Abstract
This data descriptor reports on the upload to a public repository (GNPS) of the IQAMDB, IsoQuinoline and Annonaceous Metabolites Data Base, comprising 320 tandem mass spectra. This project originated from our in-house collection of isoquinolines. The diversity of compounds included in this database was further extended through the contribution of two additional laboratories involved in isoquinoline alkaloids research: University of Angers and University of Manaus. The generated MS/MS data were processed and annotated on an individual basis to promote their straightforward reuse by natural product chemists interested in either the description of new isoquinoline alkaloids or the dereplication of isoquinoline-containing samples. The interest of the current repertoire for dereplication purposes has been validated based on the molecular networking of the well-investigated plant model Annona montana against the IQAMDB‐implemented GNPS.
Measurement(s) | electrospray ionization |
Technology Type(s) | Ultra High-performance Liquid Chromatography • Tandem Mass Spectrometry |
Similar content being viewed by others
Background & Summary
As one of the largest groups of plant alkaloids, isoquinolines include a significant number of well-known drugs and lead-compounds, such as morphine, codeine, noscapine, papaverine, D-tubocurarine, berberine, emetine or higenamine. These illustrious leads tantalized many laboratories to explore the phytochemistry of isoquinoline-producing plants, so that more than 3000 BenzylIsoQuinolines (BIQ) alkaloids are currently catalogued1, displaying varied and often significant bioactivities2. From a phylogenetic standpoint, (tetrahydro)benzylisoquinoline alkaloids are reported to occur in more than 40 different families with the most represented producers being Amaryllidaceae, Annonaceae, Berberidaceae, Fumariaceae, Hernandiaceae, Lauraceae, Menispermaceae, and Papaveraceae3. In the quest for alternative industrial processes of natural product supplies, a few isoquinoline alkaloids of outstanding interest succumbed to biotechnological manufacturing, such as morphine4 (and codeine as its biosynthetic intermediate) or noscapine5. The very last years witnessed tremendous advances in the metabolic engineering of BIQ with the development of optimized yeast strains synthesizing a considerable amount of (S)-reticuline, a biosynthetic pillar in the TetraHydroIsoQuinoline (THIQ) series, but also a diversity of unprecedented BIQ derivatives1. These advances in THIQ metabolite production guided Courdavault et al. to identify this family of compounds as holding “an eminent biosynthetic potential in the field of drug discovery”6.
Our continuous efforts towards the improvement of the molecular-networking pipeline efficiency through the upload of MS/MS data related with collections of structurally homogeneous natural products led us to implement two different databases today incorporated into the Global Natural Products Social Molecular Networking (GNPS) libraries7. A first contribution to these public repositories was dedicated to Monoterpene Indole Alkaloids8 (so-called MIADB), implying an international consortium of eight different natural product laboratories that resulted in a collection of 172 initial entries, now reaching more than 220 compounds. Our second initiative in the field, the LDB (Lichen Data Base), garnered 241 structurally diverse lichen phenolic compounds based on a collaborative project with University of Rennes 1 and the Berlin Garden and Botanical Museum9. We foresaw the possibility of building and deploying a third MS/MS database on the basis of the large diversity of isoquinoline alkaloids that have been isolated in Paris-Saclay10.
Described by Jussieu in 1789, the Annonaceae family comprises more than 120 genera and about 2100 species, all occurring in tropical and subtropical regions11. From a phytochemical standpoint, the annonaceous species have been vastly investigated, with a pronounced emphasis on their alkaloidal components12. Heretofore, more than 800 alkaloids have been reported from Annonaceous source12. Annonaceous alkaloids are vastly dominated by isoquinolines, mostly falling into benzyl- and bisbenzyltetrahydroisoquinolines, protoberberines, tetrahydroprotoberberines, proaporphines, aporphines, 7-substituted aporphines, oxoaporphines and phenanthrenes, all deriving from benzylisoquinoline precursors. A few non-isoquinoline alkaloids extend the diversity of Annonaceous metabolites through unusual structures, such as canangine, a napthyridine alkaloid13, original pyrimidine β-carboline structures referred to as annomontines14,15, a few prenylindoles and some indolosesquiterpenes16. It seems that the discovery rate of new chemical entities from Annonaceous sources decreases across decades as most of the recently published investigations in the field report known metabolites17,18. Although now uncommon, some original structures are still being unearthed from these plants, such as a few aristolactam alkaloids recently obtained from the thailandese Dasymaschalon dasymaschalum (Blume) I.M. Turner19 and new isoquinolines, including an unprecedented 8-oxohomoaporphines from the Amazonian Duguetia surinamensis R. E. Fries20. It is tempting to hypothesize that the quest for new structures from these plants is made difficult by the occurrence of important amounts of recurrent structures in Annonaceous plants that are hard to obviate in the course of untargeted phytochemical investigations. In such context, molecular networking strategies have proved useful to select natural material deserving deeper chemical studies as well as to streamline the isolation of compounds of interest through a rational, hypothesis-driven workflow21. This provides new opportunities to focus on minor, interesting compounds from deeply-dug plant material22,23. Nevertheless, the success of this dereplicative strategy primarily depends on the availability of reference tandem mass spectra uploaded to the GNPS spectral libraries7. Regarding the specific example of Annonaceous plants, some molecular networking-based investigations undertaken by some of us, involving an in-house collection of alkaloid standards, efficiently assisted the targeting of new compounds for purification and subsequent structure elucidation20. These works gratifyingly outlined the trend of Annonaceous alkaloids to clusterize in a scaffold-dependent manner. Some fragmentation patterns were even proposed by the same authors24,25, leading to extend the annotation provided by this dereplication to untagged nodes based on the cursory examination of MS/MS spectra20.
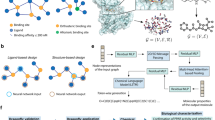
We felt that our in-house collection of isoquinolines could set the ground for building a MS/MS database dedicated to Annonaceous metabolites. The core constitution of this Annonaceous and isoquinoline database was enlarged by inviting prominent natural products researchers in the field to share their collections with us. This collaborative work took advantage of additional Annonaceous isoquinolines included in the collections of Federal University of Amazonas and Amazonas State University (Manaus, Brazil). To broaden the applicability of our database, this combined collection was further extended to structurally-related isoquinolines from non-Annonaceous plant sources through the incorporation of compounds obtained from University of Angers (France) which comprised isolates from Menispermaceous, Hernandiaceous, Lauraceous and Papaveraceous species26,27,28. These contributions led us to constitute the largest MS/MS dataset of isoquinolines to date, namely the IQAMDB (IsoQuinolines and Annonaceous Metabolites DataBase) (https://gnps.ucsd.edu/ProteoSAFe/gnpslibrary.jsp?library = IQAMDB) (Fig. 1 and Table S1)29. This database should help pinpointing new chemical entities from isoquinoline-producing plants. We also hope that the upload of this database will be of interest to increase the reliability of the dereplication workflow in the frame of the very dynamic metabolic engineering efforts currently devoted to THIQ alkaloids. This data descriptor reports on the deposition of IQAMDB in the GNPS libraries and on its subsequent technical validation.
Methods
Sample preparation
The different standard samples were dissolved in HPLC-MS grade methanol at 0.5 mg/mL and placed into 1500 μL vials for analysis. For the alkaloidic extract, 1 g of dried and milled Annona montana Macfad. root bark was extracted using 50 mL of 0.25 M H2SO4 for one hour. The phase was alkalinized to a pH of 12 and counter-extracted three times using 35 mL of methylene chloride. The combined organic phases were evaporated in vacuo and dissolved in analytical grade methanol at a concentration of 0.5 mg/mL for UPLC/HRMS² analysis. Solvents were purchased from Sigma-Aldrich.
Data acquisition
Samples were analyzed using an Agilent 6546 Accurate-Mass Q-TOF hyphenated with a 1290 Agilent Infinity II LC system. The chromatographic system was fitted with a Zorbax RRHD Eclipse Plus C18 column (2.1 × 50 mm, 1.8 μm). Elution solvents used were Milli-Q water + 0.1% formic acid (A) and acetonitrile + 0.1% formic acid (B), eluted using the following gradient: 5% B at 0 min; 100% B at 8 min, linear gradient; 8–12 min 100% B; 12–16 min 5% B. The flow rate was 500 µL.min−1. Mass spectrometry settings were as follows: capillary temperature at 320 °C, source voltage at 3.5 kV, and a sheath gas flow rate at 10 L/min. The mass spectrometer operated in positive polarity. Mass spectrometric acquisitions were divided into four scan events: positive MS with a window from m/z 100 to 1200, followed by three data-dependent MS/MS scans of the three most intense ions detected through the first scan event. Tandem mass spectrometric parameters were set as follows: collision energy = 50 eV, default charge of 1, isolation width of m/z 1.3. Dynamic exclusion was disabled. Purine (C5H4N4, m/z 121.050873), and HP-0921 (hexakis(1H, 1H, 3H-tetrafluoropropoxy)-phosphazene C18H18F24N3O6P3, m/z 922.009798 were used as internal lock masses. Full scans were acquired at a resolution of 60.000 (m/z 922) and 35.000 (m/z 121).
Database constitution
The analysis of all these substances resulted in 320 files with the standard Agilent.d format. The list of features detected in every sample was generated following the auto MS/MS data mining process implemented in MassHunter software on every single file. Within the list of detected features, the exact mass of the feature of interest was identified and the other features were filtered out. The MS/MS data related to the signal of interest were subsequently converted into a .mgf file using a tailored intensity threshold thanks to the dedicated “Export” option of the MassHunter software.
Molecular networking parameters (using IQAMDB standards as an input)
Every single MS/MS spectrum related to the IQAMDB standards was curated as indicated in the Database constitution section. A molecular network was then created using all the .mgf files as an input, using the online molecular networking workflow (version release_28.2) at GNPS7 (http://gnps.ucsd.edu) with a parent mass tolerance of 0.02 Da and a MS/MS fragment ion tolerance of 0.02 Da. The data were not clustered with MS‐Cluster. A network was then created where edges were filtered to have a cosine score above 0.6 and more than 6 matched peaks. Further edges between two nodes were kept in the network if and only if each of the nodes appeared in each other’s respective top 10 most similar nodes. The spectra in the network were then searched against GNPS spectral libraries. All matches kept between network spectra and library spectra were required to have a score above 0.6 and at least 6 matched peaks. The molecular networking data were analyzed and visualized using Cytoscape (ver. 3.6.0)30.
Feature-based molecular networking parameters (dereplication of A. montana)
The MS2 data files related to the alkaloidic extract of Annona montana were converted from the .d (Agilent) standard data-format to .mzXML format using the MSConvert software, part of the ProteoWizard package31. The .mzXML file was further processed using MZmine 2 v5332. Mass detections were performed using a noise level threshold at 2E3 in MS1 and at 1.5E1 in MS2. The ADAP chromatogram builder used a minimum group size of scans of 2,a group intensity threshold of 2E3, a minimum highest intensity of 2E3 and a m/z tolerance of 10 ppm33. The chromatogram deconvolution used the Local Minimum Search algorithm with the following settings: chromatographic threshold = 1%, search minimum in RT range (min) = 0.05, minimum relative height = 2%, minimum absolute height = 1E3, min ratio of peak top/edge = 0.9, peak duration range (min) = 0.00–1. MS2 scans were paired using a m/z tolerance range of 0.02 Da and RT tolerance range of 0.15 min. Isotopes were grouped using the isotopic peaks grouper algorithm with a m/z tolerance of 10 ppm and a RT tolerance of 0.5 min keeping the most intense peak as the representative isotope. The peak list was filtered to keep only rows with MS2 features. The .mgf and .csv files were exported using the MZmine2 built-in “Export/Submit to GNPS /FBMN” option. The molecular network was finally created using the online FBMN workflow (version release_28.2) at GNPS (http://gnps.ucsd.edu) with a parent mass tolerance of 0.02 Da and a MS/MS fragment ion tolerance of 0.02 Da, where edges were filtered to have a cosine score above 0.6 and more than 6 matched peaks. Further edges between two nodes were maintained only if each of the nodes appeared in each other’s respective top 10 most similar nodes. The spectra in the network were then searched against GNPS spectral libraries. Matches between network spectra and library spectra were required to have a cosine score above 0.6 and at least 6 matched peaks. The molecular networking data were analyzed and visualized using Cytoscape (ver. 3.6.0)30. The obtained molecular network can be accessed at: https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task = 3ff5f1048c7948ff96627438ac905acd.
Data Records
Data reported in this article have been uploaded to the GNPS platform. Each MS² spectrum of the 320 compounds is assigned an individual accession number on the GNPS. The .mgf files were deposited and are publicly available at the MassIVE repository (MSV000088909) (https://doi.org/10.25345/C55D8NG3T)34. The spectral collection is available for download from the library webpage of the GNPS (https://gnps.ucsd.edu/ProteoSAFe/libraries.jsp)29.
Technical Validation
Spectroscopic validation of IQAMDB compounds
The structure elucidation of the alkaloids implemented in the IQAMDB relied on an extensive set of spectroscopic techniques comprising at least nuclear magnetic resonance spectroscopy and high-resolution mass spectrometry. Further analyses were carried out whenever needed to ensure unambiguous structure assignment. These analyses were performed in the laboratory where the product had been isolated. The manual curation of each mass spectrometric file revealed the expected elemental composition, confirming that the samples had not degraded since they were isolated.
Retained strategies for IQAMDB validation
The validation of the IQAMDB repertoire was achieved following two distinct strategies. At first, the topology of the molecular network obtained using the IQAMDB as an input was inspected to assess whether the uploaded MS/MS data could outline structural similarities between the standards included in the database, as a first indicator of the quality of IQAMDB spectrometric data. Then, the dereplication efficiency of the IQAMDB-implemented GNPS libraries was estimated by annotating the molecular network obtained from the alkaloidic extract of the thoroughly studied Annona montana Macfad.
Topology of the molecular network obtained using the IQAMDB as an input
The molecular network generated using the tandem mass spectra of the IQAMDB as an input is disclosed in Fig. 2, with its nodes being colored according to their phytochemical class. Generally speaking, the topology of the molecular network does not deserve to be too thoroughly studied as it vastly depends on a wealth of different parameters, ensuring a low degree of reproducibility (mass spectrometric acquisition settings, molecular networking parameters, and even the diversity of compounds included in a molecular network that could instigate edges between some nodes or not). While having in mind these limits, we nevertheless felt relevant to outline a few clustering trends which may be of use to strengthen putative identifications against the IQAMDB-implemented GNPS. Chemical structures reported below to illustrate some rational clustering trends are provided in Figure S1 (Supporting Information).
The disclosed molecular network tended to cluster according to compounds structural class. The most striking examples of scaffold-based clustering are represented by tetrahydroprotoberberines (green nodes), benzylisoquinolines (light blue nodes) and bisbenzylisoquinolines (navy blue nodes)35. However, each of these scaffolds did not result in a unique and homogeneous cluster in the retained settings, sometimes conveying additional pieces of structural information. An example of rational structure-dependent clustering is that of bisbenzylisoquinolines which were split into three main clusters (A, B and C). As the largest group of isoquinoline alkaloids, bisbenzylisoquinolines are usually being classified based on their assembly modes, so that 26 different subtypes had been identified by Shamma and Moniot36. The benzylisoquinoline building blocks can indeed be joined by one or more ether bridges, sometimes accompanied by carbon-to-carbon biphenyl linkages or methyleneoxy junctions. Cluster A, exclusively, consisted of bisbenzylisoquinolines featuring two connections between their benzylisoquinoline components. Most nodes of this cluster corresponded to bisbenzylisoquinolines featuring a head-to-head ether connection and a tail-to-tail ether linkage between their monomeric units (berbamine, oxyacanthine, thalicberine and thalidasine subtypes). The rodiasine-type antioquine, featuring a head-to-head ether connection and a carbon-to-carbon biphenyl linkage as a tail-to-tail junction, connected with many representatives of the aforementioned subtypes. Notably, the southernmost part of cluster A comprised curine and isochondodendrine representatives, both featuring two head-to-tail ether connectivities. Cluster B connected bisbenzylisoquinolines featuring three intermonomeric connections. Most of the compounds fit the so-called trilobine-isotrilobine-micranthine type consisting of two ether bridges as head-to-head connections and an additional ether tail-to-tail linkage. The westernmost part of this cluster comprised tiliacorine and tiliacorinine disclosing two head-to-head ether connections and a biphenyl carbon-to-carbon bond tethering their tails. At last, four representatives loosely attached to the tetrahydroprotoberberine cluster (sub-cluster C). These molecules only featured one connection between their isoquinolines building blocks as a tail-to-tail junction (viz. daurisoline subtype). Likewise, the cursory examination of the clustering behaviour of other isoquinoline series revealed some tendencies which were seemingly related to some sharp structural features. Cluster D was composed of benzylisoquinolines substituted with a OH group on their benzyl component and by two oxygenated functionalities on their isoquinoline part. Benzylisoquinolines containing more than two methoxy groups on their isoquinoline component and a further methoxy functionality on their benzyl ring were detected in cluster E. At last, cluster F only collated structures disclosing a reticuline-type substitution pattern (a methoxy and a phenolic function on both isoquinoline and benzyl parts).
Validation of the IQAMDB against the phytochemically-defined Annona montana alkaloidic extract
The validation of the IQAMDB repertoire was based on the dereplication of Annona montana, commonly known as the mountain soursop, as a deeply-dug Annonaceous plant model. An alkaloidic extract was prepared from the historical sample of A. montana root bark, which had been phytochemically investigated by Lebœuf and co-workers in our laboratory in the 1980s14,37 and was further analyzed in positive polarity by UPLC-HRMS². To capture the chemical diversity of this extract, the UPLC−HRMS² data were processed using the feature-based molecular networking workflow38, and subsequently dereplicated against the IQAMDB, hosted by GNPS7. This pipeline resulted in dereplicating 27 unique and structurally-diverse compounds (Table 1). Notably, all these hits were exclusive to the IQAMDB, indicating the limited number of former Annonaceous compounds references in GNPS spectral libraries prior to the upload of the IQAMDB. Among those hits, 7 were previously reported from this plant source: Annomontine, methoxyannomontine14, argentinine39, atherosperminine37, coreximine37, liriodenine39, and reticuline37. Besides being closely related to some of these putative structures, 13 further hits had already been reported from other Annona species: benzyltetrahydroisoquinolines anomuricine and anomurine (both known from A. muricata31), N-methylcoclaurine (from A. sericea40), reticuline N‐oxide (A. salzmanni41) and tembetarine (tentatively known from A. salzmanni41); proaporphines such as glaziovine and stepharine, both reported from A. purpurea42,43; tetrahydroprotoberberines kikemanine (A. glabra44), pessoine (from A. spinescens45), and stepholidine (A. cherimolia46); aporphines such as nornuciferine (A. muricata47), obovanine (A. coriacea48) and roemerine (from both A. squamosa49 and A. senegalensis50). Most other hits corresponded to metabolites occurring in other Annonaceous genera. Among all the hits, only nandigerine was hitherto unknown from Annonaceous source. Even though this aporphine alkaloid is mainly related to Hernandiaceae12 and Lauraceae51, one of its derivatives, N-methylnandigerine N-oxide, had been reported from the Annonaceous Polyalthia longifolia52.
Metadata
The MS/MS spectra of the IQAMDB library are associated to a variety of details including: LC-MS/MS acquisition parameters, RT (see Supporting Information), instrument details, smiles and InChi codes, structures, and chemical formula. These metadata are available on the GNPS website.
Code availability
The LC-MS feature detection software (MassHunter®) used in this work is commercially available from Agilent®.
References
Pyne, M. E. et al. A yeast platform for high-level synthesis of tetrahydroisoquinoline alkaloids. Nature Commun. 11, 1–10 (2020).
Nguyen, V. K. & Kou, K. G. The biology and total syntheses of bisbenzylisoquinoline alkaloids. Org. Biomol. Chem. 19, 7535–7543 (2021).
Bisset, N. G. Plants as a source of isoquinoline alkaloids. in The Chemistry and Biology of Isoquinoline Alkaloids 1–22 (Springer, 1985).
Galanie, S., Thodey, K., Trenchard, I. J., Interrante, M. F. & Smolke, C. D. Complete biosynthesis of opioids in yeast. Science 349, 1095–1100 (2015).
Li, Y. et al. Complete biosynthesis of noscapine and halogenated alkaloids in yeast. Proc. Natl Acad. Sci. 115, E3922–E3931 (2018).
Kulagina, N., Papon, N. & Courdavault, V. Microbial Cell Factories for Tetrahydroisoquinoline Alkaloid Production. ChemBioChem 22, 639–641 (2021).
Wang, M. et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 34, 828–837 (2016).
Ramos, A. E. F. et al. Collected mass spectrometry data on monoterpene indole alkaloids from natural product chemistry research. Sci. Data 6, 1–6 (2019).
Olivier-Jimenez, D. et al. A database of high-resolution MS/MS spectra for lichen metabolites. Sci. Data 6, 1–11 (2019).
Lebœuf, M., Cavé, A., Bhaumik, P. K., Mukherjee, B. & Mukherjee, R. The phytochemistry of the annonaceae. Phytochemistry 21, 2783–2813 (1980).
Hutchinson, J. The genera of flowering plants. Dicotylédones. Vol. 1. The genera of flowering plants. Dicotylédones. Vol. 1. (1964).
Lúcio, A. S. S. C., de Almeida JR., da-Cunha, E. V. L., Tavares, J. F. & Barbosa Filho, J. M. Alkaloids of the Annonaceae: Occurrence and a Compilation of Their Biological Activities. in The Alkaloids: Chemistry and Biology vol. 74 233–409 (Elsevier, 2015).
Lebœuf, M. & Cavé, A. Alkaloids of Annonaceae. XIV. Identification of canangine and eupolauridine, new alkaloids with a naphthyridine nucleus. Lloydia 39, 459–460 (1976).
Lebœuf, M. et al. Alkaloids of the Annonaceae. Part 33. Annomontine and methoxyannomontine, two new pyrimidine-β-carboline-type alkaloids from Annona montana. J. Chem. Soc., Perkin Trans. 1205–1208 (1982).
Costa, E. V. et al. Full NMR analysis of annomontine, methoxy-annomontine and N-hydroxyannomontine pyrimidine-β-carboline alkaloids. Magn. Reson. Chem. 46, 69–74 (2008).
Cave, A., Guinaudeau, H., Leboeuf, M., Ramahatra, A. & Razafindrazaka, J. Alcaloïdes des Annonacées XVIII1: Alcaloïdes des Ecorces de tronc du Polyalthia suaveolens Engl. et Diels. Planta Med. 33, 243–250 (1978).
de Souza Araújo, M., da Silva, F. M. A., Koolen, H. H. F. & Costa, E. V. Isoquinoline-derived alkaloids from the bark of Guatteria olivacea(Annonaceae). Biochem. Syst. Ecol. 92, 104105 (2020).
Nardelli, V. B. et al. Isoquinoline-derived alkaloids and one terpene lactone from the leaves of Duguetia pycnastera(Annonaceae). Biochem. Syst. Ecol. 94, 104206 (2021).
Suthiphasilp, V. et al. Dasymaschalolactams A–E, Aristolactams from a Twig Extract of Dasymaschalon dasymaschalum. J. Nat. Prod. 82, 3176–3180 (2019).
Paz, W. H. et al. Structure-Based Molecular Networking for the Target Discovery of Oxahomoaporphine and 8-Oxohomoaporphine Alkaloids from Duguetia surinamensis. J. Nat. Prod. 82, 2220–2228 (2019).
Fox Ramos, A. E., Evanno, L., Poupon, E., Champy, P. & Beniddir, M. A. Natural products targeting strategies involving molecular networking: Different manners, one goal. Nat. Prod. Rep. 36, 960–980 (2019).
Fouotsa, H. et al. Voatriafricanines A and B, Trimeric Vobasine-Aspidosperma-Aspidosperma Alkaloids from Voacanga africana. J. Nat. Prod. 84, 2755–2761 (2021).
Fouotsa, H. et al. Pyrrovobasine, hybrid alkylated pyrraline monoterpene indole alkaloid pseudodimer discovered using a combination of mass spectral and NMR-based machine learning annotations. Org. Biomol. Chem. 20, 98–105 (2022).
da Silva, F. M. et al. Positive electrospray ionization ion trap mass spectrometry and ab initio computational studies of the multi-pathway fragmentation of oxoaporphine alkaloids. Int. J. Mass Spectrom. 418, 30–36 (2017).
Neto, F. C. et al. Characterization of aporphine alkaloids by electrospray ionization tandem mass spectrometry and density functional theory calculations. Rapid Commun. Mass Spectrom. 34, e8533 (2020).
Richomme, P., Lavault, M., Jacquemin, H. & Bruneton, J. Etude des Hernandiacées VI (1): lignanes et alcaloïdes de Hernandia guianensis. Planta Med. 50, 20–22 (1984).
Chalandre, M.-C., Bruneton, J., Cabalion, P. & Guinaudeau, H. Alcaloïdes de Gyrocarpus americanus. J. Nat. Prod. 49, 101–105 (1986).
Lavault, M. et al. Alcaloïdes bisbenzylisoquinoléiques de Albertisia cf. A. papuana. Can. J. Chem. 65, 343–347 (1987).
IsoQuinoline and Annonaceous Metabolites Data Base (IQAMDB), GNPS, https://gnps.ucsd.edu/ProteoSAFe/gnpslibrary.jsp?library=IQAMDB
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Chambers, M. C. et al. A cross-platform toolkit for mass spectrometry and proteomics. Nature Biotechnol. 30, 918–920 (2012).
Pluskal, T., Castillo, S., Villar-Briones, A. & Orešič, M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinformatics 11, 1–11 (2010).
Myers, O. D., Sumner, S. J., Li, S., Barnes, S. & Du, X. One Step Forward for Reducing False Positive and False Negative Compound Identifications from Mass Spectrometry Metabolomics Data: New Algorithms for Constructing Extracted Ion Chromatograms and Detecting Chromatographic Peaks. Anal. Chem. 89, 8696–8703 (2017).
Le Pogam, P. GNPS_IQAMDB Isoquinolines and other Annonaceous Metabolites DataBase. MassIVE https://doi.org/10.25345/C55D8NG3T (2022).
Shamma, M. The bisbenzylisoquinolines. in The Isoquinoline Alkaloids. Chemistry and Pharmacology. 115–152 (Academic Press, 1972).
Shamma, M. & Moniot, J. The Systematic Classification of Bisbenzylisoquinolines. Heterocycles 4, 1817–1824 (1976).
Lebœuf, M. et al. Alcaloides des annonacées XL: Etude chimique et pharmacologique des alcaloïdes de l’Annona montana. Plant. Medic. Phytother. 3, 169–184 (1982).
Nothias, L.-F. et al. Feature-based molecular networking in the GNPS analysis environment. Nature Methods 17, 905–908 (2020).
Wu, Y.-C., Chang, G.-Y., Chang-Yih, D. & Shang-Kwei, W. Cytotoxic alkaloids of Annona montana. Phytochemistry 33, 497–500 (1993).
Campos, F. R. et al. Isoquinoline alkaloids from leaves of Annona sericea (Annonaceae). Biochem. Syst. Ecol. 36, 804–806 (2008).
Lima, J. M., Leme, G. M., Costa, E. V. & Cass, Q. B. LC-HRMS and acetylcholinesterase affinity assay as a workflow for profiling alkaloids in Annona salzmannii extract. J. Chromatogr. B 1164, 122493 (2021).
Chang, F.-R., Wei, J.-L., Teng, C.-M. & Wu, Y.-C. Antiplatelet Aggregation Constituents from Annona purpurea. J. Nat. Prod. 61, 1457–1461 (1998).
Sonnet, P. E. & Jacobson, M. Tumor inhibitors II: cytotoxic alkaloids from Annona purpurea. J. Pharm. Sci. 60, 1254–1256 (1971).
Chang, F.-R., Chen, C.-Y., Hsieh, T.-J., Cho, C.-P. & Wu, Y.-C. Chemical Constituents from Annona glabra III. J. Chinese Chemical Soc. 47, 913–920 (2000).
Queiroz, E. F., Roblot, F., Cavé, A., de Q. Paulo, M. & Fournet, A. Pessoine and spinosine, two catecholic berbines from Annona spinescens. J. Nat. Prod. 59, 438–440 (1996).
Villar, A., Mares, M., Rios, J. L. & Cortes, D. Alkaloids from Annona cherimolia leaves. J. Nat. Prod. 48, 151–152 (1985).
Hasrat, J. A., De Bruyne, T., De Backer, J.-P., Vauquelin, G. & Vlietinck, A. J. Isoquinoline derivatives isolated from the fruit of Annona muricata as 5-HTergic 5-HT(1A) receptor agonists in rats: Unexploited antidepressive (lad) products. J. Pharm. Pharmacol. 49, 1145–1149 (1997).
Nogueira da Silva Avelino Oliveira Rocha, G. et al. Chemical constituents from the leaves and branches of Annona coriacea Mart. (Annonaceae). Biochem. Syst. Ecol. 97, 104297 (2021).
Bhakuni, D. S., Tewari, S. & Dhar, M. M. Aporphine alkaloids of Annona squamosa. Phytochemistry 11, 1819–1822 (1972).
You, M. et al. (-)-Roemerine, an aporphine alkaloid from Annona senegalensis that reverses the multidrug-resistance phenotype with cultured cells. J. Nat. Prod. 58, 598–604 (1995).
Teles, M. M. R. S. et al. Alkaloids of the Lauraceae. The Alkaloids: Chemistry and Biology 82, 147–304 (2019).
Wu, Y. C. Azafluorene and aporphine alkaloids from Polyalthia longifolia. Heterocycles 29, 463–475 (1989).
Lebœuf, M. et al. Alcaloïdes des Annonacées XXIX: Alcaloïdes de l’Annona muricata L. Planta Med. 42, 37–44 (1981).
Costa, E. V. et al. Benzylated Dihydroflavones and Isoquinoline-Derived Alkaloids from the Bark of Diclinanona calycina (Annonaceae) and Their Cytotoxicities. Molecules 26, 3714 (2021).
Lu, S.-T., Wu, Y.-C. & Leou, S.-P. Alkaloids of formosan Fissistigma and Goniothalamus species. Phytochemistry 24, 1829–1834 (1985).
Chaves, M. H., De Santos, L. A., Lago, J. H. G. & Roque, N. F. Alkaloids from Porcelia macrocarpa. J. Nat. Prod. 64, 240–242 (2001).
Rasamizafy, S., Hocquemiller, R., Cavé, A. & Fournet, A. Alcaloïdes des Annonacées, 78. Alcaloïdes des Écorces d’un Duguetia spixiana de Bolivie. J. Nat. Prod. 50, 674–679 (1987).
Acknowledgements
The authors would like to dedicate this article to Pr. Michel Lebœuf, whose lifetime work was the basis for the chemical library of isoquinoline alkaloids digitized herein, and who supported the current project, and to the memory of Prof. André Cavé. The authors are also indebted to former researchers of the University of Angers, mostly Prof. Jean Bruneton and Prof. Hélène Guinaudeau, who contributed to the isolation of many other compounds comprised in this database.
Author information
Authors and Affiliations
Contributions
P.L.P. conceived the project. S.A.A. isolated some compounds included in the IQAMDB repertoire. S.A.A., T. O., Y.A.K., A.J. and P.L.P. contributed to sample preparation. P.L.P. performed the acquisition of the LC-MS/MS data. M.A.B and P.L.P. performed the technical validation of the IQAMDB. E.V.C., F.M.A.d.S. and H.H.F.K. isolated and collected Brasil compounds. D.B., A.M.L.R.-R. and P.R. isolated and collected compounds from Université d’Angers. P.C. and P.L.P advised on the extraction aspects of the work conducted by S.A.A. P.L.P. supervised the work at Université Paris-Saclay and wrote the manuscript. All the authors read and commented the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Agnès, S.A., Okpekon, T., Kouadio, Y.A. et al. Implementation of a MS/MS database for isoquinoline alkaloids and other annonaceous metabolites. Sci Data 9, 270 (2022). https://doi.org/10.1038/s41597-022-01345-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-022-01345-y