Abstract
Bean flower thrips Megalurothrips usitatus is a staple pest of cowpea and other legumes and causes dramatic economic losses. Its small size allows for easy concealment, and large reproductive capacity easily leads to infestations. Despite the importance of a genome in developing novel management strategies, genetic studies on M. usitatus remain limited. Thus, we generated a chromosome-level M. usitatus genome using a combination of PacBio long read and Hi-C technologies. The assembled genome was 238.14 Mb with a scaffold N50 of 13.85 Mb. The final genome was anchored into 16 pseudo-chromosomes containing 14,000 genes, of which 91.74% were functionally annotated. Comparative genomic analyses revealed that expanded gene families were enriched in fatty acid metabolism and detoxification metabolism (ABC transporters), and contracted gene families were strongly associated with chitin-based cuticle development and sensory perception of taste. In conclusion, this high-quality genome provides an invaluable resource for us to understand the thrips’ ecology and genetics, contributing to pest management.
Similar content being viewed by others
Background & Summary
Bean flower thrips Megalurothrips usitatus is a highly harmful pest of leguminous crops in the genera Glycine, Arachis, and Vigna1,2,3,4. The insect lays eggs in plant tissue and feeds on leaves, flowers and pods, causing economic losses worldwide, particularly in southern China, India, Japan, the Philippines, and Australia1,3,5,6. Its small body size, cryptic behavior, and fast transmission present difficulties in pest control6,7.
Attempts to mitigate agricultural damage have largely involved chemical insecticides8,9,10,11,12. However, excessive pesticide usage leaves residues that risk consumer health and also induce resistance in pest insects. Understanding the evolution of pesticide resistance is necessary for developing novel management strategies, but the genetics of M. usitatus remains poorly understood. Filling this knowledge gap will benefit our efforts at pest control.
In this study, we assembled a chromosome-level genome of M. usitatus using a combination of PacBio long read, Illumina short-read sequencing, and chromosome conformation capture (Hi-C) technologies. We compared the genomic features of M. usitatus with those of other insects to explore the genomic signatures of resistance. The high-quality reference genome of the bean flower thrips obtained in this study will lay the foundation for future investigations on the ecology of thrips and provide valuable genetic information for its management.
Methods
Sample preparation and genomic DNA sequencing
Megalurothrips usitatus samples were collected from Wanning, Hainan province, and reared for approximately 100 generations in the laboratory. Adults were fed Lablab purpureus and kept at 25 ± 1 °C, 70 ± 5% relative humidity, and 14:10 light:dark cycle. Stages were confirmed under a light microscope and verified with pictorial keys13. Individuals were then quickly placed into collection tubes, flash-frozen in liquid nitrogen, and stored at −80 °C until use.
We prepared approximately 2,000 mixed-sex M. usitatus individuals for genome sequencing. Genomic DNA was extracted using the CTAB method, followed by purification using a Blood and Cell Culture DNA Midi Kit (QIAGEN, Germany). The purity and concentration of extracted DNA were determined with 0.75% agarose gel electrophoresis and a Qubit 2.0 Fluorometer (Thermo Fisher Scientific, USA), respectively. The library constructed from the extracted DNA was approximately 10–20 Kb in size. A PacBio Sequel sequencer (Pacific Biosciences, Menlo Park, USA) was used for long DNA fragments, and Illumina Novoseq 6000 was used to generate 150 bp paired-end short reads. The sequencing yielded 98.30 Gb (412.78 × coverage) of long-reads with an N50 length of 14,475 bp and an average length of 10,352.68 ± 2.46 bp (mean ± S.E.). The Illumina platform sequenced 58.80 Gb raw data, of which adapters and low-quality short reads were removed using Fastp version 0.21.014 with default parameters, resulting in a total of 55.86 Gb (234.57 × coverage) clean data (Table 1).
Hi-C library preparation and sequencing
Chromosome conformation capture (Hi-C) sequencing used fresh tissues from 1,500 mixed-sex M. usitatus individuals. The samples were cross-linked with a 2% formaldehyde isolation buffer and then treated with DpnII (NEB) to digest nuclei. Biotinylated nucleotides were used to repair the tails, and the ligated DNA was split into fragments of 350 bp in length. The resulting Hi-C library was sequenced in Illumina Novoseq. 6000 with paired-end 150 bp. After applying the same filter criteria for short reads, a total of 53.90 Gb (226.34 × coverage) of clean data was generated (Table 1).
Transcriptome sequencing
A pooled M. usitatus sample was prepared using 30 eggs, 20 pseudo-pupae, 10 females, and 10 males. Total RNA was extracted using the TRIzol reagent (Thermo Fisher Scientific, USA). A paired-end library was constructed using the TruSeq RNA Library Preparation Kit (Illumina, USA) and sequenced on an Illumina Novoseq 6000 platform. It resulted in a total of 5.61 Gb RNA-seq clean data (Table 1). Additionally, total RNA (1 µg) was used to construct a full-length transcript isoform library using the SMRT bell Express Template Prep Kit 2.0 (Pacific Biosciences, USA). Target-size sequences were generated using the PacBio sequel II platform. A total of 47.67 Gb full-length transcriptome data was obtained (Table 1).
Estimation of genomic characteristics
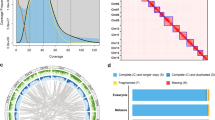
Genomic characteristics were determined based on 55.86 Gb of short-read data using a K-mer-based statistical analysis in JELLYFISH version 2.1.315 with the following parameters: ‘count -m 17 -C -c 7 -s 1 G -F 2’. Genome heterozygosity and genome size were estimated in GenomeScope version 2.016 with default parameters. Based on 17-mer depth analysis, genome size and heterozygosity were estimated to be 255.81 Mb and 0.85%, respectively (Fig. 1).
Genome assembly
We assembled a draft genome using wtdbg2 version 2.5 with default parameters17. We then had it polished using RACON version 1.4.1318 with parameters ‘-m 8 -x −6 -g −8 -w 500 -u’ and Pilon version 1.1419 with default parameters based on 98.30 Gb long reads and 55.86 Gb short reads.
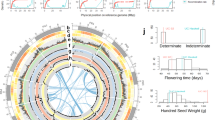
A scaffolding pipeline based on Durand (2016)20 was used to generate a high-quality chromosome-scale genome. Initially, Hi-C data were mapped to the contig assembly using BWA-MEM version 0.7.1721 with the following parameters: ‘mem -SP5M’. Next, the DpnII sites were generated using the ‘generate_site_positions.py’ script in Juicer version 1.520. The 3D-DNA pipeline (-r 2) was subsequently employed to order, orient, and cluster the contig22. After viewing Hi-C contact maps, the chromosome-scale genome was assembled in Juicebox version 1.11.08 (https://github.com/aidenlab/Juicebox). The genome assembly was screened for contaminant sequences by using the “Contamination in Sequence Databases” in NCBI. A total of 33 sequences were labeled as contaminant and removed (available in Figshare). To identify the mitochondrial genome, we amplified the cytochrome oxidase subunit 1 (COI) gene fragment with primer pairs LCO1490 and HCO2198, and obtained a DNA barcode sequence of approximately 610 bp23. We then used BLAST version 2.2.2824 (-evalue 1e-5) to find assembly sequences of a high similarity to the COI fragment (>98%), and identified one unplaced sequence (scaffold46) as mitochondrial sequence. The resulting chromosome-level genome was 238.14 Mb with a scaffold N50 of 13.85 Mb, maximum length of 20.88 Mb, and GC rate of 55.90% (Table 2). 91.89% of the genome was anchored to 16 pseudo-chromosomes (Table 2), which were well-distinguished from each other based on the chromatin interaction heatmap (Fig. 2).
Predicting repeats
Repeat sequences were annotated in Extensive de novo TE Annotator (EDTA) version 1.9.425. In brief, LTR retrotransposons were identified in LTR FINDER version 1.0726, LTRharvest27, and LTR retriever version 2.9.028 with default parameters. Next, TIR Learner29 and HelitronScanner30 were used to classify DNA transposons with default parameters. RepeatMasker version 4.0.7 (-gff -xsmall -no_is)31 and RepeatProteinMasker version 4.0.7 (-engine wublast) were utilized to identify repeat sequences based on RepBase edition 2017012732. Repeats were masked with de novo predictions using RepeatModeler version 2.0.1 with parameters ‘-engine ncbi -pa 4’. Additionally, Tandem Repeats Finder33 was used to annotate tandem repeats with parameters ‘2 7 7 80 10 50 500 -f -d -m’. Overall, 20.20% of the assembled genome was classified as repetitive sequences in the M. usitatus genome (Table 3). Tandem repeat elements were found to be the most abundant (8.42%), followed by the terminal inverted repeat category (5.39%) (Table 3).
Gene and functional predictions
Genes in the assembled genome were predicted using a combination of homology-based, transcriptome-based, and ab initio methods. Homology-based predictions involved downloaded sequences of peptides and transcripts from Aptinothrips rufus (http://v2.insect-genome.com/Organism/87), Frankliniella occidentalis (https://ftp.ncbi.nlm.nih.gov/genomes/all/GCF/000/697/945/GCF_000697945.3_Focc_3.1), and Thrips palmi (https://ftp.ncbi.nlm.nih.gov/genomes/all/GCF/012/932/325/GCF_012932325.1_TpBJ-2018v1). The IsoSeq version 3.4.0 workflow was utilized to generate 28,608 high-quality transcripts from the full-length transcriptome data, with quality parameters of 0.99 (https://github.com/PacificBiosciences/IsoSeq). Next, RNA-seq short data were mapped to the reference genome using HISAT2 version 2.2.134 with the parameter ‘-k 2’. The mapped reads were then assembled into transcripts using StringTie version 2.4.035 with default parameters. Homologous proteins and transcripts were aligned using Exonerate version 2.4.0 with default parameters to train the gene sets. Meanwhile, a sorted and mapped bam file of RNA-seq data was transferred to a hints file using the bam2hints program in AUGUSTUS version 3.2.336 with the parameter ‘–intronsonly’. The trained gene sets and hint files were combined as inputs for AUGUSTUS version 3.2.336, which predicted coding genes from the assembled genome with default parameters. Finally, homology-based, de novo-derived, and transcript genes were merged in MAKER version 2.31.10 to generate a high-confidence gene set37. It resulted in the annotation of 14,000 M. usitatus genes. The average transcript length was 2,243.30 bp with an average length of coding sequence (CDS) of 1,588.94 bp. The average exon number per gene was 7.38, and the average exon length was 303.85 bp (Table 4).
Gene structure and annotations were determined through several methods, including eggnog-mapper38 (-m diamond–tax_scope auto–go_evidence experimental–target_orthologs all–seed_ortholog_evalue 0.001–seed_ortholog_score 60–query-cover 20–subject-cover 0 –override), InterProscan version 5.039 (-iprlookup -goterms -appl Pfam -f TSV), BLAST version 2.2.2824 (-evalue 1e-5), and HMMER version 3.3.240 (–noali–cut_ga Pfam-A.hmm). These methods were used to search against multiple public databases, including NCBI non-redundant protein (Nr), Gene Ontology (GO), Clusters of Orthologous Groups of Proteins (COG), Kyoto Encyclopedia of Genes and Genomes (KEGG), Swiss-Prot, and Pfam. Most genes (91.74%) were successfully annotated with at least one public database (Table 5).
Comparative genomic analysis
To identify single-copy orthologous genes, we utilized the longest protein sequence of each gene from M. usitatus and multiple other species (Table 6), including F. occidentalis41, T. palmi42, Acyrthosiphon pisum43, Triatoma rubrofasciata44, Columbicola columbae45, Aedes aegypti46, Danaus plexippus47, Tribolium castaneum48, Apis mellifera49 and Daphnia galeata50. We performed all-to-all single-copy ortholog BLAST comparisons in OrthoFinder version 2.5.451 with the parameters ‘-a blast -M msa’. We aligned the resulting single-copy orthologous genes using MAFFT version 7.487 (–auto)52 and further trimmed the poorly aligned regions using Gblocks version 0.91b53 (-t = p -b4 = 5). We maintained the genes that met the stationary, reversible and homogeneous (SRH) assumptions54 using IQ-TREE version 2.2.055 with a p-value cut-off of 0.05. We finally obtained 1,573 single-copy genes under these criteria. Next, We used FASconCAT-G version 1.05.156 to concatenate the genes to form a supermatrix, which was used for subsequent phylogenetic analysis.
We performed a maximum likelihood analysis of concatenated sequences in IQ-TREE version 2.2.055 with 1,000 UFBoot replicates (–bb 1,000 –model JTT + I + G4). The minimum correlation coefficient for the convergence criterion was set at 0.99 (-bcor 0.99). The age of each node was estimated using a correlated rates clock in MCMCTREE of PAML version 4.457. To estimate the divergence times, we selected fossil records listed in Table 7.
Gene-family expansion and contraction were estimated using CAFÉ version 4.2 with parameters ‘lambda -s -t’, based on maximum likelihood and reduction methods58. Phylogenetic tree topology and branch lengths were considered when inferring the significance of changes to gene-family size in each branch. The results revealed 684 expanded gene families and 1,639 contracted gene families in M. usitatus (Fig. 3). Next, functional enrichment analysis (GO enrichment and KEGG pathway) was performed in KOBAS version 3.059. Significantly enriched GO terms were those with an adjusted p < 0.05 under Fisher’s exact test. Expanded gene families were enriched in cAMP signaling pathway, fatty acid metabolism, detoxification metabolism (ABC transporters) and the ionotropic glutamate receptor pathway (Fig. 4a, available in Figshare). Contracted gene families were enriched in chitin-based cuticle development, sensory perception of taste and NADP + activity (Fig. 4b, available in Figshare).
A time-calibrated phylogenetic tree inferred from 1,573 single-copy orthologs using IQ-TREE version 2.2.0 was shown. The upper panel in wheat represents Paraneoptera insects and the lower panel in light-blue represents Holometabola. The divergence between M. usitatus and F. occidentalis diverged 73.93 Mya (Million years ago). Bootstrap support values based on 1,000 replicates are equal to 100 (orange dot). The number of expanded (+red) and contracted (−blue) gene families are shown for each lineage.
(a) Expanded genes. (b) Contracted genes. Each row represents an enriched function, and the bar length represents the enrichment ratio (input gene number/background gene number). Bar colors represent different clusters. If any cluster has more than five terms, the top five with the highest enrichment ratio are displayed.
Data Records
Genomic PacBio sequencing data were deposited in the Sequence Read Archive at NCBI under accession number SRR2213748560.
Genomic Illumina sequencing data were deposited in the Sequence Read Archive at NCBI under accession SRR2213748261.
RNA-seq data were deposited in the Sequence Read Archive at NCBI under accession number SRR2213748462.
Full-length transcript isomer sequencing data were deposited in the Sequence Read Archive at NCBI under accession number SRR2213748363.
Hi-C sequencing data were deposited in the Sequence Read Archive at NCBI under accession number SRR2213748164.
The final chromosome assembly was deposited in GenBank at NCBI under accession number JAPTSV00000000065.
The contaminant file, single-copy orthologous genes, gene-family expansion and contraction, gene function annotation, and repeat annotation are available in Figshare66.
Technical Validation
DNA integrity
The integrity of extracted genomic DNA was determined using 0.75% agarose gel electrophoresis and analyzed with an Agilent 2100 Bioanalyzer (Agilent Technologies, USA). DNA concentration was measured using a Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, USA) and Qubit 2.0 (Thermo Fisher Scientific, USA). Absorbance at 260/280 nm was approximately 1.8.
Assessment of genome assemblies
We assessed the accuracy of the final genome assembly by mapping Illumina short reads to the M. usitatus genome with BWA-MEM version 0.7.1721. The analysis showed that 96.52% of short reads were successfully mapped to the M. usitatus genome (Table 8). We further assessed the base quality of genome assembly by estimating the quality value score (QVS) using Merqury version 1.167, which showed a high QVS of 32.65 (Table 8). These findings indicate that the quality of our assembled genome is high.
Furthermore, we evaluated the completeness of the final genome assembly using Benchmarking Universal Single-Copy Orthologs (BUSCO version 3.0.2) insecta_odb1068, which includes 1,367 orthologous genes. The analysis revealed a high completeness of 97.40% for the M. usitatus genome with only 0.60% of BUSCO genes being fragmented, 2.00% being missing, and 0.40% being duplicated (Table 8). These BUSCO results were comparable to the completeness for other thrips genomes, such as T. palmi (97.20%), F. occidentalis (98.50%), and A. rufus (95.00%) (Table 9).
Code availability
No specific codes or scripts were used in this study. All software used is in the public domain, with parameters clearly described in the Methods section.
References
Oparaeke, A. M. The sensitivity of flower bud thrips, Megalurothrips sjostedti Trybom (Thysanoptera: Thripidae), on cowpea to three concentrations and spraying schedules of Piper guineense Schum. & Thonn. extracts. Plant Prot. Sci. 42, 106, https://doi.org/10.17221/2757-PPS (2006).
Tillekaratne, K., Edirisinghe, J., Gunatilleke, C. & Karunaratne, W. Survey of thrips in Sri Lanka: a checklist of thrips species, their distribution and host plants. Ceylon J. Sci. 40, 89–89, https://doi.org/10.4038/cjsbs.v40i2.3926 (2011).
Tang, L.-D. et al. The life table parameters of Megalurothrips usitatus (Thysanoptera: Thripidae) on four leguminous crops. Fla. Entomol. 2, 620–625, https://doi.org/10.1653/024.098.0235 (2015).
Zafirah, Z. & Azidah, A. A. Diversity and population of thrips species on legumes with special reference to Megalurothrips usitatus. Sains Malays. 47, 433–439, https://doi.org/10.17576/jsm-2018-4703-02 (2018).
Duraimurugan, P. & Tyagi, K. Pest spectra, succession and its yield losses in mungbean and urdbean under changing climatic scenario. Legume Res. 37, 212–222, http://www.arccjournals.com/journal/legume-research-an-international-journal/LR-3023 (2014).
Yasmin, S., Ali, M., Rahman, M. M., Akter, M. S. & Latif, M. A. Biological traits of bean flower thrips, Megalurothrips usitatus (Thysanoptera: Thripidae) reared on mung bean. Herit. Sci. 5, 29–33, https://doi.org/10.26480/gws.02.2021.29.33 (2021).
Liu, P. et al. The male‐produced aggregation pheromone of the bean flower thrips Megalurothrips usitatus in China: identification and attraction of conspecifics in the laboratory and field. Pest Manage. Sci. 76, 2986–2993, https://doi.org/10.1002/ps.5844 (2020).
Peter, C. & Govindarajulu, V. Management of blossom thrips, Megalurothrips usitatus on pigeonpea. Int. J. Pest Manage. 36, 312–313, https://doi.org/10.1080/09670879009371495 (1990).
Hossain, M. A. Efficacy of some insecticides against insect pests of mungbean (Vigna radiata L.). Bangladesh J. Agric. Res. 40, 657–667, https://doi.org/10.3329/bjar.v40i4.26940 (2015).
Sujatha, B. & Bharpoda, T. Evaluation of insecticides against sucking pests grown during Kharif. Int. Curr. Microbiol. App. Sci. 6, 1258–1268, https://doi.org/10.20546/ijcmas.2017.610.150 (2017).
Yasmin, S., Latif, M., Ali, M. & Rahman, M. Management of thrips infesting mung bean using pesticides. SAARC J. Agric. 17, 43–52, https://doi.org/10.3329/sja.v17i2.45293 (2019).
Maradi, R. M. et al. Evaluation of bio-efficacy of newer molecules of different insecticides against thrips, Aphis craccivora in yard long bean, Vigna unguiculata subsp. sesquipedalis. J. Entomol. Zool. Stud. 15, 189–192, https://doi.org/10.55446/IJE.2021.360 (2020).
Khan, R., Seal, D. & Adhikari, R. Bean flower thrips Megalurothrips usitatus (Bagnall) (Insecta: Thysanoptera: Thripidae). EDIS, 1–7, https://doi.org/10.32473/edis-IN1352-2022 (2022).
Chen, S, Zhou, Y., Chen, Y. & Gu, J. Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, 884–890, https://doi.org/10.1093/bioinformatics/bty560 (2018).
Marcais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770, https://doi.org/10.1093/bioinformatics/btr011 (2011).
Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204, https://doi.org/10.1093/bioinformatics/btx153 (2017).
Ruan, J. & Li, H. Fast and accurate long-read assembly with wtdbg2. Nat. Methods 17, 155–158, https://doi.org/10.1038/s41592-019-0669-3 (2020).
Vaser, R., Sović, I., Nagarajan, N. & Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746, http://www.genome.org/cgi/doi/10.1101/gr.214270.116 (2017).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963, https://doi.org/10.1371/journal.pone.0112963 (2014).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98, https://doi.org/10.1016/j.cels.2016.07.002 (2016).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760, https://doi.org/10.1093/bioinformatics/btp324 (2009).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95, https://doi.org/10.1126/science.aal3327 (2017).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 3, 294–299 (1994).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinform. 10, 421–429, https://doi.org/10.1186/1471-2105-10-421 (2009).
Ou, S. et al. Benchmarking transposable element annotation methods for creation of a streamlined, comprehensive pipeline. Genome Biol. 20, 1–18, https://doi.org/10.1186/s13059-019-1905-y (2019).
Ou, S. & Jiang, N. LTR_FINDER_parallel: parallelization of LTR_FINDER enabling rapid identification of long terminal repeat retrotransposons. Mobile DNA 10, 48–48, https://doi.org/10.1186/s13100-019-0193-0 (2019).
Ellinghaus, D., Kurtz, S. & Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinform. 9, 1–14, https://doi.org/10.1186/1471-2105-9-18 (2008).
Ou, S. & Jiang, N. LTR_retriever: a highly accurate and sensitive program for identification of long terminal repeat retrotransposons. Plant Physiol. 176, 1410–1422, https://doi.org/10.1104/pp.17.01310 (2017).
Su, W., Gu, X. & Peterson, T. TIR-Learner, a new ensemble method for TIR transposable element annotation, provides evidence for abundant new transposable elements in the maize genome. Mol. Plant 12, 447–460, https://doi.org/10.1016/j.molp.2019.02.008 (2019).
Xiong, W., He, L., Lai, J., Dooner, H. K. & Du, C. Helitronscanner uncovers a large overlooked cache of Helitron transposons in many plant genomes. Proc. Natl. Acad. Sci. USA 111, 10263–10268, https://doi.org/10.1073/pnas.1410068111 (2014).
Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinformatics 5, 1–14, https://doi.org/10.1002/0471250953.bi0410s25 (2004).
Jurka, J. et al. Repbase update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467, https://doi.org/10.1186/s13100-015-0041-9 (2005).
Benso, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580, https://doi.org/10.1093/nar/27.2.573 (1999).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360, https://doi.org/10.1038/nmeth.3317 (2015).
Kovaka, S. et al. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol. 20, 1–13, https://doi.org/10.1186/s13059-019-1910-1 (2019).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, 435–439, https://doi.org/10.1093/nar/gkl200 (2006).
Cantarel, B. L. et al. MAKER: an easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res. 18, 188–196, http://www.genome.org/cgi/doi/10.1101/gr.6743907 (2008).
Cantalapiedra, C. P., Hernández-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggNOG-mapper v2: functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 38, 5825–5829, https://doi.org/10.1093/molbev/msab293 (2021).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240, https://doi.org/10.1093/bioinformatics/btu031 (2014).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, 29–37, https://doi.org/10.1093/nar/gkr367 (2011).
Rotenberg, D. et al. Genome-enabled insights into the biology of thrips as crop pests. BMC Biol. 18, 1–37, https://doi.org/10.1186/s12915-020-00862 (2020).
Guo, S. K. et al. Chromosome‐level assembly of the melon thrips genome yields insights into evolution of a sap‐sucking lifestyle and pesticide resistance. Mol. Ecol. Resour. 20, 1110–1125, https://doi.org/10.1111/1755-0998.13189 (2020).
Consortium, I. A. G. Genome sequence of the pea aphid Acyrthosiphon pisum. PLoS Biol. 8, e1000313, https://doi.org/10.1371/journal.pbio.3000029 (2010).
Liu, Q. et al. A chromosomal-level genome assembly for the insect vector for Chagas disease, Triatoma rubrofasciata. GigaScience 8, giz089, https://doi.org/10.1093/gigascience/giz089 (2019).
Baldwin-Brown, J. G. et al. The assembled and annotated genome of the pigeon louse Columbicola columbae, a model ectoparasite. G3 11, jkab009, https://doi.org/10.1093/g3journal/jkab009 (2021).
Nene, V. et al. Genome sequence of Aedes aegypti, a major arbovirus vector. Science 316, 1718–1723, https://doi.org/10.1126/science.1138878 (2007).
Mongue, A. J., Nguyen, P., Voleníková, A. & Walters, J. R. Neo-sex chromosomes in the monarch butterfly, Danaus plexippus. G3 7, 3281–3294, https://doi.org/10.1534/g3.117.300187 (2017).
Richards, S. et al. The genome of the model beetle and pest Tribolium castaneum. Nature 452, 949–955, https://doi.org/10.1038/nature06784 (2008).
Consortium, H. G. S. Insights into social insects from the genome of the honeybee Apis mellifera. Nature 443, 931–949, https://doi.org/10.1038/nature05260 (2006).
Nickel, J. et al. Hybridization dynamics and extensive introgression in the Daphnia longispina species complex: new insights from a high-quality Daphnia galeata reference genome. Genome Biol. Evol. 13, evab267, https://doi.org/10.1093/gbe/evab267 (2021).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 1–14, https://doi.org/10.1186/s13059-019-1832-y (2019).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780, https://doi.org/10.1093/molbev/mst010 (2013).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552, https://doi.org/10.1093/oxfordjournals.molbev.a026334 (2000).
Naser-Khdour, S., Minh, B. Q., Zhang, W., Stone, E. A. & Lanfear, R. The prevalence and impact of model violations in phylogenetic analysis. Genome Biol. Evol. 11, 3341–3352, https://doi.org/10.1093/gbe/evz193 (2019).
Nguyen, L.-T., Schmidt, H. A., Von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274, https://doi.org/10.1093/molbev/msu300 (2015).
Kück, P. & Longo, G. C. FASconCAT-G: extensive functions for multiple sequence alignment preparations concerning phylogenetic studies. Front. Zool. 11, 1–8, https://doi.org/10.1186/s12983-014-0081-x (2014).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591, https://doi.org/10.1093/molbev/msm088 (2007).
De Bie, T., Cristianini, N., Demuth, J. P. & Hahn, M. W. CAFE: a computational tool for the study of gene family evolution. Bioinformatics 22, 1269–1271, https://doi.org/10.1093/bioinformatics/btl097 (2006).
Bu, D. et al. KOBAS-i: intelligent prioritization and exploratory visualization of biological functions for gene enrichment analysis. Nucleic Acids Res. 49, 317–325, https://doi.org/10.1093/nar/gkab447 (2021).
NCBI Sequence Read Archive, https://identifiers.org/ncbi/insdc.sra:SRR22137485 (2022).
NCBI Sequence Read Archive, https://identifiers.org/ncbi/insdc.sra:SRR22137482 (2022).
NCBI Sequence Read Archive, https://identifiers.org/ncbi/insdc.sra:SRR22137484 (2022).
NCBI Sequence Read Archive, https://identifiers.org/ncbi/insdc.sra:SRR22137483 (2022).
NCBI Sequence Read Archive, https://identifiers.org/ncbi/insdc.sra:SRR22137481 (2022).
Ma, L., Liu, Q., Li, H. & Cai, W. Megalurothrips usitatusgenome sequencing and assembly, GenBank, https://identifiers.org/ncbi/insdc:JAPTSV000000000 (2022).
Ma, L. et al. Chromosome-level genome assembly of bean flower thrips Megalurothrips usitatus, Figshare, https://doi.org/10.6084/m9.figshare.c.6603697.v1 (2023).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 245, https://doi.org/10.1186/s13059-020-02134-9 (2020).
Simao, F. A., Waterhouse, R. M., loannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212, https://doi.org/10.1093/bioinformatics/btv351 (2015).
Mita, K. et al. The genome sequence of silkworm, Bombyx mori. DNA Res. 11, 27–35, https://doi.org/10.1093/dnares/11.1.27 (2004).
Adams, M. D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195, https://doi.org/10.1126/science.287.5461.2185 (2000).
Rehm, P. et al. Dating the arthropod tree based on large-scale transcriptome data. Mol. Phylogen. Evol. 61, 880–887, https://doi.org/10.1016/j.ympev.2011.09.003 (2011).
Wang, Y.-h. et al. Fossil record of stem groups employed in evaluating the chronogram of insects (Arthropoda: Hexapoda). Sci. Rep. 6, 38939, https://doi.org/10.1038/srep38939 (2016).
Krzeminski, W. & Krzeminska, E. Triassic Diptera: descriptions, revisions and phylogenetic relations. Acta Zool. Cracov. 46, 153–184 (2003).
Nikolajev, G. & Ren, D. The oldest fossil Ochodaeidae (Coleoptera: Scarabaeoidea) from the middle Jurassic of China. Zootaxa 2553, 65–68, https://doi.org/10.11646/zootaxa.2553.1.4 (2010).
Grimaldi, D. & Engel, M. S. Evolution of the Insects. (Cambridge University Press, 2005).
Nel, A. et al. The earliest known holometabolous insects. Nature 503, 257–261, https://doi.org/10.1038/nature12629 (2013).
Johnson, K. P. et al. Phylogenomics and the evolution of hemipteroid insects. Proc. Natl. Acad. Sci. USA 115, 12775–12780, https://doi.org/10.1073/pnas.1815820115 (2018).
Acknowledgements
We thank Prof. Wangpeng Shi and Dr. Mingyue Feng for their assistance with sample collection, and Prof. Feng Zhang and Dr. Yingqi Liu for their help with divergence-time estimation. This work was supported by the National Natural Science Foundation of China (No. 31922012), Sanya Yazhou Bay Science and Technology City (No. SYND-2022-04), and the 2115 Talent Development Program of China Agricultural University.
Author information
Authors and Affiliations
Contributions
H.L. and W.C. conceived the project. L.M. and Q.L. collected samples and extracted genomic nucleotides. L.M. and H.L. performed data analysis and wrote the manuscript. S.W., S.L., L.T., F.S. and Y.D. contributed to data analyses. All authors contributed to revising the manuscript. All authors have read and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ma, L., Liu, Q., Wei, S. et al. Chromosome-level genome assembly of bean flower thrips Megalurothrips usitatus (Thysanoptera: Thripidae). Sci Data 10, 252 (2023). https://doi.org/10.1038/s41597-023-02164-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02164-5
This article is cited by
-
Chromosome-level genome of spider Pardosa pseudoannulata and cuticle protein genes in environmental stresses
Scientific Data (2024)
-
A chromosome-level genome for the flower thrips Frankliniella intonsa
Scientific Data (2024)
-
Chromosome-level genome assembly of the flower thrips Frankliniella intonsa
Scientific Data (2023)