Abstract
Rice blast caused by Pyricularia oryzae (syn., Magnaporthe oryzae) was one of the most destructive diseases of rice throughout the world. Genome assembly was fundamental to genetic variation identification and critically impacted the understanding of its ability to overcome host resistance. Here, we report a gapless genome assembly of rice blast fungus P. oryzae strain P131 using PacBio, Illumina and high throughput chromatin conformation capture (Hi-C) sequencing data. This assembly contained seven complete chromosomes (43,237,743 bp) and a circular mitochondrial genome (34,866 bp). Approximately 14.31% of this assembly carried repeat sequences, significantly greater than its previous assembled version. This assembly had a 99.9% complement in BUSCO evaluation. A total of 14,982 genes protein-coding genes were predicted. In summary, we assembled the first telomere-to-telomere gapless genome of P. oryzae, which would be a valuable genome resource for future research on the genome evolution and host adaptation.
Similar content being viewed by others
Background & Summary
Pyricularia oryzae (syn., Magnaporthe oryzae), an ascomycete fungal pathogen, causes rice blast, one of the most destructive diseases of rice throughout the world1,2. The pathogen is an important and long-established model species for understanding fungal-plant interactions3,4. Previously, we sequenced and assembled the first genomes of field strains (P131 and Y34) and performed a comparative analysis between the laboratory and field strains, which demonstrated that translocation of transposable elements (TEs), gain or loss of isolate-specific genes and gene family expansion are essential factors, delimiting genomic plasticity and adaptability of P. oryzae5. Although these assemblies had facilitated the understanding of the genome characteristics of P. oryzae, the genome of the two strains were highly fragmented to more than one thousand scaffolds, for Sanger (2-fold) and 454 (18-fold) sequencing technologies were used in the previous study. Recently, over 50 genomes of different strains of P. oryzae have been available in public genome databases. These genomes were sequenced on the next-generation sequencing platforms, such as second-generation sequencing platforms (e.g., Illumina sequencers) and/or third-generation sequencing platforms [e.g., Pacific Biosciences (PacBio)], which facilitated the genetic studies of genomic changes and pathogenicity variation within P. oryzae6,7,8. However, currently most of these assemblies are fragmented and contain a large number of unplaced contigs and/or gaps owing to the presence of repetitive DNA elements in the P. oryzae genomes, which prevented the dissection of molecular mechanisms of adaptive evolution. Since the importance of genome assembly completeness in genomic analysis, we re-assemble the genome of P. oryzae stain P131 by combining Illumina, PacBio sequencing and high throughput chromatin conformation capture (Hi-C) mapping, which was the first telomere-to-telomere gapless assembly of the P. oryzae genome.
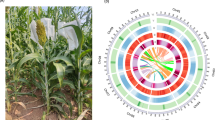
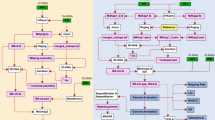
A total of 10.03 Gb PacBio long-read sequencing data (~250x genome coverage) and 4.44 Gb Illumina short-read sequencing data were generated (Table 1). Hi-C library was prepared, sequenced and generated 5.57 Gb sequencing data (~140x genome coverage). The long reads were de novo assembled and corrected. The short reads were used to polish the assembly. Redundant genomic contigs or mitochondrial contigs were then removed. The Hi-C sequencing data were used to anchor and refined remained contigs. The mitochondrial genome was assembled independently by Mitochondrial Long-read Iterative Assembly (MLIA) pipeline9. The final polishing of the complete genome was performed. Finally, seven gapless chromosomes (43,237,743 bp with a contig N50 of 7.05 Mb; Fig. 1a) and a circular mitochondrial genome (34,866 bp; Fig. 1b) were constructed in the final assembly (Fig. 2). The new assembly represented a significant improvement over the previous version GCA_000292605.15,10 (1,823 assembled contigs and contig N50 = 12.3 kb; see Table 2 and Fig. 3).
The P. oryzae strain P131 genome assembly. (a) Nuclear genome: Track 1 illustrates the seven assembled nuclear chromosomes with the indicated sizes. The red arrows at the end of chromosomes indicate telomeric repeat sequences (TTAGGG)n. Tracks 2 through 4 show transposon distribution, gene density, and GC density on the seven chromosomes, respectively. (b) Mitochondrial genome: Track 5 depicts a circular illustration of the mitochondrial genome carrying genes (represented by different color blocks), including genes encoding 14 standard fungal core PCGs (nad1-nad6, nad4L, cob, cox1-cox3, atp6, atp8, and rps3), ribosomal subunits (rns and rnl) and 27 tRNA genes. Tracks 6 and 7 display the introns present in the above genes and GC density of the mitochondrial genome, respectively.
The nuclear genome was annotated by Braker2 pipeline11. The mitochondrial genome was annotated by MFannot12 using genetic code 4. In conclusion, the nuclear genome is predicted to contain 14,968 genes (including 20,797 transcripts), and the mitochondrial genome is likely to carry 14 conserved protein-coding genes (Table 3). A total of 99.9% of the BUSCOs were mapped onto the P131 genome assembly. Approximately 14.31% of the genome carried repeat sequences, most of which were TEs, which was significantly greater than the previous version (Table 4).
The telomere repeat sequence (TRS) (TTAGGG)n was presented on both ends of chromosomes 2, 4, 5, 6, and 7 and one end of chromosomes 1 and 3 in our assembly. We then compared the TRS in the published near-complete assembled genome of P. oryzae strains with the genome assembly generated in this study. Interestingly, minority deficiency and telomere variability of TRSs in P. oryzae were extensively observed, which may play subtle roles in pathogenic adaptation13,14,15. In summary, we assembled the first telomere-to-telomere gapless genome of P. oryzae, which can be instrumental in understanding the genome evolution and host adaptation in the rice blast fungus.
Methods
Sampling and DNA extraction
The P. oryzae strain P131 was grown and maintained on oatmeal tomato agar (OTA) plates16. Conidia were produced on OTA plates and harvested from 7-day culture plates grown at 25 °C under constant fluorescent light. Hyphae were collected from 2-day-old cultures in complete medium shaken at 150 rpm at 25 °C. Genomic DNA extracted from vegetative mycelia using cetyltrimethylammonium bromide (CTAB) protocol was used for genome sequencing17.
Illumina, PacBio and Hi-C sequencing
Genome sequencing was conducted on Pacific Biosciences Sequel (PacBio, Menlo Park, CA) at CapitalBio Technology Co., Ltd (Beijing, China). Qualified genomic DNA was fragmented with G-tubes (Covaris, Woburn, MA, USA) and end-repaired to prepare SMRTbell DNA template libraries (with fragment size of >10 kb selected). Library quality was detected by Qubit dsDNA HS Assay Kit (Thermo Fisher Scientific, Waltham, MA, USA, Q33230). The average fragment size was estimated on Bioanalyzer 2100 (Agilent, Santa Clara, CA). SMRT sequencing was performed on the Pacific Biosciences RSII sequencer (PacBio, Menlo Park, CA) according to standard protocols using the P4-C2 chemistry. A total of 10.03 Gb PacBio sequencing data with a subread N50 of 14.5 kb. In addition, Illumina HiSeq X Ten sequencer using paired-end technology was also used to perform genome sequencing and 4.44 Gb sequencing data (150 bp paired-end reads) were yielded at CapitalBio Technology Co., Ltd (Beijing, China).
Hi-C library was prepared from cross-linked chromatins of fungal mycelia by Novogene Co., Ltd (Beijing, China). In brief, the tissue was ground and then cross-linked with 4% formaldehyde solution. After the sample of crosslinking reaction and cell lysis, nuclei were digested with 4-cutter restriction enzyme DpnII. Subsequently, ligated DNA was purified and fragmented into 300 bp size on average. The constructed Hi-C library was sequenced by Illumina NovaSeq 6000. 5.57 Gb paired-end sequencing data (150-bp length) were generated. The Hi-C maps from raw data were performed by Juicer (v1.6)18, followed by using a manually correction with Juicebox (v2.13.07)19.
RNA sequencing and analysis
Total RNA was extracted from conidia and hyphae with the Trizol reagent (Invitrogen, Carlsbad, CA, USA, 15596026) and then enriched by RNeasy Pure mRNA Bead Kit (Qiagen, Germany), respectively. High-throughput cDNA libraries were prepared according to the Illumina whole transcriptome library preparation protocol and sequenced on the Illumina GA platform by the BGI Genomics (Shenzhen, China)20. Quality control was performed by FastQC (v0.11; https://github.com/s-andrews/FastQC). RNA-Seq data were mapped to P. oryzae by HISAT2 (v2.2.1)21, and SAMTools (v1.12)22 were used to evaluate read alignments.
Genome assembly
The de novo long-read assembler Canu v2.1.123 (parameters: genomeSize = 44 m corOutCoverage = 200 corMinCoverage = 2 minReadLength = 4000 minOverlapLength = 800 correctedErrorRate = 0.050) was used to assemble PacBio reads to generated draft contigs, which were then corrected by GCpp v1.9 (https://github.com/PacificBiosciences/gcpp; parameters:–algorithm = arrow -x 5 -X 200 -q 40) using PacBio long-reads. The polishing step was performed by Pilon v1.2324 (parameters:–changes–vcf) using the Illumina short reads. Contigs were considered redundant if they aligned concordantly (identity >99%) with another contig, and the redundant contigs, along with mitochondrial contigs, were removed, resulting in a total of 13 contigs. The Hi-C sequencing data were used to anchor all 13 contigs using Juicer v1.618, resulting in 7 scaffolds, which were further refined using Juicebox v2.13.0719. Gaps within the scaffolds were filled using LR_Gapcloser25. We then manually checked whether long reads aligned the bridging cross the gaps, or whether overlapping contig ends (>20 kb length and 99.9% sequence identity) existed.The mitochondrial genome was assembled independently by Mitochondrial Long-read Iterative Assembly (MLIA) pipeline9. The final polishing of the complete genome was performed again using Pilon v1.2324.
Gene model and function annotations
Repetitive sequences of P. oryzae strain P131 was firstly de novo identified via RepeatModeler (v2.0.1)26 and masked by RepeatMasker (v4.1.1)27 (parameters: -e rmblast -pa 30 -xsmall -nolow -norna -gff -a). The nuclear genome was annotated by Braker2 pipeline11 (parameters: –softmasking –gff3 –fungus –gth2traingenes –prg = gth), combining three aspects evidences: ab initio prediction, homologous proteins, and RNA-Seq evidences. The AUGUSTUS v3.4.028 and Genemark-EP+29 was used as ab initio prediction tools in the pipeline. All proteins of the genus Pyricularia in the Uniref100 database30 were collected and the 100% non-redundant protein dataset was built by cd-hit31 (parameters: -c 1.00 -aS 1.00 -aL 1.00 -n 5 -M 20000), which was used as the protein-based training evidence. The GenomeThreader v1.7.332 was used as the alignment tool. RNA-Seq data previous used20 (i.e. SRR1517063833, SRR1517063734 and SRR1517063635) were aligned by HISAT2 (v2.2.1)21 (parameters: -t -dta). The mitochondrial genome was annotated by MFannot12 with genetic code 4.
Data Recodes
The raw genomic sequencing data used and/or analyzed during the current study are available at NCBI Sequence Read Archive database (Accession number SRR2489091036, SRR2489091137 and SRR2489091238). The assembled genome was deposited under the same BioProject with P. oryzae strain P131 at NCBI (Accession number: GCA_000292605.239; BioProject ID: PRJNA82693; BioSample ID: SAMN31867770). The accession numbers from Chr1 to Chr7 chromosome sequences were CP114135 to CP114141, respectively. And the accession number corresponding to the mitochondrial genome sequence was CP114142.
Technical Validation
DNA sample quality
The DNA quality was detected using Qubit (Thermo Fisher Scientific, Waltham, MA) and Nanodrop (Thermo Fisher Scientific, Waltham, MA).
Sequencing data assessment
The short read data were assessed by fastp v0.2340. The genomic short sequencing reads had 49.75% GC content. The Q20 and Q30 percentages were 97.1% and 92.06%, respectively. The Hi-C sequencing data had 50.5% GC content, and had quality scores of 97.67% (Q20) and 93.64% (Q30), respectively.
Evaluation of the genome assembly
The genome assembly quality was evaluated through the Benchmarking Universal Single-Copy Orthologs (BUSCO) tool with the “fungi_odb10” lineage as a reference dataset. The results showed that 99.9% of all 758 BUSCO markers were assembled, implying a high level of completeness of the assembly. In addition, the results generated from “ascomycota_odb10” lineage showed 99.4% of all 1706 BUSCO markers were include (Table 2).
Code availability
The published softwares used in this work were cited in the Methods section. If no detailed parameters were mentioned for the software, default parameters were applied.
References
Valent, B. & Chumley, F. G. Molecular genetic analysis of the rice blast fungus, magnaporthe grisea. Annu Rev Phytopathol 29, 443–67 (1991).
Talbot, N. J. On the trail of a cereal killer: Exploring the biology of Magnaporthe grisea. Annu Rev Microbiol 57, 177–202 (2003).
Ebbole, D. J. Magnaporthe as a model for understanding host-pathogen interactions. Annu Rev Phytopathol 45, 437–56 (2007).
Dean, R. A. et al. The genome sequence of the rice blast fungus Magnaporthe grisea. Nature 434, 980–6 (2005).
Xue, M. et al. Comparative analysis of the genomes of two field isolates of the rice blast fungus Magnaporthe oryzae. PLoS Genet 8, e1002869 (2012).
Zhang, H., Zheng, X. & Zhang, Z. The Magnaporthe grisea species complex and plant pathogenesis. Mol Plant Pathol 17, 796–804 (2016).
Bao, J. et al. PacBio Sequencing Reveals Transposable Elements as a Key Contributor to Genomic Plasticity and Virulence Variation in Magnaporthe oryzae. Mol Plant 10, 1465–1468 (2017).
Wang, Y. et al. Genome Sequence of Magnaporthe oryzae EA18 Virulent to Multiple Widely Used Rice Varieties. Molecular Plant-Microbe Interactions 35, 727–730 (2022).
Ji, X. et al. Mitochondrial characteristics of the powdery mildew genus Erysiphe revealed an extraordinary evolution in protein-coding genes. Int J Biol Macromol 230, 123153 (2023).
Xue, M. et al. Genome assembly MoP131_2.0. GenBank https://identifiers.org/ncbi/GCA_000292605.1 (2013).
Brůna, T. et al. BRAKER2: automatic eukaryotic genome annotation with GeneMark-EP+ and AUGUSTUS supported by a protein database. NAR Genom Bioinform 3, lqaa108 (2021).
Valach, M. et al. Widespread occurrence of organelle genome-encoded 5S rRNAs including permuted molecules. Nucleic Acids Res 42, 13764–77 (2014).
Farman, M. L. Telomeres in the rice blast fungus Magnaporthe oryzae: the world of the end as we know it. FEMS Microbiol Lett 273, 125–32 (2007).
Peng, Z. et al. Effector gene reshuffling involves dispensable mini-chromosomes in the wheat blast fungus. PloS Genet 15, e1008272 (2019).
Rehmeyer, C. et al. Organization of chromosome ends in the rice blast fungus, Magnaporthe oryzae. Nucleic Acids Res 34, 4685–701 (2006).
Peng, Y.-L. & Shishiyama, J. Temporal sequence of cytological events in rice leaves infected with Pyricularia oryzae. Botany 66, 730–735 (1988).
Liu, X. et al. Prp19-associated splicing factor Cwf15 regulates fungal virulence and development in the rice blast fungus. Environ Microbiol. 10, 5901–5916 (2021).
Durand, N. C. et al. Juicer Provides a One-Click System for Analyzing Loop-Resolution Hi-C Experiments. Cell Syst 3, 95–8 (2016).
Durand, N. C. et al. Juicebox Provides a Visualization System for Hi-C Contact Maps with Unlimited Zoom. Cell Syst 3, 99–101 (2016).
Li, Z. et al. Transcriptional Landscapes of Long Non-coding RNAs and Alternative Splicing in Pyricularia oryzae Revealed by RNA-Seq. Front Plant Sci 12, 723636 (2021).
Kim, D. et al. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol 37, 907–915 (2019).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–9 (2009).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res 27, 722–736 (2017).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PloS One 9, e112963 (2014).
Xu, G. C. et al. LR_Gapcloser: a tiling path-based gap closer that uses long reads to complete genome assembly. Gigascience 8, giy157 (2019).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc Natl Acad Sci USA 117, 9451–9457 (2020).
Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr Protoc Bioinformatics 4, Unit 4.10 (2004).
Keller, O. et al. A novel hybrid gene prediction method employing protein multiple sequence alignments. Bioinformatics 27, 757–63 (2011).
Brůna, T., Lomsadze, A. & Borodovsky, M. GeneMark-EP+: eukaryotic gene prediction with self-training in the space of genes and proteins. NAR Genom Bioinform 2, lqaa026 (2020).
Suzek, B. E. et al. UniRef clusters: a comprehensive and scalable alternative for improving sequence similarity searches. Bioinformatics 31, 926–32 (2015).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–9 (2006).
Gremme, G. et al. Engineering a software tool for gene structure prediction in higher organisms. Information and Software Technology 47, 965–978 (2005).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR15170638 (2021).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR15170637 (2021).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR15170636 (2021).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR24890910 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR24890911 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR24890912 (2023).
Li, Z. et al. Genome assembly PoP131. GenBank https://identifiers.org/ncbi/insdc.gca:GCA_000292605.2 (2023).
Chen, S. et al. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 34, 884–890 (2018).
Acknowledgements
This study was funded by China Agricultural Research System (Grant No. CARS-01-43). The funders had no roles in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Zhigang Li, Jun Yang, Xiaobei Ji, Jintao Liu, and Changfa Yin performed the experiments. All authors analyzed the data. You-Liang Peng, Zhigang Li, and Jun Yang designed the study. You-Liang Peng, Zhigang Li, Jun Yang, and Vijai Bhadauria wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, Z., Yang, J., Ji, X. et al. First telomere-to-telomere gapless assembly of the rice blast fungus Pyricularia oryzae. Sci Data 11, 380 (2024). https://doi.org/10.1038/s41597-024-03209-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03209-z