Abstract
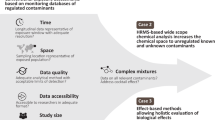
Direct monitoring of chemical concentrations in different environmental and biological media is critical to understanding the mechanisms by which human and ecological receptors are exposed to exogenous chemicals. Monitoring data provides evidence of chemical occurrence in different media and can be used to inform exposure assessments. Monitoring data provide required information for parameterization and evaluation of predictive models based on chemical uses, fate and transport, and release or emission processes. Finally, these data are useful in supporting regulatory chemical assessment and decision-making. There are a wide variety of public monitoring data available from existing government programs, historical efforts, public data repositories, and peer-reviewed literature databases. However, these data are difficult to access and analyze in a coordinated manner. Here, data from 20 individual public monitoring data sources were extracted, curated for chemical and medium, and harmonized into a sustainable machine-readable data format for support of exposure assessments.
Measurement(s) | occurrence of chemicals in environmental and biological media |
Technology Type(s) | mass spectrometry (curation of public data) |
Sample Characteristic - Organism | Homo sapiens • aquatic invertebrates • aquatic vertebrates/mammals • birds • fish • terrestrial invertebrates/worms • terrestrial vertebrates • livestock |
Sample Characteristic - Environment | surface water • drinking water • groundwater • ambient air • indoor air • indoor dust • landfill leachate • personal air • precipitation • wastewater • soil • sediment • sludge • consumer products • food products • raw agricultural commodities |
Similar content being viewed by others
Background & Summary
Chemical exposure can be defined as the degree of contact between a chemical and a human or ecological target receptor (i.e., the person, population, or thing that is being exposed). EPA’s Exposure Forecasting (ExpoCast)1 project is charged with collecting exposure-relevant information for thousands of chemicals. This information feeds integrated datasets and predictive models that support risk-related decisions. The gold-standard method for quantifying occurrence to support exposure assessment is the analytical measurement of a chemical in the fluids or tissues of an organism (biomonitoring) or in an environmental medium such as air, water, or soil (environmental monitoring). These data, known collectively as chemical monitoring data, are used to assess exposures and ultimately risks in research and regulatory applications.
Despite their value, there are many challenges associated with the collection and use of chemical monitoring data. One key issue is that data are generated by many different government, academic, and commercial bodies, with each institution having unique methods of analysis and reporting. These differences contribute to variations in data quality and formatting, which complicates data synthesis. Chemical synonymy is a notable challenge; chemicals may be reported under many different names and associated identifiers including Chemical Abstract Service (CAS) Registry Numbers, European Community (EC) numbers etc., making it difficult to correctly assemble all related data for a given substance. Another notable challenge is data sparsity; biological and environmental monitoring studies are expensive and time consuming, and monitoring data simply do not exist for many chemicals used in commerce (tens-of-thousands). Many exposure assessments instead rely on predictive models that consider chemical uses, releases, and fate and transport. A recent report from the National Academies of Sciences on improving risk-related evaluations2 emphasizes the need to integrate measured and modelled data to improve confidence in exposure assessments.
The goal of the current effort is to develop a harmonized and well-curated (in terms of the specific chemicals and media in which they were monitored) database of chemical monitoring data to support predictive modelling efforts and efficient exposure assessments. This manuscript describes the collection and curation of a large amount of publicly available chemical monitoring data from various sources. The scope of the current effort includes data and reports made publicly available on the web by government agencies, academic groups, or others. A de novo search of the open literature was not performed here and is the focus of ongoing work. The general approach used in this study was to download (either manually or using standard scripting methods) individual data records and to compile them into a database containing both the raw data (i.e., stored using the original variable names from the individual sources) and a harmonized version. In the harmonized version the raw data were curated and assigned new standard variable names. The harmonized variables include, for example, information that describes the chemicals monitored; the media (i.e., the type of biological or environmental sample) in which chemicals were measured; temporal and geographic information; and analytical results including any reported concentrations, instrumentation information, detection/quantification limits, or quality assurance (QA) data.
These data have several potential uses. The first and primary use is to provide an accessible resource for drawing existing monitoring data into chemical assessments. The database described here provides a means to search existing data using standardized media names and chemical identifiers. This allows for the efficient development of geographic or temporal summaries when needed. In addition, the database will support development of data-driven exposure models. New data mining, cheminformatic, and machine-learning techniques have the potential to extract meaningful patterns from large chemical datasets. One initial application of the data described herein is the development of machine learning models for estimating the likelihood of occurrence of any chemical in a medium, based on the chemical’s structure and/or known use(s). The harmonized monitoring data provides a rich training set for such models. These data can also provide evaluation information for existing process-based models of chemical release, fate, and transport.
Methods
The Multimedia Monitoring Database (MMDB) was compiled from existing reputable monitoring databases using a combination of automated and manual curation approaches. Datasets that are currently included in the database are listed in Table 1. These data sources were readily available monitoring databases that were reviewed to confirm that they met the following criteria:
-
1.
Accuracy-reliability – Source is reputable, defined as government entity, or an entity with documented credentials regarding a particular topic.
-
2.
Applicability – Source contains quantitative monitoring data for environmental samples and not spiked samples for the purpose of method validation or development.
-
3.
Representativeness – Sample size must be greater than 5 measurements. Preference for data based on large surveys or studies as opposed to case studies or studies based on 1 or a few sites.
-
4.
Accessibility- Datasets must be freely available, generally on publicly available websites, and preferably “FAIR” data3.
Data sources were categorized as either “single-sample”, where each record was a single analytical measurement, or “summary” (or “aggregate”), where each record was a summary metric (e.g., a mean, median, or specific percentile) for a group of measurements.
Data were collected from these data sources in three data collection phases (dictated by EPA funding cycles and generally occurring during the years 2017, 2018, and 2019). Some sources were simply updated in the 2018 and 2019 cycles and while some sources were newly added. Both automated and manual processes were used to curate unique data sources into the format of the multimedia monitoring database. Each data source was unique in format. Where necessary, data were obtained from the original sources using R or Python scripts. Biomonitoring data for the IPCHEM data source were provided directly by the IPCHEM team in CSV format. For data sources in PDF form, a combination of manual extraction and automated extraction was used to generate the dataset (method varied depending on source; all methods and scripts were retained). In sources that had tables that could be directly exported, the data were saved and manually reformatted to CSV format. Detailed descriptions of the source data, including the number of records and location/availability of metadata, the original form of raw data, type of data extraction (e.g., manual or script) and phase(s) of data collection are provided in Supplementary Table S1.
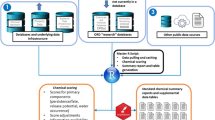
A MySQL4 relational database was designed to store both the raw and harmonized data and source metadata. The database entity-relationship diagram is provided in Fig. 1. All tables are described below in Data Records; all variable and table definitions are provided in Supplementary Table S2. The general workflow for populating the database is shown in Fig. 2 and described in detail below. In brief, raw data extracted from the data sources were pre-processed, loaded into the database, and then harmonized to standard variable names via an automated mapping process. Media and chemical identifiers were then also harmonized and secondary variables generated. Quality assurance (QA) of the raw data and the final database is described in the Technical Validation section.
MMDB Entity Relationship Diagram. See Supplementary Table S2 for a full description of database variables.
Data extraction
Data were extracted manually, via script, or through direct download, dependent on the data source. Details per database and per primary reference, where appropriate, are provided in Supplementary Table S1. Online databases often required multiple queries to obtain data efficiently and files were concatenated or pre-processed to join various outputs. Details of queries and pre-processing are provided for each source in the “Data and Curation Details” field in Table S1.
Initial processing
The raw data files underwent initial processing to prepare them for loading into the raw data tables of the database. In some cases, metadata from the raw source that would facilitate curation (e.g., chemical identifier information) were included in separate raw data files or downloads. In these cases, files were merged on appropriate raw variables (e.g., an internal chemical ID) included in both files. In the case of summary studies, it was desirable to organize the data into single variable “reported statistic” (e.g., a “mean” or “median”) and a “value” of that rather than a wide table of many metrics. This standardized the format across the raw summary studies and simplified later processing and reporting. Therefore, the raw data were “melted” using tools in the “reshape” package within the R environment prior to loading. Note that this step does not remove or alter any of the raw data, rather it just reshapes it into an optimal form. Note that aggregated summary statistics reported in MMDB were as reported in the original data sources (none were calculated by the authors); different sources may have handled values under the limit of quantification (LOQ) differently and thus the original source metadata should be consulted. Flags were added to the database (where possible) to indicate records associated with values <LOQ.
Loading of raw data
Once the intermediate raw data files (CSV) were extracted, quality checked, and processed, they were loaded into MMDB using an R script and an input control file. The control file contains information about every data source including directory and file paths for the intermediate raw data files. First, a unique entry for the data source was created in the source table, with accompanying source ID, name, data type (e.g., single-sample or summary), and other identifiable data. Next, the unique file information (e.g., filename, directory location, row count) for each data source’s raw data files were added to the files table, with relational reference to the source ID from the source table. Once file information was prepped, the file data were transformed into long form (to standardize storage) and written to the corresponding raw data table based on data source type. The raw data, from both single-sample data summary sources, were fed into a raw data table in this long form, with each record containing a variable name and a value, to allow storage of all original source data regardless of number of variables or format. Each row entry was also tagged with its corresponding file ID, source ID, and record ID. Finally, if a data source has additional supplemental or data documents, their file information was loaded into the documents table in a similar manner as for the file table. At every loading step, binary indicator variables track if a specific step was completed. This ensures that if a script fails or has an error, the workflow can pick up exactly where it left off. This also saves computational resources if a new data source is added, deleted, or updated.
Mapping to harmonized variables
Once the raw data source files were loaded to the raw table, their variables and values were ready to be transformed into a harmonized form across data sources. This was performed using an R script and data source variable map. The variable map, given in Supplementary Table S3, is a file containing every raw data field from each data source, mapped to the corresponding harmonized variable names. First, empty fields were added to the corresponding harmonized table for a data source, by data type, based on the file ID, source ID, and record ID within the raw table. Next, the raw data were transformed back into wide form. Then, both the raw variable name values and the mapped name values were processed to unify character case, white space, and remove punctuation, to ensure mapping was not inhibited, before the raw variable names were renamed for harmonization. The harmonized raw data, now in wide form, was then written to the harmonized table with accompanying source ID, file ID, record ID, and new harmonized ID.
Secondary processing
After the raw data were harmonized, further secondary processing was performed to enrich data analysis. That is, additional harmonized variable values were created by further processing raw variables (e.g., populating a flag indicating the value was a non-detect from a character appended to a raw analytical result) or entering study metadata (e.g., adding a reported medium of “ambient air” for an ambient air-related data source). Key additions were location variables and a detection indicator variable. Location data were unified into “country”, “US State”, and “US County” fields from the harmonized tables’ “reported location” field. For most, this was a duplicate field, while others required mapping from country abbreviations, country codes, or sampling site ID. A particularly useful harmonized variable was a flag indicating whether an observation was associated with detection of the chemical (if it could be discerned from available data). For the detection variable, an R script was created to assign a 1 or 0 value to a harmonized record based on the “reported result”, “LOD”, “LOQ”, and other QA flags. Additional variables derived from secondary processing may be added in the future.
Media and chemical extraction and curation
The original reported chemical substances and media underwent curation efforts to harmonize them for modeling. To assist with harmonizing chemicals and media, a unique list of values from each source was extracted from the harmonized tables. This included reported chemical name and CAS number, and reported media and reported species, for chemical and media data respectively. The reported chemical names and CAS Registry Numbers (CASRN) were curated into EPA’s Distributed Structure-Searchable Toxicity (DSSTox) Database using an automated curation workflow described elsewhere5. Prior to automated curation, parenthetical names were parsed to provide an additional name for potential automapping. The resulting curation process produced a chemical list including unique identifiers (DSSTox substance identifier, DTXSID) for each substance. The automated mapping process assigns a DTXSID with a given QC flag level (indicating confidence in the curation). Whether or not a given record was curated depended multiple factors including the previous confirmed curation of the identifier into DSSTox and assignment to substance IDs to the identifier. Not all identifiers will have corresponding IDs; a common case in this database were measurements associated with mixtures. Manual curation of the list of the identifiers included in this dataset by a trained curation team is ongoing as resources allow. The reported sample media (e.g., a species and tissue name or water sample type) were mapped to a set of 32 unique media (listed in Table 2). Mappings of all reported chemicals and media identifiers to their harmonized values are provided in Supplementary Tables S4 and S5. Once mapped, the unique values were written to the substance and media tables with unique substance and media ID values. In addition, observations in the harmonized tables not associated with chemical measurements were removed, as these were sometimes reported in the same fields as chemical measurements in various data sources. This included many environmental condition or weather measurements in the USGS data source, and physiological measurements on studied species in the ICES data source.
Data Records
The monitoring data compiled and harmonized here are stored in a MySQL relational database maintained by the USEPA and available via Figshare6. An export of the MySQL database is archived; this file contains a set of SQL statements that can be executed to reproduce the original database object definitions and all table data. Users may install MySQL and download the file from Figshare for manipulation and data extraction. Addition of new data, updating of harmonized variables from raw variables, and updates to the underlying monitoring dataset are ongoing. Versioned updates of the database will be provided as available, e.g., as more raw chemical identifiers in the raw data are curated or as reported concentration data are curated to standard formats and units. Details presented below represent data records archived in the MySQL database V1.0.
Within the MySQL database, there are tables containing data records and tables including metadata. The data records may be raw monitoring data from the original source or harmonized data records. Metadata tables contain descriptions and information which relate to all data records (or large subsets), as compared to record-specific data which is specific to a single data record (e.g., a single analytical measurement or study metric). Metadata may include information about individual data sources or downloaded files, or information about the chemicals or media referenced in the data tables. Metadata tables are linked to the data record tables by a set of database IDs referencing a specific data source, downloaded raw file, medium, or chemical.
There are 9 tables in the MySQL database. A full description of all variables included in the database tables is included in Supplementary Table S2.
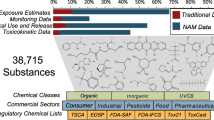
The multimedia database contains 63,768,583 individual harmonized data records (54,520,407 single-sample records and 9,248,176 aggregate records.) A total of 9,956 unique raw reported chemical (substance) identifiers (name and/or CASRN) were identified; 8,757 these could be mapped in the DSSTox database to one of 3,271 unique DTXSIDs. The mapped chemicals represent a wide range of chemical substance and use classes, including metals, pesticides, flame retardants, polychlorinated biphenyls, pharmaceuticals, and both consumer and industrial use chemicals. Counts of database observations (single-sample and summary) associated with each curated DTXSID are included in Supplementary Table S6. A summary of the chemicals and media represented in each of the database sources is given in Table 3. Figure 3 provides a summary of the location (US State, European Country, and worldwide occurrence) associated with each single-sample record in the database; counts of samples in countries outside the U.S. and Europe were small compared to these locations.
Technical Validation
The main quality objectives for the multimedia database were to ensure that data included in the raw data tables accurately reflected the data as provided in the raw data source, and that variables and data were accurately mapped to harmonized variables. Efforts to curate the dataset have not focused on checking for errors that may have been made by the original provider of the data; however, QA of raw data for suspected issues and QA of the harmonized tables was performed. Organizations or individuals who use this database are encouraged to perform QA consistent with data quality objectives for their assessments.
Raw data quality assurance
Raw datasets were divided into two categories: (i) tabular datasets and (ii) manually extracted data from an existing report or pdf. For tabular data, an R script was developed that reviewed each line of data within the set to determine if it met criteria to be flagged for a specific QA concern. The script appended the QA concern to the row of data if it met one or any of the criteria for any of the QA flags. One row of data could have multiple QA concerns associated. Separate R scripts were developed for each dataset, since they contained different variable names, however the scripts were based on the same QA flag criteria. An independent reviewer ran the R script and verified (i) proper file selection and loading of files via file name, row counts, and column counts and (ii) appropriate assigned of column headers to variables used in flagging code.
Counts for each individual flag created by the code were conducted. Datasets that were small enough to be opened in Excel were filtered by various variables in Excel to determine a manual count of the number of rows expected to be flagged for each QA concern. For datasets that exceeded the data limit in Excel, logic in R was used to obtain a count of the expected number of rows of data to be flagged for each QA concern. These independent, expected flag counts were then compared to the number of rows flagged by the QA R script to determine if the R script accurately flagged the correct number of rows. If there was a discrepancy, the script was reviewed and revised until the counts matched.
For manually extracted datasets, data were extracted from the original source into a standard Excel template with standard variable names. The datasets were then independently reviewed to verify the transcription accuracy. An R script was developed to append the QA codes to the extracted data (see Supplementary Table S7). The resulting flagged datasets were then reviewed by an independent reviewer to confirm the script identified all QA concerns accurately.
For datasets where the sampling year, unit of measurement, or media was missing, a manual review of the database documentation was performed to determine if the missing data could be identified. If the missing information could be identified, it was added to the dataset through the R script.
QA of harmonized data
Data entries within all database tables were checked in an automated QA process using three separate code scripts. These scripts checked key points in the workflow for data loading and processing. The first checked for missing and extraneous data source ID, file ID, and document ID values for associated table row entries. It also checked if the correct number of data records matched between the raw files and raw and harmonized tables. The script reported any discrepancies found between the raw CSV files and raw data table. The second checked for harmonized variable mappings between raw and harmonized tables. The third checked the mapping between media and chemical map tables to harmonized table entries. The output of this script could be visually inspected for any obvious errors in mapping (especially in the media mappings). The mappings of chemical records from raw chemical identifiers to DTXSID were obtained using an established semi-automated chemical curation workflow. Mappings are assigned QC levels based on the method and confidence of the mapping. Refinement of algorithms used in the semi-automated mapping process, and additional manual verification of mappings by the trained DSSTox curation team are always ongoing.
Usage Notes
MMDB is currently released as a MySQL “dump” file containing MySQL Statements that can be used to recreate the database objects (e.g., tables) and all data. MySQL is a free, open-source database management system (DBMS). It can be installed on Windows or Linux machines and provides both a server application (MySQL server) for creating, updating, and hosting databases (such as MMDB) and client application (MySQL client) for querying functionality. Once MySQL is installed and configured, the MMDB MySQL file can be run using MySQL server to create an exact copy of the original database. Note that when uncompressed, the MMDB file is very large (over 300GB), as is the database, so this may be a limitation on some systems. Once the database is built, a user can query MMDB using standard SQL commands (e.g., using MySQL client) or other scripting methods. The R programming language has packages that provide a direct interface to MySQL (“RMySQL”) and allow one to query a MySQL database (and transform the data) using simple syntax (“dplyr”). An example R script for querying MMDB by chemical or media using the RMySQL and dplyr packages is provided by the authors (see Code Availability).
The MySQL DBMS system and our robust standardized data loading and mapping procedures support the straightforward addition of other sources of monitoring data in the future. We further note that the database will be supported and maintained (as resources allow) under EPA’s Chemical Safety for Sustainability Research Program. In the future, it is planned that this data or summaries from MMDB will be incorporated into EPA ORD’s data infrastructure and be surfaced via the CompTox Chemicals Dashboard7 (https://comptox.epa.gov/dashboard) or other public-facing systems as ORD continues to work to make exposure-relevant data accessible to stakeholders. In addition, curation of the existing data, for updating or addition of other useful variables (e.g., flags indicating handling of non-detects in original summary sources) or harmonization of reported concentrations to standardized units, may continue.
Though data included in the database can be used in many ways for future analysis, users should be aware of limitations of the dataset, and appropriate usage of the data. Although a wide range of chemicals are included in the database, all the data here are the result of targeted analytical studies, wherein one or more chemicals were identified a priori for inclusion. Recent suspect-screening and non-targeted analysis of environmental and biological samples provides evidence that the true number of man-made or naturally-occuring chemicals present in biological and environmental media may be greater than what is currently included in this database. Therefore, this database does not contain an exhaustive list of chemicals found in organisms or the environment. In addition, there are still a significant number of raw chemical identifiers present that have not been mapped to harmonized identifiers (for several reasons, including their absence from the DSSTox database of synonyms for known substances or the inability to be considered a single chemical substance (e.g., mixtures of co-eluting PCBs). However, the raw reported identifiers are included in the database (and listed in Supplementary Table S4) and thus could be further addressed by end-users.
Due the size of this database, it is expected to provide a useful training set for machine-learning models that predict the occurrence of chemicals in different media. Other chemical information, such as chemical structure, chemical properties, and information about how chemicals are used are potential descriptors for such models. These models will complement analogous models developed by the ExpoCast project which predict chemical functional use8,9 and exposure pathway10. Within the ExpoCast project, new efforts are underway to identify thousands of chemicals present in environmental media using new non-targeted analytical (NTA) methods11. This database and subsequent models built upon it provide a useful resource for confirming tentative identifications in NTA studies.
Code availability
All scripts used to obtain raw data, clean or process raw data, perform QA, and construct the database are available in the MMDB Processing Scripts folder at https://doi.org/10.23645/epacomptox.16674298. Various versions of R and python were used in different project stages; the primary version for both data cleaning and building the database was R version 3.6.2. An example R script containing sample queries of MMDB by chemical and media is maintained in the Sample Queries folder. We will also maintain an SQL script (to be run in MySQL immediately after the MMDB dump file) to correct any identified curation mistakes in official MMDB releases in the MMDB Correction Scripts folder.
References
Cohen Hubal, E. A. et al. Advancing exposure characterization for chemical evaluation and risk assessment. J. Toxicol. Environ. Health. B Crit. Rev. 13, 299–313 (2010).
Using 21st century science to improve risk-related evaluations. (National Academies of Sciences, 2017).
FAIR Principles, https://www.go-fair.org/fair-principles/ (2021).
MySQL Open Source Database, https://www.mysql.com/ (2019).
Grulke, C. M., Williams, A. J., Thillanadarajah, I. & Richard, A. M. EPA’s DSSTox database: History of development of a curated chemistry resource supporting computational toxicology research. Comp. Tox. 12, 100096 (2019).
Isaacs, K. K. et al. Multimedia Monitoring Database (MMDB), figshare https://doi.org/10.23645/epacomptox.17065024.v1 (2021).
Williams, A. J. et al. The CompTox Chemistry Dashboard: a community data resource for environmental chemistry. J. Cheminform. 9, 61 (2017).
Isaacs, K. K. et al. Characterization and prediction of chemical functions and weight fractions in consumer products. Toxicology Reports 3, 723–732 (2016).
Phillips, K. A., Wambaugh, J. F., Grulke, C. M., Dionisio, K. L. & Isaacs, K. K. High-throughput screening of chemicals as functional substitutes using structure-based classification models. Green Chem. 19, 1063–1074 (2017).
Ring, C. L. et al. Consensus modeling of median chemical intake for the U.S. population based on predictions of exposure pathways. Environmental Sci. Tech. 53, 719–732 (2018).
Sobus, J. R. et al. Integrating tools for non-targeted analysis research and chemical safety evaluations at the US EPA. J. Expo. Sci. Environ. Epidemiol. 28, 411–426 (2018).
Stout, D. M. et al. American Healthy Homes Survey: a national study of residential pesticides measured from floor wipes. Environmental Sci. Tech. 43, 4294–4300 (2009).
American Healthy Homes Survey-Lead and Arsenic Findings. (U.S. Department of Housing and Urban Development Office of Healthy Homes and Lead Hazard Control, 2011).
Vidrio, E., Wofford, P., Segawa, R. & Schreider, J. Air Monitoring Network Results for 2013 - Volume 3. Report No. AIR 14-01 (California Environmental Protection Agency, Sacramento, CA, 2014).
King, K. D., Vidrio, E., Wofford, P. & Segawa, R. Air Monitoring Network Results for 2016 - Volume 6. Report No. AIR 17-01 (California Environmental Protection Agency, Sacramento, CA, 2017).
Tuli, A., Vidrio, E., Wofford, P. & Segawa, R. Air Monitoring Network Results for 2014 – Volume 4. Report No. AIR 15‐02 (California Environmental Protection Agency, Sacramento, CA, 2015).
Tuli, A., Vidrio, E., Wofford, P. & Segawa, R. Air Monitoring Network Results for 2015 – Volume 5. Report No. AIR 16 01 (California Environmental Protection Agency, Sacramento, CA, 2017).
Vidrio, E., Wofford, P., Segawa, R. & Schreider, J. Air Monitoring Network Results for 2012 - Volume 2. Report No. AIR 13-02 (California Environmental Protection Agency, Sacramento, CA, 2013).
Vidrio, E., Wofford, P., Segawa, R. & Schreider, J. Air Monitoring Network Results for 2011 - Volume 1. Report No. AIR 13-01 (California Environmental Protection Agency, Sacramento, CA, 2013).
Indoor air pollution in California: report to the California Legislature. (California Environmental Protection Agency, Air Resources Board, 2005).
Davis, A. P. et al. The Comparative Toxicogenomics Database: update 2019. Nucleic Acids Res. 47, D948–D954 (2019).
Comparative Toxicogenomics Database. http://ctdbase.org/ (2018).
The National Study of Chemical Residues in Lake Fish Tissue. Report No. EPA-823-R-09-006 (U.S. Environmental Protection Agency, Office of Water, 2009).
Targeted National Sewage Sludge Survey Statistical Analysis Report. (U.S. Environmental Protection Agency Office of Water, 2009).
Acknowledgements
The authors would like to thank Caroline Ring and Katherine Phillips of EPA ORD for their technical review of this manuscript, and Cathy Fehrenbacher and Andrea Pfahles-Hutchens of the EPA Office of Pollution Prevention and Toxics for their support of this project. The information in this document has been funded wholly or in part by the US Environmental Protection Agency. It does not signify that the contents necessarily reflect the views of the U.S. EPA or U.S. CPSC, nor does mention of trade names or commercial products constitute endorsement or recommendation for use. The paper has been subjected to the U.S. EPA review process and approved for publication.
Author information
Authors and Affiliations
Contributions
K.K.I. drafted the manuscript text and figures, designed and built the MySQL database, obtained and extracted raw data source data, mapped raw source variables to harmonized variables, wrote database workflow code to populate the database with processed harmonized data, and wrote code for generating other database variables. J.T.W. extracted raw data source data (including writing scripts for data extraction), generated select table data and maps, optimized the programmatic workflow for populating the database with processed harmonized data, created raw data files, wrote code for generating database variables, developed QA routines, and contributed to text. A.R.W., C.A.F., J.M. and K.A.H. obtained and extracted raw data source data (including writing scripts for data extraction), created raw data files, harmonized media identifiers, performed quality assurance (QA) of raw and harmonized data, and contributed to text. J.R.S. consulted on the design of the MySQL database, collection and interpretation of raw analytical data, and chemical and media curation procedures. E.U. consulted on the design of the MySQL database, collection and interpretation of raw analytical data, and chemical and media curation procedures. D.L. consulted on the design on the MySQL database and methods for importing raw data and generated the public data file from the database. K.L.D. managed the database infrastructure and curation of chemicals, designed QA procedures and tools, and contributed to text. A.J.W. leads the development of the CompTox Chemicals Dashboard and chemical curation methods and tools, and contributed to text. C.G. curated reported chemical identifiers. C.B. conceptualized the creation of a monitoring database, developed quality criteria for data sources, initiated the collection of all raw monitoring data, identified all data sources, managed ICF authors in the creation of raw data files, and contributed to text.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Isaacs, K.K., Wall, J.T., Williams, A.R. et al. A harmonized chemical monitoring database for support of exposure assessments. Sci Data 9, 314 (2022). https://doi.org/10.1038/s41597-022-01365-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-022-01365-8
This article is cited by
-
A harmonized Danube basin-wide multi-compartment concentration database to support inventories of micropollutant emissions to surface waters
Environmental Sciences Europe (2024)
-
Screening for drinking water contaminants of concern using an automated exposure-focused workflow
Journal of Exposure Science & Environmental Epidemiology (2024)
-
The chemical landscape of high-throughput new approach methodologies for exposure
Journal of Exposure Science & Environmental Epidemiology (2022)