Abstract
Generation of multiple mRNAs by alternative splicing is well known in the group of cytokines and has recently been reported for the human erythropoietin (EPO) gene. Here, we focus on the alternatively spliced EPO transcript characterized by deletion of exon 3 (hEPOΔ3). We show co-regulation of EPO and hEPOΔ3 in human diseased tissue. The expression of hEPOΔ3 in various human samples was low under normal conditions, and distinctly increased in pathological states. Concomitant up-regulation of hEPOΔ3 and EPO in response to hypoxic conditions was also observed in HepG2 cell cultures. Using LC-ESI-MS/MS, we provide first evidence for the existence of hEPOΔ3 derived protein EV-3 in human serum from healthy donors. Contrary to EPO, recombinant EV-3 did not promote early erythroid progenitors in cultures of human CD34+ haematopoietic stem cells. Repeated intraperitoneal administration of EV-3 in mice did not affect the haematocrit. Similar to EPO, EV-3 acted anti-apoptotic in rat hippocampal neurons exposed to oxygen-glucose deprivation. Employing the touch-screen paradigm of long-term visual discrimination learning, we obtained first in vivo evidence of beneficial effects of EV-3 on cognition. This is the first report on the presence of a naturally occurring EPO protein isoform in human serum sharing non-erythropoietic functions with EPO.
Similar content being viewed by others
Introduction
The cytokine erythropoietin (EPO) was firstly identified as the haematopoietic factor responsible for the proliferation and maturation of erythroid progenitor cells by preventing their apoptosis1. Meanwhile, EPO has been described as a protein with multifaceted properties. Besides stimulating erythropoiesis, EPO acts as growth and survival factor1 and is part of an endogenous tissue protection system2,3,4,5. Neuroprotective actions of EPO have been extensively shown in vitro (e.g. 6,7,8) and in vivo (e.g. 9,10,11) and efficacy in clinical trials using EPO for the treatment of a variety of degenerative disorders has been reported (e.g. 12,13,14,15,16).
Although the long scientific history of EPO – it was isolated in 197717 from human urine and cloning of its gene was only several years later18 – alternatively spliced EPO transcripts were only recently described19: An EPO transcript variant with complete loss of exon 3 (denoted hS3) and a second variant with a partial loss of exon 4 (denoted hS4) were detected in samples of human kidney and foetal brain. The generation of multiple mRNAs by alternative splicing is well known in the group of cytokines, for example for IL-220, IL-421, IL-1022, IL-1223 and G-CSF24. Several splice variants are also known for EPOR 25. While some splice variants act as competitive inhibitors of the full-length protein20, 21, other acquire different biological functions26. For the recombinant protein products derived from the two alternatively spliced transcripts of EPO, hS3 and hS4 respectively, neuroprotective properties and in parallel, a lack of erythropoietic function were described19.
In the present study, we focussed on the splice variant hS3 (herein denoted as hEPOΔ3) and extended the panel of human samples for the detection of the spliced transcript according to the reported EPO expression27,28,29,30. Elevated levels of EPO were found in patients with chronic liver disease31, 32 and in individual samples of clear cell renal cell carcinoma (ccRCC)33,34,35. In order to test whether the splice variant hEPOΔ3 is also regulated under those pathological conditions, we established a realtime-PCR assay to quantify EPO and hEPOΔ3 transcript levels in human diseased (liver cirrhosis, ccRCC) and normal tissues. The primary factor for up-regulation of EPO expression is hypoxia (for review see 36) and the human hepatocellular carcinoma cell line HepG2 is known to express EPO37. Therefore, we used HepG2 cells to study hypoxia-related regulation of hEPOΔ3 compared to EPO in a model system.
Not all splice variants are functionally relevant and are often products of errors of the splicing process38. We expected a functionally relevant spliced EPO transcript to be translated into protein and secreted into the blood circulation. EPO itself is only present in picomolar concentrations in blood39. Further, the results obtained from the experiments mentioned above pointed to a low expression of the splice variant hEPOΔ3 under basal conditions. Therefore, in order to be able to detect the anticipated low levels of hEPOΔ3 derived protein (EV-3) and allow clear separation from EPO in serum of healthy human volunteers, we applied a multistep processing protocol and analysed the samples by LC-ESI/MS-MS. Based on the cDNA sequence of hEPOΔ319 the corresponding protein EV-3 is supposed to lack amino acids encoded by exon 3. The peptide sequence spanning the last three amino acids encoded by exon 2 and the first six amino acids encoded by exon 4 is uniquely contained in EV-3 and therefore, allows its identification as well as differentiation to EPO in human serum samples.
For functional studies, EV-3 was produced by transient expression in CHO cells, the commonly employed cell line for production of recombinant EPO. Orienting our studies with EV-3 on previously described functional activity of EPO, we here focused on erythropoietic, neuroprotective and potential cognitive effects of the recombinant produced EV-3 in comparison to EPO. The results indicate that neuroprotective and beneficial effects on cognition are preserved in EV-3, whereas erythropoietic effects of EPO are not mediated by the EV-3 protein. Together with the observed co-regulation of the transcripts of EPO and EV-3 in human specimen we speculate that tissue protective and/or regenerative effects are part of the biological function of the here described EV-3 protein detected in the human serum.
Methods
PCR
Total RNA from human tissues (adult kidney, foetal brain) was purchased from Biocat (Germany). Total RNA from HepG2 cells was extracted using peqGold RNAPure (VWR, Germany) according to standard protocols. The Access-RT-PCR system from Promega (Germany) was used for reverse transcription and PCR amplification of EPO transcripts using RNA (3 µg) and EPO-specific forward and reverse primers (FWD: 5′-GATGGGGGTGCACGAAT GTCCTGC-3′; REV: 5′-CACACCTGGTCATCTGTCCCCTGTC-3′) according to manufacturer’s instructions. This PCR product was used for a second PCR amplification round using internal EPO primers (FWD: 5′-TCCTGTCCCTGCTGTCGCTC-3′; REV: 5′-CTGCAGGCCTCCCCTGTGTA-3′) and Pfu polymerase (Promega). PCR products were separated on agarose gels, purified using the QIAquick Gel Extraction Kit (Qiagen, Germany) and subcloned into the Zero Blunt TOPO vector (Thermo Fisher Scientific, Germany) for sequencing.
Quantitative PCR analysis in human tissues
Human tissue samples were handled in accordance with the ethical rules of the Klinikum Rechts der Isar, Technical University Munich, for research use of surplus diagnostic materials. Written informed consent was obtained from all patients. Total RNA from formalin fixed paraffin-embedded (FFPE) human tissues was isolated using the High Pure RNA Paraffin Kit from Roche. In brief, tumour cells from clear cell renal cell carcinoma (ccRCC; n = 12) samples (Fuhrmann-grading G2 and G3) were macrodissected and compared to normal adjacent renal tissue. Cirrhotic liver tissues were compared to normal liver samples (n = 10). Total RNA from fresh human tissues (adult kidney, foetal brain, adult brain, liver and cirrhotic liver) was purchased from Biocat (Germany) and Amsbio (UK). RNA reverse transcription was performed on RNA (2 µg) with Superscript II, RNase Out, random primers and dNTPs. Primers specific for EPO transcript (SPEC_EPO; FWD: 5′-GTGCTGAACACTGCAGCTTG-3′; REV: 5′-CAGACTTCTACGGCCTGCTG-3′) and specific for hEPOΔ3 transcript (SPEC_EV-3; FWD: 5′-CACCACGCCTCATCTGTGAC-3′; REV: 5′-CTGCTGCCCGACCGTGA-3′) were ordered from Eurofins MWG Synthesis GmbH (Germany). QuantiTect primer assays for Alas1 (Hs_Alas1_1_SG), SRSF4 (Hs_SRSF4_1_SG) and UBC (Hs_UBC_1_SG) were purchased from Qiagen. Real-time PCR reactions were performed in the Lightcycler 480 apparatus using Light Cycler 480 SYBR Green I Master (Roche) with the following cycling: 5 min at 95 °C; 45 cycles: 10 sec at 95 °C, 17 sec at 55 °C, 10 sec at 72 °C; acquisition at 55 °C. SPEC_EV3 acquisition was done at 82 °C to avoid primer dimer generation, which significantly affects low copy EV-3 detection. Melting curve analysis was done to check for unspecific products. Efficiency of SPEC_EPO and SPEC_EV-3 assays was tested using dilution series of cDNA. Serial dilutions of plasmids containing EPO full-length and hEPOΔ3 inserts were used to test for primer specificity.
In silico homology modelling
Homology modelling of EV-3 was performed at the SWISS-MODEL Protein Modelling Server database40,41,42,43 accessible via ‘http://swissmodel.expasy.org/’ using the EPO structure 1cn4.1.C.pdb as template.
Recombinant proteins
Recombinant EV-3 and EPO proteins were produced by IBA GmbH (Germany) as Twin-Strep-Tag fusion proteins via transient transfection in CHO cells. Proteins were purified from cell culture supernatants using Strep-Tactin® Superflow® high capacity, dialyzed against D-PBS and sterile filtered. Silver staining of recombinant proteins after separation by SDS-PAGE revealed purity of >95%. The endotoxin contamination ranged from 8.3–15.1 EU/mg protein (Limulus Amebocyte Lysate Test, Endosafe nexgen PTS, Charles River), resulting in an endotoxin level of <0.01 ng/ml injection solution used for in vivo experiments.
HepG2 culture and hypoxia
HepG2 cells were maintained in complete high-glucose DMEM medium (DMEM supplemented with 10% FCS and 1% penicillin/streptomycin). To study effects of hypoxia on EPO mRNA expression, cells were cultured in 6-well plates (1 × 106 cells/well). After 24 hours, medium was exchanged and cultures were exposed to hypoxia (oxygen concentration <1%) or normoxia for 4 hours. Immediately thereafter, cells were lysed and RNA was extracted (miRNeasy Kit, Qiagen) and DNase digested. Reverse transcription was performed on RNA (1 µg) with Superscript III, RNase Out, random hexamers, oligo-dT primers and dNTPs. PPIA (Hs_PPIA_1_SG) primer assay was purchased from Qiagen. Real-time PCR reactions were performed in the Lightcycler 480 apparatus using the Power SYBR Green PCR Master Mix from Thermo Fisher Scientific with the following cycling: 2 min at 50 °C, 10 min at 95 °C; 45 cycles: 15 sec at 95 °C, 1 min at 60 °C, acquisition at 60 °C. Melting curve analysis was done to check for unspecific products. Primer dimer generation for SPEC_EV-3 was not detected in HepG2 samples.
Analysis of HepG2 cell culture supernatants
Cell culture supernatants were harvested after 3–4 days, pooled and concentrated (Vivaspin, MWCO 10,000; Sartorius, Germany). For immunoprecipitation, biotinylated anti-EPO antibody (0.5 µg; BAF959, R&D) was added to concentrated supernatant (750 µl) and incubated over-night at 4 °C on a rotating shaker. Streptavidin-agarose was added and incubated for 1 hr at room-temperature. After three PBS-washing and centrifugation (700 g, 1 min, 4 °C) steps, agarose was re-suspended in SDS sample buffer and heated for 5 min at 99 °C. For subsequent analysis in mass spectrometry, proteins were eluted using 0.1 M glycine (pH 2.6) and neutralized with 2 M Tris (pH 8.5). Western Blotting was performed using a polyclonal rhEPO-antibody (H-162, Santa-Cruz) according to standard protocols; the ImageJ-program was used for densitometry analysis. Wheat germ affinity (WGA) chromatography was performed according to manufacturer’s instructions (Thermo Fisher Scientific) on concentrated HepG2 culture supernatants. Eluates from immunoprecipitation and WGA purification were desalted, concentrated and transferred to PNGase F (New, England Biolabs, Frankfurt, Germany) and endoprotease digestion (Asp-N/Glu-C) for analysis using multiple reaction monitoring (see below).
Serum fractionation
Sera (n = 2; 20 ml of serum 1 and 100 ml of serum 2) of healthy human donors were fractionated via size exclusion chromatography. To this end, Superdex 75 columns for high resolution separation of biomolecules in the range of 3–70 kDa with different loading volume were used (XK 26/60: 2.6 cm diameter, 318 ml Superdex 75, flow rate set to 2.5 ml/min; BGP 100/950: 10 cm diameter, 5.2 L Superdex 75, flow rate set to 20 ml/min; GE Healthcare Life Sciences). The elution time of EPO was predicted from a calibration run (see Supplementary Figs S3a and S4a) using BSA (66 kDa) or a solution containing BSA and Azurin (30 kDa). Serum 1 (20 ml) was loaded on the XK 26/60 column and fractions of interest, having same elution times as Azurin in the calibration run, were pooled. The pool was concentrated using VivaFlow 50 (MWCO 10,000; Sartorius, Germany) and re-run twice on the XK 26/60 column until obtaining a narrow peak for serum albumin (Supplementary Fig. S3b). Serum 2 (100 ml) was first loaded on the BPG 100/950 column. Eluate fractions of interest according to the calibration run following the albumin peak, were pooled, concentrated and re-applied to the column. Elution fractions of the second run were chromatographed twice on the XK 26/60 column until obtaining a clear separation of proteins smaller than albumin (see Supplementary Fig. S4b). Final eluate fractions of serum 1 (C3 – C12, D2) and serum 2 (C4– C12, D1) were analysed separately. Using a commercial ELISA (Roche Diagnostics, Germany), EPO-positive fractions were identified and further purified using WGA chromatography. Eluates were PNGase F and endoprotease digested (Asp-N/Glu-C) for analysis using multiple reaction monitoring (see below).
Multiple reaction monitoring (nano-LC-ESI-MS/MS)
Through a theoretical enzymatic digest peptides were searched which allow differentiation of EPO and EV-3 within one sample. Processing of the samples included digestion with PNGase F which not only removes N-linked oligosaccharides from glycoproteins but also deaminates glycosylated N-residues to D-residues (N → D). The subsequent cleavage of de-glycosylated proteins with endoproteases Asp-N (cuts before D) and Glu-C (cuts after E) generated EV-3 and EPO specific peptides: ‘DITVGQQAVE’ is specific for EV-3 and ‘DITTGCAE’ is specific for EPO (Table 1).
10 μg of PNGase treated sample was reduced with Dithiothreitol (DTT), alkylated with iodacetamide (IAA), then first digested with Asp-N (0.2 μg) at 37 °C over-night, then with Glu-C (0.2 μg) at room temperature for 4 hours. After completion of digest, the sample was lyophilized, re-dissolved in 0.1% formic acid (100 μl) and spiked with each isotopically labelled peptide (DITVGQQAVE; DITTGCAE). For nano-LC-ESI-MS/MS, total digested protein (3.5 μg) was applied. HPLC-separation was done on an Ultimate 3000 system (Dionex): Peptides were loaded onto a C18-column and subsequently separated on a PepMap™ analytical column (LC PACKINGS Nano SeriesTM PepMapTM C18) with linear gradients. Mass spectrometry was done using a QTRAP 5500 mass-spectrometer (ABSciex) coupled online to the HPLC-system. Identification and quantitation of the endogenous peptides, the EV-3 specific DITVGQQAVE and the EPO specific DITTGCAE, was based on retention times of the isotopically labelled peptides spiked to the enzymatically digested samples. Quantitation utilizes 5 transitions for each peptide. Standard curves were generated with linear regression as duplicates for each transition separately using peptide concentrations of 0.001, 0.01, 0.1, 1 and 10 fmol/μl.
Colony forming unit (CFU) assay with human CD34+ cells
CD34+ cells were used in accordance with the guidelines approved by the Ethics Committee of the Dresden University of Technology. The informed consent was obtained from all donors. Granulocyte colony-stimulating factor (G-CSF) mobilized CD34+ cells from peripheral blood of two healthy donors were isolated by centrifugation over Biocoll (Biochrom) and by the use of CD34 microbeads and LS-MACS columns (Miltenyi Biotec; 94% ± 3% purity). CD34+ stem cells, for each donor separately, were seeded in methylcellulose plates (500 cells per well, 6 well plates, Stem Cell Technologies, Canada) with 1.1 ml of medium (Methocult H4035 without EPO; Stem Cell Technologies). For treatment, different concentrations (0, 0.3, 3, 10, or 30 U/ml or weight equivalent) of rEPO or rEV-3 were supplemented to Methocult H4035. As positive control, CD34+ stem cells were seeded in MethoCult H84434 (includes EPO, Stem Cell Technologies). Colony-forming units (BFU-E; colonies >50 cells, CFU-E; colonies <50 cells, CFU-G/M/GM and CFU-GEMM) were automatically analysed and counted after 14 days of incubation at 37 °C using STEMvision equipment (Stem Cell Technologies). CFUs were examined in duplicates.
Hypoxia in primary hippocampal neurons
Hippocampal neurons were prepared from new-born Wistar-Imamichi rats according to Lewczuk44 with slight modification. Briefly, hippocampi were dissected in HBSS solution supplemented with 0.1% D-Glucose, 1 mM sodium bicarbonate, 1 mM pyruvate and 10 mM HEPES, and transferred into precooled HBSS/B27 solution (0.2% B27) until trituration. Hippocampi were incubated for 30 min with papain (4 ml, 184 U/ml) at 30 °C and shaken every 5 min. 25 µl of DNase I (10 mg/ml in HBSS) were added for 2 min. The hippocampi were triturated in HBSS/B27 and the cell suspension was underlayered with a density gradient prepared in four 1 ml steps of 35, 25, 20 and 15% of Optiprep (Thermo Fisher Scientific) in HBSS/B27. The cell suspension was centrifuged for 15 min (2000 rpm, RT). Neuron-enriched fractions were collected and washed with 5 ml HBSS/B27. After centrifugation at 1100 rpm for 2 min the pellet was resuspended in growth medium (1 ml; NeurobasalA, 0.2% B27, 5 ng/ml bFGF, 0.5 mM L-glutamine, 100 U/ml penicillin and 100 µg/ml streptomycin). The neurons were plated on poly-D-lysine coated 12 mm coverslips in 4-well-plates, with a density of 8000 cells/coverslip. At day 3 the medium was replaced by starvation medium (growth medium without bFGF and B27) complemented with EPO 3 U/ml (NeoRecormon, Roche, UK), EV-3 at 0.3 U/ml and 3 U/ml weight equivalent of EPO, or placebo (solvent solution). At day 5, cultures were exposed to hypoxia or normoxia for 20 hours. Cultures were stained with 0.4% Trypan blue. Total cell number and number of dead cells were counted. Results are presented as % cell death; the hypoxia effect was set to 100%.
Animals
Juvenile (28 day-old) male C57BL/6NCrl mice were used. They were housed in groups of 5 in standard plastic cages and maintained in a temperature-controlled environment (21–22 °C) on a 12 hours light/dark cycle with food and water available ad libitum. All experiments were approved by and conducted in accordance with the regulations of the local Animal Care and Use Committee (Niedersächsisches Landesamt für Verbraucherschutz und Lebensmittelsicherheit—LAVES).
Treatment
Mice (10–11 per group) were injected intraperitoneally (i.p.) with recombinant human erythropoietin (EPO) (NeoRecormon, Roche, 5000 IU/kg body weight), EV-3 (IBA, weight equivalent) or placebo (solvent solution), each at a volume of 0.01 ml/g. Treatment started at the age of 28 days and continued every other day for 3 weeks (11 injections in total). After the last injection, the animals were adapted for 1 week to the food deprivation procedure (85% of free feeding body weight) prior to start of the touch-screen experiment.
Haematocrit determination
A separate cohort of mice either received EPO (n = 15), EV-3 (n = 14) or placebo (n = 15) injections as described above. Mice were tested for haematocrit levels 9 days after the last injection. Blood samples were collected in EDTA Microvettes, transferred into heparinized capillaries, centrifuged and the volume percentage of erythrocytes in whole blood was then determined using a haematocrit reading device. T-tests for independent samples were used to compare haematocrit levels between groups.
Cognitive testing (Touch-screen visual discrimination task)
Detailed description of the operant chambers and experimental procedures including visual stimuli have been described previously45. In brief, every mouse was subjected to the following sequence of test phases: (1) acclimation, (2) autoshaping, (3) pre-training 1–3, and (4) visual discrimination. The daily sessions were terminated after 30 min or if the mouse had accomplished the specific learning criterion of the respective test phase. Acclimation: The animal learned that food pellets are available in the food magazine. Learning criterion: Collection of 10 pellets within 30 min. Autoshaping: The animal learned to associate the presentation of visual stimuli on the touch-screen with the delivery of food pellets. Learning criterion: Collection of 30 pellets within 30 min. Pre-training 1: The animal learned that a touch response to the correct side of the screen, i.e. where a visual stimulus was presented, is rewarded by the delivery of a food pellet. Learning criterion: 30 correct responses within 30 min. Pre-training 2: Now, the animal had to learn to initiate a new trial by disrupting the infrared light beam in the magazine a second time (after it had collected the food pellet). Pre-training 3: During this stage the animal had to memorize that an incorrect response resulted in a 5 sec timeout and the initiation of a correction trial. Learning criterion: At least 27 correct responses out of 30 trials. Visual discrimination: Here, the animal had to discriminate between a rewarded visual stimulus S+ (correct response rewarded with food pellet) and a non-rewarded stimulus S- (incorrect response resulted in timeout). Learning criterion: At least 27 correct responses out of 30 trials or 26 correct responses out of 30 trials on 2 consecutive days, whatever appeared first. The following performance readouts were measured: 1. Days needed to achieve the pre-training criteria (1–3), 2. Days needed to achieve the learning criterion in the visual discrimination task, 3. Days needed to complete whole sequence of tasks (pre-training and visual discrimination). Statistics: Touch-screen data were analysed by means of Kaplan-Meier analysis. Two-tailed p-values lower than 0.05 were considered to be significant.
Results
Alternative splicing of EPO in human tissue confirmed
At baseline condition, EPO mRNA is expressed at very low levels46, 47. Therefore, we used a two-step PCR approach to amplify EPO mRNA from human foetal brain and human kidney total RNAs. Primer sequences for reverse transcription and first PCR amplification were taken from the original discovery description19 lying at 5′ (exon 1) and 3′ (exon 5) borders of the coding region. A second primer pair lying in exon 2 and exon 5 was used in a second PCR, using products from the first PCR as template (Fig. 1a). Electrophoretic separation of 2-step PCR amplification products revealed the expected PCR product at approximately 550 bp corresponding to EPO truncated by 17 nucleotides at 5′ and by 16 nucleotides at 3′ through the usage of the second primer pair lying in exon 2 and exon 5 of the EPO gene. A second PCR product at approximately 460 bp corresponding to the 495 bp product described before19 was observed in both adult kidney and foetal brain samples (Fig. 1b). The amplification products were cut from the agarose gel and purified separately. Subcloning and sequencing confirmed the previously reported results19, i.e. revealed the EPO full-length form for the long PCR products and a shorter EPO form containing an internal 87-nucleotide deletion ranging from nucleotide 160 to 246. The human EPO gene consists of five exons and four introns. The missing nucleotides in the short EPO transcript (hEPOΔ3) correspond to exon 3. The hEPOΔ3 transcript is seemingly generated through alternative splicing using the AG dinucleotide at the 3′ end of exon 3 as alternative splice acceptor site17.
Isolation of the human EPO spliced transcript hEPOΔ3. (a) Reverse transcription and PCR amplification is performed with a set of primers for full-length amplification of EPO lying 5′ and 3′ of the coding sequence in exon 1 and exon 5. Then some product is taken for re-amplification with an internally-situated set of primers lying in exon 2 and exon 5. (b) Agarose gel of EPO-PCR amplification products from human adult kidney and foetal brain reveals two products at 550 bp and 460 bp, respectively.
Expression of the hEPOΔ3 transcript is upregulated in disease conditions
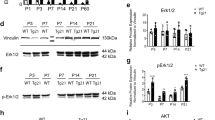
Alternative splicing is often modulated according to cell type or in response to external stimuli48. Therefore, in a pilot study we studied tissue expression of EPO and hEPOΔ3 transcripts in a heterogeneous cDNA panel derived from commercially available RNA samples using quantitative PCR. Within this panel, high EPO and hEPOΔ3 expression was found in one liver cirrhosis sample (see Supplementary Fig. S1). In order to substantiate this observation, we broadened our study to a panel of 10 healthy liver samples and 10 cirrhotic liver samples, all formalin fixed paraffin embedded (FFPE), using UBC and SRSF4 as reference genes49, 50. We designed SYBR-green-based quantitative PCR assays specific for the EPO full-length form or for hEPOΔ3. In normal liver tissue, EPO expression was detectable whereas hEPOΔ3 was below detection limit. In cirrhotic liver tissue, the analysis showed a considerably high inter-individual variation in the expression levels of both EPO and hEPOΔ3. In individual samples, the mean expression level of EPO was 2.7-fold higher in liver cirrhosis samples compared to normal liver tissue (Fig. 2a). Expression of the hEPOΔ3 form was detectable in those cirrhotic liver samples which also showed enhanced EPO expression. Suggesting therefore co-regulation of EPO and the spliced transcript hEPOΔ3 in diseased condition, we further studied expression profiles in a panel of clear cell renal cell carcinoma (ccRCC) samples and adjacent normal renal tissue. In this panel, EPO expression was enhanced in 3/12 tumour samples compared to adjacent normal tissue. Again, these samples showed also highest hEPOΔ3 expression (Fig. 2b). EPO expression is regulated by a distinctive oxygen-sensing mechanism involving the HIF pathway51. The human EPO-producing liver hepatocellular carcinoma cell line HepG237 is an established model to study hypoxia-inducible EPO transcription (e.g. 52, 53). Thus, we assessed the change in EPO and putative hEPOΔ3 transcripts in HepG2 cells exposed to hypoxia (1% O2) for 4 hours. UBC, SRSF4 and cyclophilin A (PPIA) were tested for hypoxic response. As PPIA expression was stable under hypoxic conditions, being in line with observations from others52, 54, it was used as internal control. Importantly, we did not only detect hEPOΔ3 in HepG2 cells, but EPO and hEPOΔ3 were both hypoxia inducible in HepG2 cells (Fig. 2c). In summary, our data show concomitant up-regulation of EPO and hEPOΔ3 under hypoxia and in diseased condition.
Expression analysis of EPO and hEPOΔ3 transcripts in human tissues and the human cell line HepG2 using quantitative PCR. (a) Normal and cirrhotic liver FFPE samples (n = 10 per condition) were analysed, using UBC and SRSF4 as reference genes. EPO mRNA expression is detectable in both normal (▲) and cirrhotic liver tissue (▼), individual cirrhotic samples show 3–6 fold increase compared to mean of normal tissue. hEPOΔ3 is only amplified from cirrhotic livers (□) but not from normal livers (○). (b) EPO and hEPOΔ3 expression was analysed in clear cell renal cell carcinoma FFPE samples and compared to adjacent normal tissue (n = 12 per condition). Expression levels of SRSF4 and Alas1 were constant in tumour and matched normal tissue and served as reference genes. EPO expression is enhanced in clear cell renal cell carcinoma samples 8, 10 and 11 compared to adjacent normal tissue. These samples show also highest hEPOΔ3 expression. (c) EPO and hEPOΔ3 expression is induced in HepG2 cells exposed to hypoxic conditions (1% O2) for 4 h. Cyclophilin A expression was not affected by hypoxia and used as internal control gene. (n = 4); two-tailed t-test; mean ± SEM presented.
Protein structure prediction/modelling
Human EPO is a heavily glycosylated protein consisting of one 165 amino acid chain and containing three N-linked (Asn-24, Asn-38, Asn-83) and one O-linked (Ser-126) oligosaccharide side chains55, 56 (Fig. 3). The carbohydrates contribute about 40% of the total molecular weight57. EPO and the hEPOΔ3 derived EV-3 protein differentiate by 29 amino acids19 (Fig. 3) leading to a reduction in molecular mass of unglycosylated proteins from 20 kDa for EPO to 16.8 kDa for EV-3. In the glycosylated form of the two proteins, the loss of one N-glycosylation site (Asn-38) in EV-3 leads to a more distinct difference in apparent molecular weight (~5–8 kDa). The human EPO protein has a four-helix bundle structure58, which is typical for haematopoietic growth factors. The four long α-helices (A, B, C and D) are connected by two long cross-over loops (AB and CD) and one short loop (BC) (see Supplementary Fig. S2). Near the carboxy end of the AB loop is a short alpha-helical segment (minihelix B’) which is important for binding to the EPO-receptor58. As previously described, the EV-3 protein sequence suggests loss of the AB loop19. Using automated homology modelling this is predicted to result in a considerable change in tertiary structure compared to EPO (see Supplementary Fig. S2).
Depiction of the EV-3 specific peptide used for nano-LC-ESI-MS/MS analysis. According to the obtained sequence of the two EPO-PCR amplification products from human adult kidney and foetal brain, aminoacid sequences of EPO and hEPOΔ3 derived protein EV-3 differ only by 29 aminoacids. The deleted region contains one N-glycosylation site and the AB loop including the minihelix B’. The underlined amino acids in the EV-3 sequence represent the EV-3 specific peptide used in nano-LC-ESI-MS/MS. The representation of EPO’s primary structure is taken from Boissel et al.84.
The hEPOΔ3 derived protein EV-3 is isolated from HepG2 culture supernatants using immunoprecipitation
We intended to provide evidence for the expression of the hEPOΔ3 derived EV-3 protein. Therefore, we used the HepG2 cell line which expresses EPO and hEPOΔ3 transcripts (Fig. 4a). If we find that hEPOΔ3 transcript is translated into protein, we assumed that it is secreted by the HepG2 cells similarly to EPO. Thus, supernatants of HepG2 cultures were pooled and concentrated and a biotinylated polyclonal antibody specific to the N-terminus of EPO was used for immunoprecipitation. Recombinant produced proteins, rEV-3 and rEPO, were used as positive controls for Western Blot. EPO and EV-3 migrate as diffuse bands with apparent molecular masses of approximately 40 kDa (EPO) and 35 kDa (EV-3). In the supernatants of HepG2 cultures, the polyclonal anti-EPO antibody precipitated two proteins running at the size of EPO and EV-3 (Fig. 4b,c). The identity of the two proteins was confirmed through mass spectrometry as described below.
Isolation of hEPOΔ3 from HepG2 cells and detection of hEPOΔ3 derived protein EV-3 and EPO in HepG2 cell culture supernatant. (a) The nested PCR approach amplifies EPO full-length and hEPOΔ3 transcripts from HepG2 cells. (b) Captured proteins of HepG2 supernatants (α-EPO IP, using biotinylated anti-EPO antibody BAF959) are visualized by western blotting using a polyclonal rhEPO-antibody (H-162). The Western Blot shows two EPO-antibody reactive proteins (prot. 1 and prot. 2) running at the same size of recombinantly produced EV-3 (rEV-3) and EPO (two different batches of rEPO). (c) Densitometry analysis was done on the Western Blot image (jpeg) shown in Fig. 4b. The plot profile of the HepG2 immunoprecipitate in lane 5 (ROI3) reveals a faint but distinct second peak, corresponding to the plot profile of EV-3 loaded in lane 1 (ROI1). (d) EV-3, indicated by the specific peptide ‘DITVGQQAVE’, was detected in HepG2 supernatants by nano-LC-ESI-MS/MS. Quantification of EV-3 specific peptide (fmol/µg total protein of analysed sample) in sample H1, the immunoprecipitate (α-EPO-IP) of HepG2 supernatant, or sample H2, HepG2 supernatant enriched for glycoproteins by WGA chromatography.
Confirmation of EV-3 protein in HepG2 supernatant using multiple-reaction monitoring
Systematic processing of EPO and EV-3 containing samples, as described in the method section, allowed for the generation of two peptides enabling the identification and differentiation of EPO and EV-3 within one sample. The resulting peptide with the sequence ‘DITVGQQAVE’ is contained in EV-3 but not in EPO (Fig. 3) and no other protein was found to include that sequence; therefore this peptide is the specific identifier of EV-3. Peptide sequence ‘DITTGCAE’ is absent in EV-3 and contained in human EPO only and therefore serves as the specific identifier of EPO in the processed samples (Table 1).
The protein samples obtained from HepG2 supernatants by either immunoprecipitation with an anti-EPO antibody or WGA purification were processed (PNGase F and Asp-N/Glu-C digestion). Thereby, EV-3 or EPO proteins in the sample are cleaved leading to the generation of above mentioned identifier peptides of EV-3 and EPO. For identification and quantification through multiple-reaction-monitoring (MRM), isotopically labelled EV-3 and EPO identifier peptides were spiked into the analyte (processed sample) and endogenous peptides were identified through their identical retention time. The EV-3 specific peptide DITVGQQAVE was reliably detected in the samples; also peptide DITTGCAE (specific for EPO) was detected. This finding confirms the presence of EV-3 and EPO in the supernatants of HepG2 cells as indicated by the Western Blot (Fig. 4b,c).
Quantitation of EV-3 by its specific peptide was robust, whereas quantitation of EPO by its specific peptide was unstable: DITTGCAE was detected but yielded standard curves only for 1/5 transitions and therefore, parallel quantitation of EPO in the same sample was disabled. The EV-3 specific peptide DITVGQQAVE was quantified with 5/5 transitions and revealed that the level in the immunoprecipitate (H1; 5.5 fmol/µg of total protein) was considerably lower than in the WGA purified HepG2 supernatant (H2; 127.7 fmol/µg of total protein) (Fig. 4d). Quantification is relative to the amount of total protein in the processed sample. Immunoprecipitation and lectin-affinity chromatography are two different methods to enrich for EPO and EV-3, but neither approach will result in a pure EPO/EV-3 sample. Whereas the elution step in the immunoprecipitation approach co-elutes the anti-EPO antibody, WGA purification also enriches other glycoproteins, thereby explaining the different relative levels of EV-3 specific peptide quantified in the samples. In summary, the analysis of HepG2 supernatants by MRM shows that hEPOΔ3 expressing HepG2 cells produce and secrete the corresponding EV-3 protein.
EV-3 protein is detected in human serum
The above described determination of the EPO splice variant hEPOΔ3 as well as EV-3 protein in HepG2 cells depicts a critical result of our study; however, cell lines have some limitations regarding the representation of an in vivo situation. Therefore, we investigated blood from healthy human donors for the presence of EV-3 protein. A multi-step purification strategy (Fig. 5a) including size-exclusion chromatography and glycoprotein affinity chromatography was developed for processing human serum in order to enrich for EV-3 and EPO in the fractions used for analysis (see Supplementary Figs S3 and S4). Mass spectrometry analysis of the purified and processed samples was performed according to the previously established MRM protocol. Serum samples of two healthy donors were examined and in both of them the EV-3 specific peptide ‘DITVGQQAVE’ was identified, indicating the presence of EV-3 protein (Fig. 5b). The level of EV-3 in sample 1 (donor 1), with one size exclusion chromatography included in the processing of the sample, was 0.15 fmol/µg total protein. As the applied technology allows loading of only 3.5 µg total protein/test sample, we sought to further reduce the amount of larger proteins like albumin (66.5 kDa) by additional size exclusion chromatography and thereby enrich for EV-3 and EPO in the test sample. Thus, processing of sample 2 (donor 2) included two size exclusion steps. Two different fractions of the fully processed serum sample were analysed and the levels for the EV-3 specific peptide were 1.24 fmol/µg and 2.03 fmol/µg in the respective test sample. The EPO-specific peptide DITTGCAE was also detected in the test samples but only in 1/5 transitions and therefore was unqualified for quantification.
EV-3 protein identification in human serum from healthy donors. (a) Depiction of the optimized multi-step purification strategy of serum for the detection and quantification of the EV-3 specific peptide by nano-LC-ESI-MS/MS. (b) In serum samples of the healthy human donors (S1, S2), the EV-3 specific peptide ‘DITVGQQAVE’ was detected by using nano-LC-ESI-MS/MS. Applying two size exclusion chromatography steps to serum S2 enriched EV-3 in the sample. Compared to S1, higher levels of the EV-3 specific peptide (fmol/µg total protein) were quantified in two fractions of S2.
The here described detection of EV-3 in human serum is the first proof that human hEPOΔ3 is translated into protein and secreted into the blood circulation.
EV-3 has no erythropoietic colony forming unit (CFU-E) properties in human CD34+ progenitors
CD34 is a marker of haematopoietic stem- and progenitor cells59, 60. The proliferation and differentiation of HSC depends on a variety of cytokines, the most prominent for the erythroid pathway being erythropoietin. In a semisolid colony forming unit assay, early progenitor cells will yield larger colonies referred to as burst forming units, while progenitors further down the differentiation path will yield smaller colonies (CFU-E). While complete Methocult medium (H84434, comprising EPO) yielded BFU-E and CFU-E at numbers representative for the CD34 source and content of the specimen (Fig. 6), medium that is devoid of EPO (H4035) as expected failed to show any formation of erythroid colonies. A systematic analysis of the EPO dose necessary to yield a maximum of erythroid colonies showed a clear correlation of the EPO concentration with the number and phenotype of erythroid colonies with a maximum erythroid colony stimulating capacity at 3 U/ml. Interestingly, EV-3 was not able to replace EPO in the CFU Assay. Even at concentrations 10 times higher than the optimum concentration of EPO, EV-3 did not yield any erythroid colonies (Fig. 6). Since CD34+ haematopoietic stem cells as well as BFU-E and CFU-E express the EPOR61, 62, the absence of erythroid colony forming stimulating effect of EV-3 on erythroid progenitors might therefore be due to the absence of the high affinity binding site to the EPOR in the EV-3 splice variant of erythropoietin.
EPO but not EV-3 promotes formation of erythroid progenitors from human CD34+ haematopoietic stem cells. CD34+ cells (500 cells/well) from two donors were cultured in EPO-free MethoCult medium supplemented either with EPO or EV-3 at different concentrations (0–30 U/ml) for 14 days. Analysis of the different colony-forming units (CFU) revealed no erythroid progenitors (CFU-GEMM or BFU-E) in control or EV-3 treated wells but in wells treated with EPO or positive control H84434 (medium includes EPO). n = 2 per treatment; CFU-G/M/GM, colony-forming unit-granulocyte or -macrophage or mix of granulocyte and macrophage; CFU-GEMM, colony-forming unit granulocyte, erythrocyte, monocyte, megakaryocyte; BFU-E, burst-forming unit erythroid.
Neuroprotective effects of EV-3 against hypoxia-induced cell death confirmed
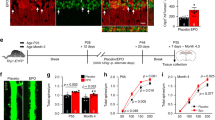
EPO’s protective effects against hypoxia-induced cell death is well described in the literature44, 63, 85. We sought to investigate previously reported conservation of neuroprotective properties in the naturally occurring EPO-isoform EV-319. Rat primary hippocampal neurons were cultured and hypoxia-induced cell death, indicated by the percentage of Trypan blue- positive cells of all cells in the respective well, under placebo conditions (control) were set as 100%. Pre-treatment with EPO (3 U/ml) lead to reduction of hypoxia-induced cell death (p = 0.03, compared to control). Pre-treatment with EV-3 (weight equivalent to 3 U/ml of EPO) also significantly protected rat primary hippocampal neurons against hypoxia-induced cell death (p = 0.03, compared to control), although the total effect seems slightly lower than that of EPO (Fig. 7). The results point towards the preservation of neuroprotective effects in EV-3.
EV-3 and EPO protect primary hippocampal neurons from cell death. Pretreatment with EPO or EV-3 reduced cell death, induced by starvation (48 h) followed by hypoxia (20 h), of rat primary hippocampal neurons (n = 5/treatment; hypoxia effect set to 100%; one-tailed paired t-test, EPO and EV-3 [U weight equivalents] treatment compared to placebo). Mean ± SEM presented.
EV-3 facilitates cognitive performance of mice and is devoid of haematocrit increasing properties
Putative in vivo actions of EV-3 were tested in mice. A learning task was chosen as enhancement of cognitive function in mice was reported for EPO as well as for its carbamoylated derivative64, 65. Cognitive performance in a visual discrimination task45, 66 of EPO, EV-3 and placebo-injected mice was tested in a touch-screen based operant system. Pretraining performance was not significantly different between the placebo, EV-3 and EPO groups (Kaplan-Meier analysis: Placebo vs. EV-3: χ2 = 2.057; p = 0.152, Placebo vs. EPO: χ2 = 1.83; p = 0.669, EV-3 vs. EPO: χ2 = 1.674; p = 0.196; Fig. 8a). During the visual discrimination task, EPO-treated mice showed significantly faster learning as compared to the placebo-treated mice (χ2 = 4.178; p = 0.041; Fig. 8b). No significant difference was found between EPO and EV-3 treated mice (χ2 = 1.344; p = 0.246). Furthermore, we investigated whether the overall performance in which pretraining and visual discrimination phases were combined was different between groups. EPO-treated mice showed a significantly better overall performance as compared to the placebo-treated mice (χ2 = 4.177; p = 0.041; Fig. 8c). Interestingly, EV-3 treated mice performed similarly to EPO treated mice, resulting in a close-to-significant difference between EV-3 and placebo-treated mice (χ2 = 3.541; p = 0.060). EV-3 mice were faster in the acquisition of the different stages of the touchscreen task compared to placebo. No such difference was found for the pair-wise comparison of the EPO and EV-3 groups (χ2 = 0.503; p = 0.478). These results suggest that touchscreen performance is facilitated by both EPO and EV-3 treatment.
Cognitive performance in a touchscreen-test and haematocrit levels in placebo, EV-3 and EPO treated mice. Kaplan-Meier presentation of group performance during (a) pretraining 1–3, (b) visual discrimination and (c) overall task (pretraining and visual discrimination). Curves represent group performance (n = 10–11 per group) and indicate the cumulative percentage of animals not reaching performance criteria of the touch-screen paradigm. The EPO- and the EV-3 treated mice finished the overall task (c) 10 days earlier than the placebo group (EV-3 vs. placebo, p = 0.060, EPO vs placebo, p = 0.041*). (d) Following 3-weeks treatment with EPO (5000 IU/kg, NeoRecormon) versus EV-3 (weight-equivalent dose) and placebo, haematocrit is significantly enhanced in the EPO-treated but not in the EV-3 treated group.
In a second batch of mice, injected according to the treatment scheme of the animals used for the cognitive testing, we investigated the erythropoietic properties of EV-3 by determination of the haematocrit. No difference between haematocrit levels of EV-3 and placebo-treated mice (Fig. 8d) was found (T[27] = 0.67; p = 0.509). Haematocrit levels were increased in EPO-treated mice as compared to both EV-3 (T[27] = 10.27; p < 0.0001; t-test for independent samples) and placebo treated mice (T[28] = 10.29; p < 0.0001). These results show that pro-cognitive effects of EV-3 on touchscreen performance are not related to changes in haematocrit levels. Further, EV-3 differs to EPO in terms of erythropoietic properties.
Discussion
The present study evaluates the existence and functionality of a naturally occurring variant of EPO. The finding of the EPO splice variant lacking exon 3 (hEPOΔ3), originally described by Meisel, Priller, Bonnas and Dirnagl (Charité Berlin)19, in human foetal brain and kidney were confirmed. We further report the co-regulation of EPO and hEPOΔ3 in the human cell line HepG2 as well as in human tissue and importantly, the detection of the hEPOΔ3 corresponding protein-product EV-3 in human serum. Investigating the biological functionality of EV-3, we provide evidence that recombinant EV-3 does not promote erythroid progenitors but shares EPO’s neuroprotective properties. Our first in vivo studies with EV-3 indicate beneficial effects on cognitive performance and simultaneously show that EV-3 is a non-erythropoietic variant of EPO expressed in humans.
The discovery of alternatively spliced variants of human EPO together with the indicated functionality of the protein product of hEPOΔ3, i.e. loss of erythropoietic but preservation of neuroprotective properties, has only recently been described. One aim of our studies was to broaden the spectrum of human samples used for expression analysis of the splice variant hEPOΔ3. Our results revealed that expression of hEPOΔ3 is in accordance with EPO expression2,3,4,5,6: The hEPOΔ3 transcript was detected in samples of kidney, liver and foetal brain but seems to be expressed at much lower levels than EPO. In a subset of human cirrhotic liver tissue samples, we see enhanced expression of both, EPO and hEPOΔ3. Although elevated blood EPO concentrations are not generally found in patients with cirrhosis32, 67, local hepatic tissue hypoxia occurs in patients with cirrhosis68. Thus, enhanced EPO and hEPOΔ3 expression in liver cirrhosis is likely explained by hypoxic mechanisms and the observed parallel enhancement in individual samples implies that EPO and hEPOΔ3 transcript expression are co-regulated in vivo. This is further supported by our finding of concomitantly enhanced EPO and hEPOΔ3 transcript expression in ccRCC samples. Several studies reported an increased expression of EPO as well as EPOR in human ccRCC, the question of whether the EPO/EPOR pathway plays a functional role and correlates to prognosis in ccRCC is under investigation by others35, 69. In line with the findings in human tissue, we detected enhanced expression of EPO and hEPOΔ3 in vitro in HepG2 cells exposed to hypoxic conditions. Oxygen-dependent regulation of the EPO gene is organ-specific (reviewed in 39, 70). In the liver, hypoxia inducible elements are located within 0.4 kb 5′ and 0.7 kb 3′ of the EPO gene and sequences required for inducible expression in the kidney are located greater than 6 kb 5′ or 0.7 kb 3′ to the gene71. Thus, from the gene structure, hypoxia inducible elements and the promoter region are not supposed to be affected by alternative splicing of exon3 in hEPOΔ3. The observed co-regulation of the EPO and the hEPOΔ3 transcript in HepG2 cells and in human tissues implicates usage of identical promoter and hypoxia inducible elements. So far, our observations together with reports regarding cytoprotective action of EPO on acute ischemia-induced injury of liver or kidney72,73,74,75,76 lead us to hypothesize that the enhanced EPO and hEPOΔ3 transcript expression is part of a response playing a crucial role for endogenous tissue protection.
The detection of a splice variant co-regulated with the origin hardly allows concluding the translation into an active protein. Therefore, we aimed at the detection of hEPOΔ3 corresponding protein EV-3 protein in human samples. Additionally, we used the HepG2 cell line, where we have detected the hEPOΔ3 transcript, for further studies. A multistep processing protocol was established which allowed the clear separation of EPO and EV-3 and the detection by means of multiple reaction monitoring. In the supernatants of HepG2 cells we detected the peptide exclusive for the protein-product EV-3 derived from the hEPOΔ3 transcript. The respective peptide sequence covers the amino acids encoded by the end of exon 2 and by the beginning of exon 4 of the EPO gene and is specific for the product of the splice variant hEPOΔ3, lacking exon 3. More importantly, we also detected EV-3 in serum of healthy humans. Here, additional purification steps needed to be included before analysing the samples by MRM because of the high abundancy of proteins in human serum. Nevertheless, we could identify the EV-3 specific peptide in both serum samples investigated. This finding allows the conclusion that the hEPOΔ3 transcript, detected in human specimen, is translated into EV-3 protein and is secreted into the blood circulation in healthy individuals. This is a strong indicator for an endogenous function of this protein identified in humans.
Similarities in the expression and regulation of EPO and hEPOΔ3 lead us to the assumption that biological functions of EV-3 and EPO might overlap to some extent. Starting to investigate the biological function of EV-3, we refer here to well documented effects of EPO and investigated the EV-3 effects upon (i) erythropoiesis, (ii) hypoxia-induced neuronal cell death7, and (iii) cognitive performance in a mouse long-term learning paradigm66. Our results in cultured human CD34+ showed that EV-3, contrary to EPO, does not promote erythroid progenitors (BFU-E or CFU-E) in the early phase of erythropoiesis. CD34+ haematopoietic stem cells as well as BFU-E and CFU-E have been reported to express the EPOR61, 62. We also observed that repeated administration of EV-3 to mice is without influence on haematocrit levels. Together, this indicates that EV-3 does not have the binding properties like EPO in order to interact with the EPOR homodimer during the crucial steps of erythrocyte differentiation. This is in line with the EV-3 protein sequence19 which suggests loss of the ‘AB’ cross-over loop present in the EPO protein (Fig. 3). The ‘AB’ loop contains a segment which has been identified as being important for binding to the EPOR and the formation of the homodimer complex58, 77. Currently, we have no information on the receptor(s) binding EV-3. On the other hand, we observed a dose-dependent protective effect of EV-3 on primary hippocampal neurons in vitro. Compared to the effects of EPO in this assay system, the efficacy of EV-3 at a similar dose as EPO seemed to be slightly lower. In terms of biological function of EV-3, we also show here that recombinant EV-3 is active in vivo and improves, again with a slightly lower effect than EPO, cognitive performance of mice in a long-term learning paradigm. The slight difference we observed in a preliminary pharmacokinetic study (see Supplementary Fig. S5) unlikely explains the somewhat weaker effects of EV-3. The fact that the statistical significance in the cognition test was just missed (p = 0.06) may be due to a still suboptimal dosing regimen of EV-3 which might be different to EPO and needs to be further investigated. Clearly, information regarding the receptor and signalling pathways will help to optimize the protocols for testing the effects of EV-3.
The results of our experiments show that EV-3 differs to EPO in terms of haematopoiesis but has functional similarities with regard to neuroprotection and cognitive enhancement. The mode of action of EV-3 is yet unknown but obviously independent of erythropoiesis. EPO is a promising candidate for the treatment of degenerative disorders78. A prominent and repeatedly reported observation for EPO in clinical studies is the beneficial effect of EPO on cognitive function79,80,81,82,83. Improvement of cognitive performance by EPO or its non-erythropoietic derivative carbamoylated EPO (CEPO) in mice65, 66 was partly explained by enhanced neurogenesis in the dentate gyrus65 and more recently by EPO-induced differentiation of precursors to neurons45. The literature provides evidence allowing refraining from the idea that EPO’s effects on memory function are merely mediated through haematopoietic action. The finding that EV-3 conserves EPO’s neuroprotective and cognition-promoting effects without stimulating erythropoiesis further supports the independency of these effects from erythropoiesis. Our study is limited by the number of EV-3 dosing regimens applied in the learning paradigm but provides novel and adequate results encouraging future studies with EV-3 to evaluate the potential of becoming a therapeutic for degenerative disorders. Simultaneously, the question regarding the mechanism of action of EV-3 arises. The binding site which mediates neuroprotective or cognitive effects of EV-3 is yet unknown and will be focus of future studies.
Taken together, our results provide evidence for EV-3 as the first example of a natural non-erythropoietic EPO protein isoform in humans. The observed co-regulation of EPO and its splicing variant hEPOΔ3 in human tissue together with the determination of the matching protein EV-3 in serum of healthy humans indicates the biological potential of our findings. Concomitant up-regulation of EPO and hEPOΔ3 under pathological conditions support our hypothesis of the involvement of both proteins in a tissue protective system and is exemplified by the neuroprotective effects against hypoxia-induced cell death of EPO and EV-3.
References
Jelkmann, W. Erythropoietin after a century of research: younger than ever. Eur. J. Haematol. 78, 183–205, doi:10.1111/j.1600-0609.2007.00818.x (2007).
Jelkmann, W. Effects of erythropoietin on brain function. Curr. Pharm. Biotechnol. 6, 65–79, doi:10.2174/1389201053167257 (2005).
Ehrenreich, H. et al. Erythropoietin: novel approaches to neuroprotection in human brain disease. Metab. Brain Dis. 19, 195–206, doi:10.1023/B:MEBR.0000043969.96895.3c (2004).
Hasselblatt, M., Ehrenreich, H. & Sirén, A.-L. The brain erythropoietin system and its potential for therapeutic exploitation in brain disease. J. Neurosurg. Anesthesiol. 18, 132–8, doi:10.1097/00008506-200604000-00007 (2006).
Brines, M. & Cerami, A. Emerging biological roles for erythropoietin in the nervous system. Nat. Rev. Neurosci. 6, 484–94, doi:10.1038/nrn1687 (2005).
Digicaylioglu, M. & Lipton, S. A. Erythropoietin-mediated neuroprotection involves cross-talk between Jak2 and NF-kappaB signalling cascades. Nature 412, 641–7, doi:10.1038/35088074 (2001).
Sinor, A. D. & Greenberg, D. A. Erythropoietin protects cultured cortical neurons, but not astroglia, from hypoxia and AMPA toxicity. Neurosci. Lett. 290, 213–5, doi:10.1016/S0304-3940(00)01361-6 (2000).
Morishita, E., Masuda, S., Nagao, M., Yasuda, Y. & Sasaki, R. Erythropoietin receptor is expressed in rat hippocampal and cerebral cortical neurons, and erythropoietin prevents in vitro glutamate-induced neuronal death. Neuroscience 76, 105–16, doi:10.1016/S0306-4522(96)00306-5 (1997).
Sirén, A. L. et al. Erythropoietin prevents neuronal apoptosis after cerebral ischemia and metabolic stress. Proc. Natl. Acad. Sci. USA 98, 4044–9, doi:10.1073/pnas.051606598 (2001).
Brines, M. L. et al. Erythropoietin crosses the blood-brain barrier to protect against experimental brain injury. Proc. Natl. Acad. Sci. USA 97, 10526–31, doi:10.1073/pnas.97.19.10526 (2000).
Bernaudin, M. et al. A potential role for erythropoietin in focal permanent cerebral ischemia in mice. J. Cereb. Blood Flow Metab. 19, 643–51, doi:10.1097/00004647-199906000-00007 (1999).
Bartels, C., Späte, K., Krampe, H. & Ehrenreich, H. Recombinant human erythropoietin: novel strategies for neuroprotective/neuro-regenerative treatment of multiple sclerosis. Ther. Adv. Neurol. Disord. 1, 193–206, doi:10.1177/1756285608098422 (2008).
Wüstenberg, T. et al. Recombinant human erythropoietin delays loss of gray matter in chronic schizophrenia. Mol. Psychiatry 16, 26–36, 1, doi:10.1038/mp.2010.51 (2011).
Najmi Varzaneh, F., Najmi Varzaneh, F., Azimi, A. R., Rezaei, N. & Sahraian, M. A. Efficacy of combination therapy with erythropoietin and methylprednisolone in clinical recovery of severe relapse in multiple sclerosis. Acta Neurol. Belg. 114, 273–8, doi:10.1007/s13760-014-0286-y (2014).
Egger, K. et al. White matter changes in patients with friedreich ataxia after treatment with erythropoietin. J. Neuroimaging 24, 504–8, doi:10.1111/jon.12050 (2014).
Jang, W. et al. Safety and efficacy of recombinant human erythropoietin treatment of non-motor symptoms in Parkinson’s disease. J. Neurol. Sci. 337, 47–54, doi:10.1016/j.jns.2013.11.015 (2014).
Miyake, T., Kung, C. K. & Goldwasser, E. Purification of human erythropoietin. J. Biol. Chem. 252, 5558–64 (1977).
Lin, F. K. et al. Cloning and expression of the human erythropoietin gene. Proc. Natl. Acad. Sci. 82, 7580–84, doi:10.1073/pnas.82.22.7580 (1985).
Meisel, A., Priller, J., Bonnas, C. & Dirnagl, U. ERYTHROPOIETIN VARIANTS. Charité Universitätsmedizin - Berlin. WIPO WO/2006/120030 (US20110008363).
Tsytsikov, V. N., Yurovsky, V. V., Atamas, S. P., Alms, W. J. & White, B. Identification and characterization of two alternative splice variants of human interleukin-2. J. Biol. Chem. 271, 23055–60, doi:10.1074/jbc.271.38.23055 (1996).
Arinobu, Y. et al. Antagonistic effects of an alternative splice variant of human IL-4, IL-4delta2, on IL-4 activities in human monocytes and B cells. Cell. Immunol. 191, 161–7, doi:10.1006/cimm.1998.1431 (1999).
Wu, S. et al. Expression of interleukin-10 splicing variants is a positive prognostic feature in relapsed childhood acute lymphoblastic leukemia. J. Clin. Oncol. 23, 3038–42, doi:10.1200/JCO.2005.00.885 (2005).
Johansson, U., Cooke, S. P. & Londei, M. Identification of a new isoform of human IL-12 p35 mRNA. Eur. J. Immunogenet. 29, 439–44, doi:10.1046/j.1365-2370.2002.00337.x (2002).
Sorg, R. et al. Rapid and sensitive mRNA phenotyping for interleukins (IL-1 to IL-6) and colony-stimulating factors (G-CSF, M-CSF, and GM-CSF) by reverse transcription and subsequent polymerase chain reaction. Exp. Hematol. 19, 882–7 (1991).
Arcasoy, M. O., Jiang, X. & Haroon, Z. A. Expression of erythropoietin receptor splice variants in human cancer. Biochem. Biophys. Res. Commun. 307, 999–1007, doi:10.1016/S0006-291X(03)01303-2 (2003).
Robinson, C. J. & Stringer, S. E. The splice variants of vascular endothelial growth factor (VEGF) and their receptors. J. Cell Sci. 114, 853–65 (2001).
Juul, S. E., Yachnis, A. T., Rojiani, A. M. & Christensen, R. D. Immunohistochemical localization of erythropoietin and its receptor in the developing human brain. Pediatr. Dev. Pathol. 2, 148–58, doi:10.1007/s100249900103 (1999).
Koury, S. T., Bondurant, M. C., Koury, M. J. & Semenza, G. L. Localization of cells producing erythropoietin in murine liver by in situ hybridization. Blood 77, 2497–503 (1991).
Koury, S. T., Bondurant, M. C. & Koury, M. J. Localization of erythropoietin synthesizing cells in murine kidneys by in situ hybridization. Blood 71, 524–7 (1988).
Lacombe, C. et al. Peritubular cells are the site of erythropoietin synthesis in the murine hypoxic kidney. J. Clin. Invest. 81, 620–3, doi:10.1172/JCI113363 (1988).
Tacke, F. et al. Analysis of factors contributing to higher erythropoietin levels in patients with chronic liver disease. Scand. J. Gastroenterol. 39, 259–66, doi:10.1080/00365520310008340 (2004).
Yang, Y.-Y. et al. Plasma erythropoietin level in patients with cirrhosis and its relationship to the severity of cirrhosis and renal function. J. Gastroenterol. Hepatol. 18, 1156–61, doi:10.1046/j.1440-1746.2003.03144.x (2003).
Wiesener, M. S. et al. Erythropoietin gene expression in renal carcinoma is considerably more frequent than paraneoplastic polycythemia. Int. J. cancer 121, 2434–42, doi:10.1002/ijc.22961 (2007).
Stoyanoff, T. R. et al. Tumor biology of non-metastatic stages of clear cell renal cell carcinoma; overexpression of stearoyl desaturase-1, EPO/EPO-R system and hypoxia-related proteins. Tumor Biol. 37, 13581–93, doi:10.1007/s13277-016-5279-4 (2016).
Morais, C., Johnson, D. W., Vesey, D. A. & Gobe, G. C. Functional significance of erythropoietin in renal cell carcinoma. BMC Cancer 13, 14, doi:10.1186/1471-2407-13-14 (2013).
Bunn, H. F. Erythropoietin. Cold Spring Harb. Perspect. Med. 3, a011619 (2013).
Goldberg, M. A., Glass, G. A., Cunningham, J. M. & Bunn, H. F. The regulated expression of erythropoietin by two human hepatoma cell lines. Proc. Natl. Acad. Sci. 84, 7972–76 (1987).
Pickrell, J. K., Pai, A. A., Gilad, Y. & Pritchard, J. K. Noisy splicing drives mRNA isoform diversity in human cells. PLoS Genet. 6, e1001236, doi:10.1371/journal.pgen.1001236 (2010).
Jelkmann, W. Regulation of erythropoietin production. J. Physiol 589, 1251–8, doi:10.1113/jphysiol.2010.195057 (2011).
Biasini, M. et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 42, W252–8, doi:10.1093/nar/gku340 (2014).
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22, 195–201, doi:10.1093/bioinformatics/bti770 (2006).
Kiefer, F., Arnold, K., Künzli, M., Bordoli, L. & Schwede, T. The SWISS-MODEL Repository and associated resources. Nucleic Acids Res. 37, D387–92, doi:10.1093/nar/gkn750 (2009).
Guex, N., Peitsch, M. C. & Schwede, T. Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: a historical perspective. Electrophoresis 30(Suppl 1), S162–73, doi:10.1002/elps.200900140 (2009).
Lewczuk, P. et al. Survival of hippocampal neurons in culture upon hypoxia: effect of erythropoietin. Neuroreport 11, 3485–8, doi:10.1097/00001756-200011090-00017 (2000).
Hassouna, I. et al. Revisiting adult neurogenesis and the role of erythropoietin for neuronal and oligodendroglial differentiation in the hippocampus. Mol. Psychiatry 21, 1752–67, doi:10.1038/mp.2015.212 (2016).
Beru, N., McDonald, J., Lacombe, C. & Goldwasser, E. Expression of the erythropoietin gene. Mol. Cell. Biol. 6, 2571–5, doi:10.1128/MCB.6.7.2571 (1986).
Tan, C. C., Eckardt, K. U. & Ratcliffe, P. J. Organ distribution of erythropoietin messenger RNA in normal and uremic rats. Kidney Int. 40, 69–76, doi:10.1038/ki.1991.181 (1991).
Faustino, N. A. & Cooper, T. A. Pre-mRNA splicing and human disease. Genes Dev. 17, 419–37, doi:10.1101/gad.1048803 (2003).
Boujedidi, H. et al. Housekeeping gene variability in the liver of alcoholic patients. Alcohol. Clin. Exp. Res. 36, 258–66, doi:10.1111/j.1530-0277.2011.01627.x (2012).
Kim, S. & Kim, T. Selection of optimal internal controls for gene expression profiling of liver disease. Biotechniques 35, 456–8, 460 (2003).
Lee, F. S. & Percy, M. J. The HIF Pathway and Erythrocytosis. Annu. Rev. Pathol. Mech. Dis. 6, 165–92, doi:10.1146/annurev-pathol-011110-130321 (2011).
Benöhr, P., Harsch, S., Proksch, B. & Gleiter, C. H. Does angiotensin II modulate erythropoietin production in HepG2 cells? Nephron. Exp. Nephrol. 98, e124–31, doi:10.1159/000081556 (2004).
Chiang, C.-K., Nangaku, M., Tanaka, T., Iwawaki, T. & Inagi, R. Endoplasmic reticulum stress signal impairs erythropoietin production: a role for ATF4. Am. J. Physiol. Cell Physiol. 304, C342–53, doi:10.1152/ajpcell.00153.2012 (2013).
Sonna, L. A., Cullivan, M. L., Sheldon, H. K., Pratt, R. E. & Lilly, C. M. Effect of hypoxia on gene expression by human hepatocytes (HepG2). Physiol. Genomics 12, 195–207, doi:10.1152/physiolgenomics.00104.2002 (2003).
Lai, P. H., Everett, R., Wang, F. F., Arakawa, T. & Goldwasser, E. Structural characterization of human erythropoietin. J. Biol. Chem. 261, 3116–21 (1986).
Broudy, V. C., Tait, J. F. & Powell, J. S. Recombinant human erythropoietin: purification and analysis of carbohydrate linkage. Arch. Biochem. Biophys. 265, 329–36, doi:10.1016/0003-9861(88)90135-X (1988).
Davis, J. M., Arakawa, T., Strickland, T. W. & Yphantis, D. A. Characterization of recombinant human erythropoietin produced in Chinese hamster ovary cells. Biochemistry 26, 2633–8, doi:10.1021/bi00383a034 (1987).
Cheetham, J. C. et al. NMR structure of human erythropoietin and a comparison with its receptor bound conformation. Nat. Struct. Biol. 5, 861–6, doi:10.1038/2302 (1998).
Stella, C. C. et al. CD34-positive cells: biology and clinical relevance. Haematologica 80, 367–87 (1995).
Sidney, L. E., Branch, M. J., Dunphy, S. E., Dua, H. S. & Hopkinson, A. Concise review: evidence for CD34 as a common marker for diverse progenitors. Stem Cells 32, 1380–9, doi:10.1002/stem.1661 (2014).
Krantz, S. B. Erythropoietin. Blood 77, 419–34 (1991).
Shinjo, K., Takeshita, A., Higuchi, M., Ohnishi, K. & Ohno, R. Erythropoietin receptor expression on human bone marrow erythroid precursor cells by a newly-devised quantitative flow-cytometric assay. Br. J. Haematol. 96, 551–8, doi:10.1046/j.1365-2141.1997.d01-2071.x (1997).
Danielyan, L. et al. Similar protective effects of BQ-123 and erythropoietin on survival of neural cells and generation of neurons upon hypoxic injury. Eur. J. Cell Biol. 84, 907–13, doi:10.1016/j.ejcb.2005.07.001 (2005).
Adamcio, B. et al. Erythropoietin enhances hippocampal long-term potentiation and memory. BMC Biol. 6, 37, doi:10.1186/1741-7007-6-37 (2008).
Leconte, C. et al. Comparison of the effects of erythropoietin and its carbamylated derivative on behaviour and hippocampal neurogenesis in mice. Neuropharmacology 60, 354–64, doi:10.1016/j.neuropharm.2010.09.025 (2011).
El-Kordi, A., Radyushkin, K. & Ehrenreich, H. Erythropoietin improves operant conditioning and stability of cognitive performance in mice. BMC Biol. 7, 37, doi:10.1186/1741-7007-7-37 (2009).
Risør, L. M., Fenger, M., Olsen, N. V. & Møller, S. Hepatic erythropoietin response in cirrhosis. Scand. J. Clin. Lab. Invest. 76, 234–9, doi:10.3109/00365513.2015.1137351 (2016).
Moreau, R. et al. Tissue hypoxia and hepatocellular insufficiency in patients with cirrhosis. Gastroentérologie Clin. Biol. 13, 978–83 (1989).
Aapro, M., Jelkmann, W., Constantinescu, S. N. & Leyland-Jones, B. Effects of erythropoietin receptors and erythropoiesis-stimulating agents on disease progression in cancer. Br. J. Cancer 106, 1249–58, doi:10.1038/bjc.2012.42 (2012).
Fandrey, J. Oxygen-dependent and tissue-specific regulation of erythropoietin gene expression. AJP Regul. Integr. Comp. Physiol. 286, R977–88, doi:10.1152/ajpregu.00577.2003 (2004).
Semenza, G. L., Dureza, R. C., Traystman, M. D., Gearhart, J. D. & Antonarakis, S. E. Human erythropoietin gene expression in transgenic mice: multiple transcription initiation sites and cis-acting regulatory elements. Mol. Cell. Biol. 10, 930–8, doi:10.1128/MCB.10.3.930 (1990).
Luo, Y.-H., Li, Z.-D., Liu, L.-X. & Dong, G.-H. Pretreatment with erythropoietin reduces hepatic ischemia-reperfusion injury. Hepatobiliary Pancreat. Dis. Int 8, 294–9 (2009).
Liu, Q.-S. et al. Erythropoietin pretreatment exerts anti-inflammatory effects in hepatic ischemia/reperfusion-injured rats via suppression of the TLR2/NF-κB pathway. Transplant. Proc. 47, 283–9, doi:10.1016/j.transproceed.2014.10.045 (2015).
Vesey, D. A. et al. Erythropoietin protects against ischaemic acute renal injury. Nephrol. Dial. Transplant 19, 348–55, doi:10.1093/ndt/gfg547 (2004).
Johnson, D. W. et al. Delayed administration of darbepoetin or erythropoietin protects against ischemic acute renal injury and failure. Kidney Int. 69, 1806–13, doi:10.1038/sj.ki.5000356 (2006).
Song, Y. R. et al. Prevention of acute kidney injury by erythropoietin in patients undergoing coronary artery bypass grafting: a pilot study. Am. J. Nephrol. 30, 253–60, doi:10.1159/000223229 (2009).
Syed, R. S. et al. Efficiency of signalling through cytokine receptors depends critically on receptor orientation. Nature 395, 511–6, doi:10.1038/26773 (1998).
Maiese, K. Regeneration in the nervous system with erythropoietin. Front. Biosci. (Landmark Ed.) 21, 561–96 (2016).
Ehrenreich, H. et al. Improvement of cognitive functions in chronic schizophrenic patients by recombinant human erythropoietin. Mol. Psychiatry 12, 206–20, doi:10.1038/sj.mp.4001907 (2007).
Ehrenreich, H., Bartels, C., Sargin, D., Stawicki, S. & Krampe, H. Recombinant human erythropoietin in the treatment of human brain disease: focus on cognition. J. Ren. Nutr. 18, 146–53, doi:10.1053/j.jrn.2007.10.029 (2008).
Kristensen, P. L. et al. Influence of erythropoietin on cognitive performance during experimental hypoglycemia in patients with type 1 diabetes mellitus: a randomized cross-over trial. PLoS One 8, e59672, doi:10.1371/journal.pone.0059672 (2013).
Ohls, R. K. et al. Cognitive outcomes of preterm infants randomized to darbepoetin, erythropoietin, or placebo. Pediatrics 133, 1023–30, doi:10.1542/peds.2013-4307 (2014).
Miskowiak, K. W. et al. Effects of erythropoietin on memory-relevant neurocircuitry activity and recall in mood disorders. Acta Psychiatr. Scand. 134, 249–59, doi:10.1111/acps.12597 (2016).
Boissel, J. P., Lee, W. R., Presnell, S. R., Cohen, F. E. & Bunn, H. F. Erythropoietin structure-function relationships. Mutant proteins that test a model of tertiary structure. J. Biol. Chem. 268, 15983–93 (1993).
Miljus, N. et al. Neuroprotection and endocytosis: erythropoietin receptors in insect nervous systems. Journal of Neurochemistry, 141(1), 63–74 (2017).
Acknowledgements
We like to thank Dennis Niermeier, Dr. Uwe Carl, Dr. Christos Gouloudis, Kristian Stanar (all IBA GmbH – solutions for life sciences), Thomas Pohl (Epomedics) and the employees from TopLab for profound discussions and support. Further, we are grateful to Dr. Ronny Milde for critically reading the manuscript. Work performed at Epomedics was partially funded by a grant from the German BMBF (code 0316160B). Epomedics provided material to collaborating institutes, otherwise no financing was provided by Epomedics. Further, Epomedics was without influence on conduct or data analysis of experiments of collaborating institutes. H.E., L.W., E.D. and D.W. are supported by the Max Planck Society, the Max Planck Förderstiftung, and the DFG (CNMPB).
Author information
Authors and Affiliations
Contributions
C.B., L.W., D.W., R.K.-W., K.S., T.T., H.E., H.S., I.S. conceived and designed the experiments; C.B., L.W., D.W., R.K.-W., H.S., performed the experiments; C.B., L.W., D.W., R.K.-W., E.D., M.B., K.S., T.T., I.S. analysed the data; C.B., H.E., I.S. wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
C.B. is listed as co-inventor of EPO-variant patents and performed the experiments during her employment at Epomedics; L.W., D.W., E.D., K.S., M.B., R.K.-W. and T.T. report no potential competing interest; H.E. has submitted/holds use patents for EPO in stroke, schizophrenia and Multiple Sclerosis; H.S. is CEO of and holds stocks in Epomedics GmbH, and IS is employee of Epomedics GmbH.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bonnas, C., Wüstefeld, L., Winkler, D. et al. EV-3, an endogenous human erythropoietin isoform with distinct functional relevance. Sci Rep 7, 3684 (2017). https://doi.org/10.1038/s41598-017-03167-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-03167-0
This article is cited by
-
Protection of insect neurons by erythropoietin/CRLF3-mediated regulation of pro-apoptotic acetylcholinesterase
Scientific Reports (2022)
-
Introducing the brain erythropoietin circle to explain adaptive brain hardware upgrade and improved performance
Molecular Psychiatry (2022)
-
Leukemia inhibitory factor inhibits erythropoietin-induced myelin gene expression in oligodendrocytes
Molecular Medicine (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.