Abstract
Why is an amino acid replacement in a protein accepted during evolution? The answer given by bioinformatics relies on the frequency of change of each amino acid by another one and the propensity of each to remain unchanged. We propose that these replacement rules are recoverable from the secondary structural trends of amino acids. A distance measure between high-resolution Ramachandran distributions reveals that structurally similar residues coincide with those found in substitution matrices such as BLOSUM: Asn ↔ Asp, Phe ↔ Tyr, Lys ↔ Arg, Gln ↔ Glu, Ile ↔ Val, Met → Leu; with Ala, Cys, His, Gly, Ser, Pro, and Thr, as structurally idiosyncratic residues. We also found a high average correlation (\(\overline{R}\) = 0.85) between thirty amino acid mutability scales and the mutational inertia (I X ), which measures the energetic cost weighted by the number of observations at the most probable amino acid conformation. These results indicate that amino acid substitutions follow two optimally-efficient principles: (a) amino acids interchangeability privileges their secondary structural similarity, and (b) the amino acid mutability depends directly on its biosynthetic energy cost, and inversely with its frequency. These two principles are the underlying rules governing the observed amino acid substitutions.
Similar content being viewed by others
Introduction
In molecular evolution, protein stability is a solid indicator of function preservation thanks to a positive correlation between protein functionality and native stability1,2. Natural protein sequences evolved to avoid aggregation and increase functional diversity3, and once a protein fold is established, the selection pressure at most positions in the protein will preserve fold stability. Homologous families of proteins have related functions, and structures are similar although sequences have diverged4, even in regions with less than 30% sequence identity5,6. Accordingly, mutation events over time may replace a residue by another while keeping the backbone dihedral angles at that position unchanged7. These facts indicate that the amino acid sequence alone is an incomplete measure of evolutionary relationships between proteins. Indeed, structural similarities better reflect homology than sequence similarities8. Therefore, sequence variation around a conserved molecular architecture could be traced through amino acid substitution patterns fixed during protein evolution.
The intrinsic secondary structure propensities of amino acids are given by the statistics of Ramachandran distributions9,10,11. In this way, we could know the conformational bias of each amino acid towards specific secondary structures12,13. For instance, long polypeptide chains with the same backbone conformation are found exclusively in α − helix, PPII, and β strands structures14. In general, examining the frequency of occurrence of particular amino acid residues in stable secondary structures have been useful for determining protein structure, folding, and energetics15. We propose that, in addition, the statistics of the secondary structure of proteins may reveal their evolutionary information.
To confirm this assumption, we explore a combination of extensive physical quantities with the statistics of Ramachandran distributions P X (ϕ, ψ). In particular, we investigate the molecular mass as a measure of the amino acids biosynthetic cost. In addition, we use the protein geometry database (PGD 1.1)16 for obtaining high-resolution Ramachandran distributions as 2D-binned probability histograms (Fig. 1). This choice has some practical advantages, including the possibility of directly applying distance measures between the distributions. The secondary structure distance between the amino acids (Fig. 2) is the main task in our research because the emerging close-distance pairs can be straightforwardly compared to pairwise mutations. The optimal bin area (ΔϕΔψ) dividing the Ramachandran map is given by the method of Shimazaki & Shinomoto17. This is a key element in histogram binning because a very small bin size will result in noise amplification whereas a very large value will overpass important details of the distribution.
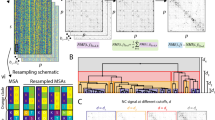
High-resolution Ramachandran probability distributions P X (ϕ, ψ) (logarithmic scale) as derived from the PGD 1.1 database at 1.895° × 1.895° bin size. Structurally similar open sets: yellow, S I = {{Arg, Lys}, {Glu, Gln}, Leu}; green, S II = {Trp, {Phe, Tyr}}; magenta, S III = {Ans, Asp}; cyan, S IV = {Val, Ile}. Ala, Met, and Ser have their first neighbor in S I ; His, Thr, and Cys are adjacent to S II . Larger images of each Ramachandran distribution are given by Supplementary Figs. S1–S20.
Distance matrix ordered according to structurally similar amino acids. The smallest distance is represented in yellow, and the largest distance in blue, with intermediate values in green. Open subsets appear, consistently, in yellow. Additionally, Gly, and Pro appear as the most distant elements, followed by Asn, Val-Ile, Ala, and Thr.
We explore the twenty amino acid distributions through some of their distinctive features such as the most probable conformation, which is given by the highest peak of each distribution. Additionally, we propose a plausible mutability parameter that combines structural information with the molecular mass of the amino acids. Our results indicate that amino acid evolutionary substitutions occur by following two optimal-efficiency principles: (a) interchangeability between amino acids occurs by preserving secondary structural propensity, and (b) the mutability of an amino acid depends directly on its mass, and inversely with its frequency. The methodology introduced here gives the basis for developing a new kind of scoring matrices involving physical quantities and secondary structure statistics. Hopefully, these future efforts will further help to improve the peptide design strategies, which can contribute to close the gap between the primary sequence and the 3D structure of proteins.
Results and Discussion
High-resolution Ramachandran Probability Distributions
We distinguish two concepts regarding the backbone dihedral angles of proteins, as suggested by Dunbrack Jr. et al.11. The first is a Ramachandran plot or Ramachandran map, which is simply a scatter plot of the ϕ, ψ values for the amino acids in a single protein structure or a set of protein structures. It provides a simple view of the conformation of a protein. The second is a Ramachandran probability distribution P(ϕ, ψ) which is a statistical representation of Ramachandran data, usually in the form of a probability density function. P X (ϕ, ψ) gives the probability of finding an amino acid conformation in a specific range of (ϕ, ψ) values.
We obtained non-parametric density estimates of P X (ϕ, ψ) for each amino acid X from 1,153,791 residues retrieved from the high-resolution protein geometry database (PGD 1.1)16. In our approach—frequentist—events have a specific probability whose determination depends on the number of observations. Therefore each distribution P X (ϕ, ψ) is given by a joint histogram. Such an approach depends on finding an optimal grid size, which can be determined with Shimazaki & Shinomoto method17. Said strategy requires a heuristic exhaustive sampling of a cost function whose minimum corresponds to an optimal binning of the distribution—see methods for details. Table 1 reports the optimal bin width for each Ramachandran probability distribution, \({{\rm{\Delta }}}_{X}^{min}\). The weighted average of these optimal bin widths gave us the bin size used (1.895°) in the present study. Thus, we obtained a grid with a total of 190 × 190 bins (36,100), each one covering an area of 1.895° × 1.895° of the dihedral space (Fig. 1), which is a significant improvement on the resolution of Ramachandran distributions previously reported.
For comparison, the 3D representation of the Ramachandran distributions for the first version of PGD uses a grid of 20.0° × 20.0° (i.e. a total of 324 bins), from a dataset containing 72,376 residues10. In another approach, the predicted protein backbone torsion angles from NMR chemical shifts made by the TALOS+ program uses an identical bin size (20.0° × 20.0°)18,19, other studies on folding trends uses a resolution of 10.0° × 10.0° (i.e 1,296 bins)11. An early report on detailed Ramachandran distributions used bin widths of 4.0° × 4.0° (i.e. 90 × 90 bins), involving 237,384 amino acids from 1,042 proteins20. Our distributions have a resolution 4.5 times higher, which translates into a higher accuracy in the distance computations between the set of distributions P X (ϕ, ψ). This high resolution was possible thanks to the fact that at least 84% of the structures reported at the protein data bank (PDB) were obtained during the last decade alone, most of which have atomic resolution.
Figure 1 reports the 3D plots of the twenty Ramachandran distributions determined for the present study; the dihedral angles are given in degrees, while the percentage probability per bin is given on a logarithmic scale. All the plots have the same height to facilitate their comparison. Larger plots are included in Supplementary Figs. S1–S20. While most distributions look similar one to another, there are some key differences. The probability distribution of glycine is very symmetrical and occupies all the allowed regions of the Ramachandran map. It is the only residue having a maximum at the left-handed α-helix conformation with a peak almost as high as the one at the α-helix region; these features are a consequence of its lack of a side chain21. On the other hand, proline—an imino acid—has two highly-populated states, with a slightly higher probability at the PPII conformation than at the α-helix conformation. It belongs to the set of structurally restricted amino acids composed by {Ile, Pro, Thr, Val}, which have an extremely low probability of occupying the right-hand side of the Ramachandran map. Indeed, the corresponding plots (Fig. 1) show few points within the quadrants I and IV (ϕ > 0). The conformational restrictions of proline arise from its pyrrolidine ring, whose flexibility is coupled to the backbone22. Isoleucine, threonine, and valine are the only amino acids with C-β branching, which means that they have more bulkiness near to the protein backbone than the rest of amino acids23. They also have a local maximum within the β-sheet region—shown as red shaded peaks in Fig. 1—a feature only shared with the three aromatic residues, Phe, Tyr, Trp, and Leu. The remaining amino acids occupy the allowed regions in a generic fashion20,24, whose distributions agree with the original Ramachandran and co-workers explanation in terms of steric clashes25.
All these observations point to the qualitative aspects of the distributions. However, a systematic comparison of the twenty Ramachandran distributions requires the use of a quantitative evaluation of their similarities. In the following subsection, we show a distance matrix accounting for dissimilarities between the secondary-structural trends of amino acids.
Secondary-structural vs BLOSUM replacements
A quantitative assessment of the similarities between the twenty distributions P X (ϕ, ψ) requires a distance measure. We used the city-block distance, which can be used to assess the differences in discrete frequency distributions. It gives more weight to the most probable dihedral conformations of the Ramachandran distributions.
Each amino acids X has a set of twenty distances, D X , including with itself, (in which case ||P X − P X || = 0):
The most plausible secondary-structural replacement to X is that amino acid Y having the smallest positive distance to X, or the minimum positive value from the set of distances: \({min}_{+}\{{D}_{X}\}\). That \({{\rm{\min }}}_{+}\{{D}_{X}\}=||{P}_{X}-{P}_{Y}||\) does not imply necessarily that \({{\rm{\min }}}_{+}\{{D}_{Y}\}=||{P}_{Y}-{P}_{X}||\). In other words, the structural replacement is not always a reciprocal operation; hence if Y is the replacement of X, we denote this by X → Y. In the case of a reciprocal replacement, we denote it by X ↔ Y.
The secondary-structural distance matrix between the amino acids is shown in Fig. 2. The proximity between amino acids is given by a color scheme: the smallest distance is represented in yellow, and the largest distance in blue, with intermediate values in green. We found open subsets by a nearest-neighbor criterion: any element within an open subset has exactly the remaining elements of said subset as its nearest neighbors—the procedure is explained in the methods section. For instance, the simplest open subset is composed by two elements for which the other one is the closest element—i.e. those elements for which D min (P X , P Y ) = D min (P Y , P X ) or, equivalently, X ↔ Y.
We found the following open sets (Fig. 3): a five-member set including a couple of two-member subsets: S I = {{Arg, Lys}, {Glu, Gln}, Leu}—in yellow; a three-member set containing a two-member set, S II = {Trp, {Phe, Tyr}}—in green; and a pair of two-member sets: S III = {Val, Ile}, and S IV = {Asn, Asp}—in cyan and magenta, respectively. Within this topology, Met appears as a boundary element of the first set S I ; Fig. 3 shows that Met first five neighbors are exactly the elements of S I . In turn, every residue in S I has Met as the fifth neighbor but Glu, which has Ala closer; this proximity may result from Ala and Glu being the strongest α-helix formers, as their respective \({P}_{X}^{{\rm{\max }}}\) values indicate (Table 1). The S I group includes aliphatic saturated side chains, while S II contains the aromatic residues. Adjacent to these two major sets we found residues sharing their physiochemical characteristics—as shown by their close distances to the main groups in the distance matrix (Fig. 2). Specifically, four residues have their nearest neighbor within a major open set: Ala have its first neighbor in S I , whereas His, Thr, and Cys have their first neighbor in S II . Those amino acids outside an open set or its boundaries were considered structurally idiosyncratic: Ala, Cys, His, Gly, Ser, Pro, and Thr. Gly and Pro are the farthest ones from any other residue, as the last column of Fig. 3 shows. Certainly, these amino acids populate the Ramachandran map in a unique way. The Ramachandran distribution of glycine is widespread over the allowed regions; while Pro is the most structurally restricted. Alanine has twice the probability of forming an α-helix (\({P}_{Ala}^{{\rm{\max }}}=\mathrm{0.437 \% }\) from Table 1) than any other residue (\({P}_{aver\ne Ala}^{{\rm{\max }}}=0.214 \% \)). The Ramachandran distribution of Thr has four peaks around the β and π regions unlike any other residue, including the C-β branched amino acids (Fig. 1). While Thr is chemically similar to Ser26, they have different structural propensities. According to our distance matrix (Fig. 2), Thr is closer to Tyr & Phe, while Ser is closer to His & Arg. A recent study shows that the phosphorylation of Ser increases its propensity of forming PPII, whereas that of Thr has the opposite effect27. This result indicates that Ser and Thr are far from being ideal secondary structural replacements. In summary, our classification reflects the intrinsic structural trends of amino acids; in particular, the S I set and its adjacent elements Met and Ala are the same alpha formers found by Fujiwara et. al.28. Within the same scale, the aromatic set, S II , and its adjacent elements (Cis, Thr) and S III are beta formers. The remaining amino acids are turn/bend formers, including S IV and Gly, Ser, and Pro, most of which have the lowest \({P}_{X}^{max}\) values in Table 1.
Rows ordered according to the cityblock distance. Open sets are indicated by the same color code used in Fig. 1. The shadowed boxes contain the BLOSUM100 pair replacements. The procedure for determining an open set consists on finding rows with the same set of first neighbors. For instance, the first neighbor of Arg (top row) is Lys; after placing the Lys row under the top row, we see that they share the seven first neighbors (up to Trp). The third row corresponds to Arg second neighbor, i.e. Glu, which also shares the same first neighbors with the previous ones up to Trp. The fourth row corresponds to Arg third neighbor, i.e. Gln, whose fifth neighbour is Ala, unlike the previous rows. The fifth row corresponds to Arg fourth neighbor, i.e. Leu, which has all the previous rows as its first neighbors. In this way, the yellow box includes those elements whose first four neighbors are completely contained within the set. Methionine is a frontier element of this set: its first five neighbors are exactly the elements of the whole closed set; however, Glu does not include Met within its first five neighbours and for that reason Met is not contained in the set. The remaining open sets S II to S IV were obtained in the same way. Notice that Pro and Gly are the farthest residues from any other one, as a consequence of their structural propensity uniqueness.
More importantly, nevertheless, is the fact that an unexpected pattern emerged: our structurally similar pairs of amino acids matches with most BLOSUM matrices pair replacements29, which are shown as shadowed boxes in Fig. 3. More details about the substitution matrices are in the methods section. Our list of structural replacements is: Asn ↔ Asp, Phe ↔ Tyr, Lys ↔ Arg, Gln ↔ Glu, Ile ↔ Val, Met → Leu. In BLOSUM matrices, Thr and Ser are replacements. For all BLOSUM matrices, Gly, Pro, Cys, His, and Ala are idiosyncratic residues. In general, our set of structurally-similar amino acids coincide with most canonical residue substitutions given by scoring matrices such as BLOSUM62 and BLOSUM10029, and consensus replacements30. This is a remarkable finding considering the extremely low probability of randomly finding six out of seven replacement pairs: less than one in a 681 million, as detailed in the methods section. In consequence, our result reveals an underlying correlation between mutation matrices and structural propensities. Hence, the replacement rules implied by the secondary structure distance (Fig. 2) may be directly used for for exploring structural amino acid replacements in peptide design strategies.
We conclude that during evolution, mutational replacements occurred between structurally similar amino acids. Hence, mutations followed a process that privileges structure and hence preserves function. But BLOSUM and PAM substitution matrices give additional information about the mutational trends of amino acids. The diagonal of these matrices determine how easy is for an amino acid to be replaced. A large value means more resistance to change. However, our distance matrix (Fig. 2) has a diagonal of zeros. For studying the mutability, we explored a parameter that combines the statistical information at the \({P}_{X}^{{\rm{\max }}}\) with a basic extensive property.
Molecular mass and optimum evolutionary cost
Molecular mass is a fundamental extensive property that might have played a central role in defining the actual protein landscape. Previously, our group revealed a very high correlation (R = 0.98) between mass and the electronic energy of amino acids—excluding the two sulfur-containing side chains31. In the present study, we found a complex relationship between the amino acids mass M X and the structural trends via the probability at the most frequent conformational state, \({P}_{X}^{{\rm{\max }}}\); this quantity is given by the highest peak of each Ramachandran distribution—max(P X (ϕ, ψ)). \({P}_{X}^{{\rm{\max }}}\) corresponds to the most frequent conformation and, therefore, it is an indicator of structural persistence32.
The α-helix conformation is the highest peak for all amino acids (but proline) with alanine at the top as the strongest helix former. While mass has an overall poor correlation with \({P}_{X}^{{\rm{\max }}}\) (R = 0.05), we identified two main and opposite trends delimited by separate ranges of \({P}_{X}^{{\rm{\max }}}\): (a) \({P}_{X}^{{\rm{\max }}} > \mathrm{0.200 \% }\) defines the set of strong helix formers {Ala, Glu, Gln, Ile, Met, Leu, Lys, Arg, Val} (in descending order), with a negative correlation R = −0.61; and, (b) \({P}_{X}^{{\rm{\max }}}\le \mathrm{0.200 \% }\) defines the weak helix formers: {Trp, Asp, Phe, Tyr, Thr, His, Cys, Asn, Ser, Gly, Pro}, with a positive correlation of R = 0.76. The small set of C-β branched amino acids ({Ile, Thr, Val}) plus proline shows a correlation of R = 0.78 between mass and \({P}_{X}^{{\rm{\max }}}\). After excluding these four elements from the two main sets, their respective correlations rise to R = −0.87 for the strong helix formers, and to R = 0.87 for the set of weak helix formers. In strong helix formers, the negative correlation between \({P}_{X}^{{\rm{\max }}}\) and the molecular mass indicates that light side chains have a better chance of forming an alpha helix than heavy ones. These three correlations reveal a direct involvement of the molecular mass on the α-helical propensities of the amino acids.
A recent observation by Lehmann et. al. reports a negative correlation between the background frequency and codon degeneracy of amino acids with mass33. Seligmann already observed that the evolutionary rate of amino acid replacements correlates negatively with mass34. Accordingly, heavier amino acids are less frequent, which suggests that the genomes preserve a fundamental distribution ruled by simple energetics. Inverse correlations between the average amino acid biosynthetic cost and the levels of gene expression are consistent with natural selection to minimize costs35. Seligmann also shows a positive correlation (R = 0.80) between the molecular mass M X and the total energetic cost per amino acid (in ATPs)34, as reported by Akashi & Gojobori36. According to Lehmann et al., highly expressed proteins tend to use amino acids with relatively low synthetic costs33. Therefore, heavy amino acids are less frequent because they are biosynthetically more expensive. We found a further confirmation of this statement: the molecular mass grows with the number of biosynthetic steps, as shown in Fig. 4. The values proposed by Davis37, are included in Table 1 as B X . The number of biosynthetic steps has been proposed as a natural way of determining the evolutionary history of amino acids38, and so does the amino acids molecular mass. We found a correlation of R = 0.64 between mass and biosynthetic steps, which rises up to R = 0.88 after excluding the set of outliers {Asn, Asp, Gln, Glu} (Fig. 4).
Correlation between the molecular mass of the amino acids M X and their energetic cost as accounted by the number of biosynthetic steps B X proposed by Davis37. The outliers {Asn, Asp, Gln, Glu} are excluded from the Pearson’s correlation and from the linear interpolation.
In summary, we found a high correlation—by parts—between the molecular mass and the probability at the most frequent conformational state (\({P}_{X}^{{\rm{\max }}}\)). We also found a high correlation between mass and the number of biosynthetic steps (B X ). These correlations are consistent with the fact that evolution privileges energetically optimal costs34,39. Thus, in the quest for a physical quantity that can explain amino acid’s mutability, mass is irreplaceable as a fundamental measure of energetic cost.
Mass over the frequency at the most probable conformation correlates with mutability
The background frequency or natural abundance of amino acids, N X , may be indicative of their evolutionary age: more abundance reflects an early adoption in molecular evolution40. The values of N X were obtained from the PGD 1.1 database (Table 1). The quantity \({W}_{X}={P}_{X}^{{\rm{\max }}}\times {N}_{X}\) is an estimator of the maximum observations at the most frequent conformation. In this way, W X combines the probability at the most probable conformation with the background frequency. In the previous section we showed that an amino acid has less probability to be changed if it is more energetically expensive, and therefore mass directly measures the resistance to be changed. Additionally, less frequent amino acids are also less replaceable, indicating an inverse correlation with the mutability. Under these considerations, we define a “replacement inertia” as the mass M X weighted by W X : I X = M X /W X . It summarizes the energetic cost per number of observations at the most probable conformation. We hypothesize that I X might reflect the mutability of amino acids—i.e. the diagonal of substitution matrices (see more details in the Methods).
In order to test if I X reflects the mutability of amino acids, we selected thirty replacement matrices reported by the AAindex41: twenty-seven that were built from sequence alignments—including a selection of six PAM and eight BLOSUM matrices; two more that were crafted from force fields (THREADER and SAUSAGE)42; and a last one that was obtained from replacements at the genetic code level43. Supplementary Table S1 contains the list of matrices used in our survey. We computed the Pearson correlation coefficient between I X and each mutability, which is shown in Fig. 5; in this figure, the correlation with alignment-derived matrices is colored in blue; the correlation with force-field derived appears in purple; and the correlation with the genetic code based matrix is plotted in green.
Pearson correlation coefficients between the replacement inertia I X (Table 1) and the mutability of thirty replacement matrices. Alignment derived matrices are shown in blue, force field derived matrices in purple, and the genetic code derived matrix in green. See Supplementary Table S1 for the abbreviations.
We found a very strong average correlation between I X and the whole mutability set of \({\overline{R}}_{30}=0.85\). This average value can be explained by the strong correlation found between I X and the mutability of matrices derived from sequence alignments, which have values R > 0.78, as Fig. 5 shows. For the family of BLOSUM matrices, R values were obtained between 0.90 and 0.96, with an average correlation of \({\overline{R}}_{B}=0.92\). For PAM matrices, the correlation was lower with an average value of \({\overline{R}}_{P}=0.82\) for the six PAM matrices included in our survey.
On the other hand, the correlation between I X and the mutability of the THREADER substitution matrix was the lowest we found, R THREADER = 0.52. The second lowest correlation for was with the matrix based on the genetic code (R BENNER = 0.64). The other force field derived matrix gave a correlation of R SAUSAGE = 0.68. These low correlations may have an interesting explanation: while force field based substitution matrices do not include evolutionary information, BENNER matrix, on the other hand, assumes that the genetic code is the only determinant of amino acid substitutions. As a consequence, the underlying factors controlling these matrices are poorly reflected on I X . Therefore, we must conclude that the very high correlation between I X and the mutability of matrices derived from sequence alignments implies that molecular mass, abundance, and the most probable secondary structure conformation may have played a decisive role on shaping the molecular evolution of proteins.
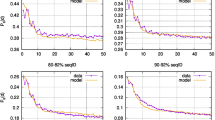
However, how significant an average correlation of \(\overline{R}=0.85\) between I X and the mutability set is? We evaluated the correlation coefficients between the mutability of all the substitution matrices, which yields a total of 430 correlations for the thirty matrices considered. The average value for these correlations is \({\overline{R}}_{430}=0.84\). This value differs little from \(\overline{R}\), which means that I X describes amino acids mutability as well as any the mutability of the accepted mutation matrices. The correlation matrix with significance levels for I X and the mutability of the whole set of matrices is shown in Supplementary Fig. S1. An excerpt of this plot is shown in Fig. 6, which includes the following matrices: BLOSUM30, BLOSUM62, BLOSUM100, PAM40, PAM160, and PAM250. This plot reveals that the correlations between PAM and BLOSUM fall within 0.70 and 0.83. Expectedly, correlations between matrices of the same family are higher, up to 0.96 for BLOSUM and up to 0.97 for PAM. It is surprising that I X had better simultaneous correlations with both matrix families than they have with each other. This observation holds for the eight BLOSUM and six PAM matrices included in our study, as shown in Supplementary Fig. S21.
Correlation matrix plot with significance levels between the replacement inertia (I X ) and the mutability of a representative set of BLOSUM and PAM matrices. The lower triangular matrix is composed by the bivariate scatter plots with a fitted smooth line. The upper triangular matrix shows the Pearson correlation plus significance level (as stars). Each significance level is associated to a symbol: p-values 0.001 (***), 0.01 (**), 0.05 (*). This plot was generated with the Performance Analytics package in R program57. The correlation matrix for the complete mutability set is plotted in Supplementary Fig. S1.
Our results indicate that amino acids mutability may be an evolutionary invariant that depends on the biosynthetic cost per amino acid and on the background frequency. These observations might have relevant consequences for future developments and improvements of the actual scoring matrices, as well on structure prediction and design.
Conclusions
Our study provides compelling evidence about the physiochemical nature of the substitution matrices. Taylor’s early work44 on evolutionary biochemistry45 proposes an integrative amino acid classification schema based on Dayhoff’s PAM matrix and properties such as volume and polarity. In a complementary way, our approach puts the evolutionary concepts closer to physiochemical properties, which might be helpful for treating proteins as integrated physical and historical wholes.
The main findings of the present work agree with accepted ideas about the molecular evolution of proteins. In the first place, we claim that secondary structural similarities resemble to a great extent the canonical replacements given by substitution matrices (Figs 2 and 3). We interpret this result as a manifestation of an underlying structural preservation principle according to which amino acids interchangeability is highly determined by their secondary structural similarity. It might be a consequence of the fact that less structurally important parts of a protein evolve faster than more important ones. In this way, conservative substitutions occur more frequently in evolution than more disruptive ones. Our result agrees with Koonin & Wolf view according to which the primary causes of protein evolution could have more to do with fundamental principles of protein folding than with unique biological functions46. In the second place, we showed that amino acids mutability is correlated with the replacement inertia I X (Fig. 5). Therefore, amino acids mutability depends on the biosynthetic cost, the most probable conformation, and the background frequency. Davis proposes that the timeline of genetically encoded amino acids correlates with the number of chemical reactions required to synthesize each amino acid37,38,47. As a consequence, the correlation between mass and biosynthetic steps (Fig. 4) indicates that the mutability of amino acids might be a timeline of protein evolution as well.
Undeniably, the biosynthetic cost, structural preservation, and frequency distribution of amino acids, all played a significant role in the molecular evolution of proteins. Indeed, two main selective factors determining the evolution of proteins are structural robustness against misfolding, and energy-cost efficiency46,48,49. Protein synthesis is very error-prone in comparison to DNA replication, and hence many folding-recognition mechanisms seem to have evolved to minimize costs of erroneous protein synthesis49. This energy-cost efficiency may explain why highly expressed proteins evolve slowly and at rates largely unrelated to their functions48.
We can summarize our two main findings in similar terms with the following optimal-efficiency principles: (a) amino acids interchangeability occurs by preserving the secondary structural propensity, and (b) the amino acid mutability depends directly on its biosynthetic energy cost, and inversely with its frequency at the most probable conformation. We believe that these two principles are the underlying rules governing the observed amino acid substitutions. They provide a unified interpretation to mutation matrices, outside the statistical realm alone. Our results also indicate that amino acids mutability might be an invariant scale that differs little from one substitution matrix to another (Supplementary Fig. S21). These results may offer a new understanding of the evolutionary processes determining the structure of proteins.
Finally, the statistical similarities between secondary structural propensities used here offer a viable methodology for systematically exploring amino acid structural replacements. For instance, one can determine a structural distance matrix limited to the β-strand region, which may differ from the one of the whole Ramachandran map. With this type of sectoral statistics one can envision new rules for the design of polypeptide chains.
Methods
Data source
We calculated the Ramachandran distributions from the protein geometry database PGD 1.1, retrieved in June 201616. We selected crystallized protein geometries with resolution equal or less than 2Å, a R-factor equals to 0.2, and a R-free maximum of 0.3. In order to avoid over-representation bias of some protein families, we used 7,398 proteins with a maximum identity of 25%. A total of 1,153,791 residues were considered.
Data analysis
The statistical analysis of the present work was implemented in Python 2.7 programming language50,51. A Python routine extracts the observed (ϕ, ψ) values from the PGD database for each amino acid (PGDread.py). The 2D optimization process was done with a routine that computes the cost function by changing the bin width equally for both dihedral variables Δ = Δϕ = Δψ, (MISE.py). The Ramachandran distribution histograms were computed and plotted with Matplotlib libraries (3DRamadistr.py)52. The cityblock distance was taken from the SCIPY package. A total of 600 code lines were written for the complete analysis shown here. The Python codes are available upon request.
Histogram optimization
Histograms are a type of non-parametric density estimates for which the number of parameters equals the number of data points53. A different approach uses analytic functions for obtaining smooth distributions that minimize low resolution and outliers effects54. The discrete (histogram) representation of the joint probability distribution P X (ϕ i , ψ j ) depends on the bin width of the dihedral variables, i.e. Δϕ and Δψ. A coarse binning size decreases the data noise but it might overpass relevant details of the structural information. On the other hand, a very fine grain bin size might highlight underlying statistical noise. The mean integrated squared error (MISE) can be estimated from the data through a cost function C(Δ). A histogram with the bin size that minimizes the MISE is optimal17. This method guarantees that a substantial increasing in the observations will further increase the accuracy of the histogram representation of probability distributions even more. The main assumption underlying this method is that the distribution can be represented by a smooth continuum function. Previous works have proven that Ramachandran distributions obey such assumption11. We assumed a regular partitioning of the Ramachandran maps i.e having the same bin size Δ for both dihedral variables: Δ = Δϕ = Δψ. The cost function for two variables is therefore given by
where the mean n and the variance v of the number of occurrences are given, respectively, by \(n=\frac{1}{N}{\sum }_{i}^{N}{n}_{i}\), and \(v=\frac{1}{N}{\sum }_{i}^{N}{({n}_{i}-n)}^{2}\). The obtained optimal bin value for each amino acid is Δ X (Table 1). We used the weighted average as the bin with for all the Ramachandran distributions: \(\overline{{\rm{\Delta }}}={\sum }_{X}^{20}{N}_{X}{{\rm{\Delta }}}_{X}/{\sum }_{X}^{20}{N}_{X}\). From the obtained Δ X values, \(\overline{{\rm{\Delta }}}={1.887}^{^\circ }\), which was approximated by the integer fraction 360°/190 ≃ 1.895°, i.e. we used 190 bins in each angular coordinate, for a total of 190 × 190 = 36,100.
Amino acid classification
We classified the amino acids according to the city-block (Manhattan) distance. Our grouping method takes advantage of the fact that a metric induces a topology on a set. Accordingly, we determined the topology induced by the city-block distance over the set of amino acids. The increasing distance between a given element X and the remaining ones determines an ordered list. Therefore, for the present case, we have twenty ordered lists, one for each amino acid. The intersection between the first neighbors of these lists gave us open subsets. An open subset consists on those elements such that, for every element within the subset, its neighbors belong to the same subset. Figure 3 reports the twenty ordered lists with an example about how to obtain open sets.
Substitution matrices and mutability
The most common method of evaluating the amino acid substitution patterns is through substitution matrices such as PAM55 or BLOSUM29. A typical substitution matrix has 20 × 20 elements, in which non-diagonal pairwise scores (log odds) represent the probability of one amino acid could be substituted by other in protein evolution. The diagonal scores of the matrix are estimators of amino acid mutability. For each amino acid, a greater score implies lesser possibilities to be substituted, on the other hand, lesser scores implies a greater chance to be substituted55,56. We used a set of thirty substitution matrices reported in the AAindex41 and NCBI (ftp://ftp.ncbi.nih.gov/blast/matrices/).
Probability of randomly finding six out of seven sets
Substitution matrices, such as BLOSUM62 & BLOSUM100, define seven replacement pairs of amino acids. Our structural similar pairs do coincide with six of them. We need an assessment of the probability for correctly obtaining six out of seven pairs. The probability of obtaining the first element of a pair is the number of elements of such pair (2) divided by the total of elements (14). Then, the probability of finding the match is the number of pair elements still in the set (1) divided by the total left (13). Hence, the combined probability of randomly finding the first pair out of seven is P1 = 2/14 × 1/13. By a similar reasoning, the probability of obtaining a second pair is P2 = 2/12 × 1/11, and so on. Therefore, the probability of simultaneously finding six out of seven pairs is \({\prod }_{i\mathrm{=1}}^{6}{P}_{i}\), or equivalently, \({\prod }_{k\mathrm{=2}}^{7}\frac{2}{2k(2k-1)}=\mathrm{1/681,080,400}=1.468\times {10}^{-9}\). In other words, there is a chance of one in 681 million of simultaneously obtaining six correct pairs from a set of seven pairs.
Change history
06 March 2018
A correction to this article has been published and is linked from the HTML and PDF versions of this paper. The error has been fixed in the paper.
References
Sikosek, T. & Chan, H. S. Biophysics of protein evolution and evolutionary protein biophysics. Journal of The Royal Society Interface 11, 20140419 (2014).
Bloom, J. D., Labthavikul, S. T., Otey, C. R. & Arnold, F. H. Protein stability promotes evolvability. Proceedings of the National Academy of Sciences 103, 5869–5874 (2006).
Yu, J.-F. et al. Natural protein sequences are more intrinsically disordered than random sequences. Cellular and Molecular Life Sciences 15, 2949–2957 (2016).
Worth, C. L., Gong, S. & Blundell, T. L. Structural and functional constraints in the evolution of protein families. Nature Reviews Molecular Cell Biology 10, 709–720 (2009).
Levy, E. D., Erba, E. B., Robinson, C. V. & Teichmann, S. A. Assembly reflects evolution of protein complexes. Nature 453, 1262–1265 (2008).
Yang, Y. et al. Sixty-five years of the long march in protein secondary structure prediction: the final stretch? Briefings in Bioinformatics bbw 129 (2016).
Orengo, C. A. & Thornton, J. M. Protein families and their evolution—a structural perspective. Annu. Rev. Biochem. 74, 867–900 (2005).
Dokholyan, N. V. & Shakhnovich, E. I. Scale-free evolution. In Power Laws, Scale-Free Networks and Genome Biology, 86–105 (Springer, 2006).
Ramachandran, G. t. & Sasisekharan, V. Conformation of polypeptides and proteins. Advances in protein chemistry 23, 283–437 (1968).
Hollingsworth, S. A. & Karplus, P. A. A fresh look at the Ramachandran plot and the occurrence of standard structures in proteins. Biomolecular concepts 1, 271–283 (2010).
Ting, D. et al. Neighbor-dependent Ramachandran probability distributions of amino acids developed from a hierarchical Dirichlet process model. PLoS computational biology 6, e1000763 (2010).
Levitt, M. Conformational preferences of amino acids in globular proteins. Biochemistry 17, 4277–4285 (1978).
Koehl, P. & Levitt, M. Structure-based conformational preferences of amino acids. Proceedings of the National Academy of Sciences 96, 12524–12529 (1999).
Hollingsworth, S. A., Berkholz, D. S. & Karplus, P. A. On the occurrence of linear groups in proteins. Protein Science 18, 1321–1325 (2009).
DeBartolo, J., Jha, A., Freed, K. F. & Sosnick, T. R. Local Backbone Preferences and Nearest-Neighbor Effects in the Unfolded and Native States. Protein and Peptide Folding, Misfolding, and Non-Folding 79–98 (2012).
Berkholz, D. S., Krenesky, P. B., Davidson, J. R. & Karplus, P. A. Protein Geometry Database: a flexible engine to explore backbone conformations and their relationships to covalent geometry. Nucleic Acids Res. 38, D320–D325 (2010).
Shimazaki, H. & Shinomoto, S. A method for selecting the bin size of a time histogram. Neural Computation 19, 1503–1527 (2007).
Shen, Y., Delaglio, F., Cornilescu, G. & Bax, A. Talos+: a hybrid method for predicting protein backbone torsion angles from nmr chemical shifts. Journal of biomolecular NMR 44, 213–223 (2009).
Shen, Y. & Bax, A. Protein structural information derived from nmr chemical shift with the neural network program talos-n. In Artificial Neural Networks, 17–32 (Springer, 2015).
Hovmöller, S., Zhou, T. & Ohlson, T. Conformations of amino acids in proteins. Acta Crystallographica Section D: Biological Crystallography 58, 768–776 (2002).
Ho, B. K. & Brasseur, R. The ramachandran plots of glycine and pre-proline. BMC structural biology 5, 14 (2005).
Ho, B. K., Coutsias, E. A., Seok, C. & Dill, K. A. The flexibility in the proline ring couples to the protein backbone. Protein Science 14, 1011–1018 (2005).
Betts, M. J. & Russell, R. B. Amino acid properties and consequences of substitutions. Bioinformatics for geneticists 317, 289 (2003).
Ho, B. K., Thomas, A. & Brasseur, R. Revisiting the ramachandran plot: Hard-sphere repulsion, electrostatics, and h-bonding in the α-helix. Protein Science 12, 2508–2522 (2003).
Ramachandran, G. & Ramakrishnan, C. t. & Sasisekharan, V. Stereochemistry of polypeptide chain configurations. Journal of molecular biology 7, 95 (1963).
Bohórquez, H. J. et al. Electronic energy and multipolar moments characterize amino acid side chains into chemically related groups. The Journal of Physical Chemistry A 107, 10090–10097 (2003).
Kim, S.-Y., Jung, Y., Hwang, G.-S., Han, H. & Cho, M. Phosphorylation alters backbone conformational preferences of serine and threonine peptides. Proteins: Structure, Function, and Bioinformatics 79, 3155–3165 (2011).
Fujiwara, K., Toda, H. & Ikeguchi, M. Dependence of α-helical and β-sheet amino acid propensities on the overall protein fold type. BMC structural biology 12, 18 (2012).
Henikoff, S. & Henikoff, J. G. Amino acid substitution matrices from protein blocks. Proceedings of the National Academy of Sciences 89, 10915–10919 (1992).
Bordo, D. & Argos, P. Suggestions for “safe” residue substitutions in site-directed mutagenesis. Journal of molecular biology 217, 721–729 (1991).
Bohórquez, H. J., Cárdenas, C., Matta, C. F., Boyd, R. J. & Patarroyo, M. E. Methods in biocomputational chemistry: a lesson from the amino acids. Quantum Biochemistry 403–421.
Chatterjee, P. & Sengupta, N. Effect of the a30p mutation on the structural dynamics of micelle-bound α synuclein released in water: a molecular dynamics study. European Biophysics Journal 41, 483–489 (2012).
Lehmann, J., Libchaber, A. & Greenbaum, B. D. Fundamental amino acid mass distributions and entropy costs in proteomes. Journal of Theoretical Biology 410, 119–124 (2016).
Seligmann, H. Cost-minimization of amino acid usage. Journal of molecular evolution 56, 151–161 (2003).
Raiford, D. W. et al. Do amino acid biosynthetic costs constrain protein evolution in saccharomyces cerevisiae? Journal of molecular evolution 67, 621–630 (2008).
Akashi, H. & Gojobori, T. Metabolic efficiency and amino acid composition in the proteomes of escherichia coli and bacillus subtilis. Proceedings of the National Academy of Sciences 99, 3695–3700 (2002).
Davis, B. K. Evolution of the genetic code. Progress in biophysics and molecular biology 72, 157–243 (1999).
Griffiths, G. Cell evolution and the problem of membrane topology. Nature Reviews Molecular Cell Biology 8, 1018–1024 (2007).
Guilloux, A. & Jestin, J.-L. The genetic code and its optimization for kinetic energy conservation in polypeptide chains. Biosystems 109, 141–144 (2012).
Brooks, D. J., Fresco, J. R., Lesk, A. M. & Singh, M. Evolution of amino acid frequencies in proteins over deep time: inferred order of introduction of amino acids into the genetic code. Molecular Biology and Evolution 19, 1645–1655 (2002).
Kawashima, S. & Kanehisa, M. Aaindex: amino acid index database. Nucleic acids research 28, 374–374 (2000).
Dosztanyi, Z. & Torda, A. E. Amino acid similarity matrices based on force fields. Bioinformatics 17, 686–699 (2001).
Benner, S., Cohen, M. A. & Gonnet, G. H. Amino acid substitution during functionally constrained divergent evolution of protein sequences. Protein Engineering 7, 1323–1332 (1994).
Taylor, W. R. The classification of amino acid conservation. Journal of theoretical Biology 119, 205–218 (1986).
Harms, M. J. & Thornton, J. W. Evolutionary biochemistry: revealing the historical and physical causes of protein properties. Nature Reviews Genetics 14, 559–571 (2013).
Koonin, E. V. & Wolf, Y. I. Constraints, plasticity, and universal patterns in genome and phenome evolution. In Evolutionary Biology–Concepts, Molecular and Morphological Evolution, 19–47 (Springer, 2010).
Davis, B. K. Molecular evolution before the origin of species. Progress in biophysics and molecular biology 79, 77–133 (2002).
Drummond, D. A., Bloom, J. D., Adami, C., Wilke, C. O. & Arnold, F. H. Why highly expressed proteins evolve slowly. Proceedings of the National Academy of Sciences of the United States of America 102, 14338–14343 (2005).
Drummond, D. A. & Wilke, C. O. The evolutionary consequences of erroneous protein synthesis. Nature Reviews Genetics 10, 715–724 (2009).
van Rossum, G. & de Boer, J. Linking a stub generator (ail) to a prototyping language (python). In Proceedings of the Spring 1991 EurOpen Conference, Troms, Norway, 229–247 (1991).
Python Software Foundation. Python language reference. URL http://www.python.org.
Hunter, J. D. Matplotlib: A 2d graphics environment. Computing In Science & Engineering 9, 90–95 (2007).
Shapovalov, M. V. & L., D. J. R. Non-Parametric Statistical Analysis Of The Ramachandran Map. Biomolecular Forms and Functions: A Celebration of 50 Years of the Ramachandran Map 76 (2013).
Lovell, S. C. et al. Structure validation by C α geometry: ϕ, ψ and C β deviation. Proteins: Structure, Function, and Bioinformatics 50, 437–450 (2003).
Dayhoff, M. O. & Schwartz, R. M. A model of evolutionary change in proteins. In In Atlas of protein sequence and structure (Citeseer, 1978).
Valdar, W. S. Scoring residue conservation. Proteins: Structure, Function, and Bioinformatics 48, 227–241 (2002).
Peterson, B. G. et al. Performanceanalytics: Econometric tools for performance and risk analysis. r package version 1.4. 3541 (2014).
Acknowledgements
We would like to thank Professor Mario Amzel for his insightful comments on the paper.
Author information
Authors and Affiliations
Contributions
C.F.S. and H.J.B. proposed the project and developed the methodology of the study. H.J.B. wrote the Python codes. C.F.S. and H.J.B. carried out computations. C.F.S. and H.J.B. analyzed the data. M.E.P. supervised the project. H.J.B. wrote the manuscript whose final version include contributions by all authors.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A correction to this article is available online at https://doi.org/10.1038/s41598-018-21981-y.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bohórquez, H.J., Suárez, C.F. & Patarroyo, M.E. Mass & secondary structure propensity of amino acids explain their mutability and evolutionary replacements. Sci Rep 7, 7717 (2017). https://doi.org/10.1038/s41598-017-08041-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-08041-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.