Abstract
The assembly of W and Y chromosomes poses significant challenges in vertebrate genome sequencing and assembly. Here, we successfully assembled the W chromosome of Verasper variegatus with a length of 20.48 Mb by combining population and PacBio HiFi sequencing data. It was identified as a young sex chromosome and showed signs of expansion in repetitive sequences. The major component of the expansion was Ty3/Gypsy. The ancestral Osteichthyes karyotype consists of 24 protochromosomes. The sex chromosomes in four Pleuronectiformes species derived from a pair of homologous protochromosomes resulting from a whole-genome duplication event in teleost fish, yet with different sex-determination systems. V. variegatus and Cynoglossus semilaevis adhere to the ZZ/ZW system, while Hippoglossus stenolepis and H. hippoglossus follow the XX/XY system. Interestingly, V. variegatus and H. hippoglossus derived from one protochromosome, while C. semilaevis and H. stenolepis derived from another protochromosome. Our study provides valuable insights into the evolution of sex chromosomes in flatfish and sheds light on the important role of whole-genome duplication in shaping the evolution of sex chromosomes.
Similar content being viewed by others
Background & Summary
Although the assembly of W and Y chromosomes presents one of the most daunting challenges in sequencing and assembling vertebrate genomes, it remains a crucial area of research1. The analysis of W and Y chromosomes can provide valuable insights into the specific evolutionary trajectory for females and males2. Sex chromosomes evolved from a pair of autosomes3,4, and with the accumulation of antagonistic sites around the sex determining gene, a recombination inhibition region was formed and gradually expanded, eventually leading to the difference between X and Y (or Z and W) chromosomes5,6. Although stable sex-determination systems controlling male and female differentiation were formed in a variety of vertebrate taxa, such as mammals and birds7,8,9, sex chromosomes have evolved independently many times throughout the tree of life, resulting in the existence of a diversity of sex chromosomes, especially in fish and amphibians10,11,12. Bony fish are the most species-rich group of vertebrates, containing nearly 30,000 species, accounting for approximately 98% of ray-finned fish and 50% of vertebrate species13. However, less than 10% of fish sex chromosomes are heteromorphic in karyotype, and most of them are young sex chromosomes, that is, in the early stage of differentiation14. Studying young sex chromosomes provides an opportunity to study the initiation process of sex chromosome recombination inhibition15. However, the low degree of differentiation and high sequence similarity between young sex chromosomes is bringing some challenges to chromosome phasing and assembly16. The development of sequencing technologies, especially high-precision PacBio HiFi sequencing, has brought opportunities for genome haplotype assembly17.
V. variegatus, H. hippoglossus and H. stenolepis belong to the family Pleuronectidae, and they have evolved different sex-determination systems. The sex-determination system of the H. stenolepis and V. variegatus is ZZ/ZW18,19, and the sex-determination system of H. hippoglossus is XX/XY20. The sex-determining genes of H. hippoglossus and H. stenolepis are also different. To date, there has been little research on the evolution of sex chromosomes and the transformation mechanism of the sex determination system in Pleuronectidae. Here, by combining population and PacBio HiFi sequencing data, we successfully assembled the W chromosome of V. variegatus, and preliminarily discussed the origin of the sex chromosome of V. variegatus through collinearity analysis, which provided rich resources for follow-up research on the mechanism of sex chromosome evolution in flatfish.
Materials and Methods
V. variegatus samples and sequencing
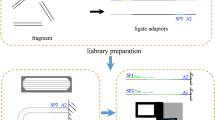
For PacBio CLR and HiFi sequencing, we used a female adult V. variegatus for DNA sequencing. DNA extraction was performed using the SDS-based method, the ground tissue cells were lysed with hot SDS, and high concentrations of KAc were added to remove proteins and polysaccharide impurities by incubating at 0 °C. Finally, precipitation was performed using ethanol or isopropanol. After fragmentation, the BluePippin Size Selection system (Sage Science) was used to screen DNA fragments of approximately 20 kb. Then, PacBio CLR and HiFi libraries were constructed according to the PacBio standard library construction process and sequenced using the PacBio SEQUEL platform. We obtained 53.71 Gb of female V. variegatus PacBio continuous long read (CLR) sequencing data.
For Illumina sequencing, after DNA extraction and fragmentation from 18 female and 17 male fish, 300 bp pair-end Illumina libraries were constructed according to the Illumina standard library construction process and then sequenced using the Illumina NovaSeq. 6000 platform. We obtained 36.38 Gb Illumina sequencing data.
Genome assembly
To obtain high-quality female V. variegatus genome sequences, we used the process shown in Supplementary Fig. 1 to assemble the genome. Canu v1.821 (-correct) was first used to correct raw PacBio data. Then, we used Flye v2.622 (–pacbio-corr) to obtain a draft genome with the corrected PacBio data. The draft genome may have partially redundant sequences, so, we used purge_dups v1.0.023 to remove redundant sequences. Then we performed two rounds of genome polishing using PacBio and resequencing data. The detailed process for polishing the genome was as follows: (1) the PacBio data were aligned to the genome by pbmm2 (SMRT Link v8.0) with default parameters, and then gcpp (SMRT Link v8.0) with default parameters was used to polish the genome. (2) the resequencing data were aligned to the genome by bwa v0.7.1724 with default parameters, and pilon v1.2325 with default parameters was used to polish the genome. After two rounds of genome polishing with software gcpp and pilon. Finally, combining PacBio and Illumina data, we obtained 543.90 Mb female V. variegatus contigs, of which the contig N50 was 17.26 Mb (Supplementary Table 1).
To obtain the chromosome-level genome, we first aligned Hi-C data (SRP263299) to contigs using juicer v1.6.226 with default parameters and then anchored contigs to chromosomes using 3D-DNA27 (-r 0). 99.89% of female sequences were anchored to 23 pseudochromosomes. Both female and male fish have 23 mounted chromosomes, which means no additional W chromosome was assembled in the female genome. Then we used nucmer to compare female and male genomes and found that the genome identity was more than 99.7%. It may be that the Z and W chromosomes may have a low degree of differentiation and the PacBio CLR data with a relatively high error rate cannot be used for the correct phase of Z and W chromosomes, causing the female Z and W chromosomes to mix-assemble into one chromosome.
W chromosome assembly
To obtain the V. variegatus W chromosome, we assembled it by combining PacBio HiFi sequencing data and whole genome resequencing data (Supplementary Fig. 1B). We first used the whole genome resequencing data to obtain female specific kmers according to the following steps: (1) The 17-kmer dataset of each individual was obtained separately using meryl count28. (2) For individual female 17-kmer datasets, kmers with frequency less than 5 were removed using meryl greater-than 5. (3) Individual 17-kmer datasets of 18 female fish were combined into a total datasest using meryl union-sum, considering only the k-mers that were present in all 18 individuals. (4) All kmer data of 18 male fish were combined into a total set using meryl union-sum. (5) Select kmers that exist in all female fish but do not exist in any male fish were identified as specific kmers using meryl difference. After obtaining female-specific kmers, we used meryl-lookup to screen PacBio HiFi sequencing reads containing female-specific kmers. A total of 573,403 female-specific kmers were identified by comparing the resequencing data of female and male fish. Subsequently, by leveraging female-specific kmers, we were able to filter out female-specific data from PacBio HiFi and Hi-C sequencing data, enabling us to conduct a more in-depth analysis of the genetic differences unique to female flatfish. In total, 515.87 Mb female-specific PacBio HiFi data and 189.05 Mb of Hi-C data were screened out. These HiFi data were used to assemble W candidate sequences using Flye with default parameters, a total of 20.48 Mb genome sequence with a contig N50 of 5.16 Mb was obtained. Then we used juicer and 3D-DNA to anchor contigs to chromosomes using the female-specific Hi-C data.
After obtaining the W chromosome, we used the W chromosome information to identify Z chromosomes in the female and male genomes29, respectively. After downloading the male V. variegatus genome from the NCBI GenBank database (GCA_013332515.1), we used MashMap v2.030 (–noSplit) to map the W chromosome to the male genome and found more than 95% sequence identity between chromosome LG21 and the W chromosome. Additionally, we found that the sequence identity of chromosomes 18 and W in the female assembled genome was more than 96%. Therefore, chromosomes LG21 and chromosome 18 were candidate Z chromosomes for the male and female genomes, respectively. Then, we examined the distribution of female-specific kmers in the female and male assembled genomes using meryl-lookup. In male genomes, almost all female-specific kmers were distributed only on the W chromosome (Fig. 1A), while in female genome sequences, female-specific kmers were distributed abundantly on the Z chromosomes in addition to the W chromosome (Fig. 1B). The above results showed that candidate Z chromosomes in the female genome have mixed W chromosome sequences. Then, we used nucmer31 to align the W and Z chromosomes, and dnadiff to show the sequence identity between the W and Z chromosomes was more than 96%. Furthermore, we used bwa to align 18 male and 18 female resequencing data to the genome of male V. variegatus. If the W and Z chromosomes of the V. variegatus genome were fully differentiated, the depth of the Z chromosome in the male sequencing data should be twice that of the female sequencing data (\({\rm{EZ}}{:\log }_{2}\frac{{\rm{M}}}{{\rm{F}}}\approx 1\)), with the decrease in differentiation degree, the coverage of male fish and female fish tended to be the same (\({\log }_{2}\frac{{\rm{M}}}{{\rm{F}}}\approx 0\)). We found that the sequencing depth of all chromosomes did not show significant differences (\({\log }_{2}\frac{{\rm{M}}}{{\rm{F}}}\approx 0\)), that is, there were no significant differences in genome sequences between Z and W chromosomes (Fig. 1C). This indicates that the V. variegatus W and Z chromosomes may be homomorphic sex chromosomes. To further analyze the characteristics of V. variegatus sex chromosomes, we used freebays32 to perform SNP calling on 18 female and 18 male fish found that except for the Z chromosome, male and female fish have a similar number of heterozygous SNPs. The number of heterozygous SNPs on the Z chromosome in females is between 63875–64986, with an average of 64201, while the number of heterozygous SNPs on the Z chromosome in males is between 10585–14598, with an average of 13598 (Fig. 1D), in other words the number of heterozygous SNPs on the Z chromosome was significantly higher in females than in males (Fig. 1D), indicating that Z and W chromosomes have started to differentiate. Previous studies have found that homomorphic sex chromosomes could accumulate SNPs before the sex chromosome decays, and differences in heterozygosity between males and females could be detected even if the sex chromosomes do not show differences in coverage33. Therefore, the V. variegatus W chromosome is a nascent sex chromosome, that could provide valuable resources for studying early sex chromosome differentiation and understanding the initial process of recombination inhibition.
Analysis of basic characteristics of W chromosome. (A) Distribution of female specific Kmer in male genome, chromosome 1 to 23 belonged to male genome, chromosome W belonged to female genome assembled by combing population and PacBio HiFi data. (B) Distribution of female specific Kmer in female genome, chromosome 1 to 23 belonged to female genome assembled by PacBio CLR data, chromosome W was assembled by combing population and PacBio HiFi data. (C) Expected genome depth in females relative to males (log2(M/F)), EZ was the expected genome depth in males relative to females (log2(M/F) ≈ 1), chromosome 1 to 23 belonged to male genome. (D) Distribution of heterozygous SNPs in females and males.
We used EDTA v2.0034 (–sensitive 1–anno 1) to identify repeated sequences. The content of repeated sequences on the W chromosome was 1.75 Mb, which was higher than that on the Z chromosome (1.44 Mb), and the expanded repeated sequences on the W chromosome were mainly Ty3/Gypsy (Supplementary Table 2). Compared with the degenerated and smaller W/Y chromosomes of mammals and birds, the W/Y chromosomes of many fish are nascent sex chromosomes and larger than the Z/Y chromosomes2,35,36. Repeated sequences played an important role in the formation of sex determining regions of nascent W/Y chromosomes and could lead to an increase in chromosome size by accumulating repetitive sequences at the early stage of differentiation37,38,39,40. It has been proven that the transposable element Ty3/Gypsy is closely related to sex specific and sex related regions, which means that Ty3/Gypsy may play a role in the process of sex chromosome differentiation and the formation of new sex determination related regions41,42,43,44.
Then we used MAker245 to predict genes in W chromosome. To predict genes, MAker2 used SNAP46 and Augustus47 for de novo gene prediction and Exonerate48 to align non-redundant proteins from flatfish and V. variegatus RNA-seq data (SRP263299) to the genome for homologous gene prediction. Finally, EVM49 was used to integrate information from de novo and homologous gene predictions to obtain the final gene structure. We annotated the W chromosome gene structure with MAKER2, and 1,033 protein coding genes were annotated.
Finally, we improved the autosomes and Z chromosome of V. variegatus using the female and male genome sequences. Briefly, we used genome puzzle master (GPM)50, an integrated pipeline for building and editing pseudomolecules from fragmented sequences, to integrate the female and male genome sequences and obtained a 543.54 Mb V. variegatus genome sequence, whose contig N50 and scaffold N50 sizes were 22.76 Mb and 24.82 Mb, respectively. Then, Liftoff51 (v1.6.1) with default parameters was used to map the annotations of male genome to the newly assembled genome.
Collinearity analysis of sex chromosomes
Seven teleosts genomes, including Mastacembelus armatus from Synbranchiformes, Gasterosteus aculeatus from Perciformes, Poecilia reticulata from Cyprinodontiformes, and C. semilaevis, V. variegatus, H. hippoglossus and H. stenolepis from Pleuronectiformes, each with specific sex chromosome information, were selected for collinearity analysis using WGDI52. Through genome collinearity analysis, the sex chromosomes (Z and W) of V. variegatus had the best collinearity relationship with the sex chromosome (chromosome 12) of H. hippoglossus, which indicates that they came from the same paleochromosome. From the dotplot, we also found that the sex chromosomes of V. variegatus had an ancient collinearity relationship with the autosome (chromosome 9) of H. hippoglossus (Fig. 2A). The comparative analysis of the collinearity between V. variegatus and H. stenolepis showed that the sex chromosomes (Z and W) of V. variegatus had the best collinear relationship with the autosome (chromosome 18) of H. stenolepis, and had an ancient collinearity relationship with the sex chromosome (chromosome 9) of H. stenolepis (Fig. 2B). Furthermore, we compared the collinearity between V. variegatus and C. semilaevis, and found that their collinear relationship was similar to that of H. stenolepis, and the Z and W chromosomes of V. variegatus had the best collinear relationship with the autosome (chromosome 3) of C. semilaevis, but had an ancient collinear relationship with the sex chromosomes (Z and W) of C. semilaevis (Fig. 2C). Then we further compared the collinearity between C. semilaevis and H. stenolepis and found that the sex chromosomes of C. semilaevis and H. stenolepis had the best collinear relationship, which indicates that they came from the same paleochromosome (Fig. 2D). In addition, by integrating all the dotplot information, we also found that the sex chromosomes of V. variegatus and H. hippoglossus and the sex chromosomes of C. semilaevis and H. stenolepis had an ancient collinear relationship, respectively. V. variegatus, C. semilaevis, H. hippoglossus and H. stenolepis did not experience an additional whole genome duplication event after the common whole genome duplication event of teleost fish53,54. In other words, the paleochromosomes of V. variegatus and H. hippoglossus and the paleochromosomes of C. semilaevis and H. stenolepis came from the same ancient chromosome. It also provides the evidence for the emergence of young sex chromosomes from autosomes and the exceptional evolutionary instability of sex chromosomes in fish55.
Collinearity analysis of four species of flatfish. (A) Collinearity analysis between V. variegatus and H. hippoglossus. (B) Collinearity analysis between V. variegatus and H. stenolepis. (C) Collinearity analysis between V. variegatus and C. semilaevis. (D) Collinearity analysis between C. semilaevis and H. stenolepis.
We applied the workflow (https://github.com/SunPengChuan/wgdi-example/blob/main/Karyotype_Evolution.md) to identify protochromosomes and resulted in an ancestral Osteichthyes karyotype (AOK) of 24 protochromosomes (Fig. 3). The sex chromosomes of five genomes (four from Pleuronectiformes and one from Cyprinodontiformes) were derived from a pair of homologous chromosomes (AOK3 and AOK4), except for M. armatus from AOK22 and G. aculeatus from AOK21. Meanwhile, H. hippoglossus and H. stenolepis both have 24 chromosomes without any fusion or fission, but possess different sex determination systems (XX/XY and ZZ/ZW, respectively), and the origins of the sex chromosomes are also different in these two species (the former from AOK4 and the latter from AOK3). Coincidentally, AOK4 occurred a fusion event (NCF) in C. semilaevis. These results will provide valuable resources for follow-up research on the mechanism of sex chromosome evolution and the transformation mechanism of sex determination system of flatfish.
Data Records
The genome assembly related data was submitted to the NCBI SRA database with accession number SRP39616156. The genome sequence has been submitted to the GenBank database, GenBank assembly accession: GCA_026259375.157. Gene structure and transposition factor annotation files have been deposited in Figshare58.
Technical Validation
To assess genome assembly quality, we evaluated it from different aspects. We first used BUSCO 5.3.259 (-l actinopterygii_odb10) to assess the assembly completeness and found that the genome completeness reached 98.1%, and only 1.4% of BUSCO groups were not found in the genome. We then used several different methods to assess genome assembly quality. We assessed genome assembly quality based on the kmer strategy, to estimate the assembly accuracy to be 99.98%. In addition, we used freebays to search for homozygous SNPs that may be assembled base errors and found 7,215 homozygous SNPs, indicating that the base accuracy rate was as high as 99.998%. Finally, we used Illumina sequencing data to assess the sequence support of the genome. After aligning the Illumina sequence data back to the genome using bwa v0.7.17 with default parameters, we then used SAMtools60 flagstat to calculate basic statistics and found that 99.48% of the reads could be mapped back to the genome, of which 98.14% of the reads had the correct pairing orientation. We used SAMtools depth to count genome base coverage and found that 99.93% of bases were covered by at least 5 reads. Overall, the genome had good integrity and assembly quality.
Code availability
The data analysis methods, software and associated parameters used in present study are described in the section of Methods. If no detail parameters were described for software used in this study, default parameters were employed. No custom scripts were generated in this work.
References
Tomaszkiewicz, M., Medvedev, P., Makova, K. D. Y. & Chromosome, W. Assemblies: Approaches and Discoveries. Trends Genet 33, 266–282 (2017).
Hughes, J. F. et al. Chimpanzee and human Y chromosomes are remarkably divergent in structure and gene content. Nature 463, 536–539 (2010).
Bachtrog, D. et al. Are all sex chromosomes created equal? Trends Genet 27, 350–357 (2011).
Charlesworth, D., Charlesworth, B. & Marais, G. Steps in the evolution of heteromorphic sex chromosomes. Heredity (Edinb) 95, 118–128 (2005).
Bergero, R. & Charlesworth, D. The evolution of restricted recombination in sex chromosomes. Trends Ecol Evol 24, 94–102 (2009).
Wright, A. E., Dean, R., Zimmer, F. & Mank, J. E. How to make a sex chromosome. Nat Commun 7, 12087 (2016).
Pennell, M. W., Mank, J. E. & Peichel, C. L. Transitions in sex determination and sex chromosomes across vertebrate species. Mol Ecol 27, 3950–3963 (2018).
Zhou, Q. et al. Complex evolutionary trajectories of sex chromosomes across bird taxa. Science 346, 1246338 (2014).
Cortez, D. et al. Origins and functional evolution of Y chromosomes across mammals. Nature 508, 488–493 (2014).
Kobayashi, Y., Nagahama, Y. & Nakamura, M. Diversity and plasticity of sex determination and differentiation in fishes. Sex Dev 7, 115–125 (2013).
Nakamura, M. Is a sex-determining gene(s) necessary for sex-determination in amphibians? Steroid hormones may be the key factor. Sex Dev 7, 104–114 (2013).
Bachtrog, D. et al. Sex determination: why so many ways of doing it? PLoS Biol 12, e1001899 (2014).
Ravi, V. & Venkatesh, B. The Divergent Genomes of Teleosts. Annu Rev Anim Biosci 6, 47–68 (2018).
Devlin, R. H. & Nagahama, Y. Sex determination and sex differentiation in fish: an overview of genetic, physiological, and environmental influences. Aquaculture 208, 191–364 (2002).
Charlesworth, D. Young sex chromosomes in plants and animals. New Phytol 224, 1095–1107 (2019).
Rhie, A. et al. Towards complete and error-free genome assemblies of all vertebrate species. Nature 592, 737–746 (2021).
Duan, H. et al. Physical separation of haplotypes in dikaryons allows benchmarking of phasing accuracy in Nanopore and HiFi assemblies with Hi-C data. Genome Biol 23, 84 (2022).
Jasonowicz, A. J. et al. Generation of a chromosome-level genome assembly for Pacific halibut (Hippoglossus stenolepis) and characterization of its sex-determining genomic region. Mol Ecol Resour 22, 2685–2700 (2022).
Ma, H. et al. Isolation of sex-specific AFLP markers in Spotted Halibut (Verasper variegatus). Environmental Biology of Fishes 88, 9–14 (2010).
Einfeldt, A. L. et al. Chromosome level reference of Atlantic halibut Hippoglossus hippoglossus provides insight into the evolution of sexual determination systems. Molecular Ecology Resources 21, 1686–1696 (2021).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res 27, 722–736 (2017).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nat Biotechnol 37, 540–546 (2019).
Guan, D. et al. Identifying and removing haplotypic duplication in primary genome assemblies. Bioinformatics 36, 2896–2898 (2020).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv: Genomics (2013).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One 9, e112963 (2014).
Durand, N. C. et al. Juicer Provides a One-Click System for Analyzing Loop-Resolution Hi-C Experiments. Cell Syst 3, 95–98 (2016).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95 (2017).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol 21, 245 (2020).
Xu, X.-w. et al. Chromosome-level genome assembly of the Verasper variegatus provides insights into left eye migration. Frontiers in Marine Science 9 (2022).
Jain, C., Koren, S., Dilthey, A., Phillippy, A. M. & Aluru, S. A fast adaptive algorithm for computing whole-genome homology maps. Bioinformatics 34, i748–i756 (2018).
Marçais, G. et al. MUMmer4: A fast and versatile genome alignment system. PLoS Comput Biol 14, e1005944 (2018).
Garrison, E. P. & Marth, G. T. Haplotype-based variant detection from short-read sequencing. arXiv: Genomics (2012).
Pucholt, P., Wright, A. E., Conze, L. L., Mank, J. E. & Berlin, S. Recent Sex Chromosome Divergence despite Ancient Dioecy in the Willow Salix viminalis. Mol Biol Evol 34, 1991–2001 (2017).
Ou, S. et al. Benchmarking transposable element annotation methods for creation of a streamlined, comprehensive pipeline. Genome Biol 20, 275 (2019).
Bellott, D. W. et al. Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 508, 494–499 (2014).
Matsubara, K. et al. Evidence for different origin of sex chromosomes in snakes, birds, and mammals and step-wise differentiation of snake sex chromosomes. Proc Natl Acad Sci USA 103, 18190–18195 (2006).
Mawaribuchi, S. et al. Sex chromosome differentiation and the W- and Z-specific loci in Xenopus laevis. Dev Biol 426, 393–400 (2017).
Śliwińska, E. B., Martyka, R. & Tryjanowski, P. Evolutionary interaction between W/Y chromosome and transposable elements. Genetica 144, 267–278 (2016).
Kondo, M. et al. Genomic organization of the sex-determining and adjacent regions of the sex chromosomes of medaka. Genome Res 16, 815–826 (2006).
Schemberger, M. O. et al. DNA transposon invasion and microsatellite accumulation guide W chromosome differentiation in a Neotropical fish genome. Chromosoma 128, 547–560 (2019).
Suntronpong, A. et al. Implications of genome-wide single nucleotide polymorphisms in jade perch (Scortum barcoo) reveals the putative XX/XY sex-determination system, facilitating a new chapter of sex control in aquaculture. Aquaculture 548, 737587 (2022).
Nguyen, D. H. M. et al. Genome-wide SNP analysis suggests male heterogamety in bighead catfish (Clarias macrocephalus, Günther, 1864). Aquaculture 543, 737005 (2021).
Nguyen, D. H. M. et al. Genome-Wide SNP Analysis of Hybrid Clariid Fish Reflects the Existence of Polygenic Sex-Determination in the Lineage. Front Genet 13, 789573 (2022).
Nguyen, D. H. M. et al. An Investigation of ZZ/ZW and XX/XY Sex Determination Systems in North African Catfish (Clarias gariepinus,). Front Genet 11, 562856 (2020).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics 12, 491 (2011).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Keller, O., Kollmar, M., Stanke, M. & Waack, S. A novel hybrid gene prediction method employing protein multiple sequence alignments. Bioinformatics 27, 757–763 (2011).
Slater, G. S. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics 6, 31 (2005).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol 9, R7 (2008).
Zhang, J. et al. Genome puzzle master (GPM): an integrated pipeline for building and editing pseudomolecules from fragmented sequences. Bioinformatics 32, 3058–3064 (2016).
Shumate, A. & Salzberg, S. L. Liftoff: accurate mapping of gene annotations. Bioinformatics (2020).
Sun, P. et al. WGDI: A user-friendly toolkit for evolutionary analyses of whole-genome duplications and ancestral karyotypes. Mol Plant (2022).
Woods, I. G. et al. A comparative map of the zebrafish genome. Genome Res 10, 1903–1914 (2000).
Taylor, J. S., Braasch, I., Frickey, T. & Meyer, A. & Van de Peer, Y. Genome duplication, a trait shared by 22000 species of ray-finned fish. Genome Res 13, 382–390 (2003).
Volff, J. N., Nanda, I., Schmid, M. & Schartl, M. Governing sex determination in fish: regulatory putsches and ephemeral dictators. Sex Dev 1, 85–99 (2007).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP396161 (2022).
NCBI GenBank https://identifiers.org/ncbi/insdc.gca:GCA_026259375.1 (2022).
Xi-wen, X. The study of Veraspe variegatus W chromosome provides valuable insights into the evolution of sex chromosomes in flatfish and sheds light on the important role of whole-genome duplication in shaping the evolution of sex chromosomes. figshare https://doi.org/10.6084/m9.figshare.21791975.v4 (2023).
Manni, M., Berkeley, M. R., Seppey, M., Simão, F. A. & Zdobnov, E. M. BUSCO Update: Novel and Streamlined Workflows along with Broader and Deeper Phylogenetic Coverage for Scoring of Eukaryotic, Prokaryotic, and Viral Genomes. Mol Biol Evol 38, 4647–4654 (2021).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10 (2021).
Acknowledgements
This work was supported by the National Key R&D Program of China (2018YFD0900201), Shandong Key R&D Program (For Academician team in Shandong, 2023ZLYS02), the Key Research and Development Project of Shandong Province (2021LZGC028), Central Public-interest Scientific Institution Basal Research Fund, CAFS (2020TD20), and Shandong Taishan Scholar Climbing Project.
Author information
Authors and Affiliations
Contributions
S.C. and X.X. applied, designed and supervised the project. X.X., C.G. and W.Z. prepared the samples for whole genome sequencing and analyzed the data. P.S. finished karyotypic analyses. X.X., P.S., C.G., W.Z. and S.C. wrote and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xu, Xw., Sun, P., Gao, C. et al. Assembly of the poorly differentiated Verasper variegatus W chromosome by different sequencing technologies. Sci Data 10, 893 (2023). https://doi.org/10.1038/s41597-023-02790-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02790-z