Abstract
Prunus campanulata is an important flowering cherry germplasm of high ornamental value. Given its early-flowering phenotypes, P. campanulata could be used for molecular breeding of ornamental species and fruit crops belonging to the subgenus Cerasus. Here, we report a chromosome-scale assembly of P. campanulata with a genome size of 282.6 Mb and a contig N50 length of 12.04 Mb. The genome contained 24,861 protein-coding genes, of which 24,749 genes (99.5%) were functionally annotated, and 148.20 Mb (52.4%) of the assembled sequences are repetitive sequences. A combination of genomic and population genomic analyses revealed a number of genes under positive selection or accelerated molecular evolution in P. campanulata. Our study provides a reliable genome resource, and lays a solid foundation for genetic improvement of flowering cherry germplasm.
Similar content being viewed by others
Background & Summary
The genus Prunus (family Rosaceae) contains many economically important plant species, such as peach, plum, apricot, almond, and cherry, grown for food and landscaping purposes. The subgenus Cerasus is classified within the genus Prunus with a corymbose inflorescence, comprises approximately 57 species of flowering trees or shrubs1,2. Cerasus has a worldwide distribution, with most species occurring mainly in the temperate zone of the northern hemisphere3. The subgenus Cerasus is believed to have originated in East Asia and then spread to West Asia2. A number of species in the subgenus Cerasus are economically and commercially important fruit crops, such as sweet cherry (Prunus avium), sour cherry (Prunus cerasus), and Chinese cherry (Prunus pseudocerasus), whose fruit can be either consumed raw or used for the production of jam or liquor4. Many Cerasus are of high ornamental value, owing to their graceful tree shape and attractive flowers, and are thus used for commercial and residential landscaping purposes.
Flowering cherries have been cultivated for over 1,000 years5. Centuries of propagation and cultivation of flowering cherries have produced a variety of natural and artificial hybrids, most of which are derived from crosses among 10 diploid species, including P. apetala, P. campanulata, P. incisa, P. jamasakura, P. leveilleana, P. maximowiczii, P. nipponica, P. sargentii, P. spachiana and P. speciosa6,7. Although most wild flowering cherries are distributed in China, modern flowering cherry cultivars are mainly derived from native Japanese taxa and their hybrids. Only two wild species native to China, P. campanulata and P. pseudocerasus, are believed to have contributed to modern cherry cultivars5,8,9.
Prunus campanulata (2n = 2x = 16), one of the main parents of flowering cherry cultivars, is considered as one of the four major ornamental cherry species, together with P. yedoensis, P. subhirtella var. pendula, and P. cerasoides10. Prunus campanulata is a typical early-flowering species, which usually blooms from January to March, and has a long flowering period (ca. 50 days). Thus, this species flowers much earlier and longer than P. yedoensis (April, 15–20 days) and P. serrulata (April–May, 11–14 days)1. Its attractive pink to magenta flowers and earlier blooming period make P. campanulata a popular choice for landscaping11. Unlike most Cerasus species, P. campanulata grows primarily in subtropical and tropical regions, showing adaptation to warmer climates. Therefore, P. campanulata possesses some desirable traits, such as early and prolonged flowering, anti-pollution effect, and heat tolerance, which could be used for breeding flowering cherry cultivars12,13. However, the lack of genome sequence information hinders our understanding of the mechanisms underlying heat tolerance and early flowering in P. campanulata.
Here, we report a chromosome-level genome assembly of P. campanulata. PacBio HiFi reads (~97 × coverage) were used to assemble the genome yielding a contig assembly of ~282.6 Mb, with contig N50 value of 12.04 Mb (Table 1). The assembled contig size was close to the estimated genome size of 282.8 Mb based on k-mer estimates (Fig. 1a). With the aid of Hi-C sequencing (~176 × coverage) technologies, 92.3% of the contigs were anchored and oriented onto eight pseudomolecules, with a scaffold N50 length of 30.65 Mb (Fig. 1b, Table 1). We traced the evolutionary dynamics of genomes and gene families for P. campanulata. Applying comparative and evolutionary genomics approaches, we identified a number of genes that underwent positive selection or accelerated molecular evolution in P. campanulata. Among them, five candidate genes (VIL1, PUB14, FD, DDL and SR45A) have previously been demonstrated to be involved in the regulation of flowering time in other species, suggesting their potential association with the early-flowering traits of P. campanulata. Our results provide genetic resources for the genetic improvement and optimization of ornamentally and agriculturally important Cerasus species.
Methods
Library construction and genome sequencing
For whole-genome sequencing, fresh young leaves were collected from a mature plant of P. campanulata grown at South China Agricultural University (Guangzhou, China) (23.1557° N, 113.3537° E). Genomic DNA was extracted from leaf tissue using a modified CTAB method14. Short-read sequencing libraries with an insert size of 350 bp were constructed and used for paired-end (PE) 150 bp sequencing on the Illumina NovaSeq 6000 platform. Reads with adapters, with > 10% unidentified nucleotides (N), and paired reads with more than 20% of base quality ≤ 5 in either paired read were filtered out. A total of 27.13 Gb of clean data was produced and used for the genome survey. For PacBio SMRT sequencing, the PacBio Sequel II platform was first used to generate sub-reads, and the sub-reads were then filtered by the ccs software using the parameter “min-passes = 3, min-rq = 0.99” to obtain 27.50 Gb of HiFi reads. A Hi-C library was constructed by chromatin crosslinking, restriction enzyme digestion (DpnII), end filling and biotin labeling, DNA purification and shearing, and extraction of biotin-containing fragments after sonication interruption. The Hi-C sequencing library was sequenced on Illumina PE150. The resulting sequencing data were filtered using the same filtering criteria as the short reads, retaining 49.75 Gb of clean data.
Five tissues including leaves, branches, flowers, fruits and roots were collected from the same P. campanulata tree for transcriptome sequencing. RNA-seq libraries were prepared and then subjected to PE150 sequencing on the Illumina NovaSeq 6000 platform.
Genome size estimation, genome assembly and quality assessments
To estimate the genome size, heterozygosity and repeat content of P. campanulata, we performed k-mer frequency analysis based on the 17 k-mers depth distribution with GCE15 using Illumina short reads. Based on the k-mer analysis, the size of the P. campanulata genome was estimated to be ~282.8 Mb, with heterozygosity of 0.61% and repeat content of 47.7%.
The PacBio HiFi reads were assembled into the initial set of contigs using hifiasm v0.816 with default parameters. The contig assembly had a total size of ~282.6 Mb, with a contig N50 value of 12.04 Mb. Genome completeness was assessed using the Benchmarking Universal Single-Copy Orthologs (BUSCO v4.1.2)17 program with the embryophyta_odb10 database and the Core Eukaryotic Genes Mapping Approach (CEGMA v2.5)18, which yielded 99.1% of the complete BUSCO genes and 95.6% of the core eukaryotic genes (Table 1). In addition, the filtered short reads were mapped against the assembled genome using the BWA-MEM v0.7.819 algorithm to assess the accuracy of the assembly, and the mapping rate and coverage of the Illumina short reads were 99.02% and 99.92%, respectively. To achieve chromosome-level assembly, the ALLHiC algorithm20 was used to group, adjust the order and orientation of contigs and anchor the assembled contigs into eight pseudomolecules based on Hi-C data. After ALLHiC scaffolding, Hi-C interaction heat map was constructed using HiC-Pro v3.1.021 and visualized using HiCPlotter22. Finally, a total of eight pseudomolecules were obtained, which contained 92.3% of the contigs. Telomere sequences (CCCTAAA/TTTAGGG repeats) were identified by searching the chromosome-level assembly using Telomere Identification toolKit (tidk, https://github.com/tolkit/telomeric-identifier). These repeat arrays were identified at both distal ends of pseudomolecules 2, 3, and 8, and at one distal end of pseudomolecules 1, 4, 5, 6, and 7 (Fig. 2). To evaluate the assembly continuity, the long terminal repeat (LTR) assembly index (LAI) value was employed using LTR_retriever23 by estimating the percentage of intact LTR elements. The LAI value of the genome assembly was 19.3, which almost reached the “gold standard” (LAI value > 20) of genome assembly proposed by Ou et al.24. Collectively, these results indicate a high quality of the P. campanulata genome assembly, thus ensuring the reliability of our subsequent analyses.
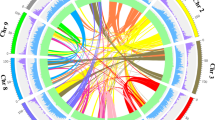
The genomic features of P. campanulata. Tracks from the outermost to the innermost circle separately represent the assembled eight pseudochromosomes (I), guanine-cytosine (GC) content (II), gene density (III), tandem duplication (IV), transposable element density (V), and long-terminal repeat density (VI). Black squares at the end of each chromosome in (I) represent telomeres. Photo credit: Ganbiao Xian.
Genome annotations
For repeat sequence annotation, we used a combined strategy of homology-based search and de novo prediction. The homology-based search was based on the Repbase database using RepeatMasker (http://www.repeatmasker.org/) and RepeatProteinMask (http://www.repeatmasker.org/) to search for interspersed repeat elements. The de novo prediction was based on a species-specific repeat database generated by LTR_FINDER25, Piler (http://www.drive5.com/piler/), RepeatScout (http://www.repeatmasker.org/) and RepeatModeler (http://www.repeatmasker.org/RepeatModeler.html). Using this library, we identified de novo involved repeats with RepeatMasker and predicted tandem repeats with TRF (http://tandem.bu.edu/trf/trf.html). Repetitive sequence annotation detected approximately 148.2 Mb of repetitive sequences in the P. campanulata genome, accounting for 52.4% of the total genome size (Table 2). In addition, we used RepeatMasker to mask the repetitive sequences as input for gene structure prediction.
For protein-coding gene structure prediction, we used a comprehensive approach that integrates homology-based prediction, de novo prediction, and RNA-Seq-based prediction. For homology-based prediction, Blast26 and Genewise27 were used to align the amino acid sequences from the Malus domestica, P. serrulata, Vitis vinifera, A. thaliana, and P. salicina genomes to the assembled P. campanulata sequences. Augustus28, GlimmerHMM29, SNAP30, GeneID31, and GenScan32 were used to predict de novo gene models. Cufflinks33 and PASA34 were applied to predict the gene models in the RNA-Seq-based prediction study. The results of the above three approaches were further integrated to generate a final non-redundant gene model set using EVidenceModeler35 and modified using PASA. We predicted 24,861 protein-coding gene models from the repeat-masked P. campanulata genome, of which 24,749 were functionally annotated against public databases (Table 3, Table 4). The distribution of genes and transposable elements (TEs) along each chromosome revealed that genes are more densely packed at the distal ends of chromosomes, whereas TEs are clustered around the centromeric regions of chromosomes (Fig. 2), thus following the typical distribution of monocentric plant genomes36. To annotate the putative function of the genes, we used BLASTP to perform homologous alignments against several public databases, including SwissProt, TrEMBL, the non-redundant protein database of NCBI (NR), GO, and KEGG37. In addition, we used InterProScan to predict gene motifs and domains, and obtained gene ontologies from InterPro38. For the annotation of non-coding RNAs, tRNAscan-SE (http://lowelab.ucsc.edu/tRNAscan-SE/) was employed to identify transfer RNAs (tRNAs). Ribosomal RNAs (rRNAs) were identified using BLASTN with an E-value of 1e-5 against rRNA sequences from related species. MicroRNAs (miRNAs) and small nuclear RNAs (snRNAs) were predicted using INFERNAL (http://infernal.janelia.org/) and searched against the Rfam database.
Comparative genomic analysis across Cerasus species
We collected the protein sequences of the longest transcripts from P. campanulata, the double-flower cultivar P. campanulata ‘Plena’39 and four other species of Cerasus, including P. avium13, P. serrulata40, P. yedoensis41, and Cerasus × yedoensis7. Cerasus × yedoensis was proposed to be an interspecific hybrid of P. spachiana and P. speciosa, and Shirasawa et al.7 assembled two haplotype phased genome sequences of this interspecific hybrid with SMRT sequencing. In this study, we used these two parental genome sequences to represent the genome assembly of P. spachiana and P. speciosa, respectively. We classify orhologues and orthogroups from the above six species using OrthoFinder v2.5.442 with the parameter ‘-S diamond’. Based on the orthogroup clustering of six Cerasus species, we identified a total of 31,421 orthogroups, of which 9,773 were core orthogroups shared by all seven species, and 2,791 were single-copy orthogroups with only one gene per species (Fig. 3a). Prunus campanulata had 54 species-specific orthogroups, containing a total of 977 genes. Gene Ontology (GO) enrichment analysis performed using the R package topGO v2.38.143 revealed that these genes were mainly involved in signaling processes such as regulation of transport, cell communication, and photoperiodism.
The evolutinary analysis of Cerasus genomes. (a) Flower plot showing the number of core orthogroups (center), orthogroups in a subset of species (annulus), and species-specific orthogroups (petals) for Cerasus species. (b) Estimation of the phylogeny, divergence times, and gene families of seven Prunus species. (c) Collinearity patterns among Cerasus species assembled to chromosomes.
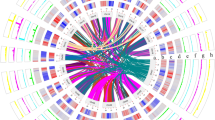
To construct the phylogenetic tree of P. campanulata and representative Cerasus species, we first performed orthogroup classification using the above-mentioned seven Cerasus species and P. persica44, and obtained 2,685 single-copy genes. Then, we used the parallel Alignment and back-Translation (paraAT) tool45 to align the protein sequences of 2,685 single-copy orthologous genes in parallel, and back-translated the aligned sequences into the corresponding aligned coding sequences (CDS). We constructed the maximum-likelihood (ML) phylogenetic tree for each gene using IQ-TREE v1.6.1246 with the parameter “-bb 1000”, setting P. persica as the outgroup. We then inferred the final species tree by summarizing a set of gene trees using ASTRAL-II v5.7.747. We conducted the divergence time estimation using Bayesian Evolutionary Analysis Sampling Trees (BEAST) v2.6.648, with two reported divergence times set as secondary calibrations: the ancestral node of P. persica and the subgenus Cerasus (mean: 58.2 MYA, Std dev: 4.3 MYA); the divergence of P. avium and P. serrulata (mean: 48.8 MYA, Std dev: 6.0 MYA)2. Markov chain Monte Carlo was run for 100,000,000 generations with 1,000 steps. Based on the ultrametric species tree and the results of the gene family clustering analysis, we used CAFE v4.2.149 to identify the patterns of gene family evolution. We filtered out gene families with abnormal gene copy numbers, that is, those gene families containing more than 100 gene copy numbers in one or multiple species. The orthologous gene family expansions or contractions at each branch were considered significant at p < 0.01, and 44 gene families underwent significant expansion (p < 0.01), while 683 gene families underwent contraction during the evolution of P. campanulata (Fig. 3b). GO enrichment analysis suggested that genes in the expanded gene families were significantly enriched in oxidation-reduction processes involved in cellular respiration, such as oxidative phosphorylation and respiratory electron transport chain. Furthermore, we used MCscan (https://github.com/tanghaibao/jcvi/wiki/MCscan-(Python version) to measure collinearity between the six chromosome-level genomes of Cerasus species, which showed an overall high degree of collinearity, indicating that species within the subgenus Cerasus exhibit relatively conserved genomic synteny (Fig. 3c).
Identification of positively selected genes and rapidly evolving genes
To search for the genomic footprint of natural selection that may be involved in flowering time regulation, we performed positive selection analysis and rapidly evolving gene analysis using 2,893 single-copy orthologous genes from P. campanulata and five late-flowering Cerasus species. For the positive selection analysis, we used the branch-site model of the codeml program in the PAML v4.9 package50, which allows for differential selective pressure both among amino acid sites and between branches on the tree. We then performed a likelihood ratio test (LRT) to compare the likelihood differences between the two models used, modelA, which allows sites to be positively selected on the foreground branch (P. campanulata), and the null model, in which sites could have evolved neutrally and/or under purifying selection. Genes with a Chi-squared test p value < 0.05 were considered to be positively selected in P. campanulata.
To identify rapidly evolving genes, we used the branch model incorporated in the PAML v4.9 package, which assumes that genes evolve at an accelerated rate on the foreground branch. Similarly, we compared the null model (model = 0) and the alternative model (model = 2) using LRT, and genes with p value < 0.05 and a higher ω value for the foreground branch than the background branch were considered to be evolving significantly faster in P. campanulata.
Data Records
The raw data of Illumina PE150 reads, PacBio HiFi reads and Hi-C reads described in this manuscript have been submitted to the National Center for Biotechnology Information (NCBI) with accession number SRR2207152051, SRR2644689952, SRR2501970853 under BioProject accession number PRJNA895162. The RNA-seq data for different tissues are also under PRJNA895162. The genome assembly has been deposited at GenBank under the accession JAXCME00000000054. The genome assembly and annotation of P. campanulata have been also submitted to the Genome Database for Rosaceae (GDR; www.rosaceae.org) under the accession number tfGDR107455. The protein sequences used in the comparative genomic analyses and the output files including orthogroup clustering analysis and PAML analysis are available at Figshare with the https://doi.org/10.6084/m9.figshare.2369416856.
Technical Validation
Evaluating the quality of the genome assembly
We evaluated the quality of our assembly using three methods: (1) Performing BUSCO and CEGMA analyses. The BUSCO analysis revealed 99.10% complete BUSCO genes, of which 96.30% were single-copy genes. In addition, 0.30% of BUSCO genes were fragmented, and 0.60% were missing from the genome. The CEGMA analysis detected 95.6% of the eukaryotic core genes in the assembly; (2) Calculation of the paired-end read coverage and mapping rate by mapping the short Illumina reads to our genome assembly, which showed 99.02% genome coverage and 99.92% mapping rate; (3) Calculation of the LAI value. The LAI value was 19.3. These results confirmed the high quality of our assembly.
Evolution of genes in P. campanulata
Of the 2,893 single-copy orthologs, 248 genes were considered to be positively selected in P. campanulata with the chi-squared test p value < 0.05. Functional enrichment analysis of the positively selected genes (PSGs) revealed that they showed enrichment in some potentially resistance-related GO terms, such as potassium ion transport, DNA repair, cellular response to DNA damage stimulus, and cellular response to stress (Fig. 4a). Among the 248 PSGs, four genes (VIL1, FD, PUB14 and DDL) were previously shown to be associated with flowering time regulation in other species (Fig. 4b–e)57,58,59,60.
Evolution of PSGs in P. campanulata. (a) GO enrichment analysis of positively selected genes in the P. campanulata genome. (b–e) Alignment of deduced amino acid sequences of four positively selected genes (PSGs), VIL1, PUB14, FD and DDL. Sites marked in red were potentially under positive selection in P. campanulata according to the empirical Bayesian approach. Dots represent sequence identity with the P. spachiana genome.
We identified a total of 94 rapidly evolving genes (REGs), that showed a higher ω value in P. campanulata than in the background. GO enrichment analysis of these REGs in P. campanulata revealed that they were significantly enriched in monovalent inorganic cation transport (GO:0015672) and potassium ion transport (GO:0006813). One of the REGs, SR45A, encoding the serine/arginine-rich (SR) splicing factor SR45a, was shown to be associated with the regulation of flowering date in sweet cherry61. Our evolutionary analysis of these genes in P. campanulata provides new evidence for loci associated with flowering time regulation and provides prime candidates for future breeding of crops such as sweet cherry.
Code availability
All software used in this study was run according to the official instructions. The version and parameters of the software and the other custom codes used were described in Methods. Anything not specified in Methods was run with default parameters.
References
Li, C. L. & Bartholomew, B. in Flora of China: Pittosporaceae through Connaraceae. (ed. Wu, C.Y., Raven, P.H. and Hong, D.Y.) Cerasus (Beijing, China: Science Press & St. Louis USA: Missouri Botanical Garden, 2003).
Chin, S. W., Shaw, J., Haberle, R., Wen, J. & Potter, D. Diversification of almonds, peaches, plums and cherries - Molecular systematics and biogeographic history of Prunus (Rosaceae). Mol. Phylogenet. Evol. 76, 34–48 (2014).
Rehder, A. Manual of cultivated trees and shrubs hardy in north America exclusive of the subtropical and warmer temperate regions 2nd edn (MacMillan, New York, 1940).
Khadivi-Khub, A., Zamani, Z. & Fatahi, M. R. Multivariate analysis of Prunus subgen. Cerasus germplasm in Iran using morphological variables. Genet. Resour. Crop Evol. 59, 909–926 (2011).
Kato, S. et al. Origins of Japanese flowering cherry (Prunus subgenus Cerasus) cultivars revealed using nuclear SSR markers. Tree Genet. Genomes 10, 477–487 (2014).
Ma, H., Olsen, R. & Pooler, M. Evaluation of flowering cherry species, hybrids, and cultivars using simple sequence repeat markers. J. Am. Soc. Hortic. Sci. 134, 435–444 (2009).
Shirasawa, K. et al. Phased genome sequence of an interspecific hybrid flowering cherry, ‘Somei-Yoshino’ (Cerasus × yedoensis). DNA Res. 26, 379–389 (2019).
Kawasaki, T. The distribution of Prunus subgenus Cerasus in East-Asia and classification of Japanese wild species. Sakura Sci. 1, 28–45 (1991).
Kuitert, W. & Peterse, A. Japanese Flowering Cherries. (Timber Press, Portland Oregon, 1999).
Lu, Y., Chen, Z. & Shi, J. Research advance, prospect and breeding strategy of Cerasus campanulata Maxim. Journal of Nanjing Forestry University (Natural Sciences Edition) 30, 115–119 (2006).
Huang, K.-F., Wen, C.-H., Wang, C.-T. & Chu, F.-H. Transcriptome and flower genes analysis of Prunus campanulata Maxim. J. Hortic. Sci. Biotech. 95, 44–52 (2019).
Weng, Y. et al. The chloroplast genome of Cerasus campanulata diverges from other Prunoideae genomes. Phyton 89, 375–384 (2020).
Wang, J. et al. Chromosome-scale genome assembly of sweet cherry (Prunus avium L.) cv. Tieton obtained using long-read and Hi-C sequencing. Hortic. Res. 7, 122 (2020).
Doyle, J. J. T. & Doyle, J. L. Isolation of plant DNA from fresh tissue. Focus 12 (1990).
Liu, B. et al. Estimation of genomic characteristics by analyzing k-mer frequency in de novo genome projects. arXiv.org, arXiv: 1308.2012 (2013).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Zhang, X., Zhang, S., Zhao, Q., Ming, R. & Tang, H. Assembly of allele-aware, chromosomal-scale autopolyploid genomes based on Hi-C data. Nat. Plants 5, 833–845 (2019).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Akdemir, K. C. & Chin, L. HiCPlotter integrates genomic data with interaction matrices. Genome Biol. 16, 198 (2015).
Ou, S. & Jiang, N. LTR_retriever: a highly accurate and sensitive program for identification of long terminal repeat retrotransposons. Plant Physiol. 176, 1410–1422 (2018).
Ou, S., Chen, J. & Jiang, N. Assessing genome assembly quality using the LTR Assembly Index (LAI). Nucleic Acids Res. 46, e126 (2018).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 35, W265–W268 (2007).
Kent, W. J. BLAT-the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Genome Res. 14, 988–995 (2004).
Stanke, M., Schöffmann, O., Morgenstern, B. & Waack, S. Gene prediction in eukaryotes with a generalized hidden Markov model that uses hints from external sources. BMC Bioinform. 7, 62 (2006).
Majoros, W. H., Pertea, M. & Salzberg, S. L. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics 20, 2878–2879 (2004).
Korf, I. Gene finding in novel genomes. BMC Bioinform. 5, 59 (2004).
Guigó, R., Knudsen, S., Drake, N. & Smith, T. Prediction of gene structure. J. Mol. Biol. 226, 141–157 (1992).
Burge, C. & Karlin, S. Prediction of complete gene structures in human genomic DNA. J. Mol. Biol. 268, 78–94 (1997).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res. 31, 5654–5666 (2003).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the program to assemble spliced alignments. Genome Biol. 9, R7 (2008).
Sun, H. et al. Chromosome-scale and haplotype-resolved genome assembly of a tetraploid potato cultivar. Nat. Genet. 54, 342–348 (2022).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–462 (2016).
Quevillon, E. et al. InterProScan: protein domains identifier. Nucleic Acids Res. 33, W116–120 (2005).
Nie, C. et al. Genome assembly, resequencing and genome-wide association analyses provide novel insights into the origin, evolution and flower colour variations of flowering cherry. Plant J. 114, 519–533 (2023).
Yi, X. G. et al. The genome of Chinese flowering cherry (Cerasus serrulata) provides new insights into Cerasus species. Hortic. Res. 7, 165 (2020).
Baek, S. et al. Draft genome sequence of wild Prunus yedoensis reveals massive inter-specific hybridization between sympatric flowering cherries. Genome Biol. 19, 127 (2018).
Emms, D. M. & Kelly, S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 16, 157 (2015).
Alexa, A. & Rahnenfuhrer, J. Gene set enrichment analysis with topGO. Bioconductor Improvement 27, 1–26 (2019).
Verde, I. et al. The Peach v2.0 release: high-resolution linkage mapping and deep resequencing improve chromosome-scale assembly and contiguity. BMC Genomics 18, 225 (2017).
Zhang, Z. et al. ParaAT: a parallel tool for constructing multiple protein-coding DNA alignments. Biochem. Biophys. Res. Commun. 419, 779–781 (2012).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Mirarab, S. & Warnow, T. ASTRAL-II: coalescent-based species tree estimation with many hundreds of taxa and thousands of genes. Bioinformatics 31, i44–52 (2015).
Bouckaert, R. et al. BEAST 2.5: An advanced software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 15, e1006650 (2019).
De Bie, T., Cristianini, N., Demuth, J. P. & Hahn, M. W. CAFE: a computational tool for the study of gene family evolution. Bioinformatics 22, 1269–1271 (2006).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR22071520 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR26446899 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR25019708 (2023).
NCBI GenBank https://identifiers.org/ncbi/insdc:JAXCME000000000 (2023).
Genome Database for Rosaceae https://www.rosaceae.org/node/10813072 (2023).
Hu, Y. X. The comparative genomic analyses output files of Cerasus. figshare https://doi.org/10.6084/m9.figshare.23694168 (2023).
Sung, S., Schmitz, R. J. & Amasino, R. M. A PHD finger protein involved in both the vernalization and photoperiod pathways in. Arabidopsis. Genes Dev. 20, 3244-–8 (2006).
Romera-Branchat, M. et al. Functional divergence of the Arabidopsis florigen-interacting bZIP transcription factors FD and FDP. Cell Rep. 31, 107717 (2020).
Feke, A. M., Hong, J., Liu, W. & Gendron, J. M. A decoy library uncovers U-Box E3 ubiquitin ligases that regulate flowering time in. Arabidopsis. Genetics 215, 699–712 (2020).
Morris, E. R., Chevalier, D. & Walker, J. C. DAWDLE, a forkhead-associated domain gene, regulates multiple aspects of plant development. Plant Physiol. 141, 932-–41 (2006).
Branchereau, C. et al. New insights into flowering date in Prunus: fine mapping of a major QTL in sweet cherry. Hortic. Res. 9, uhac042 (2022).
Acknowledgements
This work was supported by Key-Area Research and Development Program of Guangdong Province (Grant No. 2022B1111230001) and Youth Innovation Promotion Association CAS (2021348).
Author information
Authors and Affiliations
Contributions
M.K. conceived the project and designed the study. Y.H. and B.W. performed the sampling and experiments. Y.H. and C.F. performed the data analysis and generated figures and tables. Y.H. and M.K. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hu, Y., Feng, C., Wu, B. et al. A chromosome-scale assembly of the early-flowering Prunus campanulata and comparative genomics of cherries. Sci Data 10, 920 (2023). https://doi.org/10.1038/s41597-023-02843-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02843-3