Abstract
The arthropod-borne Zika virus (ZIKV) is currently causing a major international public health threat in the Americas. This study describes the isolation of ZIKV from the plasma of a 29-year-old female traveler that developed typical symptoms, like rash, fever and headache upon return from Suriname. The complete genome sequence including the 5′ and 3′ untranslated regions was determined and phylogenetic analysis showed the isolate clustering within the Asian lineage, close to other viruses that have recently been isolated in the Americas. In addition, the viral quasispecies composition was analyzed by single molecule real time sequencing, which suggested a mutation frequency of 1.4 × 10−4 for this ZIKV isolate. Continued passaging of the virus in cell culture led to the selection of variants with mutations in NS1 and the E protein. The latter might influence virus binding to cell surface heparan sulfate.
Similar content being viewed by others
Introduction
Zika virus (ZIKV) is a mosquito-borne member of the Flaviviridae family. It is primarily transmitted through mosquitoes of the Aedes genus1. First isolated in 19472 and having caused only sporadic or unnoticed infections for about 60 years, the virus has received little attention until it emerged on several islands in the Pacific ocean in 20073. Since 2015, ZIKV is causing a major outbreak, affecting millions, in Brazil and other countries in South and Central America4, 5. Many infected travelers, often asymptomatic, have returned to other parts of the world and between June and August 2016 the first local mosquito-transmitted ZIKV infections occurred in the USA6.
ZIKV has a positive-stranded RNA genome that is capped, lacks a poly(A) tail and has a typical flavivirus genome organization. It contains a single open reading frame that encodes the 3 structural and 7 nonstructural proteins in the form of a polyprotein. ZIKV strains can be classified in two separate lineages, the African and Asian lineages. Both lineages have emerged from East Africa around the end of the 19th century7. The viruses that are spreading in the Americas are very similar to the African genotypes that have been isolated decades ago, but based on phylogenetic studies they were shown to be most closely related to the Asian lineage strain that circulated in French Polynesia in 20135, 8, 9.
ZIKV causes asymptomatic infections in about 80% of the cases and mild dengue-like symptoms in the majority of symptomatic patients, including fever, myalgia, headache and a macopapular rash. However, in pregnant women ZIKV infections can result in serious damage to the fetus, causing neurologic disorders and neonatal malformations, like microcephaly10. In rare cases, ZIKV-infected adults can develop a paralytic neurological complication known as the Guillain-Barré syndrome11, 12. Besides transmission by mosquitos, ZIKV can also utilize a sexual mode of transmission13,14,15. Blood or urine is generally used to diagnose ZIKV infections, but viral RNA has been found in other body fluids and tissues as well. To date, no registered vaccine or antiviral therapy is available to prevent or treat ZIKV infections.
The RNA-dependent RNA polymerases of RNA viruses lack proofreading ability, leading to high mutation rates in comparison to cellular organisms16. These high mutation rates result in a high genetic variation within the virus population, the so-called quasispecies, a set of genetically related viruses closely distributed around a consensus sequence17, 18. Next-generation sequencing (NGS) has enabled the analysis of the population of viral quasispecies with a much higher resolution than conventional sequencing. The characterization of the ZIKV quasispecies composition and (intrahost) evolutionary dynamics in relation to adaptation, virulence, and immune escape is important to better understand ZIKV infection and pathogenesis, and to aid development of vaccines and therapeutics.
In the present study, we report and analyze the complete genome sequence and quasispecies distribution of a ZIKV strain from Suriname isolated from the plasma of a 29-year-old female traveler who returned to the Netherlands and exhibited typical symptoms of ZIKV infection.
Results
Case description
A 29-year-old female medical microbiologist in training developed a low grade fever, general malaise and a retro-orbital headache related to eye movements three days after returning from Suriname. Three days later, the patient noticed a redness in her face, a non-itching maculopapular rash on her trunk and extremities, and slight conjunctival injection, which she recorded on photo (Fig. 1). On the same day the patient developed stiffness and edematous swelling of wrists, knees and ankles. Throughout the course of disease she had contact lens intolerance. The symptoms did not lead to lost working days and diminished after 7 days. On the fifth day of her illness she visited our outpatient clinic where urine and blood was collected for diagnosis and characterization of an arboviral infection. Urine and plasma tested positive for ZIKV RNA by an in-house qRT-PCR (manuscript in preparation), and the plasma also tested positive in virus culture. No infectious virus could be recovered from the urine. The patient had been bitten by mosquitos in Suriname where she had visited Paramaribo, the Brokopondo Lake and the Commewijne district.
Isolation and sequencing of ZIKV strain SL1602
Plasma of the patient (12 μL) was added to a semi-confluent culture of Vero E6 cells. After 4 days the cell culture medium, which tested positive for ZIKV-RNA in qRT-PCR, was harvested (P1 stock) and 1 ml was used to infect a fresh batch of Vero E6 cells for subsequent passaging. The P1 stock had a ZIKV titer of 9.0 × 102 pfu/ml, while the P2 stock had a titer of 8.2 × 105 pfu/ml, as determined by plaque assay on Vero cells. Attempts to isolate infectious ZIKV from urine were unsuccessful.
Viral RNA was isolated from the P2 virus stock (cell culture supernatant) and 5 partially overlapping amplicons were generated. A near complete genome sequence was first obtained by Sanger sequencing. The sequences of the 5′ and 3′ termini, which are frequently missing or incorrect in other ZIKV sequences in GenBank, were determined by rapid amplification of cDNA ends (5′- and 3′-RACE). To obtain the consensus complete genome sequence of ZIKV SL1602, we combined Sanger sequencing, 5′- and 3′-RACE data, and Pacific Biosciences RSII single molecule real time (SMRT) sequencing. The complete genome of this isolate consists of 10,807 nucleotides with an overall G/C content of 51.2%.
The quasispecies distribution was determined by subjecting the 5 overlapping amplicons to SMRT sequencing. The data from the SMRT deep-sequencing run yielded a total of 31,421 high-quality single-molecule sequencing reads, with ZIKV genome coverage ranging from 427 to 5,831 reads at single nucleotide positions, with an average of 2,300 reads (Figure S1). The minimum number of passes for each single molecule read was 5 and the maximum number of passes was 183, with a median of 12. This led to the high minimum read quality of 0.99 and an average read quality of 0.998.
Phylogenetic relationships between ZIKV SL1602 and other isolates
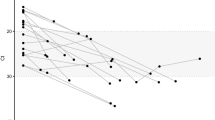
To understand the evolutionary relationship of ZIKV SL1602, a maximum likelihood phylogenetic tree was inferred using ZIKV complete genome nucleotide sequences (Fig. 2). The tree was produced for a spatiotemporal representative set of ZIKV strains for which complete genome sequences were available in GenBank. ZIKV SL1602 clusters within the Asian lineage, relatively close to viruses that were or are still circulating in French Polynesia, Brazil, Haiti, and Colombia. This suggests that the 2015/2016 ZIKV isolates in American and Asian countries form a new American clade within the Asian lineage. Although the phylogenetic distance between the isolates of the Asian lineage is very small, the Columbia/2016 isolate is the closest relative of ZIKV SL1602 with only 44 nucleotides difference thereby having a 99.5% sequence identity.
Phylogenetic tree for ZIKV SL1602 and 12 selected ZIKV isolates that represent the recognized diversity. A maximum likelihood phylogenetic tree inferred from the nucleotide sequences of full-length ZIKV genomes is shown. The country of isolation and GenBank accession numbers of the isolates that were used are: Uganda MR 766 (NC012532), Brazil-Natal (KU527068), Thailand (KU681081), Philippines (KU681082), Brazil-Paraiba (KX280026), Colombia (KX247646), Brazil-Recife (KX197192), Haiti (KX051563), Senegal (KU955591), Cambodia (KU955593), Federated States of Micronesia (EU545988) and French Polynesia (KJ776791.2). Spondweni virus (NC029055, not shown) was used as the outgroup. Bootstrap values above 70 are shown. The scale bar represents the number of nucleotide substitutions per site.
Quasispecies variation
SMRT sequencing reads were aligned to the consensus sequence and for each nucleotide position the count for each of the four bases was determined. A total of 24,815,877 nucleotides was sequenced and 3,375 single-nucleotide variants (SNV) were counted. This suggests that ZIKV has a mutation frequency of 1.4 × 10−4, which falls within the range that has been reported for other RNA viruses19.
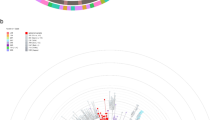
Seventy-one SNVs were mapped using coverage-corrected cut-off values (Fig. 3). Their frequencies ranged from 0.0012 to 0.0835. Out of the 71 SNVs, 27 (38%) were non-synonymous (Table 1), and 44 (62%) were synonymous mutations (Supplemental Table 2).
Four variants in the quasispecies population were found to be present at a level of over 1% of the total population. One of these variants contained a mutation in the E protein (4.1%), and the other three contained mutations in the nonstructural proteins NS2A (8.2%), NS2B (1.6%), and NS5 (1.4%) (Fig. 3).
The passage 2 (P2) ZIKV isolate that was sequenced was passaged 9 more times in Vero cells to obtain a P11 stock, which was analyzed by Sanger sequencing. The consensus sequence of the P11 virus differed at two nucleotide positions from the P2 virus. It has the G1898U, mutation, which results in a L307F substitution in the E protein, and the A3147G mutation, which leads to a M220V substitution in the NS1 protein. For both positions, SNVs were already observed in the P2 quasispecies, with a frequency of 0.0419 and 0.0046, respectively (Table 1). Sequencing of virus from earlier passages (P5 and P8 virus) revealed that both changes were fixed between passage 9 and 11. These changes likely reflect an adaptation to Vero cells.
Discussion
Here, we report the isolation and analysis of a ZIKV strain from a traveler returning from Suriname with typical symptoms of Zika. The complete genome was sequenced, a phylogenetic analysis was performed and single nucleotide variations were analyzed to characterize the quasispecies composition of this clinical isolate that caused commonly observed mild symptoms.
The analysis of the quasispecies distribution of ZIKV SL1602 was done on the Pacific Biosciences RSII, which allows the rapid sequencing of relatively large DNA molecules. Only five PCR fragments were used to sequence the complete genome, without additional library preparation steps (e.g. fragmentation). Although SMRT sequencing is prone to random high sequencing errors, every single amplicon molecule was sequenced a minimum of 5 times (Figure S2), leading to a very high accuracy of base calling (Figure S3).
Our phylogenetic analysis showed that ZIKV SL1602 clusters within the Asian lineage, close to the viruses that are currently circulating in the Americas (Fig. 1), in accordance with other studies8, 20,21,22. All of the South American isolates share a common ancestor with the French Polynesia isolate, suggesting that the currently spreading South American clade is imported from French Polynesia.
ZIKV SL1602 has a typical flavivirus genome organization with one large open reading frame that encodes the polyprotein C-prM-E-NS1-NS2A-NS2B-NS3-NS4A-NS4B-NS5 that will be processed into 3 structural and 7 nonstructural proteins. The sequence reported here includes the complete 5′ and 3′ untranslated regions, which are often lacking from other sequences that have been deposited in public databases.
RNA viruses have high mutation rates and evolve rapidly due to the lack of proofreading of the RNA-dependent-RNA-polymerase. Our NGS analysis shows that the mutation frequency of ZIKV SL1602 is 1.4 × 10−4, which is within the range commonly observed for RNA viruses (10−4 to 10−6)19. We did not observe a preferential positive selection for mutations in nonstructural proteins, as was described before23.
Repeated passaging of ZIKV SL1602 in Vero cells led to selection of a variant with an L307F substitution in the E protein and a M220V substitution in NS1. The L307F variant was already present in the P2 quasispecies with a frequency of 0.0419. Based on its structure, the flavivirus E protein can be divided into domains I, II and III24 of which domain III is believed to play a major role in antibody binding, pathogenesis and attachment to host cells. The L307F mutation is located in domain III of the ZIKV E protein. It might therefore influence cell attachment and infectivity, since domain III was shown to contribute to the tropism, infectivity, pathogenesis and cell attachment of other flaviviruses, possibly involving cell surface heparan sulfate binding25,26,27. The M220V mutation in NS1 that was fixed during repeated passaging in Vero cells is a conservative replacement. However, considering the role of NS1 in DENV particle biogenesis28, it cannot be excluded that it can contribute to increased replication or other selective advantages in cell culture.
By analyzing the complete genome sequence of a South American (Surinamese) ZIKV isolate, our study shows the quasispecies composition and minority variants of ZIKV early after isolation from a patient, and upon repeated passaging in cell culture. Insights into the evolutionary dynamics of ZIKV can aid in the development of improved diagnostics, surveillance, development of vaccines and therapeutics, and thereby in controlling further spread of ZIKV.
Methods
Cell culture, virus isolation and titration
Vero cells (ATCC CCL-81) and Vero E6 cells (ATCC CRL-1586) were cultured in Dulbecco’s modified Eagle’s medium (DMEM), supplemented with 8% fetal calf serum (FCS), 2 mM L-glutamine, 100 IU/ml of penicillin and 100 μg/ml of streptomycin at 37 °C in 5% CO2. After infection, cells were maintained in Eagle’s Minimum Essential Medium (EMEM) with 25 mM HEPES supplemented with 2% FCS, L-glutamine, and antibiotics. All experiments with infectious ZIKV were performed in the LUMC BSL-3 facility. Written informed consent was obtained from the patient to publish images that show the symptoms of infection and to use clinical samples taken for diagnosis for scientific research, including the isolation of virus. A semi-confluent monolayer of Vero E6 cells in a T-25 flask was incubated with 12 µl of plasma in a total volume of 5 ml medium. After 4 days, 1 ml of supernatant was transferred to a new T-25 flask with Vero E6 cells already containing 5 ml of medium and 8 days later the medium was harvested and stored as a P2 stock. Viral titers were determined by plaque assay on confluent monolayers of Vero cells in six-well clusters, which were infected with 10-fold serial dilutions (in duplicate) of the samples in a total volume of 0.5 ml. After a 2-h incubation at 37 °C, the inoculum was replaced with 2 ml of DMEM containing 1.2% Avicel RC-581 (FMC BioPolymer), 2% FCS, 25 mM HEPES, and antibiotics. After a 5-day incubation, cells were fixed with 3.7% formaldehyde in PBS and plaques were visualized by crystal violet staining. ZIKV RNA was detected by an internally controlled multiplex TaqMan qRT-PCR (manuscript in preparation).
ZIKV RNA isolation and generation of cDNA
ZIKV-specific primers to generate 5 overlapping amplicons were designed based on an alignment of the reference sequence ZIKV strain MR766 and other complete genomes present in GenBank in February 2016 (Table S1).
ZIKV RNA was isolated from infected cell culture medium using the Qiamp Viral mini kit (Qiagen) and cDNA was generated by reverse transcription (5 separate reactions) with Superscript III polymerase (Thermo Fisher Scientific) and primers A1R, A2R, A3R, A4R and A5R (Table S1) according to the supplier’s instructions. Subsequently, 5 µl cDNA was used in 50 µl PCR reactions with Q5 high-fidelity DNA Polymerase (NEB) with GC enhancer to generate 5 amplicons with sizes of 2.4, 3.3, 3.1, 2.5 and 0.54 kb in 5′ to 3′ order, using primer combinations A1F & A1R, A2F & A2R, A3F & A3R, A4F & A4R and A5F & A5R. An initial step of 30 s at 98 °C was followed by 35 cycles consisting of a 10 s denaturation step at 98 °C, a 30-s annealing step at 64 °C and a 140 s extension at 72 °C for amplicons 1–4, while a 66 °C annealing temperature and 40 s extension time was used to generate amplicon 5. After cleanup with the ISOLATE II PCR Kit (Bioline), DNA was quantified using the Qubit Fluorometer and dsDNA BR Assay (Thermo Fisher Scientific). It was analyzed on agarose gels and the quality was checked with a Bioanalyzer DNA 12000 Kit (Agilent).
Full genome sequencing
ZIKV cDNA amplicons were subjected to Sanger sequencing with overlapping reads that covered the amplicons on both strands (primer sequences available upon request). For sequencing of the 5′ and 3′ termini of the genome, isolated ZIKV RNA was first polyadenylated using yeast poly(A) polymerase (Affymetrix), followed by rapid amplification of cDNA ends (RACE). RACE was done with the 2nd Generation 5′/3′ RACE kit (Roche) using primers SP1, SP2, SP3, SP5 (Table S1) and the primers included in the kit, according to the manufacturer’s instructions. The 5′ and 3′ amplicons were sequenced using primers 5RACE-seq1 and 3RACE-seq, respectively (Table S1).
For NGS, the 5 amplicons were gel-purified, pooled in equimolar concentrations and library preparation was performed according to the Pacific Biosciences Amplicon Template Preparation protocol. Sequencing was performed on a single SMRT cell using a 6-hour runtime and the C4-P6 chemistry on the RSII platform. Reads-of-inserts with a minimum of 5 full passes and minimum predicted accuracy of 99% were subsequently mapped to the ZIKV SL1602 reference sequence using SMRT Analysis software version 2.3. For each amplicon, primer sequences were removed and nucleotide counts were generated for each genomic position.
Mutation frequency
All reads derived from SMRT sequencing were aligned to the ZIKV SL1602 consensus sequence. From this alignment, a table was created that contained for each genome position the counts for each of the four nucleotides. The ‘total number of mutations’ was the sum of all counts of the nucleotides that differed from the consensus sequence. The ZIKV mutation frequency was calculated by dividing the total number of mutations by the total number of nucleotides sequenced.
Phylogenetic reconstruction
Nucleotide sequences were aligned using the ClustalW software running within the BioEdit (version 7.2.5) program29. This resulted in a 10,302 nt alignment. Maximum likelihood phylogenetic trees with 1000 bootstrap replicates were estimated under the general time-reversible model (GTR)+G+I+F using PhyML 3.0 software30.
Sequence accession number
The ZIKV SL1602 consensus sequence, based on PacBio and Sanger sequencing (RACE) was submitted to GenBank under accession number KY348640.
References
Grard, G. et al. Zika virus in Gabon (Central Africa)–2007: a new threat from Aedes albopictus? PLoS neglected tropical diseases 8, e2681, doi:10.1371/journal.pntd.0002681 (2014).
Dick, G. W., Kitchen, S. F. & Haddow, A. J. Zika virus. I. Isolations and serological specificity. Transactions of the Royal Society of Tropical Medicine and Hygiene 46, 509–520, doi:10.1016/0035-9203(52)90042-4 (1952).
Duffy, M. R. et al. Zika virus outbreak on Yap Island, Federated States of Micronesia. The New England journal of medicine 360, 2536–2543, doi:10.1056/NEJMoa0805715 (2009).
Campos, G. S., Bandeira, A. C. & Sardi, S. I. Zika Virus Outbreak, Bahia, Brazil. Emerging infectious diseases 21, 1885–1886, doi:10.3201/eid2110.150847 (2015).
Zanluca, C. et al. First report of autochthonous transmission of Zika virus in Brazil. Mem Inst Oswaldo Cruz 110, 569–572, doi:10.1590/0074-02760150192 (2015).
Government, F. S. Gov. Scott: with likely mosquito-borne Zika cases, states will use full resources to protect Floridians., http://www.flgov.com/2016/07/29/gov-scott-with-likely-mosquito-borne-zika-cases-state-will-use-full-resources-to-protect-floridians/ (2016).
Gatherer, D. & Kohl, A. Zika virus: a previously slow pandemic spreads rapidly through the Americas. The Journal of general virology 97, 269–273, doi:10.1099/jgv.0.000381 (2016).
Enfissi, A., Codrington, J., Roosblad, J., Kazanji, M. & Rousset, D. Zika virus genome from the Americas. Lancet (London, England) 387, 227–228, doi:10.1016/s0140-6736(16)00003-9 (2016).
Lanciotti, R. S., Lambert, A. J., Holodniy, M., Saavedra, S. & Signor Ldel, C. Phylogeny of Zika Virus in Western Hemisphere, 2015. Emerging infectious diseases 22, 933–935, doi:10.3201/eid2205.160065 (2016).
Driggers, R. W. et al. Zika Virus Infection with Prolonged Maternal Viremia and Fetal Brain Abnormalities. The New England journal of medicine 374, 2142–2151, doi:10.1056/NEJMoa1601824 (2016).
Dos Santos, T. et al. Zika Virus and the Guillain-Barre Syndrome - Case Series from Seven Countries. The New England journal of medicine 375, 1598–1601, doi:10.1056/NEJMc1609015 (2016).
Parra, B. et al. Guillain-Barre Syndrome Associated with Zika Virus Infection in Colombia. The New England journal of medicine 375, 1513–1523, doi:10.1056/NEJMoa1605564 (2016).
D’Ortenzio, E. et al. Evidence of Sexual Transmission of Zika Virus. The New England journal of medicine 374, 2195–2198, doi:10.1056/NEJMc1604449 (2016).
Hills, S. L. et al. Transmission of Zika Virus Through Sexual Contact with Travelers to Areas of Ongoing Transmission - Continental United States, 2016. MMWR. Morbidity and mortality weekly report 65, 215–216, doi:10.15585/mmwr.mm6508e2 (2016).
Musso, D. et al. Potential sexual transmission of Zika virus. Emerging infectious diseases 21, 359–361, doi:10.3201/eid2102.141363 (2015).
Drake, J. W. & Holland, J. J. Mutation rates among RNA viruses. Proceedings of the National Academy of Sciences USA 96, 13910–13913, doi:10.1073/pnas.96.24.13910 (1999).
Eigen, M. Viral quasispecies. Sci Am 269, 42–49, doi:10.1038/scientificamerican0793-42 (1993).
Holland, J. J., De La Torre, J. C. & Steinhauer, D. A. RNA virus populations as quasispecies. Curr Top Microbiol Immunol 176, 1–20, doi:10.1007/978-3-642-77011-1 (1992).
Sanjuan, R., Nebot, M. R., Chirico, N., Mansky, L. M. & Belshaw, R. Viral mutation rates. Journal of virology 84, 9733–9748, doi:10.1128/JVI.00694-10 (2010).
Faria, N. R. et al. Zika virus in the Americas: Early epidemiological and genetic findings. Science (New York, N.Y.) 352, 345–349, doi:10.1126/science.aaf5036 (2016).
Wang, L. et al. From Mosquitos to Humans: Genetic Evolution of Zika Virus. Cell host & microbe 19, 561–565, doi:10.1016/j.chom.2016.04.006 (2016).
Ye, Q. et al. Genomic characterization and phylogenetic analysis of Zika virus circulating in the Americas. Infection, genetics and evolution: journal of molecular epidemiology and evolutionary genetics in infectious diseases 43, 43–49, doi:10.1016/j.meegid.2016.05.004 (2016).
Sironi, M., Forni, D., Clerici, M. & Cagliani, R. Nonstructural Proteins Are Preferential Positive Selection Targets in Zika Virus and Related Flaviviruses. PLoS neglected tropical diseases 10, e0004978, doi:10.1371/journal.pntd.0004978 (2016).
Rey, F. A., Heinz, F. X., Mandl, C., Kunz, C. & Harrison, S. C. The envelope glycoprotein from tick-borne encephalitis virus at 2 A resolution. Nature 375, 291–298, doi:10.1038/375291a0 (1995).
Hung, J. J. et al. An external loop region of domain III of dengue virus type 2 envelope protein is involved in serotype-specific binding to mosquito but not mammalian cells. Journal of virology 78, 378–388, doi:10.1128/JVI.78.1.378-388.2004 (2004).
McAuley, A. J. et al. Recovery of West Nile Virus Envelope Protein Domain III Chimeras with Altered Antigenicity and Mouse Virulence. Journal of virology 90, 4757–4770, doi:10.1128/JVI.02861-15 (2016).
Roehrig, J. T. et al. Mutation of the dengue virus type 2 envelope protein heparan sulfate binding sites or the domain III lateral ridge blocks replication in Vero cells prior to membrane fusion. Virology 441, 114–125, doi:10.1016/j.virol.2013.03.011 (2013).
Scaturro, P., Cortese, M., Chatel-Chaix, L., Fischl, W. & Bartenschlager, R. Dengue Virus Non-structural Protein 1 Modulates Infectious Particle Production via Interaction with the Structural Proteins. PLoS pathogens 11, e1005277, doi:10.1371/journal.ppat.1005277 (2015).
Hall, T. A. BioEdit: a user-friendly biological sequence alignment Editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98 (1999).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Systematic Biology 59, 307–321, doi:10.1093/sysbio/syq010 (2010).
Author information
Authors and Affiliations
Contributions
S.v.B., A.T., I.C.A. and M.J.v.H. performed experiments. R.v.G. and M.C.F. contributed essential reagents. S.v.B., A.T., S.Y.A., I.C.A., M.C.F., P.J.B. and M.J.v.H. analyzed the data. M.P.B. was the treating physician. S.v.B., S.Y.A., R.v.G., M.C.F., and M.J.v.H. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
van Boheemen, S., Tas, A., Anvar, S.Y. et al. Quasispecies composition and evolution of a typical Zika virus clinical isolate from Suriname. Sci Rep 7, 2368 (2017). https://doi.org/10.1038/s41598-017-02652-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-02652-w
This article is cited by
-
African ZIKV lineage fails to sustain infectivity in an in vitro mimetic urban cycle
Brazilian Journal of Microbiology (2023)
-
Comparative analysis of a Thai congenital-Zika-syndrome-associated virus with a Thai Zika-fever-associated virus
Archives of Virology (2020)
-
A natural polymorphism in Zika virus NS2A protein responsible of virulence in mice
Scientific Reports (2019)
-
A yellow fever–Zika chimeric virus vaccine candidate protects against Zika infection and congenital malformations in mice
npj Vaccines (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.