/schistosoma_haematobium/PRJNA78265/schistosoma_haematobium.PRJNA78265.WBPS8.genomic.fa.gz) and S. japonicum (http://ftp.ebi.ac.uk/pub/databases/wormbase/parasite/releases/WBPS8/species/schistosoma_japonicum/PRJEA34885/schistosoma_japonicum.PRJEA34885.WBPS8.genomic.fa.gz). The most recent genomic annotation of nearly 12,000 PC genes (Smps) was used21.
We have used 88 S. mansoni RNA-Seq libraries as input for our computational pipeline (more details on the next topic): 52 from public studies already deposited in the Sequence Read Archive (SRA) NCBI database, which encompass several developmental stages (Supplementary Table S6), and 36 in-house separate male (Accession numbers SAMN06221530-SAMN06221541 and SAMN06221554-SAMN06221559) and female libraries (Accession numbers SAMN06221542-SAMN06221553 and SAMN06221560-SAMN06221565).
Assembly of RNA-Seq reads
Trinity de novo assembly32 was applied to three different RNA-Seq datasets: (1) trin_strA dataset, comprised of ~300 million raw paired-end reads (SRA accession SAMN06221530-SAMN06221553) obtained in house from 24 samples of adult worm couples exposed in vitro to TH65 (compound 13I) (ref. 65) or to vehicle (controls); (2) trin_strB dataset, comprised of ~300 million raw paired-end reads (SRA accession SAMN06221554-SAMN06221565), again obtained in house from 12 samples of adult worm couples exposed in vitro to GSK343 (ref. 66) or to vehicle (controls); (3) trin_sra dataset, which encompasses 52 RNA-Seq libraries (Supplementary Table S6) (~2 billion raw reads) from S. mansoni at several different life cycle stages (both untreated samples or samples treated with different compounds), which were downloaded from the SRA-NCBI public repository. A second assembly approach using the Tuxedo tools (Tophat2 (ref. 33), Cufflinks and Cuffmerge34) was applied to a subset of the above RNA-Seq libraries that included 44 out of the 52 SRA RNA-Seq libraries comprised exclusively of control untreated samples, plus 12 in house-derived libraries from both strA and strB comprised exclusively of control samples (total of ~2.2 billion raw reads). Each of those 56 samples (44 + 12) was used as an independent input file for both tophat2 and cufflinks, and then all 56 independently assembled transcript sets were merged onto a single non-redundant gtf file through cuffmerge execution. See further details in the Supplementary Materials section.
Retrieving long non-coding RNAs from the assembled transcripts’ dataset
After having all the assembled transcripts mapped to the reference genome (bed12 and gtf formats for de novo and genome-guided assemblies, respectively), a series of filtering steps was applied as described below, aimed at both removing unwanted protein-coding transcripts and rescuing putative lncRNAs from the entire transcriptome datasets (red rectangles section from the pipeline depicted on Fig. 1). The tools that were used and their respective parameters were all placed in one single PERL script and adapted for automation; the script can be downloaded from https://github.com/eltonjrv/Smansoni.lncRNAs. See further details in the Supplementary Materials section.
Searching for evidence supporting S. mansoni lincRNAs as functional genes
H3K4me3 marks
In order to search for the presence of histone H3 lysine 4 trimethylation (H3K4me3) on the transcription start site (TSS) of S. mansoni genes as an epigenetic mark for transcriptional activation, we relied on four ChIP-Seq assays publicly available at the SRA-NCBI database: SRR1107840 and SRR2530135 were obtained from adult worms, whereas SRR2120359 and SRR2120360 from schistosomula parasites. Those data were all generated by the same group and are part of three different publications42, 67, 68. See further details in the Supplementary Materials section.
Evolutionary conservation by whole genomes’ comparison
In order to mask both repeats and low complexity regions present within Schistosoma spp. genomes, we ran RepeatMasker (http://repeatmasker.org) (-e crossmatch -pa 20 -q -xsmall -gff -norna -lib RepBasePerpignanSma52.fasta) on each of the three whole genomes compared herein. Pairwise alignments of S. mansoni repeat-masked genome against S. haematobium and S. japonicum ones were performed using lastz algorithm (https://www.bx.psu.edu/~rsharris/lastz/), an improved version of blastz69. See further details in the Supplementary Materials section.
Differential Expression and correlation analyses across five developmental stages
Fifteen RNA-Seq libraries from five different developmental stages of S. mansoni (biological triplicate each) were selected for investigation of gene expression profile of both PC and lncRNA genes: cercariae (ERR022872, ERR022877 and ERR022878), somula 3 h (ERR022874, ERR022876 and ERR022879), somula 24 h (ERR022880, ERR022881 and ERR022882), male (SAMN06221530, SAMN06221531 and SAMN06221532) and female (SAMN06221542, SAMN06221543 and SAMN06221544) adults. See further details in the Supplementary Materials section.
Co-expression network construction and analyses
Based on the general gene expression correlation analysis among S. mansoni five developmental stages (described on the topic just above), we were able to retrieve lincRNA-PC gene pairs that are either positively or negatively correlated by selecting arbitrary r thresholds (see r cutoffs established by us further in this topic). We used Unix/Shell tools on the output of the ad-hoc correlation’s R script in order to prepare the simple interaction formats (.sif) to feed Cytoscape software70 for both network visualization and further analyses within the network. For each correlated genes’ pair we assigned a “pos” or “neg” edge name regarding whether the correlation between the genes is positive or negative, respectively. See further details in the Supplementary Materials section.
Statistical analyses and charts plotting
Statistical analyses and charts plotting were performed within the R environment (version 3.3.2) with limma, edgeR, gplots and ggplot2 libraries loaded. Gene expression line charts were generated using matplotlib in Python. The intersection diagram in the UpSet format71 was plotted using Intervene (https://asntech.shinyapps.io/intervene/) (doi:10.1101/109728).
Parasite materials
All parasite material was from a BH isolate of S. mansoni maintained by passage through golden hamster (Mesocricetus auratus) and Biomphalaria glabrata snails. See further details in the Supplementary Materials section.
RNA extraction, quantification and quality assessment
Total RNA from cercariae (C) and schistosomula (3S and 24S) was extracted using a protocol based on Roquis et al., 2015 (ref. 42). Briefly, 25,000 cercariae or schistosomula were ground with glass beads in liquid nitrogen for 5 minutes. Then the Qiagen RNeasy Micro Kit (Cat number 74004) was used for RNA extraction and purification according to the manufacturer’s instructions, except for the DNase I treatment: the amount of DNase I was doubled and the time of treatment was increased to 45 minutes. Male or female worms were first disrupted in Qiagen RLT buffer using glass potters and pestles. RNA from males and females was then extracted and purified using the Qiagen RNeasy Mini Kit (Cat number 74104), according to the manufacturer’s instructions, except for the DNase I treatment, which was the same used for cercariae and schistosomula RNA extraction. All the RNA samples were quantified using the Qubit RNA HS Assay Kit (Q32852, Thermo Fisher Scientific) and the integrity of RNAs was verified using the Agilent RNA 6000 Pico Kit (5067-1513 Agilent Technologies) in a 2100 Bioanalyzer Instrument (Agilent Technologies). Four biological replicates were assessed for each life cycle stage.
RT-qPCR assays
The reverse transcription (RT) reaction was performed with 100 ng of each total RNA sample using the SuperScript IV First-Strand Synthesis System (18091050, Life Technologies) and random hexamer primers in a 20 μL final volume. The obtained complementary DNAs (cDNAs) were diluted 10 times in water and quantitative PCR was performed using 2.5 μL of each diluted cDNA in a total volume of 10 μL containing 1X LightCycler 480 SYBR Green I Master Mix (04707516001, Roche Diagnostics) and 800 nM of each primer in a LightCycler 480 System (Roche Diagnostics). Primers for selected transcripts (Supplementary Table S5) were designed using the Primer 3 tool (http://biotools.umassmed.edu/bioapps/primer3_www.cgi), and each real-time qPCR was run in three technical replicates. The results were analyzed by comparative Ct method72. The reference gene Smp_092920 was chosen from twelve genes that showed no differential expression in the RNA-Seq data along the five stages. Data from the RT-qPCR expression values of the twelve genes across the five stages were analyzed with RefFinder73 (http://150.216.56.64/referencegene.php) using three tools (BestKeeper, NormFinder and GeNorm) in order to choose the most stable gene for qPCR. Real-time data were normalized according to the expression level of the Smp_092920 reference gene, and p-values were determined by one-way analysis of variance (ANOVA) and Tukey post-hoc tests.
Data Availability
The RNA-seq datasets generated during the current study are available in the Sequence Read Archive (SRA) NCBI repository under Accession numbers SAMN06221530-SAMN06221541, SAMN06221554-SAMN06221559, SAMN06221542-SAMN06221553 and SAMN06221560-SAMN06221565. A GTF file with the S. mansoni lincRNAs identified with our pipeline can be downloaded from http://verjolab.usp.br/tracks/schMan/schMan1/. All other public RNA-seq data analyzed during the current study are from the SRA repository and their Accession numbers are listed in Supplementary Table S6.
References
Mattick, J. S. & Makunin, I. V. Non-coding RNA. Hum Mol Genet 15 Spec No 1, R17–29 (2006).
Quinn, J. J. & Chang, H. Y. Unique features of long non-coding RNA biogenesis and function. Nat Rev Genet 17, 47–62 (2016).
Khalil, A. M. et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci USA 106, 11667–11672, doi:10.1073/pnas.0904715106 (2009).
Mercer, T. R. & Mattick, J. S. Structure and function of long noncoding RNAs in epigenetic regulation. Nat Struct Mol Biol 20, 300–307 (2013).
Mondal, T., Rasmussen, M., Pandey, G. K., Isaksson, A. & Kanduri, C. Characterization of the RNA content of chromatin. Genome Res 20, 899–907, doi:10.1101/gr.103473.109 (2010).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res 22, 1775–1789, doi:10.1101/gr.132159.111 (2012).
Nakaya, H. I. et al. Genome mapping and expression analyses of human intronic noncoding RNAs reveal tissue-specific patterns and enrichment in genes related to regulation of transcription. Genome Biol 8, R43, doi:10.1186/gb-2007-8-3-r43 (2007).
Kapusta, A. & Feschotte, C. Volatile evolution of long noncoding RNA repertoires: mechanisms and biological implications. Trends Genet 30, 439–452 (2014).
Ulitsky, I. Evolution to the rescue: using comparative genomics to understand long non-coding RNAs. Nat Rev Genet 17, 601–614, doi:10.1038/nrg.2016.85 (2016).
Kornienko, A. E. et al. Long non-coding RNAs display higher natural expression variation than protein-coding genes in healthy humans. Genome Biol 17, 14 (2016).
Liu, S. J. et al. CRISPRi-based genome-scale identification of functional long noncoding RNA loci in human cells. Science. doi:10.1126/science.aah7111 (2016).
Liu, S. J. et al. Single-cell analysis of long non-coding RNAs in the developing human neocortex. Genome Biol 17, 67 (2016).
Li, Y., Syed, J. & Sugiyama, H. RNA-DNA Triplex Formation by Long Noncoding RNAs. Cell Chem Biol (2016).
Mondal, T. et al. MEG3 long noncoding RNA regulates the TGF-beta pathway genes through formation of RNA-DNA triplex structures. Nat Commun 6, 7743, doi:10.1038/ncomms8743 (2015).
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323, doi:10.1016/j.cell.2007.05.022 (2007).
Grote, P. et al. The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev Cell 24, 206–214, doi:10.1016/j.devcel.2012.12.012 (2013).
Kalwa, M. et al. The lncRNA HOTAIR impacts on mesenchymal stem cells via triple helix formation. Nucleic Acids Res 44, 10631–10643, doi:10.1093/nar/gkw802 (2016).
Martianov, I., Ramadass, A., Serra Barros, A., Chow, N. & Akoulitchev, A. Repression of the human dihydrofolate reductase gene by a non-coding interfering transcript. Nature 445, 666–670, doi:10.1038/nature05519 (2007).
WHO. In Media centre - Fact sheet (2016).
Schistosoma japonicum Genome, S. & Functional Analysis, C. The Schistosoma japonicum genome reveals features of host-parasite interplay. Nature 460, 345–351, doi:10.1038/nature08140 (2009).
Protasio, A. V. et al. A systematically improved high quality genome and transcriptome of the human blood fluke Schistosoma mansoni. PLoS Negl Trop Dis 6, e1455 (2012).
Young, N. D. et al. Whole-genome sequence of Schistosoma haematobium. Nat Genet 44, 221–225 (2012).
Copeland, C. S. et al. Homology-based annotation of non-coding RNAs in the genomes of Schistosoma mansoni and Schistosoma japonicum. BMC Genomics 10, 464 (2009).
Anderson, L. et al. Schistosoma mansoni Egg, Adult Male and Female Comparative Gene Expression Analysis and Identification of Novel Genes by RNA-Seq. PLoS Negl Trop Dis 9, e0004334, doi:10.1371/journal.pntd.0004334 (2015).
Oliveira, K. C., Carvalho, M. L., Maracaja-Coutinho, V., Kitajima, J. P. & Verjovski-Almeida, S. Non-coding RNAs in schistosomes: an unexplored world. An Acad Bras Cienc 83, 673–694 (2011).
Chen, J. et al. Evolutionary analysis across mammals reveals distinct classes of long non-coding RNAs. Genome Biol 17, 19, doi:10.1186/s13059-016-0880-9 (2016).
Sun, L., Liu, H., Zhang, L. & Meng, J. lncRScan-SVM: A Tool for Predicting Long Non-Coding RNAs Using Support Vector Machine. PLoS One 10, e0139654, doi:10.1371/journal.pone.0139654 (2015).
Sun, L. et al. Prediction of novel long non-coding RNAs based on RNA-Seq data of mouse Klf1 knockout study. BMC Bioinformatics 13, 331, doi:10.1186/1471-2105-13-331 (2012).
Zhao, J., Song, X. & Wang, K. lncScore: alignment-free identification of long noncoding RNA from assembled novel transcripts. Sci Rep 6, 34838, doi:10.1038/srep34838 (2016).
Broadbent, K. M. et al. Strand-specific RNA sequencing in Plasmodium falciparum malaria identifies developmentally regulated long non-coding RNA and circular RNA. BMC Genomics 16, 454 (2015).
Nam, J. W. & Bartel, D. P. Long noncoding RNAs in C. elegans. Genome Res 22, 2529–2540 (2012).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29, 644–652, doi:10.1038/nbt.1883 (2011).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14, R36, doi:10.1186/gb-2013-14-4-r36 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28, 511–515, doi:10.1038/nbt.1621 (2010).
Kong, L. et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res 35, W345–349 (2007).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat Protoc 8, 1494–1512 (2013).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Research 25, 3389–3402 (1997).
Kadakkuzha, B. M. et al. Transcriptome analyses of adult mouse brain reveal enrichment of lncRNAs in specific brain regions and neuronal populations. Frontiers in cellular neuroscience 9, 63, doi:10.3389/fncel.2015.00063 (2015).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837, doi:10.1016/j.cell.2007.05.009 (2007).
Cosseau, C. et al. Native chromatin immunoprecipitation (N-ChIP) and ChIP-Seq of Schistosoma mansoni: Critical experimental parameters. Mol Biochem Parasitol 166, 70–76 (2009).
Roquis, D. et al. The Epigenome of Schistosoma mansoni Provides Insight about How Cercariae Poise Transcription until Infection. PLoS Negl Trop Dis 9, e0003853 (2015).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res 15, 1034–1050 (2005).
Blanchette, M. et al. Aligning multiple genomic sequences with the threaded blockset aligner. Genome Res 14, 708–715, doi:10.1101/gr.1933104 (2004).
Necsulea, A. et al. The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature 505, 635–640 (2014).
Wiberg, R. A. et al. Assessing Recent Selection and Functionality at Long Noncoding RNA Loci in the Mouse Genome. Genome Biol Evol 7, 2432–2444, doi:10.1093/gbe/evv155 (2015).
Parker-Manuel, S. J., Ivens, A. C., Dillon, G. P. & Wilson, R. A. Gene expression patterns in larval Schistosoma mansoni associated with infection of the mammalian host. PLoS Negl Trop Dis 5, e1274 (2011).
Li, J., Xu, Y., Xu, J., Wang, J. & Wu, L. Dynamic co-expression network analysis of lncRNAs and mRNAs associated with venous congestion. Mol Med Rep 14, 2045–2051 (2016).
St Laurent, G. et al. Functional annotation of the vlinc class of non-coding RNAs using systems biology approach. Nucleic Acids Res 44, 3233–3252, doi:10.1093/nar/gkw162 (2016).
Lepesant, J. M. et al. Combination of de novo assembly of massive sequencing reads with classical repeat prediction improves identification of repetitive sequences in Schistosoma mansoni. Exp Parasitol 130, 470–474 (2012).
Berriman, M. et al. The genome of the blood fluke Schistosoma mansoni. Nature 460, 352–358 (2009).
Buske, F. A., Bauer, D. C., Mattick, J. S. & Bailey, T. L. Triplexator: detecting nucleic acid triple helices in genomic and transcriptomic data. Genome Res 22, 1372–1381 (2012).
Zhang, J. D. & Wiemann, S. KEGGgraph: a graph approach to KEGG PATHWAY in R and bioconductor. Bioinformatics 25, 1470–1471 (2009).
Sandoval, R. et al. A mutant allele of BARA/LIN-9 rescues the cdk4−/− phenotype by releasing the repression on E2F-regulated genes. Exp Cell Res 312, 2465–2475 (2006).
Cabezas-Cruz, A., Lancelot, J., Caby, S., Oliveira, G. & Pierce, R. J. Epigenetic control of gene function in schistosomes: a source of therapeutic targets? Front Genet 5, 317 (2014).
Krautz-Peterson, G., Bhardwaj, R., Faghiri, Z., Tararam, C. A. & Skelly, P. J. RNA interference in schistosomes: machinery and methodology. Parasitology 137, 485–495 (2010).
Piao, X. et al. Genome-wide transcriptome analysis shows extensive alternative RNA splicing in the zoonotic parasite Schistosoma japonicum. BMC Genomics 15, 715, doi:10.1186/1471-2164-15-715 (2014).
Dunham, I. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Mattick, J. S. RNA regulation: a new genetics? Nat Rev Genet 5, 316–323 (2004).
Eddy, S. R. Computational analysis of RNAs. Cold Spring Harb Symp Quant Biol 71, 117–128, doi:10.1101/sqb.2006.71.003 (2006).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res 22, 1760–1774, doi:10.1101/gr.135350.111 (2012).
Lizio, M. et al. Update of the FANTOM web resource: high resolution transcriptome of diverse cell types in mammals. Nucleic Acids Res 45, D737–D743, doi:10.1093/nar/gkw995 (2017).
Figueiredo, B. C. et al. Kicking in the Guts: Schistosoma mansoni Digestive Tract Proteins are Potential Candidates for Vaccine Development. Front Immunol 6, 22 (2015).
Amit-Avraham, I. et al. Antisense long noncoding RNAs regulate var gene activation in the malaria parasite Plasmodium falciparum. Proc Natl Acad Sci USA 112, E982–991 (2015).
Heimburg, T. et al. Structure-Based Design and Synthesis of Novel Inhibitors Targeting HDAC8 from Schistosoma mansoni for the Treatment of Schistosomiasis. J Med Chem 59, 2423–2435 (2016).
Verma, S. K. et al. Identification of Potent, Selective, Cell-Active Inhibitors of the Histone Lysine Methyltransferase EZH2. ACS Med Chem Lett 3, 1091–1096 (2012).
Lepesant, J. M., Boissier, J., Climent, D., Cosseau, C. & Grunau, C. Female biased sex-ratio in Schistosoma mansoni after exposure to an allopatric intermediate host strain of Biomphalaria glabrata. Exp Parasitol 135, 350–356 (2013).
Roquis, D. et al. Exposure to hycanthone alters chromatin structure around specific gene functions and specific repeats in Schistosoma mansoni. Front Genet 5, 207 (2014).
Schwartz, S. et al. Human-mouse alignments with BLASTZ. Genome Res 13, 103–107 (2003).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13, 2498–2504 (2003).
Lex, A., Gehlenborg, N., Strobelt, H., Vuillemot, R. & Pfister, H. UpSet: Visualization of Intersecting Sets. IEEE transactions on visualization and computer graphics 20, 1983–1992, doi:10.1109/TVCG.2014.2346248 (2014).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Xie, F., Xiao, P., Chen, D., Xu, L. & Zhang, B. miRDeepFinder: a miRNA analysis tool for deep sequencing of plant small RNAs. Plant molecular biology 80, 75–84, doi:10.1007/s11103-012-9885-2 (2012).
Acknowledgements
We thank Dr. J.C. Setubal for access to the computational facilities of the Bioinformatics Laboratory of Instituto de Química, Universidade de São Paulo (USP); Dr. Robert Harris (Pennsylvania State University) and Dr. Minmei Hou (Northern Illinois University) for helping on computational issues related to running blastz/lastz and TBA programs. We also acknowledge Patricia Aoki Miyasato and Dr. Eliana Nakano, Laboratorio de Malacologia, Instituto Butantan, for maintaining the S. mansoni life cycle. We thank Dr. Roberto Hirata Jr. and Dr. Roberto Marcondes Cesar-Junior from the Instituto de Matemática e Estatística - USP for the access to the eScience Computing Network. This work was supported in part by a grant from the European Union’s Seventh Framework Programme under grant agreement no. 602080 to SVA. EJRV and ASAP were supported by fellowships from Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP 2014/24560-8 and 2016/10046-6). LFdS and GML were supported by fellowships from Conselho Nacional de Desenvolvimento Cientifico e Tecnologico (CNPq). SVA was also supported by institutional funds from Fundação Butantan and received an established investigator fellowship award from CNPq, Brasil. The eScience Computing Network at the Instituto de Matemática e Estatística -Universidade de São Paulo (USP) has been financed by FAPESP, CNPq, Comissão de Aperfeiçoamento de Pessoal do Ensino Superior (CAPES) and Núcleo de Apoio à Pesquisa eScience-USP. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
E.J.R.V. and S.V.-A. conceived the project, analyzed the data and wrote the manuscript. E.J.R.V. designed and carried out all the in silico experiments encompassing pipeline development (programming), data curation/annotation, systems approach and statistical analyses, as well as wrote the original draft of the manuscript. LFdS guided on statistical analyses and performed part of data visualization. D.S.P. gave support on computational tasks, genome browser administration and performed part of data visualization. M.S.A., A.S.A.P. and G.L.M. designed and carried out the RT-qPCR assays for the validation/confirmation of lincRNAs’ expression. M.S.A. wrote the methodology for the in vitro experiments. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vasconcelos, E.J.R., daSilva, L.F., Pires, D.S. et al. The Schistosoma mansoni genome encodes thousands of long non-coding RNAs predicted to be functional at different parasite life-cycle stages. Sci Rep 7, 10508 (2017). https://doi.org/10.1038/s41598-017-10853-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-10853-6
This article is cited by
-
Ultrasound-guided hydrodistension for adhesive capsulitis: a longitudinal study on the effect of diabetes on treatment outcomes
Skeletal Radiology (2023)
-
Characterization of glutamate carboxypeptidase 2 orthologs in trematodes
Parasites & Vectors (2022)
-
Long non-coding RNAs as possible therapeutic targets in protozoa, and in Schistosoma and other helminths
Parasitology Research (2022)
-
Full-length transcriptome analysis and identification of transcript structures in Eimeria necatrix from different developmental stages by single-molecule real-time sequencing
Parasites & Vectors (2021)
-
Assessment of reference genes at six different developmental stages of Schistosoma mansoni for quantitative RT-PCR
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.