Abstract
miRNAs present in milk are mainly found in extracellular vesicles (EVs), which are nanosized membrane vesicles released by most of the cell types to ensure intercellular communication. The majority of the studies performed so far on these vesicles have been conducted on human and cow’s milk and focused on their miRNA content. The objectives of this study were to profile the miRNA content of purified EVs from five healthy goats and to compare their miRNome to those obtained from five healthy cows, at an early stage of lactation. EV populations were morphologically characterized using Transmission Electron Microscopy and Nanoparticle Tracking Analysis. The presence of EV protein markers checked by Western blotting and the absence of contamination of preparations by milk proteins. The size distribution and concentration of bovine and goat milk-derived EVs were similar. RNA-sequencing were performed, and all sequences were mapped to the cow genome identifying a total of 295 miRNAs. This study reports for the first-time a goat miRNome from milk EVs and its validation using cow miRNomes.
Similar content being viewed by others
Background & Summary
Milk is the main source of nutrients for newborn mammals, and is thought to play an important role in the development of the young immune system1. However, the mechanisms driving milk immune influence are still poorly understood and may be mediated by cells and bioactive molecules present in milk, including microRNAs (miRNAs). miRNAs are small non-coding RNAs (17–25 nucleotides long), highly conserved across species2, that post-transcriptionally regulate the expression of at least up to 60% of human genes, thus influencing many cellular processes, such as cell differentiation, proliferation, and cell death3,4,5. Milk miRNAs are, to some extent, packaged within extracellular vesicles (EVs) that are present in large amounts in milk6,7, and are, in such a way, protected from degradation in the gastrointestinal tract6,8. Therefore, they can survive digestion, be taken up by intestinal epithelial and immunologic cells in which they may be functionally active9,10. Thus, the survival of EV miRNAs and their influence on many biological processes make them crucial actors in the young and adult milk consumer health8,10,11,12. Whereas the repertories of miRNAs (miRNome) of human and cow’s milk EV were yet rather well described13,14,15,16, little is known in contrast on the miRNome of goat milk-derived EVs17. The objectives of this study were to determine the miRNome of milk-derived EVs from goat, and compare it with that of cow.
This study reports to our knowledge the first-time a goat miRNome from milk EVs and its validation using cow miRNomes. An average of 239 and 163 miRNAs were found with more than 1 Tag Per Million (TPM) in the cow and goat milk-derived EVs, respectively. The first 36 most abundant miRNAs present in goat milk-derived EVs accounted for 95% of TPM miRNAs. The miRNomes of the two ruminant species studied were very similar and among the TOP 10 most abundant miRNAs, 7, including the first 4 (miR-148a, miR-21-5p, miR-26a and miR-30a-5p) were common. These data were compared to published data on EVs from human and cow’s milk. Several abundant miRNAs were shared between species, such as miR-148a, miR-21-5p, let-7a-5p, miR-200a-3p and miR-30d-5p.
Methods
Materials
Since we did not perform any experiment on animal, no ethical approval was required for this study. Individual milk samples (100 mL each) were collected from five primiparous Alpine goats and five Holstein Friesian cows, at early lactation stages, 26 ± 9 days postpartum and 30 ± 7 days postpartum, respectively. Animals were reared in experimental farms of the Institut National de la Recherche pour l’Agriculture, l’Alimentation et l’Environnement (INRAE) at Domaine de Galle, 18520, Avord for goats, and at Domaine du Pin-au-Haras, 61310 Exmes, for cows. Preservatives [potassium dichromate: 0.05 g/L, 10 mM ε-amino caproic acid, and 10 μM phenylmethylsulfonyl fluoride (PMSF)] were added to milk immediately after milking to prevent proteolysis. Whole milk in 50 mL Falcon tubes were stored on ice during transport to the lab (1 hour). Depletion of somatic cells and milk fat was performed by centrifugation at 2,000 g, 15 min, 4 °C. Further, the second centrifugation of milk supernatant at 3,000 g for 30 min at 4 °C (Allegra X-15R, Beckman Coulter, France) was performed after transfer of liquid phase to a new tube with objective to eliminate the cell debris. The supernatant of skimmed and cell-free milk thus obtained was transferred to another 50 mL Falcon tubes and frozen at −80 °C before subsequent analysis.
Isolation of bovine and goat milk-derived EVs using sucrose gradient ultracentrifugation
Isolation of milk-derived EVs as well as their validation and exo-RNA isolation were performed as follow. Skimmed milk samples stored previously at −80 °C were incubated in a water bath at 37 °C for 45 min. This step allows the reintegration of the free beta casein from the soluble phase into the casein micelles. The whey was obtained after acid precipitation by adding 10% (v/v) of 10% acetic acid, followed by incubation in a water bath at 37 °C for 10 min. Furthermore, an additional step specific for the treatment of goat milk characterized by a lower isoelectric point of caseins was performed by adding 1% (v/v) of 50% acetic acid to pre-treated goat milk samples. The mixture was incubated in water bath at 37 °C for 10 minutes. Coagulated milk samples were mixed shortly and 10% (v/v) of 1 M sodium acetate was added followed by incubation for 10 minutes at room temperature (RT). The major milk proteins (caseins) were pelleted by centrifugation at 1,500 g, 4 °C for 15 min. The whey fraction was filtered using vacuum-driven filtration system Steritop, 0.22 μm (Merck Millipore, France). Next, the cleared whey supernatants were concentrated using Amicon-15 100 kDa centrifugal filter units (Merck Millipore, France) at 4,000 g and 20 °C while obtaining the final volume of 6–8 mL. To pellet the EVs, the obtained concentrated retentate was ultra-centrifuged at 120,000 g for 1h10 at 4 °C using 50Ti rotor (Optima XPN-80, Beckman Coulter, France). The enriched EV’s pellets were resuspended in 500 μL of PBS 1x,then loaded at the top of 11 mL of pre-prepared sucrose gradient with decreasing sucrose concentration from 5% (on the top) to 45% (on the bottom) and ultra-centrifuged using SW41 rotor at 200,000 g for 18 h at 4 °C (Optima XPN-80, Beckman Coulter, France). Whole tube volume was fractionated into 1 mL fractions. The fractions 10–12 containing targeted EV populations were collected and diluted in 6 mL of PBS 1x,then centrifuged at 100,000 g using 50Ti rotor for 1h10 at 4 °C (Optima XPN-80, Beckman Coulter, France). The pellets were resuspended in 50 μL of PBS 1x. The resuspended pellets of fractions 10–12 were pooled and stored at −80 °C, until further analysis.
Transmission electron microscopy (TEM) with uranyl acetate negative staining
The extracellular vesicles were analyzed after deposition of 4 µL of suspension on a 300-mesh copper grid covered with a carbon film membrane for 5 min, then stained with 1% uranyl acetate after absorption of the excess contrasting liquid and dried at room temperature. The grids were observed on a Hitachi HT7700 transmission electron microscope operated at 80 kVolts (Milexia). Images were acquired using a charge-coupled device camera (AMT).
Nanoparticle Tracking Analysis (NTA) of milk-derived EVs by Nanosight NS300 (Malvern)
Particle concentration and size distribution were determined using a Nanosight NS300 instrument (Malvern, version NTA 3.2 Dev Build 3.2.16). The instrument was equipped with a Blue 405 nm laser, automatic syringe pump, sCMOS camera type and the NTA software v3.2. The capture settings were as follow: camera level of 12, Slider shutter of 1200, Slider gain of 146, and syringe pump speed of 50. Per sample, three videos of 90 seconds with a frame rate of 30 frames/sec were captured at constant acquisition temperature of 25 °C and subsequently analyzed with a threshold set up at 5. The NTA experiments were carried out with pre-diluted milk-derived EV samples in PBS 1x according to input sample concentrations, The obtained results were validated with at least 1,000 valid tracks for each triplicate.
Bradford protein assay
One microliter of each EV sample was used to measure the protein concentration with a Bradford Protein Assay kit (Thermofisher). A standard linear curve was set up using BSA (Thermofisher). The protein quantification was achieved thanks to the reaction of proteins with a Coomassie Protein Assay reagent (Thermofisher). Absorbance measurement (OD) was performed in the visible spectrum at 595 nm on a spectrophotometer UVmini-1240 (Shimadzu).
Western blot analysis
Western blotting (WB) was performed to validate the presence of known EV markers such as CD81 and hsp70 as well as to control the possible contamination by caseins. For this purpose, the presence of β-CN was also investigated. Twenty µg of EV proteins, corresponding approximately to a volume of 4-5 µL of bovine and goat EV samples were mixed with a denaturating solution of 4X Laemmli Buffer (Bio-Rad) and 10% β-mercaptoethanol prior to be heated for 10 min at 95 °C. Once cooled on ice, the protein samples were loaded onto a pre-cast 4–12% gradient polyacrylamide gel (Bio-Rad), and, after migration (15 mA/gel) in 1X TGS at 4 °C for 1 hour, subsequently transferred to a nitrocellulose membrane using Transblot Turbo system (Bio-Rad). The membrane was then blocked with 5% BSA + 0.05% Tween for 1 hour at room temperature. The membranes were thereafter exposed to anti-hsp70 and anti-CD81 rabbit primary antibodies (1:1000 in 5% BSA + 0.05% Tween, SBI) and to anti-β-casein rabbit polyclonal antibody (1:300 in 5% BSA + 0.05% Tween, Thermofisher) at 4 °C overnight. After incubation, several washing steps of the membranes with 0.05% Tween in TBS were applied. The membranes were subsequently incubated for one hour at room temperature with the goat anti-rabbit HRP secondary antibody (EXOAB-KIT, SBI) diluted 1:20000 in 5% BSA + 0.05% Tween TBS and washed. The chemiluminescence ECLPlus 2 kit (Pierce) was applied to the membranes 5 min prior imaging. Imaging was performed using ChemiDoc Touch Imaging System (Bio-Rad) equipped with CCD LAS1000 camera.
RNA isolation
Total RNAs, including small RNAs were isolated from EV samples using an optimized mirVana Total RNA Isolation Kit (Life Technologies) with some modifications: firstly, Trizol LS Reagent (Ambion) was used for initial cell disruption instead of mirVana Phenol Lysis Reagent, followed by chloroform addition, ethanol precipitation (sample: 100% ethanol ratio of 1:1.25, v/v) and mirVana kit columns (Thermofisher) fractionation. Glycogen (1 µL per 800 µL of lysate) was added to improve RNA recovery. To obtain high quality RNA, a DNAse I treatment (Qiagen) was performed on columns for 15 min at RT according to manufacturer’s instructions. Finally, the RNAs were eluted in 50 µL of Elution buffer, then quantified by Bioanalyzer 2100 instrument (Agilent) using Pico chip and ND-1000 NanoDrop™ Spectrophotometer (Thermofisher).
Small RNA library preparation and Next Generation RNA sequencing
MicroRNA NGS libraries were performed from 100 ng of total RNA using NEBNext library generation kit (New England Biolabs Inc.) according to the manufacturer’s instructions. Adaptors were ligated to the 3′ and 5′ of each RNA and converted into cDNA. A pre-amplification was carried out with specific primers for 15 cycles. After a purification using QiaQuick columns, the insert efficiency was evaluated on high sensitivity DNA chip using a Bioanalyzer 2100 instrument (Agilent). A size fractionation of obtained libraries was performed on a LabChip XT (Caliper Life Sciences) and bands corresponding to adaptors with 15–40 bp insert were excised using the manufacturer’s instructions. Sample quantifications were performed using qPCR. Based on quality and quantity of the inserts, the libraries were pooled in equimolar concentrations. An optimal concentration of the library pools determined by qPCR was used to generate the clusters on the flow cell surface before sequencing. Samples were sequenced on the Illumina NextSeq 500 system, according to the manufacturer instructions (Illumina). Sequencing data quality control (Q-score: QC), and adapter trimming were performed prior sequence mapping using miRbase 20. Because bovine is a phylogenetically related species of goat and has a higher number of miRNAs, the following results were based on the annotations obtained using Bos taurus as reference genome18.
Putative novel miRNAs were predicted from the sequences that did not map to any organism present in miRbase, or to other known RNA sequences. miRPara19 was used to analyze the potential folding of these sequences. These results were combined to identify 180 putative novel miRNAs (see raw data).
Data Records
Read data
The RNA-seq profiling was successfully completed for the 9 samples (one goat sample did not pass the RNA quality control). On average 11.6 million reads were obtained per sample with an average of 12.8 and 10.8 million of reads for Capra hircus and Bos taurus samples, respectively. The data were accessible with the GEO accession number: GSE22755920 and at Figshare number 2199710621.
Identified microRNAs
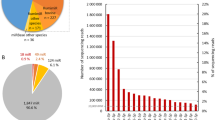
After mapping the data on the bovine genome and counting to relevant entries in miRBase 20 the numbers of known microRNAs was determined. Taking into account all 9 samples (5 cows and 4 goats) of milk-derived EVs analyzed, a total of 295 miRNAs were identified. Goat samples displayed higher call-rates (number of identified known bta-miRNAs) than cow samples, ranging between 228 and 247, and 143 and 154 for number of counts > 1 TPM and > 10 TPM, respectively (Fig. 1).
Most highly expressed miRNAs
In this study, the most widely represented miRNA (number one), with a mean TPM above 50,000 in both goats (88,925) and cows (123,755) is miR-148a. The four first miRNAs are the same in goat and cow, miR-21-5p at rank 2, positions 3 and 4 being reversed (miR-26a and miR-30a-5p). miR-146b, which ranks 5th in the goat species, is only 69th in the bovine species with a 150-fold lower level of expression (Table 1). The same is true for miR-224 and miR-378, which rank 190 (vs. 36 in goats) and 53 (vs. 17 in goats) respectively, with expression levels 800 and 100 times lower in bovine than in caprine. The data are accessible with GEO accession number: GSE22755920 and on figshare21.
Technical Validation
Morphological characterization of EVs populations
Transmission electron microscopy (TEM)
As shown in Fig. 2, milk-derived EVs appeared rather homogeneous in shape, in cow milk (A) as well as in goat milk (B), ranging in size between 30 and 150 nm. They were sometimes clustered in aggregates. Even though samples must be dehydrated before analysis no cell debris and minimal background were observed, revealing the efficiency of the purification procedure.
Nanoparticle tracking analysis (NTA) of EVs
Based on the NTA data obtained using Nanosight NS300 (Malvern), the average diameter of milk-derived EV (±standard error) was 165.00 ± 3.44 nm for cows, and 169.66 ± 3.65 nm for goats (Table 2). The distribution of mEV populations was “gaussian-like” for both species with a quite unique peak and a narrow range (Fig. 3), revealing a homogeneous population of particles both within and between species.
Western blot analysis of proteins from EVs preparations
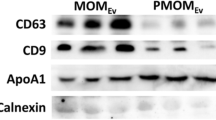
The presence of EV-associated markers (hsp70 and CD81) and the absence of milk protein contaminant were clearly demonstrated by Western Blotting (Fig. 4) both for EVs isolated from cow’s milk (1) and from goat’s milk (2).
Quality control of RNA samples
The Bioanalyzer RNA assay providing an objective measurement of RNA preparations quality and a sensitive analysis to resolve nucleic acid samples in the size range of 10 to 4000 nt was performed. However, it must be kept in mind that the RNA Integrity Number (RIN) is meaningless as long as the RNA preparation does not contain ribosomal RNA, which is an additional proof of the quality of the miRNA preparations. A typical result of quality control from goat milk-derived EV is given in Fig. 5.
Validation of miRNA sequencing
Principal component analysis plot
Principal Component Analysis (PCA) was used to explore the naturally arising sample classes based on the expression profile. Here, by including the TOP 50 miRNAs that had the largest variation across all samples, an overview of how the samples cluster based on this variance was obtained. The data was normalized with tag per million (TPM) method and converted to a log2 scale. Then all features were filtered on “expressed in all samples” criteria and the 50 features with the highest coefficient of variation (% CV) selected for the analysis. The largest component in the variation is plotted along the X-axis and the second largest is plotted on the Y-axis. As shown in Fig. 6, the samples tend to cluster based on the organism (mainly along the second axis, i.e. second component in the variation), the first factor of variation being the individual intra-species variability (X-axis). Thus, PCA separated the samples according to their origin.
Heat map and unsupervised clustering
The heat map diagram shows the result of the two-way hierarchical clustering of miRNAs and samples (Fig. 7). The data is normalized with TPM method and converted to a log2 scale. Then all features are filtered on “expressed in all samples” criteria and the 50 miRNAs with the highest coefficient of variation (%CV) selected for the analysis. Each row represents one miRNA, and each column represents one sample. The color of each point represents the relative expression level of a miRNA across all samples: This analysis led to classifying the samples according to their organism of origin.
For goats, the 10 most abundant miRNAs account for nearly 75% (ranging between 72,7 and 73,8%) of the 295 miRNAs mapped, at least in one sample, referring to the bovine genome. For cows, the 10 most abundant miRNAs account for nearly 80% (ranging between 78,8 and 80,0%) of the miRNAs mapped. The most abundant miRNAs in both species is miR-148a, which occupies almost systematically the pole position in milk-derived miRNAs profiling studies, whatever the mammalian species considered, including human22,23 and pig24. Our results are validated by the comparison with data previously reported in cow and human, showing a close pattern of abondance (Table 3). The miRNAs, which were not in the TOP 10 miRNAs of the present study, were detected with a lower rank but were present.
Code availability
No custom code was written.
References
Admyre, C. et al. Exosomes with immune modulatory features are present in human breast milk. J. Immunol. 179, 1969–1978 (2007).
Saetrom, P. et al. Conserved microRNA characteristics in mammals. Oligonucleotides 16, 115–144 (2006).
Ambros, V. The functions of animal microRNAs. Nature 431, 350–355 (2004).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Bartel, D. P. Metazoan MicroRNAs. Cell 173, 20–51 (2018).
Benmoussa, A. et al. Commercial dairy cow milk microRNAs resist digestion under simulated gastrointestinal tract conditions. J. Nutr. 146, 2206–2215 (2016).
Xu, L., Yang, B. F. & Ai, J. MicroRNA transport: A new way in cell communication. J. Cell Physiol. 228, 1713–1719 (2013).
Liao, Y., Du, X., Li, J. & Lönnerdal, B. Human milk exosomes and their microRNAs survive digestion in vitro and are taken up by human intestinal cells. Mol. Nutr. Food Res. 61, e1700082 (2017).
Arntz, O. J. et al. Oral administration of bovine milk derived extracellular vesicles attenuates arthritis in two mouse models. Mol. Nutr. Food Res. 59, 1701–1712 (2015).
Kahn, S. et al. Exosomal MicroRNAs in Milk from Mothers Delivering Preterm Infants Survive in vitro Digestion and Are Taken Up by Human Intestinal Cells. Mol. Nutr. Food Res. 62, e1701050 (2018).
Mar-Aguilar, F. et al. Evidence of transfer of miRNAs from the diet to the blood still inconclusive. Peer J. 8, e9567 (2020).
Leroux, C., Chervet, M. L. & German, J. B. Perspective: Milk microRNAs as important players in infant physiology and development. Adv. Nutr. 12, 1625–1635 (2021).
Izumi, H. et al. Bovine milk exosomes contain microRNA and mRNA and are taken up by human macrophages. J. Dairy Sci. 98, 2920–2933 (2015).
Ma, S., Tong, C., Ibeagha-Awemu, E. M. & Zhao, X. Identification and characterization of differentially expressed exosomal microRNAs in bovine milk infected with Staphylococcus aureus. BMC Genomics 20, 934 (2019).
Kirchner, B., Buschmann, D., Paul, V. & Pfaffl, M. W. Postprandial transfer of colostral extracellular vesicles and their protein and miRNA cargo in neonatal calves. PLoS ONE 15, e0229606 (2020).
López de Las Hazas, M. C. et al. Dietary bovine milk miRNAs transported in extracellular vesicles are partially stable during GI digestion, are bioavailable and reach target tissues but need a minimum dose to impact on gene expression. Eur. J. Nutr. 61, 1043–1056 (2022).
Mecocci, S. et al. Transcriptomic Characterization of Cow, Donkey and Goat Milk Extracellular Vesicles Reveals Their Anti-Inflammatory and Immunomodulatory Potential. Int. J. Mol. Sci. 22, 12759 (2021).
Mobuchon, L. et al. Annotation of the goat genome using next generation sequencing of microRNA expressed by the lactating mammary gland: comparison of three approaches. BMC Genomics 16, 285 (2015).
Wu, Y., Wei, B., Liu, H., Li, T. & Rayner, S. MiRPara: a SVM-based software tool for prediction of most probable microRNA coding regions in genome scale sequences. BMC Bioinformatics 12, 107 (2011).
Martin, P., Krupova, Z. & Leroux, C. GEO. https://identifiers.org/geo/GSE227559 (2023).
Martin, P. Goat and cow milk-derived extracellular vesicle miRNomes. Figshare https://doi.org/10.6084/m9.figshare.21997106 (2023).
Simpson, M. R. et al. Human breast milk miRNA, maternal probiotic supplementation and atopic dermatitis in offspring. PLoS One 10, e0143496 (2015).
van Herwijnen, M. J. C. et al. Abundantly Present miRNAs in Milk-Derived Extracellular Vesicles Are Conserved Between Mammals. Front. Nutr. 5, 81 (2018).
Gu, Y. et al. Lactation-related microRNA expression profiles of porcine breast milk exosomes. PLoS one 7, e43691 (2012).
Cai, M. et al. Genome-wide microRNA profiling of bovine milk-derived exosomes infected with Staphylococcus aureus. Cell Stress Chaperones 23, 663–672 (2018).
Acknowledgements
The authors warmly thank the Agence Nationale pour la Recherche (ANR) for funding the following research program: “MilkChEST” (ANR-12-BSV6-0013-04), within the framework of which this work was carried out.
Author information
Authors and Affiliations
Contributions
P.M., C.L. and Z.K. conceived, designed, and initiated the project. Z.K. and C.B. performed isolation of EVs from milk samples. C.P. ensure T.E.M. analyses. C.B. and Z.K. contributed in RNA extraction and RNA quality control analyses. Z.K. conducted N.T.A. and Western blot analyses. NGS libraries were prepared, quantified and sequenced by an external provider (Exiqon) who also mapped the sequences on both goat and cow genomes. P.M., Z.K. and C.L. wrote the original draft. P.M. coordinated and supervised the project. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Krupova, Z., Leroux, C., Péchoux, C. et al. Comparison of goat and cow milk-derived extracellular vesicle miRNomes. Sci Data 10, 465 (2023). https://doi.org/10.1038/s41597-023-02347-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02347-0