Abstract
The classic brown body (bwb) mutation in the housefly Musca domestica impairs normal melanization of the adult cuticle. In Drosophila melanogaster, a reminiscent pigmentation defect results from mutations in the yellow gene encoding dopachrome conversion enzyme (DCE). Here, we demonstrate that the bwb locus structurally and functionally represents the yellow ortholog of Musca domestica, MdY. In bwb Musca strains, we identified two mutant MdY alleles that contain lesions predicted to result in premature truncation of the MdY open reading frame. We targeted wildtype MdY by CRISPR-Cas9 RNPs and generated new mutant alleles that fail to complement existing MdY alleles, genetically confirming that MdY is the bwb locus. We further found evidence for Cas9-mediated interchromosomal recombination between wildtype and mutant bwb alleles. Our work resolves the molecular identity of the classic bwb mutation in Musca domestica and establishes the feasibility of Cas9-mediated genome editing in the Musca model.
Similar content being viewed by others
Introduction
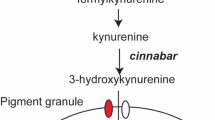
The so-far unidentified brown body (bwb) locus in the housefly Musca domestica was named after a recessive loss-of-function phenotype in which the adult cuticle manifests in a brown color rather than the wildtype black pigmentation (Fig. 1A). The absence of black coloration in mutant Musca has been proposed to result from impaired synthesis and incorporation of the black pigment melanin during pupal stages1. In insects, melanization of the cuticle is common and contributes to the diverse coloration patterns that are the most visible features of the outer morphology. Most insights into the pathway that produces and incorporates melanin into the insect cuticle comes from studies in Drosophila melanogaster 2,3,4. The melanin pathway starts with conversion of tyrosine to Dihydroxyphenylalanine (DOPA) by tyrosine hydroxylase (TH). DOPA in turn is converted to dopamine by dopa carboxylase (DDC). Both substrates are used as precursors for production of black melanin. In Drosophila, the dopachrome conversion enzyme (DCE) catalyzes steps downstream of TH and DCC in melanin production; however, whether DCE is involved in only in DOPA, only in dopamine conversion, or in both, remains unclear. Loss of function in the yellow gene (y) that encodes Drosophila DCE causes a lack of melanin incorporation and results in a yellowish overall appearance of the cuticle. Similar phenotypes have been observed in other insects such as the lepidopteran Bombyx mori, Papilio xuthus, and the coleopteran Tribolium castaneum 5,6,7,8,9. Loss of activity of the corresponding yellow orthologs in these species drastically lowers the synthesis of melanin, causing regions of the body that are normally black to display a lighter coloration.
bwb phenotype is associated with nonsense mutations in MdY. (A) Phenotypes are displayed from left to right: multimarked aabys male, bwb wildtype male from M III strain and bwb mutant female from M III strain. Genotypes are indicated; bwb: brown body, pw: pointed wings (notches along the edge of the wing, see arrow), w: white eyes. (B) Schematic drawing of the MdY locus in the bwb mutant aabys strain and the Siat bwb wildtype strain. The aabys allele, MdY a1, contains a 1.5 kb insertion in the 5′ UTR and an additional 4 bp insertion in the ORF of exon 1. This frame-shift leads to a premature TAA stop in the 5′ end of exon 2. The Siat MdY allele has an intact ORF (boxed yellow). (C) Genomic amplification with flanking primers Y-GAP1-F1 and Y-EXON1-R show that the 1.5 kb insertion is present in males and females of the bwb mutant aabys strain, but not in the bwb wildtype Siat flies. (D) Affected part of the coding region (codon position 63 to 80) of the nonsense alleles of MdY. Deviations from the wildtype sequence (Siat) are marked in red and translational stops in bold. Location of intron is indicated with a triangle.
In recessive bwb-mutant houseflies, the loss of normal black coloration affects all body parts. This phenotype closely resembles the phenotype observed in yellow-mutant Drosophila. As no other gene in the network of melanin genes is known to manifest this phenotype, we sought to investigate whether the bwb gene in the housefly is the structural and functional homolog of the yellow (DCE) gene in Drosophila. In addition to the correspondence in phenotypes, the bwb locus has been mapped to Chromosome III in Musca which corresponds to Muller element A, the X chromosome in Drosophila that harbors the y locus10, 11.
Here, we identified the locus mutated in bwb as the Musca domestica ortholog of the Drosophila gene yellow and refer to the gene as MdY. The MdY gene shows a high degree of similarity at the level of both protein sequence and gene structure. We find three mutant alleles of the MdY gene in bwb-mutant flies: the alleles MdY a2 and MdY b feature sequence disruptions of the coding sequence, while a third allele MdY a1 is a compound allele featuring the coding lesion of MdY a2 and an additional 1.5 kb sequence insertion in the 5′ UTR. Using CRISPR-Cas9, we generated a series of de novo loss-of-function alleles of MdY that all fail to complement original bwb alleles; these experiments mark, to our knowledge, the first report of the successful application of Cas9-based mutagenesis in Musca domestica. Altogether, we conclude that the bwb phenotype in Musca is caused by a lack of DCE activity normally provided by the MdY gene. Our findings clarify the molecular lesions in the classic bwb mutation and further underline the notion that yellow plays a conserved role in the melanin production pathway in dipteran species.
Results
Characterization of the yellow ortholog in Musca domestica
Based on phenotype resemblance and mapping position, we hypothesized that bwb affects the so-far undescribed DCE ortholog in Musca domestica. To identify sequences homologous to the Drosophila yellow gene in Musca, we performed BLAST searches against the recently published genome of the multi-marked aabys strain that shows the bwb phenotype12. We recovered an annotated mRNA sequence (NCBI RefSeq XM_011292650.1) with a high degree of sequence similarity to Drosophila yellow (Supplementary data Fig. 1). Annotation of the aabys genome called this gene a pseudogene based on the lack of an intact open reading frame. Indeed, we detected a frame-shift starting 67 codons downstream of the first AUG start codon (Fig. 1B)12. Hence, the molecular nature of this allele already suggested that the bwb-mutant aabys strain carries a non-functional yellow variant. The mutated putative Musca yellow gene, which we named MdY, is located on Scaffold18750 (502 kb) and is composed of two exons separated by a 35.6 kb-spanning intron (Fig. 1B). A Musca homolog of the acheate (ac) gene is present on the same scaffold separated by 156 kb, revealing a relatively close linkage of yellow and ac orthologs in Musca. This coupling is conserved in Drosophila melanogaster and sibling species, and previous work proposed that evolutionary variation of the y-ac region is reduced due to the selective fixation of one or more advantageous mutations in this region13. We next isolated MdY sequences from a wildtype strain Siat that displays a normal melanization pattern; the MdY sequence in Siat has an intact ORF of 522 amino acids, markedly lacking the 4 bp insertion that causes a frame-shift in the aabys-derived allele (Fig. 1B and D). The predicted MdY protein shares a high level of similarity with YELLOW proteins of other dipterans (Supplementary Figure 2a): as expected, MdY shares the highest level of identity (88%) with the predicted YELLOW protein in Stomoxys calcitrans, a close relative of the same family Muscidae (Supplementary data Fig. 2b).
To extend our analysis of mutant bwb alleles, we included the M III strain that carries the male determining factor on Chromosome III tightly linked to wildtype alleles of bwb and pointed wings (pw), while females of this strain are homozygous mutant for bwb and pw (Fig. 1A). Analysis of MdY sequences isolated from M III females unveiled that again the bwb allele that contains a 4 bp insertion at amino acid position 67. Unexpectedly, we also reproducibly detected a second mutant MdY allele in this strain, a nonsense mutation at amino acid position 65 (Fig. 1D). Therefore, the M III laboratory strain carries two different loss-of-function alleles of MdY.
When mapping wildtype genomic MdY fragments against the corresponding aabys genomic sequences that harbor the 4 bp insertion in the coding sequence, we additionally identified a structural difference in the 5′ UTR region upstream of the putative start codon (Fig. 1B). Sequence analysis revealed the presence of a 1.5 kb insertion in the aabys-derived allele that is absent in the wildtype Siat strain (Fig. 1B and C). A BLAST search against the Musca genome shows that this aabys-specific insertion shares 77% of sequence identity to an incomplete gene complement of the nicotinic acetylcholine receptor subunit-encoding Mdalpha2 14. This 5′ UTR insertion in the mutant MdY allele of the aabys strain consequently represents a third, compound mutant allele of MdY.
Altogether, we identified two distinct coding-frame alleles in bwb-mutant Musca domestica strains, and a third compound lesion that introduced a 1.5 kb insertion into the 5′ UTR of one of the putatively inactive MdY loci. We refer to the original aabys-derived compound allele as MdY a1, and to the two M III-derived alleles as MdY a2 and MdY b (Fig. 1D, Supplementary data Fig. 3). From our analysis of these mutants, we conclude that the bwb phenotype is associated with nonsense mutations in the MdY gene. These observations support the notion that the housefly DCE homolog is involved in the melanin production pathway.
CRISPR-Cas9-mediated disruption of MdY confirms causative association with bwb
We next performed targeted disruption of the wildtype MdY locus to confirm its predicted role in melanization of the housefly cuticle and to corroborate whether the bwb mutations are indeed loss-of-function alleles of MdY. To this end, we selected two target sites in the second exon of MdY for non-homologous end joining (NHEJ)-mediated disruption by the CRISPR-Cas9 system. Preassembled ribonucleoprotein complexes (RNPs) composed of purified Cas9 protein loaded with two different sgRNAs (sgY2 and sgY3) were injected into early syncytial embryos. Both sgY2 and sgY3 target two different sites in coding exon 2 separated by 340 bp, (Fig. 2a). As host strain, we used the M III strain that carries the male determining factor on the chromosome III linked to wildtype alleles of bwb and pw. Females of this strain are homozygous mutant for both markers and are brown coloured with pointed-wings, while males are heterozygous and phenotypically wildtype for both markers15, 16 (Fig. 2b). This genetic background facilitates detection of possible somatic effects of MdY disruption in males that carry only one wildtype allele of bwb (Fig. 1A). The two sgRNAs were preloaded separately on purified recombinant Cas9 protein17, 18 and a 1:1 mix was micro-injected into 1 h old embryos of the MIII strain19. Of 2565 embryos injected with a 1:1 mix of solubilized Cas9 RNPs containing sgY2, and sgY3, we recovered 188 surviving adult houseflies. While 106 of these adults were males, none of the males displayed any patches of brown coloration, indicating that our targeting procedure does not introduce significant somatic mutation mosaicism. We proceeded with crossing 84 of the 106 injected G0 males to bwb-mutant females of the same M III strain (2 males and 6 females per cross) to screen for possible germline effects. Screening the F1 generation, 17% of the crosses produced brown-colored M III pw + males that can only arise from paternal transmission of a mutant bwb allele (Table 1, Fig. 2c). This observation reveals that our Cas9 RNP-mediated mutagenesis protocol introduced bwb mutations by targeting the MdY gene in germ cells of the syncytial embryo. Of note, the injected males vary greatly in the proportion of potentially MdY-disrupted F1 individuals they sire (Table 1 ), revealing variable germline mosaicism resulting from Cas9 mutagenesis akin to observations in other model organisms19.
Strategy for CRISPR/Cas9 mediated disruption of MdY. (a) A schematic of the MdY gene showing the positions of the two target sites in exon 2. Sequences used for the design of the two sgRNAs, sgY3 and sgY2, are indicated. Both sequences are flanked by a PAM motif (in red) and separated by 343 bp. (b) Crossing scheme for screening mutational events affecting melanization. Injected G0 males (with a mix of sgY3-CAS9 and sgY2-CAS9) are crossed with bwb females and F1 is examined for occurrence of bwb males. (c) Right a bwb mutant F1 male from line MdY#16 which is heterozygous for a CRIPSR induced 10 bp deletion in sgY3 over MdY b. Left an unaffected bwb wildtype F1 male from the same line with the paternal genotype (MdY + over MdY b).
To characterize the putative lesions induced in the MdY gene by sgRNAs sgY2 and sgY3, we PCR-isolated genomic sequences of exon 2 from brown-bodied M III pw + F1 males by PCR and examined the targeted sites in sub-cloned and Sanger-sequenced fragments using CrispRVariants20. Overall, the position and extent of the induced lesions vary between individual lines but show a clear preference for lesions at the sgY3 target site (Table 2, Supplementary Fig. 4). In seven lines (MdY#2, MdY#9, MdY#10, MdY#13, MdY#16, MdY#33, MdY#36, and MdY#38) we found small indels exclusively in the target site of sgY3 (Table 2). In contrast, only line MdY#14 carries an allele of MdY with a lesion (2 bp deletion) in the sgY2 site (Table 2, Supplementary Fig. 4). In line MdY#19, we found evidence that both sites were targeted (Supplementary Fig. 4). The absence of a deletion spanning the two sites suggests that two dsDNA break events in our recovered alleles must have occurred at different times or with different kinetics, allowing the cellular repair system to independently join the breaks. In contrast, line MdY#40 carries a large deletion of 1038 bp which removes both target sites and sequences downstream of sgY2; MdY#40 likely had both sites simultaneously targeted by Cas9 and repaired to result in a larger deletion. Taken together, these data establish functional CRISPR-Cas9-mediated mutagenesis targeting the MdY locus in Musca domestica using in vitro-assembled Cas9-sgRNA RNPs.
To test for additional complementation, we crossed brown males of lines MdY#10 which carries a 15 bp deletion in the sgY3 site of MdY and MdY#16 containing a frame-shifting 10 bp deletion at the same site with brown females of the M III strain carrying the MdY a2 and MdY b alleles and the aabys strain, homozygous for the MdY a1 allele. In all crosses, all progeny displayed bwb phenotype. Lack of melanization in these animals is consistent with our hypothesis that MdY is required for proper pigmentation of the cuticle. Furthermore, non-complementation of the Cas9-induced MdY alleles with all mutant bwb alleles of the aabys and M III strains confirms our initial hypothesis that bwb corresponds to the yellow ortholog of Musca domestica.
Line MdY#29 was of particular interest, as we did not detect any sequence modifications at the two target sites in exon 2 (Table 2). Nonetheless, this line produced both brown M III pw + males and also pw females with normal melanization. The reciprocity of these sex-specific phenotypes suggests Cas9-mediated double-strand breaks in MdY induced an intragenic recombination event between the bwb + allele on the M III chromosome and the mutant bwb allele on the corresponding homolog. This event may have created a recombinant MdY allele with abolished activity on the chromosome that contains the M factor. In line with this hypothesis, we found brown males in line MdY#29 that are unaffected at the two Cas9 target sites, but homozygous for the MdY b signature (translational stop at position 65) (Fig. 3a). These sequences are likely the products of a reciprocal recombination event that may also have resulted in a reconstitution of a wildtype MdY allele on the non-M chromosome in females. Indeed, we found that melanized females are heterozygous for the MdY b signature in exon 1. Moreover, in the mutant bwb males, we detected heterozygosity for allele-specific polymorphisms just downstream of the target site sgY3 that correspond to the paternal genotype (Fig. 3a). We interpret these observations as evidence for Cas9-mediated DNA double-strand breaks at, or upstream of, the sgY3 site that induced an intragenic recombination event between the wildtype MdY allele and the MdY b allele in trans in germ cells of Cas9 RNP-injected heterozygous males (Fig. 3b). This data indicates that Cas9-mediated mutagenesis in Musca domestica can lead to break-point-guided recombination in injected germ cells.
Intragenic recombination in line MdY#29 mediated by CRISPR/Cas9. (a) Excerpts of chromatograms of exon 1 and 2 showing allele-specific polymorphisms. Mutant F1 bwb males are homozygous for the two variants in exon 1 (MdY b genotype) amplified with primers Y-ORF-F3 and RE1, but heterozygous for the three variants in exon 2 like in the paternal wildtype bwb G0 male amplified with primers FE4 Y-ORF-R5. Arrows point to the allele-specific polymorphisms examined. (b) This sequence analysis suggests that an intragenic recombination occurred downstream of the MdY b specific TGA stop codon in exon 1 and upstream of polymorphisms examined in exon 2. Locations of the SNPs between the two target sites (sgY3 ad sgY2) are indicated with short arrows.
Discussion
Our work reveals that the classic bwb phenotype in houseflies is caused by mutations in the Musca homolog of the DCE gene, which encodes the enzyme that in Drosophila has been implicated in the process that converts DOPA and/or dopamine into black melanin. Functional studies in Drosophila have provided evidence that the DCE-encoding gene yellow is involved in global body pigmentation21, whereas in the coleopteran Tribolium castaneum the loss of yellow only affects pigmentation of the hindwing5. Also, in the hemimetabolous Oncopeltus fasciatus, silencing of yellow affects only specific body parts such as abdomen and hindwings22. These observations led to the proposition that yellow belongs to a network of melanin synthesis genes which by differential deployment can generate a wide range of colors and spatial patterns22. Here, we present genetic evidence that, as in Drosophila, the Musca DCE homolog MdY is required for melanin production in the whole body. We base our conclusion on two major lines of evidence: First, complete absence of melanization found in two different bwb strains (M III and aabys) correlates with homozygosity for predicted nonsense alleles of MdY. In these strains, we identified three mutant alleles MdY a1, MdY a2 and MdY b, all of which result in premature stop codons. We note that MdY a1 and MdY a2 only differ with regard to a 1.5 kb insertion which is present specifically in the 5′ UTR of MdY a1. It is thus possible that these two alleles, both of which carry the same 4 bp insertion downstream of codon 67, have a common origin and that the 1.5 kb insertion has been acquired or lost later during a secondary mutational event. Second, we generated a set of new MdY alleles by adapting the CRISPR-Cas9 mutagenesis for disrupting the coding region in MdY exon 2. All of the new alleles have confined lesions at least in one of the two targeted sites. The majority of these mutations are deletions that remove parts of the coding sequence and generate frame-shifts. We hence consider these allelic variants to be enzymatically non-functional, if not null alleles of MdY. These Cas9-based MdY mutants fail to complement, and thus behave allelic to, the previously identified MdY variants found in bwb-mutant backgrounds.
The CRISPR-/Cas9 protocol using solubilized RNPs used in our study appears to be highly effective in the Musca germ line given that at least 1 of 6 RNP-injected males transmitted an MdY allele with a defined lesion at one of the targeted sites. Nonetheless, none of these F0 males displayed patches of non-melanized cuticle indicative of somatic mutation mosaicism. In Drosophila, y mutations behave cell-autonomously and even small y mutant clones can be readily detected in the cuticle: targeting y in Drosophila by injecting Cas9 mRNA and sgRNA produced strong somatic effects not only in males with one target gene but even in females with two copies of y 23. This study pointed out that the efficiency of inducing somatic clones not only depends on the concentration of sgRNA injected, but more importantly on the selection of the site in the y gene that was targeted. It is thus conceivable that the sgRNAs used in our work were able to efficiently disrupt the gene in the germline cells but not or less so in somatic cells. Germline transmission is a prerequisite for the investigation of any new mutation generated and, in this regard, our main objective was to recover fertile adults transmitting mutant MdY alleles. The lack of somatic effects in G0 adults can be a beneficial feature to avoid sterility that may be inflicted by the presence of mutant somatic tissue.
Recent work proposed that the CRISPR-Cas9 system can be used for genetic mapping by inducing targeted recombination events in meiotic and mitotic cells24. Our finding of an intragenic recombination event in line MdY#29 suggests that Cas9 induced breaks allows recombination between homologs in the germ line of housefly males. This observation suggests a possible Cas9-mediated strategy for male meiotic mapping in future studies. In addition, homologous recombination (HR)-mediated repair of Cas9 induced double-strand breaks in Musca offers the opportunity to attempt template-based editing of genomic sequences for targeted knock-ins in the future.
To our knowledge, our study is the first report showing that Cas9 can be effectively deployed for NHEJ mediated-disruption of genes in Musca domestica, an important addition to the toolkit of molecular methods that have already been established in Musca for gene function analysis. Together with transient RNAi-based gene silencing25, 26 and stable germline transformation16, 27, this new genome editing system provides a means to investigate evolutionary diversification of developmental pathways such as the polymorphic sex determination system of the housefly 15, 28. Our successful attempt promises that this genome editing technology can be used in the housefly to study the function of any candidate gene of interest.
Methods
Rearing of houseflies
Rearing of larvae and adult flies has been described in refs 29, 30. Since low density of larvae on standard medium can cause substantial decrease in survival rates, we reared injected embryos and the surviving larvae on porcine manure. To dispose of mites and other parasites and to avoid contamination with eggs or larvae from wild-type populations, manure was stored at −70 °C for at least two weeks prior to use.
Strains of Musca domestica
-
(1)
Wildtype strain was collected in Siat, Switzerland: females X/X; bwb +/bwb + and males X/Y; bwb +/bwb +
-
(2)
Multi-marked strain aabys: females X/X; ac/ac; ar/ar; bwb/bwb; ye/ye; snp/snp and males X/Y ac/ac; ar/ar; bwb/bwb; ye/ye; snp/snp 12;
-
(3)
Autosomal M III strain: females X/X; pw, bwb, w/pw, bwb, w and males X/X; pw +, M III, bwb +, w/pw, bwb, w 16.
Genomic DNA extraction
For genomic DNA extraction a single fly was collected in a 1.5 ml tube, frozen in liquid nitrogen and ground in 1 ml of extraction buffer (0.1 M Tris-HCl, pH 9; 0.1 M EDTA; 1% SDS and 1% of DMDC added freshly). After 30 min incubation at 70 °C, 140 µl of 8 M potassium acetate was added and sample was gently inverted and incubated for 30 min on ice. After 15 min of centrifugation at 4 °C at 13,000 rpm, supernatant was transferred to a new tube, and 550 µl of isopropanol was added. The mixture was centrifuged at RT for 5 min at 14,000 rpm and the supernatant was removed. The pellet was washed with 500 µl of 70% EtOH (−20 °C) and centrifuged at RT for 2 min at 14,000 rpm. The DNA pellet was finally dissolved in 30 or 50 µl of 10 mM Tris and 1 µl of RNaseA (10 mg/ml) to remove RNA. Amplifications for sequence analysis were performed with following primers. For exon 1 we used forward:
Y-ORF-F3 (5′-TGCTGTGGACATTGGCAAGA-3′) and reverse RE1: (5′-TCTCATTCACATCCACACCGT-3′).
For exon2 we used forward FE4 (5′-CAGGTATACCAGCCACATTGA-3′) and reverse Y-ORF-R5 (5′-CTAATGATGGGCGGATGTGGA-3′).
For insertion we used flanking primer forward Y-GAP1-F1 (5′-GGCCGAAGTGAGACAGAGAA-3′) and Y-EXON1-R (5′-CTAGTGGCGAAAAACCATTAA-3′).
sgRNA synthesis and RNP complex assembly
sgRNA were designed using MultiTargeter Website (http://www.multicrispr.net). Possible OFF-target sites were excluded with the program Cas-OFFinder (http://www.rgenome.net/cas-offinder/) and by directly BLASTing selected sequences against the published housefly genome sequence12. Following templates for sgRNA production were generated:
sgY2: 5′-GAAATTAATACGACTCACTATA GGCTTTGTCGCCCATTCGTT GTTTTAGAGCTAGAAATAGC-3′.
sgY3: 5′ -GAAATTAATACGACTCACTATA GGCATAGGGACAGGGGTTGG GTTTTAGAGCTAGAAATAGC-3′.
Common sgReverse (PAGE purified): 5′AAAAGCACCGACTCGGTGCCACTTTTTCAA.
GTTGATGGACTAGCCTTATTTTAACTTGCTATTTCTAGCTCTAAAAC AAC.
For synthesis of sgRNA we followed the instructions of Megatranscript T7 kit (Ambion) using 400 ng of target template with a 5′ flanking T7 promoter as starting material. After RNA synthesis template was removed by incubating with TurboDNase (Mmessage Mmachine T7 Ultra Kit, Ambion) for 15 min at 37 °C23.
Cas9 was expressed as an His-tagged protein and purified from bacteria18. The injection cocktail was prepared by mixing 1.5 µl purified Cas9 protein (9 mg/ml) with 2 µl of sgRNA in 1.36 µl of 2 M KCl in a total of 10 µl. Prior injection the mix was incubated for 10 min at 37 °C19. For injection we prepared a 1:1 mix of sgY3-preloaded Cas9 RNPs and sgY2-preloaded Cas9 RNPs.
Microinjection of sgRNA-Cas9 complexes
Embryos of the M III host strain were collected 1 hour after egg lay and chorion membrane was removed by incubating embryos in 3% sodium hypochlorite solution (NaOCl) for 1.5 min. Dechorionated embryos were then rinsed thoroughly with water and Ringer’s solution. Embryos were aligned on a cover slip with posterior ends pointing to injection site, dehydrated for 4 min in a silicagel chamber and then covered with 3 S/10 S (1:4) Voltalef oil (Prolabo). A glass needle was filled with the preloaded sgRNA-Cas9 mix which was injected into the posterior end of 0 to 1 hour old embryos. Injections were performed using a micromanipulator connected to an Eppendorf Femtojet pumping device which was set to inject approx. 5 nl per embryo. After injection excess Voltalef oil was carefully removed and cover slip were put on an agar plate overnight at 25 °C. Surviving larvae were transferred after 24 hours to small beaker filled with porcine manure. G0 male individuals were collected shortly after eclosing and crossed with untreated virgin females of the M III strain.
Genomic analysis of CRISPR-Cas9 mediated lesions in MdY
To examine for the presence of lesions in MdY caused by NHEJ, genomic DNA of bwb mutant F1 males was extracted following the protocol described above. The region encompassing the two target sites was amplified with the following primer:
forward primers FE3: TCTGGCAAACCACAACAAGT or F4 CAGGTATACCAGCCACATTGA; reverse primers Y-ORF-R1: GACGAATGCCAACAACCCAC or Y-ORF-R5 CTAATGATGGGCGGATGTGGA.
PCR products were purified with the Wizard® Genomic DNA Purification Kit (Promega), subcloned in the pGEM®-T Easy Vector (Promega) and sent for Sanger sequencing (GATA BIOTECH). CrispRVariants and the generation of panel plots was performed from primary sequencing data as previously described19, 20.
References
Malacrida, A., Grigolo, A., Gasperi, G. & Sacchi, L. Activity of the enzyme dopa-decarboxylase in brown body (bwb) mutants of Musca domestica L. Boll. Soc. Ital. Biol. Sper. 50, 1566–1569 (1974).
Wright, T. R. The genetic and molecular organization of the dense cluster of functionally related, vital genes in the DOPA decarboxylase region of the Drosophila melanogaster genome. Results Probl Cell Differ 14, 95–120 (1987).
Wittkopp, P. J., Williams, B. L., Selegue, J. E. & Carroll, S. B. Drosophila pigmentation evolution: divergent genotypes underlying convergent phenotypes. Proc Natl Acad Sci USA 100, 1808–1813 (2003).
Wittkopp, P. J. et al. Intraspecific polymorphism to interspecific divergence: genetics of pigmentation in Drosophila. Science 326, 540–544 (2009).
Arakane, Y. et al. Identification, mRNA expression and functional analysis of several yellow family genes in Tribolium castaneum. Insect Biochem Mol Biol 40, 259–266 (2010).
Futahashi, R. & Fujiwara, H. Regulation of 20-hydroxyecdysone on the larval pigmentation and the expression of melanin synthesis enzymes and yellow gene of the swallowtail butterfly, Papilio xuthus. Insect Biochem Mol Biol 37, 855–864 (2007).
Futahashi, R. et al. yellow and ebony are the responsible genes for the larval color mutants of the silkworm Bombyx mori. Genetics 180, 1995–2005 (2008).
Ito, K. et al. Yellow-e determines the color pattern of larval head and tail spots of the silkworm Bombyx mori. J Biol Chem 285, 5624–5629 (2010).
Wittkopp, P. J., Vaccaro, K. & Carroll, S. B. Evolution of yellow gene regulation and pigmentation in Drosophila. Curr Biol 12, 1547–1556 (2002).
Weller, G. L. & Foster, G. G. Genetic maps of the sheep blowfly Lucilia cuprina: linkage-group correlations with other dipteran genera. Genome 36, 495–506 (1993).
Meisel, R. P., Scott, J. G. & Clark, A. G. Transcriptome Differences between Alternative Sex Determining Genotypes in the House Fly, Musca domestica. Genome Biol Evol 7, 2051–2061 (2015).
Scott, J. G. et al. Genome of the house fly, Musca domestica L., a global vector of diseases with adaptations to a septic environment. Genome Biol 15, 1–16 (2014).
Begun, D. J. & Aquadro, C. F. Molecular population genetics of the distal portion of the X chromosome in Drosophila: evidence for genetic hitchhiking of the yellow-achaete region. Genetics 129, 1147–1158 (1991).
Gao, J.-R., Deacutis, J. M. & Scott, J. G. Characterization of the nicotinic acetylcholine receptor subunit gene Mdalpha2 from the house fly, Musca domestica. Arch. Insect Biochem. Physiol. 64, 30–42 (2007).
Dübendorfer, A., Hediger, M., Burghardt, G. & Bopp, D. Musca domestica, a window on the evolution of sex-determining mechanisms in insects. Int J Dev Biol 46, 75–79 (2002).
Hediger, M. et al. Molecular characterization of the key switch F provides a basis for understanding the rapid divergence of the sex-determining pathway in the housefly. Genetics 184, 155–170 (2010).
Lee, J.-S. et al. RNA-guided genome editing in Drosophila with the purified Cas9 protein. G3: Genes|Genomes|Genetics 4, 1291–1295 (2014).
Meccariello, A. et al. Highly efficient DNA-free gene disruption in the agricultural pest Ceratitis capitata by CRISPR-Cas9 RNPs., doi:10.1101/127506
Burger, A. et al. Maximizing mutagenesis with solubilized CRISPR-Cas9 ribonucleoprotein complexes. Development 143, 2025–2037 (2016).
Lindsay, H. et al. CrispRVariants charts the mutation spectrum of genome engineering experiments. Nature Biotechnology 34, 701–702 (2016).
Wittkopp, P. J., True, J. R. & Carroll, S. B. Reciprocal functions of the Drosophila yellow and ebony proteins in the development and evolution of pigment patterns. Development 129, 1849–1858 (2002).
Liu, J., Lemonds, T. R., Marden, J. H. & Popadić, A. A Pathway Analysis of Melanin Patterning in a Hemimetabolous Insect. Genetics 203, 403–413 (2016).
Bassett, A. R., Tibbit, C., Ponting, C. P. & Liu, J.-L. Highly efficient targeted mutagenesis of Drosophila with the CRISPR/Cas9 system. Cell Rep 4, 220–228 (2013).
Sadhu, M. J., Bloom, J. S., Day, L. & Kruglyak, L. CRISPR-directed mitotic recombination enables genetic mapping without crosses. Science 352, 1113–1116 (2016).
Hediger, M. et al. Sex determination in Drosophila melanogaster and Musca domestica converges at the level of the terminal regulator doublesex. Dev Genes Evol 214, 29–42 (2004).
McGregor, A. P. et al. Rapid restructuring of bicoid-dependent hunchback promoters within and between Dipteran species: implications for molecular coevolution. Evol Dev 3, 397–407 (2001).
Hediger, M., Niessen, M., Wimmer, E. A., Dübendorfer, A. & Bopp, D. Genetic transformation of the housefly Musca domestica with the lepidopteran derived transposon piggyBac. Insect Mol Biol 10, 113–119 (2001).
Bopp, D. About females and males: continuity and discontinuity in flies. J Genet 89, 315–323 (2010).
Hilfiker-Kleiner, D., Dübendorfer, A., Hilfiker, A. & Nöthiger, R. Genetic control of sex determination in the germ line and soma of the housefly, Musca domestica. Development 120, 2531–2538 (1994).
Schmidt, R., Hediger Niessen, M., Nöthiger, R. & Dübendorfer, A. The Mutation masculinizer (man) Defines a Sex-Determining Gene With Maternal and Zygotic Functions in Musca domestica L. Genetics 145, 173–183 (1997).
Acknowledgements
We thank Claudia Brunner for technical assistance, Raymond Grunder and Daniel Christen for help with the rearing of housefly cultures. We are grateful to Elena Chiavacci for her help with the target design and processing of CRISPR sequence data. This work was supported by the Canton of Zürich, a Schweizerischer Nationalfonds zur Förderung der Wissenschaftlichen Forschung (SNSF) professorship [PP00P3_139093] and a Marie Curie Career Integration Grant from the European Commission to C.M.; a University of Zurich URPP Translational Cancer Research Seed Grant to A.B., G.S. and A.M. were visiting researchers at University of Zurich (November 2015), supported by University Federico II of Naples (International Exchange Program to GS and by the Biology PhD program of University Federico II of Naples to AM). The nucleotide sequence of MdY has been submitted to GenBank (ID 2011210).
Author information
Authors and Affiliations
Contributions
D.B. and G.S. conceived the experiments, S.D.H., T.K., D.I., A.M. and A.B. conducted the experiment(s). All of the authors discussed the data and helped manuscript preparation. D.B., G.S., and C.M. wrote the manuscript with intellectual input from all authors.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Heinze, S.D., Kohlbrenner, T., Ippolito, D. et al. CRISPR-Cas9 targeted disruption of the yellow ortholog in the housefly identifies the brown body locus. Sci Rep 7, 4582 (2017). https://doi.org/10.1038/s41598-017-04686-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-04686-6
This article is cited by
-
Characterization of essential eggshell proteins from Aedes aegypti mosquitoes
BMC Biology (2023)
-
Disruption of duplicated yellow genes in Bactrocera tryoni modifies pigmentation colouration and impacts behaviour
Journal of Pest Science (2021)
-
Targeted delivery of CRISPR-Cas9 ribonucleoprotein into arthropod ovaries for heritable germline gene editing
Nature Communications (2018)
-
Highly efficient DNA-free gene disruption in the agricultural pest Ceratitis capitata by CRISPR-Cas9 ribonucleoprotein complexes
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.