Abstract
The ‘brain regionalization genes’ Six3/6, Otx, Pax2/5/8, Gbx, and Hox1 are expressed in a similar fashion in the deuterostome, ecdysozoan, and the cephalopod brain, questioning whether this holds also true for the remaining Mollusca. We investigated developmental Gbx-expression in representatives of both molluscan sister groups, the Aculifera and Conchifera. Gbx is expressed in the posterior central nervous system of an aculiferan polyplacophoran and solenogaster but not in a conchiferan bivalve suggesting that Gbx, together with Six3/6, Otx, Pax2/5/8, and Hox1, is involved in central nervous system regionalization as reported for other bilaterians. Gbx is, however, also expressed in the anterior central nervous system, i.e. the anlagen of the cerebral ganglia, in the solenogaster, a condition not reported for any other bilaterian so far. Strikingly, all Gbx-orthologs and the other ‘posterior brain regionalization genes’ such as Pax2/5/8 and Hox1 are expressed in the mantle that secretes shell(s) and spicules of mollusks (except cephalopods). In bivalves, the ancestral condition has even been lost, with Gbx and Pax2/5/8 not being expressed in the developing central nervous system anymore. This suggests an additional role in the formation of the molluscan shell field(s) and spicule-bearing cells, key features of mollusks.
Similar content being viewed by others
Introduction
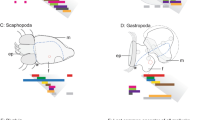
Mollusca is one of the largest bilaterian clades and accounts for an incredible diversity of body plans. Recent phylogenomic studies have supported a sister group relationship of the Aculifera and the Conchifera1,2,3. Aculiferans include the worm-shaped aplacophoran Solenogastres (Neomeniomorpha) and Caudofoveata (Chaetodermomorpha). These clades are characterized by a mantle covered with cuticle and mineralized sclerites (spicules) instead of a shell or shell plates. Polyplacophora (chitons) constitutes the third aculiferan clade and exhibits eight shell plates surrounded by cuticle and, in some cases, scales and/or spicules. All remaining molluscan class-level taxa, such as Scaphopoda, Bivalvia, Gastropoda, Monoplacophora, and Cephalopoda, belong to the Conchifera having a single shell, except for bivalves with two shell valves. While adult bivalves and scaphopods are benthic animals, many with a pronounced foot, the majority of cephalopods has internalized their shell in adaption to a life style as motile predators. Among conchiferans the phylogenetic interrelationships are far from being settled, rendering it difficult to infer how body plans have evolved4.
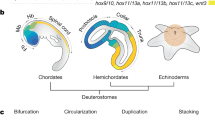
The molluscan central nervous system (CNS) is composed of two pairs of longitudinal nerve cords (tetraneurous) and it is centralized to different degrees. Among the Aculifera, polyplacophorans have rather few ganglia, while solenogastres and caudofoveates exhibit more ganglia in particular in the anterior body region5. Except for monoplacophorans that possess a CNS with few ganglia, the conchiferan CNS is ganglionated to a higher degree, most apparent in the compact cephalopod brain6,7,8. In bilaterian model organisms such as fruit fly and mouse certain homeobox genes are expressed in a staggered fashion in the CNS during ontogenesis9, 10. These genes are involved in the regionalization of the developing CNS into individual domains such as the vertebrate midbrain-hindbrain boundary9. Six3/6 and Otx are expressed in the anteriormost CNS regions, followed by an intermediate Pax2/5/8-expression domain and a posterior Gbx-expression domain with co-expression of anterior Hox genes9. Recent studies unraveled that this gene expression profile is also present in cephalopods11,12,13,14,15. Data on other mollusks, in particular with respect to the expression of the ANTP-class gene Gbx/unplugged (gastrulation brain homeobox), are scarce, but of importance to understand the evolution and development of the molluscan CNS.
In order to infer whether Gbx is also expressed in the CNS of non-cephalopod mollusks, we investigated its expression during ontogenesis in the aculiferans Acanthochitona crinita (a polyplacophoran) and Wirenia argentea (an aplacophoran solenogaster), as well as in the conchiferan protobranch bivalve Nucula tumidula.
Methods
Collection and culture of animals
Adults of the polyplacophoran Acanthochitona crinita (Pennant, 1777) were collected in summer 2012 in the intertidal zone close by the Station Biologique Roscoff (Roscoff, France). Animals were spawned and developmental stages were reared at 20 °C as described previously15. Adults of the aplacophoran Wirenia argentea Odhner, 1921 and the bivalve Nucula tumidula Malm, 1861 were collected at depths of 190–226 m and 227–312 m with a hyperbenthic sled on the silty seafloor in Hauglandsosen and Hjeltefjorden, respectively (Bergen, Norway) in March 2012, from November 2012 to January 2013, and from November to December 2013. Animals were spawned and developmental stages were reared at 7 °C in the dark as described previously15, 16.
RNA extraction and fixation of animals for in situ hybridization experiments
For all three species, several hundred individuals of different developmental stages were investigated. For the polyplacophoran Acanthochitona crinita and the bivalve Nucula tumidula developmental stages ranging from gastrulae to postmetamorphic individuals were collected. For the solenogaster Wirenia argentea adults as well as freshly hatched test-cell larvae (0–1 days after hatching (dph), early test-cell larvae (6–7 dph), mid-stage test-cell larvae (10–11 dph), and late test-cell larvae (14–15 dph)) were collected. The above-mentioned larvae were fixed for in situ hybridization experiments and used for RNA extraction as previously described15,16,17.
RNAseq and transcriptome assembly
Total RNA from pooled developmental stages of Acanthochitona crinita and Nucula tumidula, respectively, and pooled RNA from developmental stages and adults of Wirenia argentea was sequenced by Illumina technology (Eurofins, Ebersberg, Germany). Paired-end reads of an average read length of 100 bp were obtained and were subsequently filtered (rRNA removal; see refs 15 and 16 for details on transcriptome assembly). Adapter and low quality sequences were trimmed, normalized, and assembled de novo into contigs with the assembler Trinity18 or IDBA-tran, Version 1.1.1 in case of W. argentea 16, 19.
Alignment and phylogenetic analysis
Homeodomain amino acid sequences of Gbx, Dlx, Hox1, Hox3, Cdx, and Lhx2/4 were retrieved from the homeobox database (http://homeodb.cbi.pku.edu.cn/) and GenBank (accession numbers provided in Supplementary Fig. 1)20, 21. Gbx amino acid sequences were used in BLAST searches against the assembled transcriptomes of Acanthochitona crinita, Wirenia argentea, and Nucula tumidula. Predicted amino acid sequences of Acr-Gbx, War-Gbx, and Ntu-Gbx and the above-mentioned homeodomains of other homeobox sequences were aligned using MAFFT v7.123b22. The alignment was edited manually with Aliview v1.18 and used for the phylogenetic analysis. The phylogenetic analysis presented in this study includes the predicted amino acid sequences of cloned Gbx orthologs of A. crinita, W. argentea, and N. tumidula. The bayesian phylogenetic analysis was carried out with MrBayes v3.2.5 with LG model of amino acid replacement23 estimated with Prottest3 v3.4.2.24, gamma-distributed rates, 22.500.000 generations and sampling frequency of 1.000. The phylogenetic tree was manually rooted using FigTree v1.4.1.25.
Molecular isolation of RNA transcripts
First-strand cDNA synthesis of the RNA pooled from different developmental stages of Acanthochitona crinita, Wirenia argentea, and Nucula tumidula, respectively, was carried out by reverse transcription using the First strand cDNA Synthesis Kit for rt-PCR (Roche Diagnostics GmbH, Mannheim, Germany; see also15, 16 for the experimental procedure). Identified Gbx orthologs were used to design gene-specific primers and PCR products were size-fractioned by gel electrophoresis, gel bands of the expected lengths were excised and cleaned up using a QIAquick Gel Extraction Kit (QIAgen, Hilden, Germany). By insertion into pGEM-T Easy Vectors (Promega, Mannheim, Germany) cleaned-up products were cloned. Plasmid minipreps were grown overnight, cleaned-up with the QIAprep Spin MiniprepKit (QIAgen), and sent off for sequencing. A. crinita-Gbx (Acr-Gbx), W. argentea-Gbx (War-Gbx), and N. tumidula-Gbx (Ntu-Gbx) sequences were identified using the BLASTx algorithm screening the database of the NCBI. All three cloned nucleotide sequences as well as their deduced amino acid sequences were submitted to GenBank (Accession numbers: Acr-Gbx: KY500990, War-Gbx: KY500991, Ntu-Gbx: KY500992).
Probe synthesis and whole-mount in situ hybridization
The detailed experimental procedure was described previously15, 16. Riboprobe templates were amplified via standard PCR from miniprepped plasmids using M13 forward and reverse primers. In vitro transcription reactions were performed with these templates, digoxigenin-UTP (DIG RNA Labeling Kit, Roche Diagnostics), and SP6/T7 polymerase (Roche Diagnostics GmbH) for the syntheses of antisense riboprobes, according to the manufacturer’s instructions. In whole-mount in situ hybridization experiments, specimens were rehydrated into PBT (phosphate buffered saline +0.1% Tween-20) and treated with Proteinase-K at 37 °C for 10 min. Developmental stages of Acanthochitona crinita were Proteinase-K treated with 60 µg/ml in PBT and those of Wirenia argentea and Nucula tumidula with 10 µg/ml in PBT. Specimens were prehybridized in hybridization buffer for 4 h either at 65 °C (A. crinita) or at 56 °C (W. argentea, N. tumidula). Hybridization was performed at the same temperature with probe concentrations ranging between 0.5 and 1 μg/ml for 21–24 h. A DIG-labeled AP-antibody was used at a dilution of 1:5000 in blocking solution at 4 °C over night. Color development in the NBT/BCIP/Alkaline Phosphatase buffer solution took 6–24 hrs at 4 °C. For A. crinita and N. tumidula, a minimum of 40 individuals per stage was investigated, while a minimum of 20 individuals per stage were used for W. argentea. Fluorescent in situ hybridization was performed with Fast Blue (Sigma-Aldrich) on A. crinita as described previously27, 28. The majority of whole-mount preparations was cleared in a solution of benzyl-benzoate and benzyl alcohol, mounted on objective slides, and analyzed. After in situ hybridization, some specimens of A. crinita were embedded in O. C. T. medium (VWR, Vienna, Austria) and cut into 15–30 μm cryosections with a cryotome (Leica CM 3050S). For counterstains of cell nuclei, sections were stained with DAPI (Sigma- Aldrich, St. Louis, MO, USA), washed in phosphate buffered saline and subsequently mounted in Fluoromount G (Southern Biotech, Birmingham, Alabama, USA). Preparations were documented with an Olympus BX53 Microscope (Olympus, Hamburg, Germany). In addition, developmental stages were scanned with a Leica confocal SP5 II microscope (Leica Microsystems, Wetzlar, Germany) using bright-field, autofluorescence, and reflection mode scans to understand the precise location of transcripts29. If necessary, images were processed with Adobe Photoshop 9.0.2 software (Adobe Systems, Inc; San Jose, CA, USA) to adjust for contrast and brightness (Figs 1, 2, 3 and 4). Sketch drawings were created with Adobe Illustrator CC 2015.1.0 (Adobe Systems, Inc). Fluorescence labeling of filamentous F-actin with Alexa Fluor 488 was performed on 7 dph old larvae of W. argentea as described previously26.
Expression of Gbx during embryonic and early larval development of the polyplacophoran Acanthochitona crinita. Dorsal (d)-ventral (v), anterior (a)-posterior (p), and left (l)-right (r) axes indicate the orientation. Asterisks mark the blastopore/mouth opening. (a) Late gastrulae (4 hpf; ventro-posterior view) expresses Gbx in cells in the ventral (encircled) and dorsal regions (arrowheads). (b-c) In further developed early trochophore larvae (12 hpf) Gbx–expressing cells (arrowheads) are located in the ventral hyposphere (hp). (d–i) Mid-stage trochophore (35 hpf). (d,e) This dorsal (d) and more ventral optical section (e) show Gbx-expressing cells (arrowheads) in seven rows of the shell fields. Note the unspecific staining in the prototroch. (f,g) Gbx is also expressed in the ventro-lateral hyposphere in the CNS, an area that is highlighted and 3-D reconstructed in (g) (red-dashed box). Gbx is expressed in the anterior pedal nerve cord (pc) and the entire visceral nerve cord (vc). Gbx is expressed in the region of the pedal nerve cord commissure (arrowheads). (h) Note Gbx-expression in the dorsal shell fields and the nervous system of the foot (arrowhead). (i) Gbx–expression in the visceral nerve cord (arrowheads). Abbreviations: ep, episphere; pt, prototroch. Scale bars: 20 µm.
Optical section series of Gbx-expression in the metamorphic competent trochophore larva (65 hpf) of the polyplacophoran Acanthochitona crinita. Dorsal (d)-ventral (v), anterior (a)-posterior (p), and left (l)-right (r) axes indicate the orientation. Anterior faces up in all micrographs. (a–c) Sequence of three consecutive optical sections through a metamorphic competent trochophore larva. Gbx is expressed in cells of the shell fields, close to the mouth (asterisk) in the area of the pedal commissure, and in the spicule-bearing cells of the perinotum (arrowhead). Gbx-expressing cells are located in the outermost cell layer of the mantle that constitutes the shell fields. A reflection scan of the boxed region shown in (b) is highlighted in (d). (d) Reflection scan showing Gbx-expression in six rows of cells (arrowheads) in the shell fields (Dapi nuclear counterstain in blue). The seventh row is not included in this micrograph. (e) Fluorescent in situ hybridization (FISH) staining showing Gbx–expression in the spicule-bearing cells (arrowheads) in the area of the prospective perinotum (DAPI nuclear counterstain in blue). Staining in the prototroch (pt) is unspecific. (f) Cells of the pedal (pc) and visceral nerve cords (vc) express Gbx. Scale bars: 20 µm.
Expression of Gbx during ontogenesis of Wirenia argentea. Anterior (a)-posterior (p), dorsal (d)-ventral (v), and left (l)-right (r) axes indicate the orientation. Anterior faces up in all micrographs. Asterisks mark the mouth opening. (a,b) Freshly hatched larvae (0–1 dph) express Gbx in a dorso-ventral gradient with the highest expression in the ventral epidermal cells close to the pseudo-blastopore (pb). The early embryo possesses flattened and large test-cells (tc) that do not express Gbx. (c–e) Early larvae (6–7 dph) express Gbx ventral laterally in epidermal cells (2) that line the peri-imaginal space (pis). This expression domain extends towards the lateral portions of the mouth and also includes subepidermal cells (2). Two bilateral groups of Gbx-expressing cells (1) are located ventrally at the posterior pole of the outgrowing trunk. These expression domains might be associated with the developing pedal nerve cords. Note that the trunk is less retracted in (e) compared to (d) and therefore Gbx-expression domain “1” is clearly distinguishable from other expression domains. The developing cerebral ganglia (cg) show faint Gbx-expression. (f–g) Mid-stage larvae (10–11 dph) express Gbx in the developing pedal nerve cords. Gbx is still expressed in epidermal cells (2) which line the peri-imaginal space and in subepidermal cells close to the latter cells (2). Some individuals show Gbx-expression adjacent to the developing foregut (fg) (arrowheads). Gbx is also expressed in the region of the developing cerebral ganglia. (h,i) Late larvae (14–15 dph) express Gbx in the developing pedal nerve cords. Gbx is expressed ventro-laterally in epidermal cells (2) adjacent to the peri-imaginal space and in subepidermal cells close to the latter cells (2). This expression domain extends towards the lateral portions of the mouth. Gbx-expression is also visible adjacent to the developing foregut (arrowheads). Abbreviations: pc, pedal nerve cords. Scale bars: 50 µm.
Expression of Gbx during ontogenesis of the bivalve Nucula tumidula. Anterior (a)-posterior (p), dorsal (d)-ventral (v) and left (l)-right (r) axes indicate the orientation. The blastopore (asterisk) leads to the definitive mouth. (a) In early test-cell larva (3 dpf) Gbx is expressed in the ectoderm on the dorsal and lateral sides. (b) In 8 dpf old test-cell larvae Gbx is expressed in the ventral mantle margins (arrowheads). (c,d) In further developed specimens (12 dpf) Gbx is expressed laterally in the mantle. The large and flattened ciliated test-cells (tc) constitute the outer cellular layer of the larva and do not express Gbx. (e) Late test-cell larvae (12 dpf) express Gbx in the region of the ventral (lower arrowhead) and lateral mantle (upper arrowhead). Note the mantle cavity (mc), the perivisceral cavity (pc), and the stomache (stm). (f) Settled, early postmetamorphic individuals (22 hpf) express Gbx in parts of the mantle close to the foot. The test-cells have been shed during metamorphosis and will eventually be ingested. Scale bars: 30 µm (except Change “I” into “f”: 50 µm).
Results
Gbx gene orthologs and phylogenetic analysis
Acr-Gbx, War-Gbx, and Ntu-Gbx show high sequence similarity with their bilaterian orthologs as revealed by multiple amino acid sequence alignment. The phylogenetic analysis shows that all sequences cluster with their bilaterian orthologs with high support (Supplementary Figs 1 and 2).
Gbx-expression in the polyplacophoran Acanthochitona crinita
Gbx is expressed in cells that are located in ventral and dorsal regions of late gastrulae (Fig. 1a). Gastrulae subsequently develop into early trochophore larvae that are characterized by a ciliary band, the prototroch, that divides the anterior episphere from the posterior hyposphere (Figs 1b,c and 5a). Trochophore larvae possess an apical organ with a ciliary tuft in the anterior episphere, a mouth in the anterior ventral hyposphere and seven shell fields that develop on the dorsal hyposphere (Fig. 5a). In early trochophore larvae Gbx–expressing cells are only located in the ventral hyposphere (Figs 1b,c and 5a). Mid-stage trochophore larvae express Gbx in cells of the seven shell fields (Fig. 1d,e,h). Each shell field is ridge-like and all Gbx-expressing cells are located in the epidermis (Supplementary Fig. 3a,c). Gbx-transcripts are also present in the distal-most portion of spicule-bearing cells of the dorsal episphere (red-lined arrowhead in Supplementary Fig. 3a). In addition, Gbx-expressing cells are associated with the visceral and pedal nerve cords (Fig. 1f–i; Supplementary Fig. 3c–f). While cells along the entire length of the visceral nerve cords express Gbx, only cells along the anteriormost portion of the pedal nerve cord express Gbx (Fig. 1f,g,i). In the anteriormost portion of the hyposphere, Gbx is expressed in the pedal commissures (arrowheads in Fig. 1g). In metamorphic competent trochophore larvae, Gbx-expressing cells are located in seven rows in the prospective shell fields of the hyposphere as seen in mid-stage trochophores (Figs 2a,b,d and 5b; c.f. Supplementary Fig. 3a). Gbx-expressing cells are slender and located in the anterior portion of each shell field (Fig. 2d,e). Visceral and pedal nerve cords exhibit Gbx-expressing cells, however, expression is less prominent compared to earlier developmental stages. In contrast to the visceral nerve cords, the pedal nerve cords only exhibit Gbx-expressing cells in the anteriormost region as seen in mid-stage trochophores (Figs 2f and 5b; c.f. Supplementary Fig. 3c). Additional Gbx-expressing cells are located in the anteriormost portion of the hyposphere, in the region of the pedal commissures (Figs 2b,c and 5b). Spicule-bearing cells in the prospective perinotum, i.e. the areas surrounding the shell fields, also express Gbx (Fig. 2b,e). In some animals unspecific staining was observed attached to the trochoblasts (Fig. 2e).
Summary of Gbx-expression (red) during polyplacophoran, solenogaster, and bivalve development. Anterior (a)-posterior (p) axes indicate the orientation. Asterisks mark the mouth opening. (a) Early trochophore larva (12 hpf) of the polyplacophoran Acanthochitona crinita possess an episphere (ep) that is divided from the hyposphere (hp) by a prototroch (pt). The anteriormost region of the episphere exhibits an apical organ with a (ciliary) apical tuft (at). (b) Metamorphic competent individual (65 hpf) of A. crinita showing seven shell fields with each one row of Gbx-expressing cells on the dorsal side and a foot (f) on the ventral side. (c) Early test-cell larva (0–1 dph) of the solenogaster Wirenia argentea exhibit large and flattened test-cells (tc). (d) In mid-stage test-cell larva (10–11 dph) of the solenogaster W. argentea Gbx is expressed in the anlagen of the cerebral ganglia (cg), in cells (2) that line the peri-imaginal space (pis) and in cells that are associated with the pedal nerve cords (pc). (e) Late test-cell larva (12 dpf) of the bivalve Nucula tumidula possess large and flattened test-cells, a mouth, and anus (an). The shell fields (sf) give rise to both shell valves (s). (f) Postmetamorphic individuals (22 dpf) of N. tumidula shed all test-cells which are subsequently ingested. Abbreviations: mc, mantle cavity; stm, stomach. Scale bars: 50 µm.
Gbx-expression in the aplacophoran Wirenia argentea
Gastrulae develop into early test-cell larvae that are characterized by large and flattenened test-cells (Fig. 5c 30). The apical organ is associated with a tuft and the prototroch divides episphere and hyposphere (Fig. 5c). In freshly hatched larvae, Gbx is expressed in a dorsal-ventral gradient with the highest expression in the ventral epidermal cells close to the pseudo-blastopore, a posterior invagination of the larval body (Figs 3a,b and 5c). The pseudo-blastopore does not develop into the actual mouth, instead the larval body (trunk) growths out through it. Gbx-expression decreases in the dorso-lateral epidermal cells surrounding the pseudo-blastopore (Fig. 3a,b). Early larvae express Gbx latero-ventrally in epidermal cells that line the peri-imaginal space (“2” in Fig. 3c–e). The majority of these cells develop into spicule-bearing cells of the prospective trunk (Supplementary Fig. 4). The latter Gbx–expression domain extends laterally towards the mouth and includes also subepidermal cells (“2” in Fig. 3d). On each side two ventrally located spots at the posterior pole of the outgrowing trunk express Gbx (“1” in Fig. 3c–e). The developing cerebral ganglia show faint Gbx-expression (Fig. 3c,d). Mid-stage larvae express Gbx in the developing pedal nerve cords (Figs 3g and 5d), a Gbx-expression domain that might correspond to expression domain “1” of early larvae (Fig. 3e). Gbx is still expressed in epidermal cells that line the peri-imaginal space and in subepidermal cells close to the latter cells (“2” in Figs 3f,g and 5d). The former epidermal cells contribute to the mantle of the outgrowing trunk (cf. ref. 30). Some individuals show Gbx-expression adjacent to the developing foregut (arrowhead in Fig. 3g), expression domains that may correspond to the anlagen of the buccal and/or lateral ganglia30. In addition, the anlagen of the cerebral ganglia express Gbx (Figs 3f,g and 5d). Late larvae express Gbx in their developing pedal nerve cords (Fig. 3h). Gbx is still expressed latero-ventrally in epidermal cells that line the peri-imaginal space and in subepidermal cells close to the latter cells (Fig. 3h–i). Expression is strongest on both sides of the mouth opening (“2” in Fig. 3h–i) and co-localized with expression of Pax6, Paxβ, and engrailed 31 (Scherholz, unpublished data). Gbx is also expressed in cells adjacent to the developing foregut, a region where buccal or lateral ganglia may be located (arrowheads in Fig. 3h–i). In the region of the developing cerebral ganglia, faint Gbx-expression is present (Fig. 3h,i).
Gbx-expression in developmental stages of the bivalve Nucula tumidula
After gastrulation, a test-cell larva hatches that exhibits three rows of ciliated cells and an apical organ with a ciliary tuft (Fig. 5e). These rows of cilia arise from flattened and large test-cells that constitute the outermost cell layer of the larval body (Fig. 5e). The blastopore leads to the definite mouth of the larva and is located on the posterior pole together with the anus (Fig. 5e). In early test-cell larvae, Gbx is expressed in the dorsal ectoderm (Fig. 4a). In further developed larvae, Gbx is expressed in the ventral mantle margin (Fig. 4b; for details on the anatomy of the bivalve test-cell larva see ref. 32). Subsequently, larvae cease to express Gbx in the ventral mantle regions and lateral mantle areas express Gbx (Fig. 4c,d). Late test-cell larvae express Gbx again in the region of the ventral mantle and faint expression was detected in the lateral mantle areas (Fig. 4e). Early postmetamorphic individuals express Gbx in parts of the mantle close to the foot (Figs 4f and 5f).
Summary of the main Gbx-expression domains in all three molluscan species
Gbx is expressed in the posterior CNS of polyplacophoran and aplacophoran larvae but not in the CNS of bivalve larvae (Fig. 5b,d–f). The aplacophoran anteriormost CNS, i.e. the anlagen of the cerebral ganglia, also express Gbx (Fig. 5d). In polyplacophorans Gbx is expressed in spicule-bearing cells and cells of the shell fields (Fig. 5b). Spicule-bearing cells and cells of the mantle express Gbx in the aplacophoran and the bivalve, respectively (Fig. 5c–f).
Discussion
The molluscan Gbx orthologs are expressed in ectodermal domains
Our phylogenetic analysis includes Gbx-orthologs of various other bilaterian representatives and corroborates the identity of Acr-Gbx, War-Gbx, and Ntu-Gbx with high support (Supplementary material Figs 1 and 2). Our results on mollusks corroborate the finding of previous studies that Gbx is expressed in the ectoderm of bilaterians14, 33, 34. In the aculiferans Acanthochitona crinita and Wirenia argentea as well as in the conchiferan Nucula tumidula Gbx is expressed in the nervous system and/or the mantle.
Do mollusks share similar gene expression profiles in their CNS with other bilaterians?
Genes such as Six3/6, Otx, Pax2/5/8, Gbx, and the anterior Hox genes are expressed in a sequential fashion in the developing CNS of phylogenetically distantly related bilaterians9, 33. While a couple of ecdysozoans and deuterostomes have been investigated in detail, only few lophotrochozoans have been studied with respect to the expression patterns of the above-mentioned genes34. In the aculiferan mollusks Acanthochitona crinita and Wirenia argentea as well as in the conchiferan cuttlefish Sepia officinalis, Gbx is expressed in the nervous system more posteriorly to Otx- and Pax2/5/8- expression domains (this study14, 15, Wollesen, unpublished data). The polyplacophoran A. crinita expresses Gbx in the visceral nerve cords and in the anteriormost pedal nerve cords (Fig. 5b). In S. officinalis Gbx is expressed in the posteriormost brain region, i.e. in the posterior portion of the subesophageal mass (=palliovisceral ganglia) as well as in the stellate ganglia14. In W. argentea Gbx is also expressed in the posterior CNS, i.e. the pedal nerve cords, however also in the anlagen of the cerebral ganglia (Fig. 5d). The latter finding is of surprise and contrasts the situation in other bilaterians with Gbx-expression in more posterior regions, partially co-expressed with Hox genes (see above). Since W. argentea is the only bilaterian investigated so far that shows Gbx-expression in a region located thus far anterior, the most parsimonious explanation is that this expression pattern evolved secondarily. This is of interest since a recent study demonstrated that solenogasters exhibit some rather derived character states such as a secondarily simplified body plan26, 35.
Polyplacophoran anterior Hox genes are co-expressed with Gbx in the posttrochal larval region, a condition that resembles that of other bilaterians including cephalopods (this study9, 11, 34, 36,37,38). Anterior to the Gbx-expression domain, a Pax2/5/8 domain is located in the interbasal lobes of the cephalopod supraesophageal mass15. The interbasal lobes lie adjacent to the esophagus that divide the brain into a supraesophageal mass and a subesophageal mass. W. argentea also shows Pax2/5/8 expression in its cerebral ganglia and pedal nerve cords31. In contrast to cephalopods, solenogasters, and other bilaterians, Pax2/5/8 is not expressed in the developing CNS of polyplacophorans, bivalves, and gastropods15, 39.
Anterior to the Pax2/5/8 expression domain an Otx expression domain is located in cephalopods12, 15. In polyplacophoran trochophore larvae, Otx is broadly expressed in the episphere where the anlagen of the cerebral commissure are located28. Pax2/5/8 and Otx are both expressed in the region of the developing cerebral ganglia in solenogasters31 (Redl et al., unpublished results). In the gastropod Crepidula fornicata Otx is expressed in the anterior CNS of trochophore and veliger larvae40, in contrast to trochophore larvae of the patellogastropod Patella vulgata that lack Otx-expression in this region41. As in other bilaterians Six3/6 is expressed in the anteriormost region of the CNS in gastropods, cephalopods, polyplacophorans, and solenogasters13, 28, 40 (Redl et al., unpublished results).
In summary, Six3/6, Otx, and Gbx are expressed in the nervous system of the majority of mollusks investigated so far, while Pax2/5/8 is only expressed in the developing nervous system of solenogasters and cephalopods. Pax2/5/8 and Gbx are not expressed in the developing nervous system of the bivalve N. tumidula altogether. Hox1 is expressed in the cephalopod and polyplacophoran CNS.
Gbx and other brain regionalization genes have been co-opted into molluscan shell field development
Gbx is expressed in the mantle of the polyplacophoran Acanthochitona crinita, the solenogaster Wirenia argentea, and the bivalve Nucula tumidula (this study: Fig. 5). In contrast, late prehatching embryos of the cephalopod Sepia officinalis do not express Gbx in the mantle or the shell gland14. Seven horizontal rows of Gbx-expressing cells are located in the seven shell fields of the A. crinita (Fig. 5b). Polyplacophoran shell fields are characterized by a ridge-like appearance and a number of different cells give rise to the shell plates of the polyplacophoran Ischnochiton rissoa 42. “Goblet cells” are large cells that are located in the center of each shell field and anterior to these cells the slender Gbx-expressing cells are situated in A. crinita. Judging by their location and morphology, the Gbx-expressing cells may be identified as “type 2” or “type 4” cells, cells that together with the “goblet cells” secrete the shell plates in I. rissoa 42. Early settled polyplacophorans possess seven shell plates, while the eighth shell plate develops several weeks afterwards43, 44. In W. argentea Gbx is strongly expressed in the ventro-lateral mantle region of the outgrowing trunk, i.e. exactly in the same region where spicule-bearing cells are present (Scherholz, unpublished data). In N. tumidula Gbx is expressed in the ventral mantle regions and subsequently in lateral mantle regions (Fig. 5e,f). The ventral Gbx-expressing mantle regions probably contribute to the growth of both shell valves.
Interestingly, recent studies have shown that Pax2/5/8 is expressed in horizontal rows of cells in the polyplacophoran mantle that gives rise to the shell plates, as well as in the spicule-bearing cells of the aplacophoran W. argentea 15, 31. Pax2/5/8 is also expressed in the mantle of N. tumidula, the cephalopod Idiosepius notoides, and in the anterior mantle region of the veliger larva of H. asinina 15, 39. The latter expression domain corresponds to the mantle region that secretes the protoconch I, i.e. the first-formed embryonic shell. In addition to Pax2/5/8 and Gbx, Hox1 is also expressed in the shell fields of gastropods and polyplacophorans15, 37, 45,46,47.
Since Gbx, Pax2/5/8, and Hox1 are expressed in mantle domains that secrete the shell(s) of polyplacophorans and conchiferans, the most parsimonious conclusion is that these genes were already expressed in these regions in the last common ancestor of Mollusca. The precise number of shell field(s) in the last common ancestor of Mollusca remains obscure since seven shell fields were probably present at the base of the Aculifera and only one shell was present in the ground pattern of its sister group, the Conchifera1,2,3, 26, 35, 47, 48. Competing evolutionary scenarios suggest that no mineralized shell(s) but spicules were present in the last common ancestor of Mollusca47,48,49,50,51. Coleoid cephalopods probably lost Gbx and Hox1 expression in their shell field during evolution11, 14. The vast majority of coleoids secrete a slender internalized “shell” (e.g., a non-mineralized gladius or a cuttlebone), an innovation that may be correlated with the loss of Gbx and Hox1 expression in the shell field during coleoid cephalopod evolution. Recent studies on various bivalve and gastropod species demonstrate that their adult mantle secretomes are rather diverse52,53,54. Each species possesses a unique mantle secretome composed of a majority of lineage- or species-specific genes54. Although there is indication that mantle secretomes underlie significant changes with respect to their gene composition, this has not been investigated thoroughly throughout the Mollusca55,56,57. Genes that underlie the formation of the shell field(s) and spicule-bearing cells such as Gbx, Pax2/5/8, and Hox1 have been investigated to an even lesser degree.
Conclusions
This study shows that Gbx plays a role during the formation of major molluscan traits, the shells and spicules. Pax2/5/8, Gbx, and Hox1 are originally known to be involved in the regionalization of the bilaterian nervous system, however, these ‘brain regionalization genes’ were also co-opted into shell and spicule formation. With exception of Pax2/5/8, cephalopods do not express any of the above-mentioned genes in the mantle during early ontogenesis, however, all of them in the brain. In contrast, the ancestral condition has been lost with Gbx and Pax2/5/8 expressed only in the mantle but not in the nervous system in the bivalve N. tumidula. This suggests an additional role of these typical ‘brain regionalization genes’ in the formation of the shell field(s) and spicule-bearing cells. Together with paleontological evidence, this work and future studies of its kind may infer whether the last common molluscan ancestor had a single shell, multiple shell plates, or no shell at all.
References
Kocot, K. M. et al. Phylogenomics reveals deep molluscan relationships. Nature. 477, 452–456 (2011).
Smith, S. A. et al. Resolving the evolutionary relationships of molluscs with phylogenomic tools. Nature. 480, 364–369 (2011).
Smith, S. A. et al. Corrigendum: Resolving the evolutionary relationships of molluscs with phylogenomic tools. Nature. 708, 493 (2013).
Haszprunar, G. & Wanninger, A. Molluscs. Curr. Biol. 22, 13 (2012).
Sigwart, J. D., Sumner-Rooney, L. H. 18 Mollusca: Caudofoveata, Monoplacophora, Polyplacophora, Scaphopoda, and Solenogastres in Structure and Evolution of Invertebrate Nervous Systems (eds Schmidt-Rhaesa, A., Harzsch, S. & Purschke, G.) 172-189 (Oxford Univer. Press, 2016).
Voronezhskaya, E. E., Croll, R. P. 20 Mollusca: Gastropoda in Structure and Evolution of Invertebrate Nervous Systems (eds Schmidt-Rhaesa, A., Harzsch, S. & Purschke, G.) 196-221 (Oxford Univer. Press, 2016).
Wanninger, A. 19 Mollusca: Bivalvia in Structure and Evolution of Invertebrate Nervous Systems (eds Schmidt-Rhaesa, A., Harzsch, S. & Purschke, G.) 190–195 (Oxford Univer. Press, 2016).
Wollesen, T. 21 Mollusca: Cephalopoda in Structure and Evolution of Invertebrate Nervous Systems (eds Schmidt-Rhaesa, A., Harzsch, S. & Purschke, G.) 222–240 (Oxford Univer. Press, 2016).
Lichtneckert, R. & Reichert, H. Insights into the urbilaterian brain: conserved genetic patterning mechanisms in insect and vertebrate brain development. Heredity. 94, 465–477 (2005).
Steinmetz, P. R. H. Six3 demarcates the anterior-most developing brain region in bilaterian animals. EvoDevo. 1, 14 (2010).
Lee, P. N., Callaerts, P., de Couet, H. G. & Martindale, M. Q. Cephalopod Hox genes and the origin of morphological novelties. Nature. 424, 1061–1065 (2003).
Buresi, A., Baratte, S., Da Silva, C. & Bonnaud, L. Orthogenticle/otx ortholog expression in the anterior brain and eyes of Sepia officinalis (Mollusca, Cephalopoda). Gene Expr. Patt 12, 109–116 (2012).
Ogura, A. et al. Loss of the six3/6 controlling pathways might have resulted in pinhole-eye evolution in Nautilus. Sci. Reports 3, 1432, doi:10.1038/srep01432 (2013).
Focareta, L., Sesso, S. & Cole, A. G. Characterization of homeobox genes reveals sophisticated regionalization of the central nervous system in the European cuttlefish Sepia officinalis. Plos one 9, e109627 (2014).
Wollesen, T., Rodríguez Monje, S. V., Todt, C., Degnan, B. M. & Wanninger, A. Ancestral role of Pax2/5/8 in molluscan brain and multimodal sensory system development. BMC Evol. Biol. 15, 231 (2015).
Redl, E., Scherholz, M., Wollesen, T., Todt, C. & Wanninger, A. Cell Proliferation Pattern and Twist Expression in an Aplacophoran Mollusk Argue Against Segmented Ancestry of Mollusca. JEZ, Part B: Mol. Dev. Evol. 326, 422–436 (2016).
Wollesen, T., Rodríguez Monje, S. V., McDougall, C., Degnan, B. M. & Wanninger, A. The ParaHox gene Gsx patterns the apical organ and central nervous system but not the foregut in scaphopod and cephalopod mollusks. EvoDevo 6, 41 (2015).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-seqdata without a reference genome. Nat. Biotechnol. 15, 644–652 (2011).
Peng, Y. et al. IDBA-tran: a more robust de novo de Bruijn graph assembler for transcriptomes with uneven expression levels. Bioinformatics 29, i326–i334 (2013).
Zhong, Y. F., Butts, T. & Holland, P. W. H. HomeoDB: a database of homeobox gene diversity. Evol. & Dev. 10, 516–518 (2008).
Zhong, Y. F. & Holland, P. W. H. HomeoDB2: functional expansion of a comparative homeobox gene database for evolutionary developmental biology. Evol. Dev. 13, 567–568 (2011).
Katoh, K. & Standley, D. M. MAFFT: iterative refinement and additional methods. Meth. Mol. Biol. 1079, 131–146, doi:10.1007/978-1-62703-646-7_8 (2014).
Le, S. Q. & Gascuel, O. An improved general amino acid replacement matrix. Mol. Biol. Evol. 25, 1307–1320 (2008).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics. 27, 1164–1165 (2011).
Rambaut, A. FigTree v1.4.1. http://tree.bio.ed.ac.uk/figtree (2008).
Scherholz, M., Redl, E., Wollesen, T., Todt, C. & Wanninger, A. From complex to simple: myogenesis in an aplacophoran mollusk reveals key traits in aculiferan evolution. BMC Evol. Biol. 15, 201 (2015).
Lauter, G., Söll, I. & Hauptmann, G. Two-color fluorescent in situ hybridization in the embryonic zebrafish brain using differential detection systems. BMC Dev. Biol. 11, 43 (2011).
Vöcking, O., Kourtesis, I. & Hausen, H. Posterior eyespots in larval chitons have a molecular identity similar to anterior cerebral eyes in other bilaterians. EvoDevo 6, 40 (2015).
Jékely, G. & Arendt, D. Confocal detection of NBT/BCIP in situ hybridization samples by reflection microscopy. Biochemica. 4, 12–14 (2007).
Todt, C. & Wanninger, A. Of tests, trochs, shells, and spicules: Development of the basal mollusk Wirenia argentea (Solenogastres) and its bearing on the evolution of trochozoan larval key features. Front. Zool. 7, 6 (2010).
Scherholz, M. et al. Ancestral and novel roles of Pax family genes in mollusks. BMC. Evol. Bio. 17, 81 (2017).
Zardus, J. D. & Morse, M. P. Embryogenesis, morphology and ultrastructure of the pericalymma larva of Acila castrensis (Bivalvia: Protobranchia: Nuculoida). Invert. Biol. 117, 221–244 (1998).
Castro, L. F. C., Rasmussen, S. L. K., Holland, P. W. H., Holland, N. D. & Holland, L. Z. A Gbx homeobox gene in amphioxus: Insights into ancestry of the ANTP class and evolution of the midbrain/hindbrain boundary. Dev. Biol. 295, 40–51 (2006).
Steinmetz, P. R. H., Zelanda-Gonzáles, F., Burgtorf, C., Wittbrodt, J. & Arendt, D. Polychaete trunk neuroectoderm converges and extends by mediolateral cell intercalation. PNAS 104, 2727–2732 (2007).
Scherholz, M., Redl, E., Wollesen, T., Todt, C. & Wanninger, A. Aplacophoran mollusks evolved from ancestors with polyplacophoran-like features. Curr. Biol. 23, 2130–2134 (2013).
Steinmetz, P. R. H., Kostyuchenko, R. P., Fischer, A. & Arendt, D. The segmental pattern of otx, gbx, and Hox genes in the annelid Platynereis dumerilii. Evol. Dev. 13, 72–79 (2011).
Fritsch, M., Wollesen, T., de Oliveira, A. L. & Wanninger, A. Unexpected co-linearity of Hox gene expression in an aculiferan mollusk. BMC Evol. Biol. 15, 151, doi:10.1186/s12862-015-0414-1 (2015).
Fritsch, M., Wollesen, T. & Wanninger, A. Hox and ParaHox gene expression in early body plan patterning of polyplacophoran mollusks. JEZ Part B Mol. Dev. Evol. 326, 89–104 (2016).
O’Brien, E. K. & Degnan, B. M. Expression of Pax258 in the gastropod statocyst: insights into the antiquity of metazoan geosensory organs. Evol. Dev. 5, 572–578 (2003).
Perry, K. et al. Deployment of regulatory genes during gastrulation and germ layer specification in a model spiralian mollusc. Crepidula. Dev. Dyn. 244, 1215–1248 (2015).
Nederbragt, A. J., te Welscher, P., van den Driesche, S., van Loon, A. E. & Dictus, W. J. A. G. Novel and conserved roles for orthodenticle/otx and orthopedia/otp orthologs in the gastropod mollusc Patella vulgata. Dev. Genes Evol. 212, 330–337 (2002).
Kniprath, E. Ontogenetic plate and plate field development in two chitons, Middendorffia and Ischnochiton. Wilhelm Roux’s Archives 189, 97–106 (1980).
Heath, H. The Development of Ischnochiton. Zool. Jb Anat. Ontog. Tiere 12, 567–656 (1898).
Wanninger, A. & Haszprunar, G. Chiton myogenesis: perspectives for the development and evolution of larval and adult muscle systems in molluscs. J. Morphol. 251, 103–113 (2002).
Samadi, L. & Steiner, G. Involvement of Hox genes in shell morphogenesis in the encapsulated development of a top shell gastropod (Gibbula varia L.). Dev. Genes Evol. 219, 523–530 (2009).
Hashimoto, N., Kurita, Y. & Wada, H. Developmental role of dpp in the gastropod shell plate and co-option of the dpp signaling pathway in the evolution of the operculum. Dev. Biol. 366, 367–373 (2012).
Wanninger, A., Wollesen, T. Mollusca in Evolutionary Developmental Biology of Invertebrates, Vol. 2 Lophotrochozoa (Lophotrochozoa). (ed. Wanninger, A.) Springer, 103–153 (2015).
Vinther, J., Sperling, E. A., Briggs, D. G. & Peterson, K. J. A molecular palaeobiological hypothesis for the origin of aplacophoran molluscs and their derivation from chiton-like ancestors. Proc. R. Soc. B 279, 1259–1268 (2011).
Salvini-Plawen, Lv A reconsideration of systematics in the Mollusca (Phylogeny and higher classification). Malacologia 19, 249–278 (1980).
Fedonkin, M. A. & Waggoner, B. M. The late Precambrian fossil Kimberella is a mollusc-like bilaterian organism. Nature 388, 868–871 (1997).
Ax, P. Multicellular Animals. The phylogenetic system of the metazoan. Springer (2000).
Jackson, D. J. et al. Parallel evolution of nacre building gene sets in molluscs. Mol. Biol. Evol. 27, 591–608 (2010).
Kocot, K. M., Aguilera, F., McDougall, C., Jackson, D. J. & Degnan, B. M. Sea shell diversity and rapidly evolving secretomes: insights into the evolution of biomineralization. Front. Zool. 13, 23 (2016).
Aguilera, F., McDougall, C. & Degnan, B. M. Co-option and de novo gene evolution underlie molluscan shell diversity. Mol. Biol. Evol. 34, 779–792 (2017).
Jackson, D. J., Wörheide, G. & Degnan, B. M. Dynamic expression of ancient and novel molluscan shell genes during ecological transitions. BMC Evol. Biol. 7, 160 (2007).
Vinther, J., Parry, L., Briggs, D. E. G. & Van Roy, P. Ancestral morphology of crown-group molluscs revealed by a new Ordovician stem aculiferan. Nature 542, 471–474 (2017).
Wanninger, A., Wollesen, T. The evolution of mollusks. Nature Ecol. Evol. (in review).
Acknowledgements
The authors thank André Luiz de Oliveira (Vienna) for assembling the transcriptome of Wirenia argentea and for advice on the phylogenetic analysis. Oliver Vöcking (Bergen) is thanked for advice on fluorescent in situ hybridization. The authors thank Henrik Glenner (Bergen) for providing boat time and lab space, and the staff of the SBR for logistic support. The crew of the R/V Hans Brattström (Bergen) is thanked for collection of animals. While rearing developmental stages of A. crinita during his stay at the Station Biologique Roscoff (SBR) (France) T.W. was supported by an ASSEMBLE (Association of European Marine Biological Laboratories) grant (grant agreement no. 835 (SBR-1). AW was supported by a FWF (Austrian Science Fund) grant (grant number P24276-B22). We thank two reviewers for their comments on an earlier version of this manuscript.
Author information
Authors and Affiliations
Contributions
T.W. designed the project, assembled the transcriptomes of A. crinita and N. tumidula, interpreted the data, and drafted the manuscript. A.W. contributed to data interpretation and writing of the manuscript. T.W. raised developmental stages of N. tumidula and A. crinita. M.S., C.T., E.R. and T.W. raised developmental stages of W. argentea and C.T. and E.R. helped to raise developmental stages of N. tumidula. T.W. and S.V.R. cloned Acr-Gbx and Ntu-Gbx and carried out the in situ experiments on A. crinita and N. tumidula. M.S. cloned War-Gbx and carried out the experiments on W. argentea and analyzed the respective data. S.V.R. performed the phylogenetic analysis. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wollesen, T., Scherholz, M., Rodríguez Monje, S.V. et al. Brain regionalization genes are co-opted into shell field patterning in Mollusca. Sci Rep 7, 5486 (2017). https://doi.org/10.1038/s41598-017-05605-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-05605-5
This article is cited by
-
The adult shell matrix protein repertoire of the marine snail Crepidula is dominated by conserved genes that are also expressed in larvae
BMC Ecology and Evolution (2024)
-
Shell field morphogenesis in the polyplacophoran mollusk Acanthochitona rubrolineata
EvoDevo (2023)
-
The brain regulatory program predates central nervous system evolution
Scientific Reports (2023)
-
Octopod Hox genes and cephalopod plesiomorphies
Scientific Reports (2023)
-
Non-collinear Hox gene expression in bivalves and the evolution of morphological novelties in mollusks
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.