Abstract
We report on a new colorimetric DNA detection method that takes advantage of the power of polymerase chain reaction (PCR) and the simplicity of the classic litmus test. The strategy makes use of a modified set of primers for PCR to facilitate ensuing manipulations of resultant DNA amplicons: their tagging with urease and immobilization onto magnetic beads. The amplicon/urease-laden beads are then used to hydrolyze urea, resulting in the increase of pH that can be conveniently reported by a pH-sensitive dye. We have successfully applied this strategy for the detection of two hypervirulent strains of the bacterium Clostridium difficile that are responsible for the recent increase in the global incidence and severity of C. difficile infections. Furthermore, the viability of this test for diagnostic applications is demonstrated using clinically validated stool samples from C. difficile infected patients.
Similar content being viewed by others
Introduction
Polymerase chain reaction (PCR) is a popular DNA amplification technique and can create millions of amplicons of a target sequence in a short period of time1,2,3,4. This technique has been widely utilized for a variety of applications, including the detection of pathogenic bacteria5,6,7,8. PCR-based DNA detection is attractive for bacterial detection simply because specific DNA sequences can serve as reliable bacterial biomarkers and the amplification power of PCR permits the detection of a small number of bacteria before they can grow to infectious quantities5,6,7,8.
Although PCR has become a widely adopted technique in clinical laboratories, it has not become commonly used point-of-care or field tools. One significant roadblock that prevents such applications is the need of thermal cyclers that are often too expensive and bulky. However, there have been significant efforts towards miniaturizing PCR machines9,10,11,12. Another significant barrier that restricts PCR from becoming a popular field tool is the lack of simple yet effective signal transduction mechanisms that permit detection of PCR products without the use of expensive equipment (such as real-time PCR machine) and complicated process (such as DNA separation by gel electrophoresis). These issues reduce the utility of PCR as field tools. One approach to address this problem is to develop simple colorimetric assays that can be easily implemented in the field to detect DNA amplicons of PCR. Recently, our group reported an approach of adopting the classic litmus test for bacterial detection using a protein-activated DNAzyme and the protein enzyme urease13. Herein we describe a strategy for adopting the same litmus test for the detection of PCR amplicons through the use of a set of specially modified DNA primers and urease.
Our devised strategy is illustrated in Fig. 1. Two modified DNA primers are used to carry out PCR. The forward primer contains a 5′-biotin and the reverse primer contains a triethylene glycol linker that separates the target binding sequence (blue) from the sequence (purple) that is designed to hybridize with the DNA strand (green) coupled onto urease, a conjugate denoted “UrD” in this report. By this design, the PCR amplicons can be immobilized on the surface of magnetic beads containing streptavidin and the immobilized amplicons are capable of capturing UrD. The amplicon/urease-charged beads can then be used to hydrolyze urea, resulting in the increase of pH that can be reported by a pH-sensitive dye, such as phenol red, which can produce a sharp, yellow-to-pink transition when the solution pH changes from acidic to basic. Simply put, the proposed strategy converts the detection of DNA into tracking of pH increases using procedures that are easy to carry out.
To validate the proposed idea, we chose to develop a test that can be used to detect two hypervirulent strains of Clostridium difficile (C. difficile) known as 027/NAP1 and 078/NAP7. C. difficile is a gram-positive, spore-forming anaerobic bacterium that has been identified as the leading cause of diarrhea in developed countries and pseudomembranous colitis in humans14. The incidence and mortality of C. difficile infection (CDI) has increased dramatically over the past 15 years and CDI has become a common problem in hospitals across North America, Europe and some regions of Asia15, 16. This has been mostly attributed to the emergence of highly virulent strains. One such strain was characterized as ribotype 027, as North America Pulsed-field type NAP1 by pulsed field gel electrophoresis (PFGE) and was subsequently referred as 027/NAP117. The 027/NAP1 strain is known for its high-level resistance to fluoroquinolones, with markedly high toxin production and a mortality rate three times as high as that associated with less virulent strains, such as the ribotype 00117,18,19,20. Another epidemic strain of C. difficile, known as 078/NAP7, has also been described as hypervirulent since it can cause symptoms of similar severity to strain 027/NAP1. However, 078/NAP7 affects the younger population and is more frequently community-associated21,22,23,24. Identification of these epidemic strains may assist in determining relatedness of strains and recognizing transmission of C. difficile within healthcare facilities.
Current methods available for the diagnosis of CDI include cytotoxicity assays, anaerobic toxigenic culture, enzyme immunoassay (EIA) and nucleic acid amplification-based tests16, 25. These methods generally target toxins or their associated toxin genes and are unable to discriminate epidemic strains from non-epidemic strains14, 25,26,27. Meanwhile, current strain typing methods for C. difficile, such as ribotyping, restriction endonuclease analysis and PFGE, require culture of the organism, lack discriminatory power and are performed in reference laboratories, with resulting delays of days to weeks28, 29. Consequently, there is a significant need for a simple and rapid method for identifying epidemic C. difficile strains.
The virulence of C. difficile is mainly caused by toxin A and toxin B which are encoded by tcdA and tcdB, respectively, and their expression is regulated by tcdR (positive regulator) and tcdC (negative regulator)30,31,32. Interestingly, mutations in tcdC were found in both strains 027/NAP1 and 078/NAP7. A single-base deletion at nucleotide position 117 of tcdC (∆1stop-tcdC) has been found in the strain 027/NAP1 and this single nucleotide deletion results in a frame shift that introduces a stop codon at nucleotide position 19633, 34. Consequently, this generates a shortened protein of 65 residues. Similarly, strain 078/NAP7 carries a tcdC with a C184T transition (TAAstop-tcdC) leading to a truncated protein of 61 residues35. These mutations in tcdC have been proposed as a possible explanation for the increased virulence of the epidemic strains. Meanwhile, since these mutations are well conserved in strains 027/NAP1 and 078/NAP7, they can serve as molecular markers for the rapid identification of the hypervirulent C. difficile strains34,35,36.
It is well established that PCR can achieve highly specific DNA detection. For example, allele-specific PCR has been developed for Single Nucleotide Polymorphisms (SNP) detection using a DNA primer containing a single mismatch at 3′-end. This principle was used in this study to achieve the detection of mutations in tcdC genes associated with 027/NAP1 and 078/NAP7.
Through sequence analysis of tcdC (Figure S1, Supplementary Information), three forward primers (FP1, FP2 and FP3) and one reverse primer (RP) were designed for the discrimination between four possible outcomes: wild type tcdC (wt-tcdC), ∆1stop-tcdC, TAAstop-tcdC and no tcdC, as illustrated in Fig. 2a (sequences of the DNA molecules used for this work are provided as Table S1 in the Supplementary Information). We found that both ∆1stop-tcdC and TAAstop-tcdC have a thymine at position 120, whereas wt-tcdC has a cytosine at this position. P1 with 3′-thiamine was then designed to detect this difference. Similarly, P2 targets adenine at position 184 of tcdC to discriminate TAAstop-tcdC and ∆1stop-tcdC. P3 was designed to amplify all tcdC variants and could be used to identify the presence of tcdC gene and play the role of quality control during PCR. Since P1, P2 and P3 have similar Tm, they should theoretically have comparable PCR yields under same condition. The design of each primer was checked by BLAST to avoid nonspecific amplifications. Secondary structures and self-pairing of these primers were also examined.
PCR-based tests for the wildtype and two mutated tcdC genes. (a) Primer design. Primers for tcdC genes are color-coded. The mismatched 3′ terminus between forward primer and template are shown in red. Green tick and red cross denote a successful and unsuccessful PCR reaction, respectively. (b) Analysis of PCR products by 2% agarose gel electrophoresis. (c) Litmus test with PCR products. The photographs were taken at 30 minutes.
Three C. difficile strains, 027/NAP1 (with ∆1stop-tcdC), 078/NAP7 (with TAAstop-tcdC) and 001/NAP2 (with wt-tcdC) were tested to examine the performance of the designed primers. Genomic DNA from each strain was extracted based on a previous described method. 200 ng of genomic DNA was used as starting material for 28 cycles of PCR amplification. Agarose gel analysis indicates that a significant amount of PCR product was generated when primers matched the template, whereas no PCR product was observed in the absence of target gene or when using primers with mismatched 3′ termini (Fig. 2b). Based on the performances of three primer pairs toward each strain, we could successfully discriminate between epidemic C. difficile strains, 027/NAP1 and 078/NAP7 and non-epidemic C. difficile strain 001/NAP2. We also tested the performance of each primer set by changing the number of PCR cycles. Figure S2 indicates that non-specific products were generated after more than 30 cycles of PCR. This is consistent with previous findings that primer 3′ nucleotide mismatches could reduce PCR efficiency but not fully inhibit the reaction26. As a result, we proceeded to perform each assay within 30 cycles of PCR.
To achieve colorimetric detection of PCR products, we firstly conjugated urease with an oligonucleotide as reported in a previous method (see details in the methods)13. This conjugated urease-DNA (UrD) is used to hybridize to the extended binding region found in the reverse primer. Although the UrD does not contain a biotin modifier, the UrD has a tendency to non-specifically bind to magnetic beads at high concentrations and increase the background signal. To circumvent non-specific binding, varying concentrations of UrD was incubated with magnetic beads and tested with addition of phenol red (pH indicator) and urea after thorough washing of the beads. From this control experiment, we can determine the suitably diluted UrD solution to be used in subsequent experiments. As presented in Figure S3, we found that there was no observed color change after 2 hours, when 1 μL of 0.6 μM UrD in 10 μL of magnetic beads was used. Therefore, we chose to use this stock concentration for further assay development and testing.
We then carried out the colorimetric test on the PCR sample, expecting that the presence of PCR products would induce a color change. PCR positive products indeed yielded a color change from yellow to pink while samples that remained yellow indicated the absence of amplified PCR products and therefore did not capture the UrD. As shown in Fig. 2c, the results of three colorimetric reactions for each strain form a triplet pattern (no color change for absence of tcdC, one color change for wt-tcdC (non-epidemic strain), two color changes for ∆1stop-tcdC (epidemic strain 027/NAP1), and three color changes for TAA stop-tcdC (epidemic strain 078/NAP7)). These patterns can then be used to identify epidemic C. difficile strains.
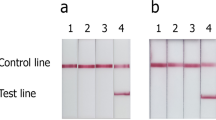
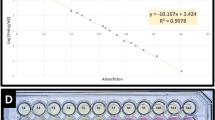
The sensitivity of the PCR-litmus assay was evaluated by testing genomic DNA isolated from serially diluted C. difficile stocks containing a known number of cells (Fig. 3a). A sharp color transition was observed for the sample containing 2 × 106 cells after incubation for 1 minute (top panel) and 2 × 105 cells after incubation for 10 minutes (middle panel). A subtle but detectable color transition, in comparison to the reference samples (without target), was observed for the sample containing 200 cells after color development of 1 hour. We compared the colorimetric test to conventional agarose gel based tests where PCR products were visualized through the staining of DNA-binding dyes. Three commonly used dyes were chosen for this experiment (SYBRSafe ; Fig. 3b), SYBRGold and GelRed (Figure S4). We found the dye-staining methods resulted a detection limit of approximately 104–105 bacteria cells. Therefore our litmus test exhibits a better detection sensitivity by as much as 100 fold. We have also performed spectroscopic analysis to quantify the detection sensitivity by plotting OD570/OD443 vs. cell numbers (Figure S5), based on the fact that the color of phenol red exhibits a gradual transition from yellow (λmax = 443 nm) to red (λmax = 570 nm) when the pH of a test solution changes from acidic to basic37. This analysis confirms that our method can indeed detect as low as 200 cells; however, the spectroscopic analysis did not lead to improved detection sensitivity.
Sensitivity of PCR-litmus test for C. difficile detection. 28 cycles of PCR was performed with genome DNA prepared from various numbers of C. difficile cells. Strain ATCC 1803, forward primer P3 and reverse primer RP were used in this test. (A) Litmus test with PCR products. The photograph was taken after a signal-producing time of 1 minute (top panel), 10 minutes (middle) and 60 minutes (bottom). (B) Analysis of PCR products by 2% agarose gel electrophoresis.
To further verify the accuracy of the assay in strain discrimination, we applied the assay towards genomic DNA extracted from 14 different strains of C. difficile (see Fig. 4). In addition, the tcdC genes from all these strains were independently sequenced to confirm their identity. It was found that our PCR-litmus test accurately identified three 027/NAP1 strains and one 078/NAP7 strain of C. difficile in reference to the tcdC sequencing analysis and the documentation provided by ATCC.
PCR-litmus test with genomic DNA from 14 different C. difficile strains. The tcdC genes from all the strains were sequenced and their nucleotide sequences were compared with database entries by using PubMLST program. Ribotype of each strain is provided by ATCC. The photographs of test were taken at 30 minutes.
Evidence of a C. difficile infection can be determined non-invasively by analyzing a patient’s stool sample. However, due to the composition and complexity of these samples, we next investigated the utility of our PCR-litmus assay in clinically relevant settings. We randomly selected stool samples of 12 CDI patients and 5 healthy donors obtained from St. Joseph’s Healthcare (Hamilton, Canada). In addition, healthy stool samples were also spiked with known strains of C. difficile cells with a concentration of 108 cells/g. Total DNA from each sample was extracted using Powerfecal DNA isolate kit (MO BIO LABORATORIES, USA). 100 ng of total DNA from each sample was used for PCR amplification. From Fig. 5, stool collected from healthy patients did not indicate color change. All 3 spiked samples revealed color patterns indicative of their respective spike strains. Of the 12 CDI patient samples, 4 presented a pattern of 027/NAP1 while the other 8 were identified as C. difficile (with wt-tcdC) but neither as 027/NAP1 nor 078/NAP7. These patterns are consistent with the tcdC sequencing results. These tests demonstrated that our approach could potentially be used for clinical diagnosis.
In summary, we have developed a simple method that can achieve colorimetric detection of DNA amplicons. To the best of our knowledge, our work represents the first example of adopting the classic litmus test for PCR-based DNA detection through the use of DNA conjugated urease. We have also shown that this method can be used to identify epidemic strains of C. difficile through the use of primers that can selectively amplify the tcdC genes in these strains. We have further illustrated the clinical usefulness of the proposed method by identifying epidemic strains of C. difficile in stool samples.
The featured PCR-litmus test can be easily extended to the detection of other gene targets simply by changing the primers, and thus we envision that this method can find wide applications in diverse fields.
Methods
Materials
All DNA oligonucleotides were prepared by automated DNA synthesis using standard phosphoramidite chemistry (Integrated DNA Technologies, Coralville, IA, USA) and purified by 10% denaturing (8 M urea) polyacrylamide gel electrophoresis (dPAGE). Their concentrations were determined spectroscopically. Deoxynucleoside 5′-triphosphates (dNTPs), SYBR Safe and SYBR Gold dyes were purchased from Thermo Scientific (Ottawa, ON, Canada). GelRed was obtained from Bitium (Fremont, CA, USA). Thermus thermophilus DNA polymerase was acquired from Biotools. Streptavidin coated magnetic beads of 1.5 μm (BioMag-SA) was purchased from Bangs Laboratories Inc. Urease powder from Canavaliaensiformis (Jack bean), maleimidobenzoic acid N-hydroxy-succinimide ester (MBS), phenol red were obtained from Sigma-Aldrich. Water was purified with a Milli-Q Synthesis A10 water purification system. All other chemicals were purchased from Bioshop Canada and used without further purification.
Bacterial strains and routine culture conditions
A panel of 14 C. difficile strains obtained from the American Type Culture Collection (ATCC; Manassas, USA) was used in this study. Cells were cultured in cooked meat broth medium (Sigma-Aldrich) under anaerobic condition with gas mixture of 80% N2, 10% CO2, 10% H2 in a Whitley Anaerobic Workstation (Don Whitley Scientific).
Total DNA extraction from bacteria strains
Cells were grown in 5 mL of cooked meat broth medium until OD600 reached ~1. 200 μL of cultures were spun down (11000 g, 5 min) in order to remove the culture medium; and the obtained pellets were suspended in 200 μL of 5% Chelex 100 (Bio-Rad) with 0.2 mg protease K. The mixture was then vortexed and incubated at 56 °C for 30 min and then 95 °C for 15 min. After centrifugation for 10 min at 10 000 g, the supernatant was transferred into a fresh tube and stored at 4 °C until PCR testing.
PCR primer design
For tcdC gene, primer design was conducted with the OligoAnalyzer 3.1(http://www.idtdna.com/calc/analyzer) after alignment of 26 tcdC_fragment gene sequences got from PubMLST (http://pubmlst.org/). All primers were checked using the alignments of sequences, and subsequently with the basic local alignment search tool (BLAST; http://www.ncbi.nlm.nih.gov/BLAST/). Secondary structures and self-pairing of all primers were also checked with the OligoAnalyzer 3.1. All primers are listed in Table S1.
PCR reaction
The PCR mixture (50 µL) contained 200 ng of DNA, 0.5 µM of forward primer (P1, P2 or P3) and reverse primer (RP), 200 µM of dNTPs (dATP, dCTP, dGTP and dTTP), 1 × PCR buffer (75 mM Tris-HCl, pH 9.0, 2 mM MgCl2, 50 mM KCl, 20 mM (NH4)2SO4) and 1 units of Thermus thermophilus (Tth) DNA polymerase. The DNA was amplified using the following thermocycling steps: 94 °C for 5 min; 28 cycles of 94 °C for 1 min, 60 °C for 1 min and 72 °C for 1 min; 72 °C for 3 min.
DNA-urease conjugation
Urease-DNA was prepared according to a previously reported method29. An MBS solution (6.4 mM) was made by dissolving 2 mg MBS (6.4 μmol) in 1 mL of dimethyl sulphoxide (DMSO). Similarly a urease solution was produced by dissolving 1.5 mg urease (2.75 nmol urease haxamer) powder in 1 mL of 1 × PBS buffer (pH 7.2). 10 nmol NH2-DNA and 3.2 μL of the MBS solution (20 nmol) were mixed and adjusted to a final reaction volume of 400 μL with 1 × PBS buffer, and allowed to react at room temperature. After 30 min, the mixture was passed through a membrane based molecular sizing centrifugal column with a molecular weight cut-off of 3,000 Daltons (NANOSEP OMEGA, Pall Incorporation) in order to remove excess MBS. The column was washed with 50 μL of 1 × PBS buffer 3 times and the DNA was resuspended in 100 μL of 1 × PBS buffer. The urease solution (1 mL, 2.75 nmol) was then added to the MBS activated DNA. The conjugation reaction was allowed to proceed at room temperature for 1 h. The mixture was filtered through a 300,000-Dalton cut-off centrifugal column. The DNA-urease conjugate (Ur-DNA) was then washed with 50 μL of 1 × PBS buffer 3 times, and resuspended in 160 μL of 1 × PBS buffer. The ratio of DNA over urease in UrD was found to be 1.54 using a previously published method38. Details are provided as the Supplementary Methods in Supplementary Information.
Litmus test
50 μL of the above PCR reaction mixture was incubated with 50 μL of binding buffer (10 mM Tris-HCl, pH 7.5, 3 M NaCl, 1 mM MgCl2, 0.01% tween 20) along with 10 μL of magnetic beads (MB) for 15 minutes. Then it was placed in a magnet holder to separate the supernatant and MB. Then the MB was suspended in 100 μL of binding buffer with 1 μL of 0.6 μM UrDNA (1 pmol). After 15 min of incubation, MB was washed with 100 μL of binding buffer four times and then resuspended in 70 μL of acetic acid buffer (0.1 mM, pH 5). Then 10 μL of 0.04% phenol red and 100 μL of substrate solution (3 M NaCl, 60 mM MgCl2, 50 mM urea) were added. Note that this substrate solution should have a starting pH of 5.0. A photograph was taken after a signal-producing time of 0–1 h according to individual experiments for Figs 2, 3, 4 and 5.
Sequencing of tcdC genes
Each tcdC gene was amplified by PCR as previously described33. The reaction mixture contained 1 × PCR buffer, 200 pmol of each dNTPs, 25 pmol of the FP-F and RP-F (Table S1), and 1 U of Tth DNA polymerase. The template was denatured for 5 min at 94 °C, and DNA was amplified for 30 cycles consisting of 1 min at 94 °C, 1 min at 50 °C, and 1 min at 72 °C. The PCR products were sequenced at the Mobix Lab (McMaster University). The nucleotide sequences were compared with database entries by using the PubMLST program.
Sensitivity test
A single colony of strain ATCC1803 from an anaerobic cooked meat broth agar plate was taken and cultured in 5 mL of cooked meat broth medium overnight. The bacterial culture was then diluted in 10-fold intervals seven times with cooked meat broth medium; 100 μL of 10−5, 10−6 and 10−7 dilutions were placed on a cooked meat broth plate and cultured for colony development in order to calculate the cell numbers (average colony-forming units) for each dilution. Meanwhile, 200 μL of each diluted cultures were used for DNA extraction and PCR-litmus test. After a signal producing time of 1 hour, measurements at 443 nm and 570 nm were performed by using a microplate scanning spectrometer (TECAN M1000). Agarose gel based tests were also performed. SYBR Safe, SYBR Gold and GelRed were used for gel staining.
Stool sample test
Stool samples from 5 healthy donors and 12 patients infected by C. difficile were obtained from St. Joseph’s Healthcare-Hamilton, with informed consent from all the healthy donors and infected patients. The experimental protocols were approved by Hamilton Integrated Research Ethics Board (HiREB), McMaster University (Hamilton, Ontario, Canada). The methods were carried out in accordance with the relevant guidelines and regulations. The three spiked samples were also prepared by adding ~2 × 107 C. difficile cells (ATCC 1870, ATCC1871 or ATCC1875) into 200 mg of stool sample from healthy donors. A Powerfecal DNA isolate kit (MO BIO LABORATORIES, USA) was used to extract total DNA from stool samples, and 100 ng of total DNA from each sample was used as template DNA for PCR-litmus test. The DNA was amplified using the following thermocycling steps: 94 °C for 5 min; 30 cycles of 94 °C for 1 min, 60 °C for 1 min and 72 °C for 1 min; 72 °C for 3 min.
Data Availability
All data generated or analysed during this study are included in this published article and its Supplementary Information.
References
Saiki, R. K. et al. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 239, 487–491 (1988).
Li, H. H. et al. Amplification and analysis of DNA sequences in single human sperm and diploid cells. Nature 335, 414–7 (1988).
Mullis, K. B. & Faloona, F. A. Specific synthesis of DNA in vitro via a polymerase-catalyzed chain reaction. Methods Enzymol. 155, 335–350 (1987).
Mullis, K. B. The unusual origin of the polymerase chain reaction. Scientific American 262(56–61), 64–65 (1990).
Law, J. W.-F., Ab Mutalib, N.-S., Chan, K.-G. & Lee, L.-H. Rapid methods for the detection of foodborne bacterial pathogens: principles, applications, advantages and limitations. Front. Microbiol. 5, 770 (2014).
Yamamoto, Y. PCR in diagnosis of infection: detection of bacteria in cerebrospinal fluids. Clin. Diagn. Lab. Immunol. 9, 508–514 (2002).
Erlich, H. A., Gelfand, D. & Sninsky, J. J. Recent advances in the polymerase chain reaction. Science 252, 1643–51 (1991).
Murakami, K. et al. Identification of methicillin-resistant strains of staphylococci by polymerase chain reaction. J. Clin. Microbiol. 29, 2240–2244 (1991).
Chan, K. et al. A rapid and low-cost PCR thermal cycler for infectious disease diagnostics. PLoS One 11, e0149150 (2016).
Wong, G., Wong, I., Chan, K., Hsieh, Y. & Wong, S. A Rapid and Low-Cost PCR Thermal Cycler for Low Resource Settings. PLoS One 10, e0131701 (2015).
Agrawal, N., Hassan, Y. A. & Ugaz, V. M. A pocket-sized convective PCR thermocycler. Angew. Chem. Int. Ed. 46, 4316–4319 (2007).
Marx, V. PCR heads into the field. Nat. Methods 12, 393–397 (2015).
Tram, K., Kanda, P., Salena, B. J., Huan, S. & Li, Y. Translating bacterial detection by DNAzymes into a litmus test. Angew. Chem. Int. Ed. 53, 12799–12802 (2014).
Leffler, D. A. & Lamont, J. T. Clostridium difficile Infection. N. Engl. J. Med. 372, 1539–1548 (2015).
Ekma, N., Yee, L. Y. & Aziz, R. A. Prevalence of Clostridium difficile infection in Asian countries. Rev. Med. Microbiol. 23, 1–4 (2012).
Rupnik, M., Wilcox, M. H. & Gerding, D. N. Clostridium difficile infection: New developments in epidemiology and pathogenesis. Nat. Rev. Microbiol. 7, 526–536 (2009).
McDonald, L. C. et al. An epidemic, toxin gene-variant strain of Clostridium difficile. N. Engl. J. Med. 353, 2433–41 (2005).
Loo, V. G. et al. A predominantly clonal multi-institutional outbreak of Clostridium difficile-associated diarrhea with high morbidity and mortality. N. Engl. J. Med. 353, 2442–9 (2005).
Warny, M. et al. Toxin production by an emerging strain of Clostridium difficile associated with outbreaks of severe disease in North America and Europe. Lancet 366, 1079–1084 (2005).
Pépin, J., Valiquette, L. & Cossette, B. Mortality attributable to nosocomial Clostridium difficile-associated disease during an epidemic caused by a hypervirulent strain in Quebec. CMAJ 173, 1037–1041 (2005).
Goorhuis, A. et al. Emergence of Clostridium difficile infection due to a new hypervirulent strain, polymerase chain reaction ribotype 078. Clin. Infect. Dis. 47, 1162–70 (2008).
Jhung, M. A. et al. Toxinotype V Clostridium difficile in humans and food animals. Emerg. Infect. Dis. 14, 1039–1045 (2008).
Rupnik, M., Widmer, A., Zimmermann, O., Eckert, C. & Barbut, F. Clostridium difficile toxinotype V, ribotype 078, in animals and humans. Journal of Clinical Microbiology 46, 2146 (2008).
Costa, M. C. et al. Prevalence and molecular characterization of Clostridium difficile isolated from feedlot beef cattle upon arrival and mid-feeding period. BMC Vet. Res. 8, 38 (2012).
Planche, T. et al. Diagnosis of Clostridium difficile infection by toxin detection kits: a systematic review. The Lancet Infectious Diseases 8, 777–784 (2008).
Longtin, Y. et al. Impact of the type of diagnostic assay on clostridium difficile infection and complication rates in a mandatory reporting program. Clin. Infect. Dis. 56, 67–73 (2013).
Bélanger, S. D., Boissinot, M., Clairoux, N., Picard, F. J. & Bergeron, M. G. Rapid detection of Clostridium difficile in feces by real-time PCR. J. Clin. Microbiol. 41, 730–734 (2003).
Citron, D. M. et al. Typing and susceptibility of bacterial isolates from the fidaxomicin (OPT-80) phase II study for C. difficile infection. Anaerobe 15, 234–236 (2009).
Brazier, J. S. Typing of Clostridium difficile. Clin. Microbiol. Infect. 7, 428–431 (2001).
Braun, V., Hundsberger, T., Leukel, P., Sauerborn, M. & Von Eichel-Streiber, C. Definition of the single integration site of the pathogenicity locus in Clostridium difficile. Gene 181, 29–38 (1996).
Thelestam, M. & Chaves-Olarte, E. Cytotoxic effects of the Clostridium difficile toxins. Curr Top Microbiol Immunol 250, 85–96 (2000).
Matamouros, S., England, P. & Dupuy, B. Clostridium difficile toxin expression is inhibited by the novel regulator TcdC. Mol. Microbiol. 64, 1274–1288 (2007).
Spigaglia, P. & Mastrantonio, P. Molecular analysis of the pathogenicity locus and polymorphism in the putative negative regulator of toxin production (TcdC) among Clostridium difficile clinical isolates. J. Clin. Microbiol. 40, 3470–3475 (2002).
Curry, S. R. et al. tcdC genotypes associated with severe TcdC truncation in an epidemic clone and other strains of Clostridium difficile. J. Clin. Microbiol. 45, 215–221 (2007).
Dingle, K. E. et al. Clinical clostridium difficile: Clonality and pathogenicity locus diversity. PLoS One 6, e19993 (2011).
Wilmer, A. et al. Polymerase chain reaction assay to detect Clostridium difficile tcdC variants is valuable in characterizing hospital epidemiology. J. Hosp. Infect. 84, 252–255 (2013).
Barbara Krajewska, B., van Eldik, R. & Brindell, M. Temperature- and pressure-dependent stopped-flow kinetic studies of jack bean urease. Implications for the catalytic mechanism. J. Biol. Inorg. Chem. 17, 1123–1134 (2012).
Kukolka, F., Lovrinovic, M., Wacker, R. & Niemeyer, C. M. Covalent coupling of DNA oligonucleotides and streptavidin. Methods Mol. Biol. 283, 181–96 (2004).
Acknowledgements
This work was supported by a grant to YL from the Natural Sciences and Engineering Research Council of Canada (NSERC).
Author information
Authors and Affiliations
Contributions
Y.L., D.C., C.H.L., B.J.S conceived the project; Y.L., D.C. and K.T. designed the project. D.C., B.L., Q.F., and Z.S. carried out the experimental Work. Y.L., B.J.S., Y.L., D.C., K.T., and B.L. wrote the manuscript with helps from all other authors.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chang, D., Tram, K., Li, B. et al. Detection of DNA Amplicons of Polymerase Chain Reaction Using Litmus Test. Sci Rep 7, 3110 (2017). https://doi.org/10.1038/s41598-017-03009-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-03009-z
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.