Abstract
Sipuncula is a class of exocoelomic unsegmented animals whose evolutionary relationships are unresolved. The peanut worm Sipunculus nudus is a globally distributed, economically important species belonging to the class Sipuncula. Herein, we present the first high-quality chromosome-level assembly of S. nudus based on HiFi reads and high-resolution chromosome conformation capture (Hi-C) data. The assembled genome was 1,427 Mb, with a contig N50 length of 29.46 Mb and scaffold N50 length of 80.87 Mb. Approximately 97.91% of the genome sequence was anchored to 17 chromosomes. A BUSCO assessment showed that 97.7% of the expectedly conserved genes were present in the genome assembly. The genome was composed of 47.91% repetitive sequences, and 28,749 protein-coding genes were predicted. A phylogenetic tree demonstrated that Sipuncula belongs to Annelida and diverged from the common ancestor of Polychaeta. The high-quality chromosome-level genome of S. nudus will serve as a valuable reference for studies of the genetic diversity and evolution of Lophotrochozoa.
Similar content being viewed by others
Background & Summary
Sipuncula (peanut worms) are unsegmented coelomate worms with bilaterally symmetrical bodies that are separated into a trunk and are retractable introverts1. Belonging to Lophotrochozoa, they are believed to form a small phylum with approximately 150 described species2. However, they are widely distributed in the world’s oceans at all depths, occupying most marine habitats, from intertidal zones to abyssal depths and polar to equatorial seas, including extreme environments. Over the past 520 million years, the typical features of extant Sipuncula have undergone only minor changes3. Therefore, Sipuncula is an exciting resource to study environmental adaptation and evolution and as an indicator of global climate change. In coastal environments, these species are critical in bioturbation to reshape the physicochemical properties and biological characteristics of the sediment4. In marine wetlands and pond aquaculture systems, Sipuncula and other taxa increase organic matter transport and improve ecosystem services5. However, gene and genome data for Sipuncula that are available in the PDB, a public database, are insufficient.
Despite the early recognition of the group, phylogenetic relationships between Sipuncula and other taxa are unclear. Sipunculus nudus was first described by Linnaeus in 1767 and was later considered to be a derived group of annelids6,7,8. Morphological and developmental characteristics suggest that Sipuncula is the sister group of Mollusca9. However, phylogenetic analyses based on mitochondrial DNA sequences as well as traits related to nervous and muscle system development indicate that Sipuncula is more closely related to Annelida than to Mollusca10,11. Torsten et al. performed phylogenomic analyses using 47,953 amino acid positions to explore the relationships among 34 annelid taxa and found that Sipuncula belongs to Annelida12. Therefore, the assignment of Sipuncula to annelids is still a controversial issue. Furthermore, the lack of segments in Sipuncula, which is different from other annelid taxa, provides a basis for understanding the mechanism underlying segment development. Genome sequence information is important for phylogenetic analyses. However, sequencing data for molluscs and annelids are limited. In Sipuncula, only one draft genome of Phascolosoma esculenta was published by Zhong et al.13. The genome data suggested that Sipuncula belonged to Annelida; however, the evolutionary relationships among Polychaeta, Oligochaeta, and Hirudinea in their reconstructed phylogenetic tree were inconsistent with previous results, making evolutionary inferences difficult. Therefore, additional genome data for Sipuncula, especially chromosome-level genome data, are needed to clarify the evolutionary relationships of lophotrochozoans and to provide genomic resources for “evo-devo” studies of body segmentation.
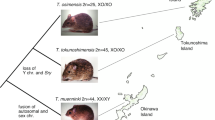
S. nudus is a cosmopolitan Sipuncula species that is distributed in temperate, subtropical, and tropical waters in all oceans (Fig. 1). In this study, we assembled the first high-quality genome of S. nudus using PacBio HiFi sequencing and high-throughput chromosome conformation capture (Hi-C). We used HiFi reads for assembly and Hi-C technology for chromosome anchoring. We obtained a contig N50 of 29.47 Mb and a scaffold N50 of 80.87 Mb for the final genome assembly, which is approximately 1,427 Mb. Using Hi-C data, 97.91% of the assembled bases were associated with the 17 chromosomes. These high-quality genomic data are expected to improve the resolution of phylogenetic analyses of Sipuncula and to provide a reference for detailed analyses of their characteristics, adaptation to complex habitats, and ecological niches.
Methods
Sample collection and DNA extraction
Male 2-year-old S. nudus samples were collected from the field of Suixi, Zhanjiang Guangdong Province, China (21°35′N, 109°81′E), and were used for whole-genome sequencing. The body wall tissue was stored in liquid nitrogen, and total genomic DNA was isolated by using the QIAGEN DNeasy Blood & Tissue Kit (QIAGEN, Shanghai, China) following the manufacturer’s instructions.
Library construction and sequencing
Three SMRTbell libraries of circular consensus sequencing (CCS) were constructed according to the standard PacBio protocol using 15–20 kb preparation solutions (Pacific Biosciences, Menlo Park, CA, USA). Five cells were sequenced on the PacBio Sequel II platform by the CCS model (Pacific Biosciences) to generate HiFi (high-fidelity) reads. The reads were produced by calling consensus from subreads that were generated by multiple passes of the enzyme around a circularized template. This resulted in a HiFi read that was both long and accurate. In total, 103.13 Gb of HiFi reads with 72.63× coverage was generated, and the N50 value was 14,008 bp (Table 1).
Hi-C libraries were prepared as previously reported14. The body wall tissue cells were fixed by using formaldehyde to keep the 3D structure of DNA intact. Cells were digested with the HindIII restriction endonuclease. Biotin-labelled bases were used for end repair. The DNA fragments maintaining interaction relationships were captured to construct the Hi-C library. Finally, 289.30 Gb of high-quality Hi-C data (Q20 > 98% and Q30 > 94%) was obtained with the BGISEQ-500 sequencing platform (Table 2).
Genome survey and assembly
The size, heterozygosity, and repeat rate of the S. nudus genome were estimated using the k-mer frequency method. Jellyfish15 and GenomeScope v.1.016 were employed to calculate the K-mer frequency (k = 21), which was based on HiFi reads, and the genome size was estimated to be 1305 Mb with a peak K-mer frequency of 66X. The heterozygosity and repeat rate were 2.03% and 39.68%, respectively (Fig. 2). We first assembled the genome using HiFi reads via HiFi-asm (v0.15.1)17 with default parameters. After preliminary assembly, we used purge_haplotigs18 to purge haplotigs. The haploid genome size was 1426.68 Mb, and the N50 length was 29.46 Mb (Fig. 3 and Table 3).
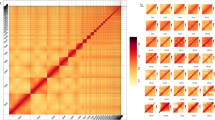
The contigs were anchored to chromosomes using Hi-C data. Juicer (version 1.6)19 was used to align the double-ended sequencing data against the assembled genome to complete the evaluation of the Hi-C library. The 3D-DNA pipeline20 under default parameters without breaking contigs was chosen to generate the final chromosome-level scaffolds. Manual checking and refinement of the draft assembly were carried out via Juicebox Assembly Tools (https://github.com/aidenlab/Juicebox, v1.1108). A heatmap of the Hi-C assembly interaction bins indicated that the quality of the genome assembly was excellent (Fig. 4). The length of the final assembled genome was 1,426,776,655 bp, with a contig N50 of 29,460,569 bp and scaffold N50 of 80,869,746 bp (Table 3 and Fig. 3). Approximately 1,397 Mb (97.91%) of the contig sequences were anchored to 17 chromosomes (Table 4), which is consistent with the known karyotype in our previously published manuscript21. Using the minimap2 (v2.17, parameters: -a -x map-pb)22 alignment results and the HiFi data, we used BamDeal (https://github.com/BGI-shenzhen/BamDeal) to evaluate the mapping rate and coverage and obtained estimates of 99.95% and 99.73%, respectively. The CIRCOS tool23 was used to visualize the 17 chromosomes, GC content, read depth and mapping depth (Fig. 5). The average depth of each chromosome was calculated and is shown in Fig. 6. Seventeen chromosomes had a comparable sequencing depth, and there was no whole chromosome with half the read depth. Therefore, XY- or ZW-type sex chromosomes did not exist in the assembled chromosomes of S. nudus. Based on 20-kb nonoverlapping sliding windows in the chromosomes to calculate the GC content and read average depth, there was a small cluster of sliding windows (a total of 11.6 Mb with 581 sequences) that exhibited relatively high GC contents ( > 48%) but with a normal sequencing depth (Fig. 7). By extracting those block sequences with high GC contents and mapping them to the NT database (Nucleotide Sequence Database) using MegaBlast (parameter: −e 1e-5), the alignments with identity >90% and coverage length >100 bp were filtered. The matched reference species in the alignments from the NT database were grouped into three categories: the S. nudus species, the species of other invertebrates, and all other species except the two mentioned above. All the matched sequences (228) could be correlated with S. nudus or other invertebrate species (Fig. 8), which demonstrated that the sequence blocks with high GC content and normal depth in chromosomes were from the S. nudus species rather than from contamination or cobionts.
Repeat annotation
Prior to gene prediction, we identified the repetitive elements in the genome of S. nudus by using a combination of homology-based and ab initio-based methods. To identify tandem repeats, we used Tandem Repeats Finder v4.0924. For the homology-based method, transposable elements were identified by RepeatMasker v4.0.7 (-nolow -no_is -norna -engine ncbi -parallel 1) and RepeatProteinMask v4.0.7 (-engine ncbi -noLowSimple -pvalue 0.0001)25 against the TE protein databases and RepBase library v21.1226. For the ab initio-based method, LTR_FINDER v1.0627 and RepeatModeler v1.0.8 (http://repeatmasker.org/RepeatModeler/) with default parameters were used to build the de novo library before RepeatMasker v4.0.7 was used to classify the different categories of repetitive elements against this library. The final repetitive elements were identified by integrating the results of these methods according to sequence overlap, revealing that nearly half of the genome consists of repetitive elements (Tables 5, 6; Fig. 5).
Gene prediction
Gene annotation was performed by integrating homology-, de novo- and transcriptome-based information. We used the annotation data from three closely related species (Caenorhabditis elegans, Capitella teleta, and Helobdella robusta) for homology prediction. The MAKER tool28 was used to integrate the annotation data from the three related species and the transcriptome data from S. nudus. Based on AED values from MAKER, 2000 genes with complete structures were selected and used to train the de novo prediction tools Augustus29 and Snap30 to construct de novo models. Finally, all data were integrated using MAKER28. The final comprehensive gene set contained 28,749 genes (Table 7).
Gene function annotation
Gene function annotation was performed based on sequence similarity and domain conservation. First, the protein-coding genes of S. nudus were aligned against the KEGG31, SwissProt32, TrEMBL33, GO34, KOG (ftp://ftp.ncbi.nih.gov/pub/COG/KOG/), and Nr (https://ftp.ncbi.nlm.nih.gov/blast/db/FASTA/nr.gz) databases by using BLASTP with an E-value threshold of 1e-5. Subsequently, the best match from the alignment was used to predict gene functions. Second, searches performed using InterProScan (51.0–55.0)35 against the following databases were used to identify the motif and domain: PANTHER36, Pfam37, PRINTS38, ProDom39, SUPERFAMILY40, and SMART41. In total, 88.75% of the predicted genes were functionally annotated (Table 8).
Data Records
The National Center for Biotechnology Information (NCBI) BioProject accession number for the sequence reported in this paper is PRJNA901211. The raw data for Hi-Fi and Hi-C sequencing were submitted to NCBI SRA (accession number SRP408321; https://identifiers.org/ncbi/insdc.sra:SRP408321)42 and deposited in the CNGB Sequence Archive (CNSA) of the China National GeneBank DataBase (CNGBdb) (accession number CNR0640303-CNR0640323; https://db.cngb.org/search/project/CNP0003624/)43. The assembled genome sequence was deposited into NCBI under accession number JAPPUL00000000044. The assembled genome, gene structure annotation, repeat predictions, gene function annotation, KEGG analysis of expanded genes and positively selected gene data were deposited in the China National GeneBank DataBase (CNGBdb) under the project with accession number CNP0003624.
Technical Validation
Genome assembly and gene prediction quality assessment
The BUSCO pipeline was used to evaluate the completeness of the genome assembly and gene set based on a benchmark of 255 conserved genes in eukaryota_odb10 (creation date: 2020-09-10, number of genomes: 70, number of BUSCOs: 255). In total, 97.7% of the 255 expected conserved genes were identified as complete, and 2% were identified as fragmented. Furthermore, we used minimap2 (v2.17, parameters: -a -x map-pb)22 to align the assembly with the HiFi data, and the mapping rate and coverage rate were estimated to be 99.95% and 99.73%, respectively. The BUSCO (v5)45 results supported the completeness of the assembly; 97.7% of 255 conserved genes were identified as complete by using eukaryota_odb10 (Table 9). The BUSCO results and alignment results indicated high genome assembly completeness and correctness.
Comparative genomic analysis
The protein-coding genes of S. nudus and 15 additional species were used to identify orthologous gene groups. The reference protein sequences of the following 15 species were obtained: Caenorhabditis elegans (Ensembl Release 10), Danio rerio (Ensembl Release 10), Homo sapiens (Ensembl Release 10), Drosophila melanogaster (Ensembl Release 10), Capitella teleta (NCBI: GCA_000328365.1), Crassostrea gigas (NCBI: GCF_902806645.1), Dimorphilus gyrociliatus (NCBI: GCA_904063045.1), Eisenia andrei (ngdc.cncb.ac.cn: PRJCA002327), Helobdella robusta (NCBI: GCF_000326865.1), Lamellibrachia satsuma (NCBI: GCA_022478865.1), Lottia gigantea (NCBI: GCF_000327385.1), Metaphire vulgaris (NCBI: GCA_018105865.1), Owenia fusiformis (NCBI: GCA_903813345.2), Phascolosoma esculenta (https://figshare.com/: PRJNA819496), and Nematostella vectensis (NCBI: GCF_932526225.1) as the outgroup. To perform the gene family analysis, orthogroups of the 16 species were identified using OrthoFinder (v2.3.11) with default parameters46. After analysis of the gene family, 416,469 genes from the 16 species were grouped into 30,677 gene families. The results revealed that 717 gene families that involved 4,217 genes were unique in S. nudus. The gene families and genome statistics of all the species are shown in Table 10. Among the orthologous genes in the 16 species, a total of 255 single-copy genes were identified. The single-copy orthologues were aligned using MUSCLE (v3.7)47 with default parameters, and then the aligned protein sequences were reverse translated into codon sequences. The alignments were then concatenated to generate a superalignment matrix for phylogenetic reconstruction based on the maximum-likelihood (ML) method using IQ-TREE (v1.6.12)48, with the best-fit evolutionary substitution model being determined using ModelFinder49. Divergence times for each node in the phylogenetic tree were estimated using MCMCtree, which is implemented in PAML package v4.8a50, under the following parameters: -nsample 100000, -rootage 800, and -burnin 500000. The calibration times were obtained from TimeTree51: 630.0–830.0 million years ago (Ma) for Caenorhabditis elegans and Homo sapiens, 424.2–440.0 Ma for Danio rerio and Homo sapiens, and 545.0–681.5 Ma for Capitella teleta and Crassostrea gigas. The phylogenetic tree representing the evolutionary relationships among Mollusca, Annelida and Sipuncula is shown in Fig. 9. Gene collinearity, which shows the preservation of ancestral genome structure in the modern genome, is an important means of unveiling genomic evolution. Thus, MCscan (Python version)52 was used for the genomic synteny analysis between S. nudus, O. fusiformis and P. esculenta. The collinearity figure was drawn based on the homologous blocks with ≥ 4 gene collinear pairs between species by JCVI (https://github.com/tanghaibao/jcvi) (Fig. 10). Regarding intergenomic gene collinearity, 109 blocks containing 508 collinear gene pairs were revealed between S. nudus and O. fusiformis, and 622 blocks containing 3248 collinear gene pairs were revealed between S. nudus and P. esculenta, showing similar collinearity between the two Sipuncula species.
The time-calibrated phylogenetic tree was used to assess gene family expansions and contractions using CAFÉ 4.2.153 with a random birth-and-death model with lambda. In total, 543 and 97 significantly expanded and contracted gene families were identified, respectively (P < 0.05). GO and KEGG enrichment analyses of the expanded gene families revealed that these families were mainly involved in pathways that are related to apoptosis, detoxification, the immune response, amino acid and fatty acid metabolism anion, oxidative stress, and energy metabolism.
PSGs (positively selected genes) were predicted using branch-site likelihood ratio tests for single-copy gene families with a conservative 10% false discovery rate (FDR) criterion54. We used proteins from S. nudus, C. teleta, E. Andrei, L. satsuma, O. fusiformis, and P. esculenta to extract 3,192 one-to-one orthologous genes using the OrthoFinder (v2.3.11) pipeline. The one-to-one orthologous genes were then used to generate multiple sequence alignments by using PRANK (v. 121002)55. The dN/dS ratios of the codons were calculated using the branch-site model of Codeml in the PAML package50, in which S. nudus was set as the foreground branch and the other five taxa as background branches. Using a likelihood ratio test (LRT) of ≤0.05 and an FDR of ≤0.05 as thresholds, 326 PSGs were identified in the S. nudus genome. These PSGs were significantly enriched in the terms “Spliceosome,” “Base excision repair,” “DNA replication,” and “Cell cycle” in the KEGG pathway enrichment analysis.
In summary, we obtained the high-quality chromosome-level genome of S. nudus, which contributes to our understanding of the evolutionary status of Sipuncula and the evolutionary relationship among the subgroups of the phylum Annelida. Gene family expansion and extraction and genomic synteny analyses revealed the potential adaptation mechanism of Sipuncula to different living environments.
Usage Notes
All analyses were run on Linux systems, and the optimal parameters are given in the Code availability section.
Code availability
No specific code or script was used in this work. Commands used for data processing were all executed according to the manuals and protocols of the corresponding software.
References
Cutler, E. B. The Sipuncula: Their Systematics, Biology, And Evolution (New York: Cornell University Press, https://doi.org/10.7591/9781501723643, 1994)
Nielsen, C. Some aspects of spiralian development. Acta Zool. 91, 20–28, https://doi.org/10.1111/j.1463-6395.2009.00421.x (2010).
Huang, D. Y., Chen, J. Y., Vannier, J. & Saiz Salinas, J. I. Early Cambrian sipunculan worms from southwest China. Proceedings of the Royal Society of London. Series B: Biological Sciences 271, 1671–1676, https://doi.org/10.1098/rspb.2004.2774 (2004).
Li, J. et al. Experimental study of bioturbation by Sipunculus nudus in a polyculture system. Aquaculture 437, 175–181, https://doi.org/10.1016/j.aquaculture.2014.12.002 (2015).
Li, J. et al. Bioturbation of peanut worms Sipunculus nudus on the composition of prokaryotic communities in a tidal flat as revealed by 16S rRNA gene sequences. MicrobiologyOpen 8, 1–12, https://doi.org/10.1002/mbo3.802 (2019).
Linnaeus, C. Systema Naturae Per Regna Tria Naturae, Secundum Classes, Ordines, Genera, Species, Cum Characteribus Differentiis, Synonymis, Locis (Editio duodecima, reformata. Holmiae, 1767).
Delle Chiaie, S. Memorie Sulla Storia Enotomia Degli Animali Senza Vertebre Del Regno Di Napoli Vol. 1–4 (Dalla stamperia de’Fratelli Fernandes, 1823).
Hyman, L. H. The Protostomatous Coelomates-Phylum Sipunculida. The Invertebrates. (New York: McGraw-Hill Press, 1959).
Scheltema, A. H. Aplacophora as progenetic aculiferans and the coelomate origin of mollusks as the sister taxon of Sipuncula. Biol Bull. 184, 57–78, https://doi.org/10.2307/1542380 (1993).
Shen, X., Ma, X., Ren, J. & Zhao, F. A close phylogenetic relationship between Sipuncula and Annelida evidenced from the complete mitochondrial genome sequence of Phascolosoma esculenta. BMC genomics 10, 1–11, https://doi.org/10.1186/1471-2164-10-136 (2009).
Wanninger, A., Koop, D., Bromham, L., Noonan, E. & Degnan, B. M. Nervous and muscle system development in Phascolion strombus (Sipuncula). Dev Genes Evol 215, 509–518, https://sci-hub.se/10.1007/s00427-005-0012-0 (2005).
Struck, T. H. et al. Phylogenomic analyses unravel annelid evolution. Nature 471, 95–98, https://sci-hub.se/10.1038/nature09864(2011).
Zhong, S. et al. The Draft Genome of Chinese endemic species Phascolosoma esculenta (Sipuncula, Phascolosomatidae) reveals the phylogenetic position of Sipuncula. Front. Genet. 13, 1–7, https://doi.org/10.3389/fgene.2022.910344 (2022).
Belton, J. M. et al. Hi-C: A comprehensive technique to capture the conformation of genomes. Methods 58, 268–276, https://doi.org/10.1016/j.ymeth.2012.05.001 (2012).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770, https://doi.org/10.1093/bioinformatics/btr011 (2011).
Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204, https://doi.org/10.1093/bioinformatics/btx153 (2017).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175, https://doi.org/10.1038/s41592-020-01056-5 (2021).
Roach, M. J., Schmidt, S. A. & Borneman, A. R. Purge Haplotigs: allelic contig reassignment for third-gen diploid genome assemblies. BMC bioinform. 19, 1–10, https://doi.org/10.1186/s12859-018-2485-7 (2018).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell syst. 3, 95–98, https://doi.org/10.1016/j.cels.2016.07.002 (2016).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95, https://doi.org/10.1126/science.aal3327 (2017).
Wang, Q. H., Du, Y. H., Lin, S. L., Du, X. & Deng, Y. Karyotype analysis of Sipunculus nudus. Chinese Journal of Zoology 43, 95–98, https://doi.org/10.13859/j.cjz.2008.05.012 (2008).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100, https://doi.org/10.1093/bioinformatics/bty191 (2018).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645, http://www.genome.org/cgi/doi/10.1101/gr.092759.109 (2009).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21, i351–i358, https://doi.org/10.1093/bioinformatics/bti1018 (2005).
Bergman, C. M. & Quesneville, H. Discovering and detecting transposable elements in genome sequences. Brief. bioinformatics 8, 382–392, https://doi.org/10.1093/bib/bbm048 (2007).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet Genome Res 110, 462–467, https://doi.org/10.1159/000084979 (2005).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 35, W265–W268, https://doi.org/10.1093/nar/gkm286 (2007).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinform. 12, 1–14, http://www.biomedcentral.com/1471-2105/12/491 (2011).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439, https://doi.org/10.1093/nar/gkl200 (2006).
Johnson, A. D. et al. SNAP: a web-based tool for identification and annotation of proxy SNPs using HapMap. Bioinformatics 24, 2938–2939, https://doi.org/10.1093/bioinformatics/btn564 (2008).
Kanehisa, M., Goto, S., Kawashima, S., Okuno, Y. & Hattori, M. The KEGG resource for deciphering the genome. Nucleic Acids Res. 32, D277–D280, https://doi.org/10.1093/nar/gkh063 (2004).
Boeckmann, B. et al. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Res. 31, 365–370, https://doi.org/10.1093/nar/gkg095 (2003).
Bairoch, A. & Apweiler, R. The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res. 28, 45–48, https://doi.org/10.1093/nar/28.1.45 (2000).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 25, 25–29, https://doi.org/10.1038/75556 (2000).
Jones, P. et al. InterProScan 5: genome scale protein function classification. Bioinformatics 30, 1236–1240, https://doi.org/10.1093/bioinformatics/btu031 (2014).
Mi, H. et al. PANTHER version 11: expanded annotation data from gene ontology and reactome pathways, and data analysis tool enhancements. Nucleic Acids Res. 45, D183–D189, https://doi.org/10.1093/nar/gkw1138 (2017).
Bateman, A. et al. The Pfam Protein Families Database. Nucleic Acids Res. 28, 263–266, https://doi.org/10.1093/nar/28.1.263 (2000).
Attwood, T. K. et al. PRINTS-S: the database formerly known as PRINTS. Nucleic Acids Res. 28, 225–227, https://doi.org/10.1093/nar/28.1.225 (2000).
Corpet, F., Gouzy, J. & Kahn, D. Recent improvements of the ProDom database of protein domain families. Nucleic Acids Res. 27, 263–267, https://doi.org/10.1093/nar/27.1.263 (1999).
Wilson, D. et al. SUPERFAMILY–sophisticated comparative genomics, data mining, visualization and phylogeny. Nucleic Acids Res. 37, D380–D386, https://doi.org/10.1093/nar/gkn762 (2009).
Schultz, J., Copley, R. R., Doerks, T., Ponting, C. P. & Bork, P. SMART: a web-based tool for the study of genetically mobile domains. Nucleic Acids Res. 28, 231–234, https://doi.org/10.1093/nar/28.1.231 (2000).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP408321 (2022).
Zheng, Z. Genome of Sipunculus nudus. China National GeneBank DataBase https://doi.org/10.26036/CNP0003624 (2022).
Zheng, Z. The first high-quality chromosome-level genome for the Sipuncula Sipunculus nudus using HiFi and Hi-C data. GenBank https://identifiers.org/ncbi/insdc:JAPPUL000000000 (2022).
Simao, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212, https://doi.org/10.1093/bioinformatics/btv351 (2015).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 1–14, https://doi.org/10.1186/s13059-019-1832-y (2019).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797, https://doi.org/10.1093/nar/gkh340 (2004).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274, https://doi.org/10.1093/molbev/msu300 (2015).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589, https://doi.org/10.1038/nmeth.4285 (2017).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591, https://doi.org/10.1093/molbev/msm088 (2007).
Kumar, S., Stecher, G., Suleski, M. & Hedges, S. B. TimeTree: A resource for timelines, timetrees, and divergence times. Mol. Biol. Evol. 34, 1812–1819, https://doi.org/10.1093/molbev/msx116 (2017).
Tang, H. et al. Synteny and collinearity in plant genomes. Science 320, 486–488, https://doi.org/10.1126/science.1153917 (2008).
Han, M. V., Thomas, G. W. C., Lugo-Martinez, J. & Hahn, M. W. Estimating gene gain and loss rates in the presence of error in genome assembly and annotation using CAFE 3. Mol. Biol. Evol. 30, 1987–1997, https://doi.org/10.1093/molbev/mst100 (2013).
Nielsen, R. et al. A scan for positively selected genes in the genomes of humans and chimpanzees. PLoS Biol. 3, el70, https://doi.org/10.1371/journal.pbio.0030170 (2005).
Loytynoja, A. & Goldman, N. Phylogeny-aware gap placement prevents errors in sequence alignment and evolutionary analysis. Science 320, 1632–1635, https://doi.org/10.1126/science.1158395 (2008).
Acknowledgements
This work was supported by the Science and Technology Department of Guangdong Province (Nos. 2021A05190 and 163-2019-XMZC-0009-02-0059).
Author information
Authors and Affiliations
Contributions
Zheng Z., Wang Q.H. and Deng Y.W. designed the study; Zheng Z., Lai Z.X., Wu B, Song X.L. and Zhao W. performed genome sequencing, data processing, and genome analysis; Zhang J.W., Jian J.B., Zhen Y. and Zhong R.Z. performed the assembly quality validation and improved gene annotation; Liao Y.S. and Yang C.Y. collected and prepared the samples; Zheng Z., Lai Z.X., and Wu B wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zheng, Z., Lai, Z., Wu, B. et al. The first high-quality chromosome-level genome of the Sipuncula Sipunculus nudus using HiFi and Hi-C data. Sci Data 10, 317 (2023). https://doi.org/10.1038/s41597-023-02235-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02235-7