Abstract
A curated database of shark and ray biological data is increasingly necessary both to support fisheries management and conservation efforts, and to test the generality of hypotheses of vertebrate macroecology and macroevolution. Sharks and rays are one of the most charismatic, evolutionary distinct, and threatened lineages of vertebrates, comprising around 1,250 species. To accelerate shark and ray conservation and science, we developed Sharkipedia as a curated open-source database and research initiative to make all published biological traits and population trends accessible to everyone. Sharkipedia hosts information on 58 life history traits from 274 sources, for 170 species, from 39 families, and 12 orders related to length (n = 9 traits), age (8), growth (12), reproduction (19), demography (5), and allometric relationships (5), as well as 871 population time-series from 202 species. Sharkipedia relies on the backbone taxonomy of the IUCN Red List and the bibliography of Shark-References. Sharkipedia has profound potential to support the rapidly growing data demands of fisheries management, international trade regulation as well as anchoring vertebrate macroecology and macroevolution.
Measurement(s) | Life history traits • Abundance time-series |
Technology Type(s) | literature review |
Factor Type(s) | Species ID • Location • Year |
Sample Characteristic - Organism | Class Chondrichthyes |
Sample Characteristic - Environment | All aquatic environments |
Sample Characteristic - Location | Global |
Similar content being viewed by others
Background & Description
“The sea has always challenged [our] minds and imagination and even today it remains the last great frontier of Earth”, a quote from Rachel Carson1, speaks to the fascination humans hold for life under the surface, as well as the difficulty in learning its secrets.
Although the sea and its residents are mysterious and diverse, scientific research into the macroecology and macroevolution of fishes has revealed a diversity of life history traits and strategies with deep convergence in their allometric and trait relationships2,3. For example, we know that fishes – to a first approximation – are cubes4, that there are three main dimensions of fish life histories5,6, and that key life history traits such as natural mortality and generation length are shaped by body size and environmental temperature7,8,9. These patterns have been found through comparative analyses of life history trait information (much of which has been compiled in FishBase, one of the first and most extensive bioinformatic databases for fishes10). We have learned this wide range of life history traits and strategies underlie a surprisingly narrow range of population dynamics11,12,13,14,15. Humans’ long history of exploitation of fishes has yielded numerous time-series of abundance of varying quality that are used, inter alia among other things, to develop fisheries stock assessments16, track the impacts of climate change17, and assess the extinction risk status of species, for example through IUCN Red List of Threatened Species18. Time-series of abundance can only be fully understood in the context of life histories. For example, in simple stock assessments the intrinsic population growth rate (r) is negatively related to the ecological carrying capacity (K), thus requiring an understanding of life histories to refine the set of plausible model scenarios19,20. While there are many vertebrate trait databases21,22 and some population trend databases23,24, we are not aware of any database that offers both traits and trends aligned to the same taxonomy.
For species with high evolutionary diversity and conservation concern, combining information on species traits with information on their population trends through time offer a unique resource for interdisciplinary science15. These data are key to prioritizing species for conservation actions through Red List assessments, managing species for sustainable fisheries, developing proposals for listing species on conservation and international trade regulations18,25,26. Further, data deficiency is a perennial problem that can be increasingly solved by imputing life histories27,28 and ‘borrowing’ information from life histories and time-series of data-rich species to infer the trajectories of data-poor species using ‘Robin Hood’ methods15,29. These methods are needed to solve key problems for fish lineages with high risk of extinction that suffer from considerable data paucity, including many shark and ray species15,30,31.
Sharks, rays, and chimaeras (class Chondrichthyes, hereafter “sharks and rays”) are one of the three classes of fishes and one of the seven classes of vertebrates32. Sharks and rays represent a pivotal point in the evolution and radiation of all vertebrate life for three reasons. First, they are the most evolutionary distinct radiation of vertebrates and represent an important lineage of jawed vertebrates (Gnathostomes) going back ~450 MY33. Second, they are the earliest radiation that exhibits the archetype of the vertebrate brain defined by the appearance of the first true cerbellum34,35. Third, while sharks and rays do not provide post-partum parental care they exhibit the greatest reproductive diversity among vertebrates; ranging in the degree of maternal investment from egg-laying to live-bearing, including oophagy, intra-uterine cannibalism, and placentotrophy36,37. This diversity of morphology, physiology, and life history makes contemporary chondrichthyans an important component of marine ecosystems38.
Many unaddressed questions in shark and ray ecology, evolution, and conservation can be best answered with a comparative approach39. Motivated by the success of FishBase and the need to continually update the taxonomy of our own personal comparative datasets, we sought a solution to set these data free both to facilitate reproducible science and to make them available for a wider scientific community. Furthermore, more than one third of sharks and rays are threatened with extinction, making them the second most threatened vertebrate lineage after amphibians33. Hence, fisheries management and conservation policy processes require access to comparative data to undertake stock assessments, ecological risk assessments, IUCN Red List assessments, and develop advisory materials for conservation policies and agreements40,41,42.
In this Data Descriptor, we introduce Sharkipedia: a curated open-source database designed for the continued updating of life history trait information and population time-series for all sharks and rays across all oceans. Sharkipedia currently hosts information on 58 life history measures related to length (n = 9), age (8), growth (12), reproduction (19), demography (5), and allometric relationships (5) assembled from 264 literature sources. Further, Sharkipedia archives 871 population time-series from 202 species and includes a browser feature to aid the selection of time-series for compilation of biodiversity indicators. This research initiative aims to make all published biological traits and population trends accessible to everyone and accelerate shark and ray research and conservation.
Methods
The data are held within the Sharkipedia database (www.sharkipedia.org)43, which is designed to hold population/stock level information on the life history traits and population abundance trends for all extant chondrichthyans. The database consists of several associated relational data frames: (1) a structural backbone of chondrichthyan taxonomy and references each uniquely identified by AuthorYear coding and linked to an external valid reference database (the Shark-References database, www.shark-references.com)44, (2) life history traits database (i.e. Traits database), and (3) population abundance trends database (i.e. Trends database) (Fig. 1). The most current taxonomic checklist was compiled from a recently published chondrichthyan phylogeny, including recent name changes and species description (www.vertlife.org/sharktree33, global checklists45, and the IUCN Red List40). The Traits and Trends databases are both related through the taxonomy and reference databases (Fig. 1) so these data streams can easily be brought together for comparative analysis. The specific methods for compiling the traits and trends databases are discussed below.
Traits
The initial schema for the Traits database (Fig. 1) was developed at a workshop in 2018, bringing together eight experts on shark and ray life history traits. The schema incorporated three requirements for data to be included in the final database: a valid reference ID, valid taxonomic name, and location information. The Traits database contains two core data tables (Observations and Measurements) used to maintain data traceability, ensure accuracy, avoid duplication, and provide a framework for quality control. Observations include metadata associated with a reference–author, year, DOI (Digital Object Identifier)–and are identified with a unique reference code. Measurements are tied to the Observation and include the information associated with a trait – species, sex, trait class, trait name, value, precision, and method (Fig. 1Observation and Measurement sections). Location information is coded by Marine Ecoregion44 and latitude and longitude when available. Retaining the spatial information for traits will capture interspecific variation in traits with geography. Measurements must be coded by ‘trait class’ (e.g., length, age, growth, reproduction, demography, and relationships), which will allow for the selection of appropriate traits (e.g., the “trait class” of growth can be populated by growth model parameters [k, L∞, L0] but not uterine fecundity). Methodological information is retained, for example, for growth trait Measurements the model used to estimate parameters is recorded to allow for filtering, selection, or as a covariate in comparative analyses (e.g., two- or three-parameter von Bertalanffy or Gompertz). Estimates of precision (range, standard error, standard deviation) can be recorded when available. Confidence in Measurements can be noted when there are known biases, for example incomplete size ranges for age and growth parameter estimation. Finally, new data can be embargoed by researchers prior to publication, to be released by editors upon researcher approval.
Trends
Time-series data on abundance were gathered from both peer-reviewed publications and grey literature, such as government reports. Further detail on the selection and use of time-series can be found here18,46. An Observation represents the source of the time-series (fishery-dependent [scientific survey, observer], fisheries, stock-assessment, other) and the trends type (Nominal or Standardized, Catch or Sightings per Unit Effort (CPUE), or abundance, or biomass), where CPUE is ‘Standardized’ by modeling Nominal data with covariates. Measurement represents the information associated with trend values – units, transformation, model used – and are tied to the Observation (Fig. 1). Location information is coded by Marine Ecoregion47,48, FAO Major Fishing Area, Ocean basin, and latitude and longitude (Fig. 3). Location information was entered using the map of the sampled area when available, and indicated in the metadata of the Observation. When a map was missing, location was determined by any information found in the text (e.g., latitude/longitude, name of locations, etc.) of the document presenting the data or searched online in the case of scientific survey. Numerical data on abundance are not always reported in documents and are often displayed graphically. We extracted the data from document figures with a digitization software commonly used by scientists (Webplotdigitizer)49. The software allows the import of a plot, asks for calibration of axes by clicking known values and then interpolates a coordinate system. Several automatic data extraction functions can be used, and each data point can manually be adjusted (using a magnification between 5 and 10 depending on the plot quality). In addition to trend values, the presence of associated estimates of precision (range, standard error, standard deviation) in the original document were noted.
Data Records
All datasets are available for viewing, downloading, and contributions through the Sharkipedia database (www.sharkipedia.org) and via Zenodo50.
Traits
We have compiled data from 264 sources, for 170 species (14% of chondrichthyan diversity) from 39 families (65%) and 12 orders (85%) (Fig. 2a). 155 species contain multiple Measurements from distinct populations and/or trait classes. Currently available data is unevenly distributed with more data available for the family of Requiem Sharks (Carcharhinidae) and orders Mackerel Sharks (Lamniformes), and Rhinorays (Rhinopristiformes). Length Measurements are the most common type with >1700 currently available, a large proportion of which are the observed maximum length (>700 Measurements; Fig. 2b).
Data Summary. Summary of traits and trends data available in the databases. (a) The taxonomic distribution of data on life history traits: (L) Length, (A) Age, (G) Growth, (R) Reproduction, (D) Demography, (C) Relationships, and (T) Trends. (b) The number of measurements within each trait class. (c) The summary of taxonomy and length of available time-series. Color of time-series indicates taxonomic order corresponding to panel a.
Trends
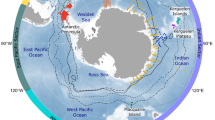
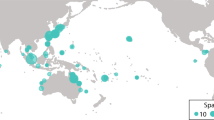
We have compiled 871 population time-series data from 337 sources (165 published and 172 unpublished), for 202 species (17% of chondrichthyan diversity), from 42 families (70%), and 14 orders (100%) (Fig. 2c). Currently available data is not evenly distributed spatially or taxonomically with more data available in the North Atlantic (e.g., Gulf Stream and North Atlantic Current) and the Southern Subtropical Front, due to the number of time-series from southern Australia (Fig. 3), and more data available for the Carcharhinidae and Rajidae families. Fishery independent data are the most common with 596 time-series currently available, and 64 time-series from stock-assessment (the highest quality of data as it integrates the catch history, abundance trends and life-history information to infer population dynamics).
Technical Validation
The database is curated on a volunteer basis, including database administrators and editors (https://www.sharkipedia.org/procedure). Volunteer curators are approved based on taxonomic or subject area expertise. Quality control of data includes:
Contributor approval
To contribute data, users must create a login ID to track data entry. Following sign-up users can contribute data in bulk (e.g., multiple Observations) using templates provided to ensure correct data formatting (https://www.sharkipedia.org/imports/new). Single Observation entries can be uploaded using the web entry GUI (Traits https://www.sharkipedia.org/observations/new; Trends https://www.sharkipedia.org/trends/new). When data is uploaded, an automated check is conducted to ensure basic requirements are met (valid reference, valid taxon, no required values missing). If basic requirements are not met, the user will be notified immediately of the specific locations (row and cell) of any issues that must be addressed. If basic requirements are met the initial upload is approved.
Editorial approval
Editors are notified when new Observations are uploaded. Editors conduct QA/QC on uploaded data to ensure all data requirements are met, and will check the accuracy of data including location, species, sex, value, and precision. An Editor can reject the upload if there are major issues (e.g., incomplete data entry, missing Measurements) with uploaded data. If revisions are required, the Editor can send data back to the contributor with the necessary changes required. Once QA/QC has been completed and all requirements are met the Editor can approve and import the data into the master database. These series of checks ensure that data is thoroughly vetted prior to final import. If issues are noted by users after the final import, core Administrators can manually correct Measurements.
User feedback
Continued population of the database will be Editor and user driven. Users can report any data issues directly to administrators (https://www.sharkipedia.org/contact).
Usage Notes
The data are available in csv-formatted files and can be accessed via bulk downloads or through species- or trait-specific queries. Instructions for data upload or use can be found online (https://www.sharkipedia.org/procedure).
Code availability
The code for the creation and deployment of the database can be found online at www.github.com/sharkipedia/sharkipedia.
References
Carson, R. The Sea Around Us. Oxford University Press, Oxford, UK 1951.
Beverton, R. J. H. & Holt, S. J. A review of the lifespans and mortality rates of fish in nature, and their relation to growth and other physiological characteristics. In: Ciba Foundation Symposium - The Lifespan of Animals (Colloquia on Ageing, Vol. 5) 142–180 (John Wiley & Sons, Ltd, 2008).
Kiørboe, T., Visser, A. & Andersen, K. H. A trait-based approach to ocean ecology. ICES Journal of Marine Science 75, 1849–1863 (2018).
Froese, R. Cube law, condition factor and weight-length relationships: History, meta-analysis and recommendations. Journal of Applied Ichthyology 22, 241–253 (2006).
Juan-Jordá, M. J., Mosqueira, I., Freire, J. & Dulvy, N. K. Life in 3-D: Life history strategies in tunas, mackerels and bonitos. Reviews in Fish Biology and Fisheries 23, 135–155 (2012).
Beukhof, E. et al. Marine fish traits follow fast-slow continuum across oceans. Scientific Reports 9 (2019).
Pauly, D. Tropical fishes: patterns and propensities. Journal of Fish Biology 53, 1–17 (1998).
Munch, S. B. & Salinas, S. Latitudinal variation in lifespan within species is explained by the metabolic theory of ecology. Proceedings of the National Academy of Sciences 106, 13860–13864 (2009).
Gislason, H., Daan, N., Rice, J. C. & Pope, J. G. Size, growth, temperature and the natural mortality of marine fish. Fish and Fisheries 11, 149–158 (2010).
Froese, R. & Pauly, D. FishBase https://fishbase.org/ (2021).
Winemiller, K. O. & Rose, K. A. Patterns of life-history diversification in North American Fishes: implications for population regulation. Canadian Journal of Fisheries and Aquatic Sciences 49, 2196–2218 (1992).
Cortés, E. Life History patterns and correlations in sharks. Reviews in Fisheries Science 8, 299–344 (2000).
Juan-Jordá, M. J., Mosqueira, I., Freire, J., Ferrer-Jordá, E. & Dulvy, N. K. Global scombrid life history data set. Ecology 97, 809–809 (2016).
Kindsvater, H. K., Mangel, M., Reynolds, J. D. & Dulvy, N. K. Ten principles from evolutionary ecology essential for effective marine conservation. Ecology and Evolution 6, 2125–2138 (2016).
Kindsvater, H. K. et al. Overcoming the data crisis in biodiversity conservation. Trends in Ecology & Evolution 33, 676–688 (2018).
Ricard, D., Minto, C., Jensen, O. P. & Baum, J. K. Examining the knowledge base and status of commercially exploited marine species with the RAM Legacy Stock Assessment Database. Fish and Fisheries 13, 380–398 (2011).
Maureaud, A. et al. Are we ready to track climate‐driven shifts in marine species across international boundaries? ‐ A global survey of scientific bottom trawl data. Global Change Biology 27, 220–236 (2020).
Sherley, R. B. et al. Estimating IUCN Red List population reduction: JARA-A decision‐support tool applied to pelagic sharks. Conservation Letters 13 (2019).
McAllister, M. K., Pikitch, E. K. & Babcock, E. A. Using demographic methods to construct Bayesian priors for the intrinsic rate of increase in the Schaefer model and implications for stock rebuilding. Canadian Journal of Fisheries and Aquatic Sciences 58, 1871–1890 (2001).
Froese, R., Demirel, N., Coro, G. & Kleisner, K. M. & Winker, H. Estimating fisheries reference points from catch and resilience. Fish and Fisheries 18, 506–526 (2016).
Jones, K. E. et al. PanTHERIA: a species-level database of life history, ecology, and geography of extant and recently extinct mammals. Ecology 90, 2648–2648 (2009).
Oliveira, B. F., São-Pedro, V. A., Santos-Barrera, G., Penone, C. & Costa, G. C. AmphiBIO, a global database for amphibian ecological traits. Scientific Data 4 (2017).
Inchausti, P. & Halley, J. Investigating Long-Term Ecological Variability Using the Global Population Dynamics Database. Science 293, 655–657 (2001).
Collen, B. et al. Monitoring change in vertebrate abundance: the Living Planet Index. Conservation Biology 23, 317–327 (2009).
Thorson, J. T., Munch, S. B., Cope, J. M. & Gao, J. Predicting life history parameters for all fishes worldwide. Ecological Applications 27, 2262–2276 (2017).
Heinicke, S. et al. Advancing conservation planning for western chimpanzees using IUCN SSC A.P.E.S.-the case of a taxon-specific database. Environmental Research Letters 14, 064001 (2019).
Horswill, C. et al. Global reconstruction of life‐history strategies: A case study using tunas. Journal of Applied Ecology 56, 855–865 (2019).
Thorson, J. T. Predicting recruitment density dependence and intrinsic growth rate for all fishes worldwide using a data‐integrated life‐history model. Fish and Fisheries 21, 237–251 (2019).
Brown, C. J. & Roff, G. Life-history traits inform population trends when assessing the conservation status of a declining tiger shark population. Biological Conservation 239, 108230 (2019).
Walls, R. H. L. & Dulvy, N. K. Eliminating the dark matter of data deficiency by predicting the conservation status of Northeast Atlantic and Mediterranean Sea sharks and rays. Biological Conservation 246, 108459 (2020).
Guy, C. S. et al. A paradoxical knowledge gap in science for critically endangered fishes and game fishes during the sixth mass extinction. Scientific Reports 11 (2021).
Compagno, L. J. V. Alternative life-history styles of cartilaginous fishes in time and space. In Alternative life-history styles of fishes 33–75 (Springer Netherlands, 1990).
Stein, R. W. et al. Global priorities for conserving the evolutionary history of sharks, rays and chimaeras. Nature Ecology & Evolution 2, 288–298 (2018).
Yopak, K. E. et al. A conserved pattern of brain scaling from sharks to primates. Proceedings of the National Academy of Sciences 107, 12946–12951 (2010).
Mull, C. G., Yopak, K. E. & Dulvy, N. K. Maternal Investment, Ecological Lifestyle, and Brain Evolution in Sharks and Rays. The American Naturalist 195, 1056–1069 (2020).
Mull, C. G., Pennel, M. W., Yopak, K. E. & Dulvy, N. K. Maternal investment evolves with larger body size and higher diversification rate in sharks and rays. BioRxiv TBC (2022).
Dulvy, N. D. & Reynolds, J. D. Evolutionary transitions among egg-laying, live-bearing, and maternal inputs in sharks and rays. Proceedings of the Royal Society B: Biological Sciences 264, 1309–1315 (1997).
Heithaus, M. R. et al. Advances in our understanding of the ecological importance of sharks and their relatives. In: Biology of sharks and their relatives, 3rd Ed. Carrier, J. C., Simpfendorfer, C. A., Heithaus, M. R., & Yopak, K. E. (Ed).
Simpfendorfer, C. A., Heupel, M. R., White, W. T. & Dulvy, N. K. The importance of research and public opinion to conservation management of sharks and rays: a synthesis. Marine and Freshwater Research 62, 518 (2011).
Dulvy, N. K. et al. Overfishing drives over one-third of all sharks and rays toward a global extinction crisis. Current Biology 31, 4773–4787.e8 (2021).
Cortés, E., Brooks, E. N. & Shertzer, K. W. Risk assessment of cartilaginous fish populations. ICES Journal of Marine Science 72, 1057–1068 (2014).
D’Alberto, B. M., Carlson, J. K., Pardo, S. A. & Simpfendorfer, C. A. Population productivity of shovelnose rays: Inferring the potential for recovery. PLOS ONE 14, e0225183 (2019).
Sharkipedia: elasmobranch traits & trends http://www.sharkipedia.org.
Bibliography Database. Shark-References http://www.shark-references.com.
Weigmann, S. Annotated checklist of the living sharks, batoids and chimaeras (Chondrichthyes) of the world, with a focus on biogeographical diversity. Journal of Fish Biology 88, 837–1037 (2016).
Pacoureau, N. et al. Half a century of global decline in oceanic sharks and rays. Nature 589, 567–571 (2021).
Spalding, M. D. et al. Marine Ecoregions of the World: A Bioregionalization of Coastal and Shelf Areas. BioScience 57, 573–583 (2007).
Spalding, M. D. et al. Pelagic provinces of the world: A biogeographic classification of the world’s surface pelagic waters. Ocean & Coastal Management 60, 19–30 (2012).
Rohatgi, A. WebPlotDigitizer. Extract data from plots, images, and maps https://automeris.io/WebPlotDigitizer/.
Mull, C. G. et al. Sharkipedia: A database of shark and ray life history traits and abundance time-series. Zenodo https://doi.org/10.5281/zenodo.6656525 (2012).
Acknowledgements
We thank H. Yan for additional data-mining of shark trends, C.L. Rigby for time-series checking, B. D’Alberto for traits data-mining, for M-J. Juan-Jordá for interpretation of stock assessments, and J. Pollerspöck for assisting with relational references. CM acknowledges support from M.A. MacNeil. This work was supported by both a Small Project Grant to CGM and HKK and a Project Grant to NKD from the Shark Conservation Fund, a philanthropic collaborative that pools expertise and resources to meet the threats facing the world’s sharks and rays. The Shark Conservation Fund is a project of Rockefeller Philanthropy Advisors. Further support was provided by a sponsorship from the Save Our Seas Foundation to CGM. Additional funding was provided by the US National Science Foundation grant DEB-1556779 to HKK, and the Department of Fish and Wildlife Conservation at Virginia Tech. NKD was supported by the Discovery and Accelerator grants from Natural Science and Engineering Research Council and the Canada Research Chair program.
Author information
Authors and Affiliations
Contributions
C.G.M., N.P., S.A.P., H.K.K. and N.K.D. conceived and led the project. C.G.M., N.P., S.A.P., L.S.R., E.G.R., B.F., A.H., H.K.K. and N.K.D. designed the project. C.G.M., N.P., S.A.P., L.S.R., E.G.R., B.F., A.H., A.J., W.V. and J.S.Y. acquired or provided data. C.G.M., N.P., M.H., H.K.K. and N.K.D. contributed essential code and analyses. C.G.M., N.P., H.K.K. and N.K.D. drafted and revised the paper. All authors provided edits and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mull, C.G., Pacoureau, N., Pardo, S.A. et al. Sharkipedia: a curated open access database of shark and ray life history traits and abundance time-series. Sci Data 9, 559 (2022). https://doi.org/10.1038/s41597-022-01655-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-022-01655-1