Abstract
We present a whole-brain in vivo diffusion MRI (dMRI) dataset acquired at 760 μm isotropic resolution and sampled at 1260 q-space points across 9 two-hour sessions on a single healthy participant. The creation of this benchmark dataset is possible through the synergistic use of advanced acquisition hardware and software including the high-gradient-strength Connectom scanner, a custom-built 64-channel phased-array coil, a personalized motion-robust head stabilizer, a recently developed SNR-efficient dMRI acquisition method, and parallel imaging reconstruction with advanced ghost reduction algorithm. With its unprecedented resolution, SNR and image quality, we envision that this dataset will have a broad range of investigational, educational, and clinical applications that will advance the understanding of human brain structures and connectivity. This comprehensive dataset can also be used as a test bed for new modeling, sub-sampling strategies, denoising and processing algorithms, potentially providing a common testing platform for further development of in vivo high resolution dMRI techniques. Whole brain anatomical T1-weighted and T2-weighted images at submillimeter scale along with field maps are also made available.
Measurement(s) | brain measurement |
Technology Type(s) | magnetic resonance imaging |
Sample Characteristic - Organism | Homo sapiens |
Machine-accessible metadata file describing the reported data: https://doi.org/10.6084/m9.figshare.14058443
Similar content being viewed by others
Background & Summary
Diffusion magnetic resonance imaging (dMRI)1 has proven to be an invaluable tool in the study of brain structural connectivity and complex tissue microarchitecture. It has also offered the ability to investigate a variety of neurological, neurodegenerative and psychiatric disorders2,3,4,5,6,7,8,9,10. In recent years, a number of dMRI data repositories11,12,13,14,15,16,17,18 have been published to enhance the understanding of human brain, including large-scale population study projects launched across the world, such as the Human Connectome Project (HCP)19,20,21,22 and the UK Biobank project23. However, most of these studies acquired in vivo dMRI datasets at a relatively low spatial resolution of 1.25–3 mm, due to technical difficulties in achieving higher spatial resolution that include: i) the inherent low SNR of dMRI acquired with smaller voxels; ii) the distortion/blurring artifacts from longer EPI readouts; and iii) the inevitable participant motion during long scan time required for obtaining multiple diffusion directions. Submillimeter isotropic resolution dMRI datasets have therefore been mostly confined to ex-vivo studies24,25,26 where there are no scan time and motion constraints. While ex vivo dMRI plays an important role in studying brain structures with its higher achievable resolution, the alteration of tissue properties due to fixation may fundamentally change the dMRI measurement and affect its ability to reflect accurate values in the living human brain. In vivo dMRI serves as a non-invasive probe of tissue structure in different brain pathologies, and can also be combined with functional and behavioral data for further investigation of structure-function relationships. The availability of submillimeter resolution in vivo dMRI datasets would enable more detailed examination of the brain’s fine-scale structures, such as probing short-association fibers at the gray-white matter boundary27 or studying the cytoarchitecture of thin cortical layers (<1–4 mm in thickness)26,28. Submillimeter resolution in vivo dMRI would also facilitate the development and optimization of new sampling, modeling and processing methods specifically targeted at recovering information about tissue microstructure from such high resolution in vivo dMRI datasets.
With the advent of advanced hardware20,29,30, acquisition and reconstruction strategies31,32,33,34,35,36, submillimeter isotropic resolution dMRI is becoming increasingly feasible37,38,39,40,41, presenting new opportunities for studying brain structure at the mesoscopic scale in vivo. In this work, we aim to create the first publicly available in vivo whole brain dMRI reference dataset acquired at submillimeter isotropic resolution of 760 μm, i.e. at 7.7 times smaller voxel size than conventional dMRI acquisition at ~1.5 mm resolution. To enable high-SNR at this extreme resolution, state-of-the-art hardware and acquisition techniques were used along with a carefully designed diffusion protocol targeting at a single-participant scan across 9 two-hour sessions. The data were acquired on the MGH-USC 3 T Connectom scanner equipped with high-strength gradient (Gmax = 300 mT/m)20,29,42 and a custom-built 64-channel phased-array coil30, using the recently developed SNR-efficient simultaneous multi-slab imaging technique termed gSlider-SMS39,40. To mitigate potential image artifacts and loss in resolution, a custom-made form-based headcase (https://caseforge.co) that precisely fit the shape of participant’s head and the coil was also used to ensure minimal motion contamination, consistency of participant positioning and same field shim settings across sessions. Advanced parallel imaging reconstruction with ghost-reduction algorithm (Dual-Polarity GRAPPA43,44) and reversed phase-encoding acquisition were also performed to mitigate ghosting artifacts and image distortion. We acquired a total of 1260 q-space points including 420 directions at b = 1000 s/mm2 and 840 directions at b = 2500 s/mm2 across the 9 sessions to ensure high angular resolution. An optimized preprocessing pipeline for the multi-session data was employed afterwards to correct for intensity drifts, distortions and motion while preserving as much high-resolution information as possible.
We envision that this reference dMRI dataset will have a broad range of investigational, educational, and clinical applications that will advance the understanding of human brain structure. In particular, the data should be useful in the exploration of mesoscale structures revealed through submillimeter in vivo dMRI, and in the evaluation of structural brain connectivity that should improve with reduced intravoxel fiber complexity using smaller voxels and with higher intravoxel fiber crossing resolution using high angular resolution45,46,47. The data may also serve as a test bed for new modeling, data sub-sampling, denoising and processing algorithms48,49,50, potentially providing a common testing platform for further technical development of in vivo high resolution dMRI.
Methods
Participant
The participant was a young healthy Caucasian man (born in 1989). He was fully instructed about the scans and gave written informed consent for participation in the study. The experiments were performed with approval from the institutional review board of Partners Healthcare.
Data acquisition
All data were acquired in 9 two-hour scan sessions on the MGH-USC 3 T Connectom scanner equipped with a maximum gradient strength of 300 mT/m and a maximum slew rate of 200 T/m/s20,29,42, using a custom-built 64-channel phased-array coil30. A custom-designed personalized motion-robust stabilizer (https://caseforge.co), that was precisely fit to the shape of the participant’s head and the inside of the coil, was used to ensure the same participant positioning across sessions and minimal motion contamination within a session (Fig. 1a). Field shim settings were kept identical across sessions to minimize differences in image distortion, with the goal of minimizing reliance on image post-processing that could introduce blurring.
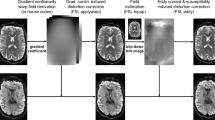
Data acquisition, reconstruction and preprocessing workflow. (a) Hardware and acquisition techniques used to acquire the data. The Connectom scanner, the custom-built 64-channel phased-array coil and the gSlider-SMS acquisition technique provide high SNR, while the personalized motion-robust stabilizer ensures consistent positioning across multiple scan sessions and minimal motion contamination within each session. (b) Data acquisition pipeline in each 2-hour session. The dMRI acquisition was divided into 4 acquisition blocks, each of which comprised 70 diffusion volumes and 8 interleaved b = 0 s/mm2 volumes. Two acquisition blocks with paired reversed phase encoding composed a data subset with a unique set of diffusion directions. 6 data subsets were acquired at b = 1000 s/mm2 in the first 3 sessions, and 12 data subsets at b = 2500 s/mm2 in the remaining 6 sessions. (c) Data reconstruction workflow. dMRI data were reconstructed using Dual Polarity GRAPPA, partial Fourier reconstruction, real value diffusion and gSlider slab-to-slice reconstruction, while field maps and structural images were reconstructed using standard GRAPPA reconstruction implemented on the scanner. Minimally preprocessed data were then provided after the gradient non-linearity correction (Data output A). (d) dMRI preprocessing with one-step resampling. The preprocessed data were obtained after intensity normalization to correct for signal drift, along with susceptibility-induced and eddy current-induced distortion correction, gradient non-linearity correction and motion correction, all of which were performed together using a one step resampling process to minimize interpolation-induced image blurring (Data output B).
To obtain high spatial resolution dMRI data, the gSlider-SMS39 sequence (Fig. 1a) was used in our study. gSlider-SMS is a recently developed self-navigated simultaneous multi-slab acquisition that combines blipped-CAIPI51 controlled aliasing with a slab-selective radio frequency encoding, which has been demonstrated to provide high SNR-efficiency for high resolution dMRI. The gSlider-SMS protocol was constructed to achieve a whole brain isotropic submillimeter acquisition at high SNR with low reconstruction errors and limited distortion and blurring, while still keeping the scan time tolerable. Motivated by this goal, the decision of the final imaging parameters was made by evaluation of data quality of pilot scans and conclusions from prior studies. The 0.76-mm isotropic resolution was chosen as it provides a good trade-off in term of resolution and scan time to achieve reasonable SNR. A gSlider encoding factor of 5 and a multi-band acceleration factor of 2 were used to achieve high SNR efficiency by acquiring 10 slices per shot across two RF-encoded slabs, where each slab contains 5 slices that can be resolved through 5 gSlider-encoded repetitions. In-plane acceleration of 3 was selected to mitigate EPI distortion and blurring by reducing the effective echo spacing from 1.02 ms to 0.34 ms, while ensuring good reconstruction performance with low noise amplification when the reconstruction was performed with Dual polarity GRAPPA to also mitigate ghosting artifacts. Despite the use of in-plane acceleration and shorter readout length, a small level of image blurring still exists along phase-encoding (PE) direction due to signal decay across EPI readout, characterized by a ~7–15% increase in the full width at half maximum of the point spread function along PE direction for normal gray and white matter (T2 around 60–100 ms, and T2* around 40–80 ms) based on theoretical simulation analysis. Before diffusion acquisition, a calibration scan was acquired using a multi-shot mono-polar SE-EPI to provide fully-sampled data, with the same effective echo spacing as the in-plane accelerated data and in both readout polarities, to calculate the dual-polarity GRAPPA kernel43,44. The detailed imaging parameters are listed in Fig. 2a. Using an actual maximum gradient amplitude of 180 mT/m and a slew rate of 125 T/m/s in our acquisition, a shorter TE (75 ms) was achieved at b = 2500 s/mm2 on Connectom scanner than other state-of-art clinical scanners (e.g., TE = 87 ms using 3 T Prisma) with all the same parameters but a lower achievable maximum gradient amplitude of 80 mT/m. Noted that the maximum gradient strength (300 mT/m) and the maximum slew rate (200 T/m/s) were not applied in Connectom scanner to avoid strong eddy current and exceeding peripheral nerve stimulation limit. The TEs and TRs of b = 1000 s/mm2 and b = 2500 s/mm2 were matched. The TR of each RF encoded volume was 3.5 s, and the use of 5 RF encoded volumes resulted in an effective volume TR of 17.5 s.
To achieve good SNR and high angular resolution, a total of 2808 volumes of dMRI data were acquired across the 9 sessions, consisting of 144 b = 0 s/mm2 images, 420 b = 1000 s/mm2 diffusion-weighted images (DWIs) and 840 b = 2500 s/mm2 DWIs, along with their paired reversed phase-encoding volumes. The two shells at b = 1000 s/mm2 and 2500 s/mm2 are shown in Fig. 2b, where the 420 directions at b = 1000 s/mm2 represented a subset of the 840 directions at b = 2500 s/mm2. The diffusion vectors were designed using the algorithm described in Emmanuel Caruyer et al.52 (publicly available tool at http://www.emmanuelcaruyer.com/q-space-sampling.php), which not only provides uniform angular coverage for each shell and for overall multishell sampling, but is also able to provide uniform incremental angular distribution. In each session, 312 dMRI volumes were acquired, including 280 DWIs (140 unique diffusion directions with reversed phase-encoding acquisition) and 32 b = 0 s/mm2 images (interleaved with every 10 diffusion images). These data were further divided and acquired in four acquisition blocks, so the calibration data could be updated every ~25 mins and the paired reversed phase-encoding volumes could be interleaved.
In each session, a low-resolution field map and a structural scan (T1 or T2-weighted image) were also acquired as time permitted. The low-resolution field maps were acquired using a two-TE gradient echo field mapping sequence: TR = 351 ms; TEs = 4.14, 6.6 ms; FOV = 220 × 220 mm2; voxel size = 2 × 2 mm2; number of slices = 38; slice thickness = 2.5 mm; slice gap = 1.3 mm; flip angle = 45°; bandwidth = 500 Hz/Pixel; total acquisition time = 79 s. The T1-weighted images were acquired with the Multi-echo Magnetization-Prepared Rapid Acquisition Gradient Echo (MEMPRAGE)53 sequence at both 1-mm isotropic and 0.7-mm isotropic resolutions. The 1-mm MEMPRAGE protocol was acquired using the following parameters: TR = 2530 ms; TI = 1100 ms; number of echoes = 4; TEs = 1.61, 3.47, 5.34, 7.19 ms; flip angle = 7°; FOV = 256 × 256 × 208 mm3; bandwidth = 650 Hz/Pixel; GRAPPA factor along phase encoding direction = 2; total acquisition time = 6 minutes. The 0.7-mm MEMPRAGE protocol: TR = 2530 ms; TI = 1110 ms; number of echoes = 4; TE = 1.29, 3.08, 4.83, 6.58 ms; flip angle = 7°; FOV = 256 × 256 × 146 mm3; bandwidth = 760 Hz/Pixel; GRAPPA factor along PE = 2; total acquisition time = 8.38 minutes. T2-weighted data were acquired with a T2-SPACE54 sequence at 0.7-mm isotropic resolution with the following parameters: TR/TE = 3200/563 ms; FOV = 224 × 224 × 180 mm3; bandwidth = 745 Hz/Pixel; GRAPPA factor along PE = 2; total acquisition time = 8.4 minutes. All T1- and T2-weighted data were acquired with image readout in the head-foot (HF) direction, PE in the anterior-posterior (AP) direction, and partition in the left-right (LR) direction. One 1-mm isotropic resolution MEMPRAGE, three 0.7-mm isotropic resolution MEMPRAGE and three 0.7-mm isotropic resolution T2-SPACE images were acquired in session 3–9, respectively.
Data reconstruction
The image reconstruction for dMRI was performed on raw k-space data in MATLAB (Mathworks, Natick, MA, USA) (Fig. 1c). First, dual-polarity-GRAPPA43,44 (DPG) was used for parallel imaging reconstruction of both in-plane and multi-band accelerations. DPG has been validated to reduce Nyquist ghost artifacts of EPI effectively by correcting for higher-order phase errors43,44. Unaliased multi-channel images were combined using SENSE 1 complex coil combination method55, where the sensitivity maps were estimated from the calibration data using ESPIRiT56. POCS57 was used for partial Fourier reconstruction to recover high spatial information from partially acquired k-space data. Diffusion background phase-corruptions were further removed using the real-valued diffusion algorithm to provide real-value data and avoid magnitude bias in subsequent post-processing steps58. Finally, gSlider’s slab-to-slice reconstruction39,40 was performed to reconstruct thick RF-encoded slabs (3.8 mm) into thin slices (0.76 mm), where T1 recovery was incorporated into the reconstruction model to mitigate slab-boundary artifacts59. The slab-to-slice reconstruction has been demonstrated to maintain image resolution with a sharp point spread function (small side lobe of 7.5%) and to provide robust performance in the presence of B1 and B0 inhomogeneity39.
The image reconstruction of the T1- and T2-weighted images and field maps was performed by the vendor’s (Siemens Healthcare, Erlangen, Germany) online reconstruction (including vendor’s implementation of parallel imaging reconstruction, fast Fourier transformation, coil combination, root mean square combination of different echo images and calculation of field maps) and was finally stored in Digital Imaging and Communications in Medicine (DICOM) format.
Data preprocessing
Preprocessing corrections across the reconstructed image series were performed to correct for signal drift, susceptibility, eddy currents, and gradient nonlinearity induced distortions, and participant motion. An optimized pipeline was used for the multi-session data, where the warp fields for susceptibility, eddy-current distortion, gradient nonlinearity and motion were estimated, combined, and applied to each image volume in a one-step resampling process to ensure minimal smoothing effects due to interpolation60,61. The FMRIB Software Library (FSL)62,63,64 was used to perform these estimations and corrections. The preprocessing pipeline is shown in Fig. 1d and described in detail below:
-
i)
First, intensity normalization and signal drift65,66 correction were performed. The non-DW images (b = 0 s/mm2) were used to normalize the signal intensity of the data acquired within and across the different acquisition blocks (~25 minutes each). The signal drift within each 25-minute acquisition block was corrected using scaling factors obtained from fitting a second-order polynomial on the mean signals of the non-DW images as a function of the scanned non-DW and DW images. Specifically, there were 8 non-DW images interleaved throughout each 25-minute acquisition block (one non-DW volume every 10 DW volumes, 3.2 minutes). The mean signals of the non-DW volumes were used to fit polynomial coefficients using MATLAB’s polyfit function, which were then fed into the polyval function to obtain scaling factors for DW volumes in between those non-DW volumes. The signal intensity differences across acquisition blocks were normalized to the mean signal intensity of the first session.
-
ii)
Susceptibility-induced and eddy current-induced distortion estimations and corrections were performed using topup62,67 and the eddy-current correction toolbox68,69,70 in the FSL62,63,64. The warp fields for each volume were obtained and stored for later use in the combined one-step resampling correction. Note that topup and eddy-current corrections were performed for each individual data subset (2 acquisition blocks with paired reversed phase encoding block), such that the field map can be updated across scans to account for possible field changes even with the use of motion stabilizer and matched B0 shimming. In addition to updating the field map, the interaction between susceptibility-induced field changes and participant motion within each data block was also considered by using a dynamic correction algorithm70 as implemented in the FSL eddy toolbox68,69,70.
-
iii)
The warp field for gradient nonlinearity correction60,71 was calculated for the diffusion data. Here, the gradient nonlinearity correction was performed after topup and eddy correction60. This is because of the consideration that the voxel displacements due to susceptibility and eddy current distortions are much larger than those due to within-block rigid motion. Therefore, to obtain better estimation of these larger voxel displacements, the original phase-encoding direction was maintained prior to the estimation and correction for susceptibility and eddy current-induced distortions60. The effects of performing gradient-nonlinearity at a later step on the accuracy of within-block rigid motion estimation were considered to be small, because we assume that the confined motion within each acquisition block would only cause small differences in image deformation caused by gradient non-linearity.
-
iv)
The next step estimated larger motion parameters across different data subsets such that the images from different acquisition blocks and sessions could be registered and aligned. This was performed using FLIRT72,73,74 in FSL62,63,64 with 6 degrees of freedom (DOF) rigid motion.
-
v)
Finally, the warp fields for susceptibility, eddy-current distortion, gradient nonlinearity and motion from steps ii-iv were combined and applied to each image volume in a one-step resampling process to ensure minimal smoothing effects due to interpolation60,61. The Jacobian matrix computed from the combined warp field was used to correct for the signal intensity affected by the transformation. The b-vectors were reoriented for each DW image after motion correction75.
All 18 data subsets from 9 sessions and their corresponding b-value and b-vector (reoriented) files were stored in compressed Neuroimaging Informatics Technology Initiative (NIfTI) format for downloading (Fig. 1d, data output B) and further analysis. Freesurfer76 functions were used to load and save NIfTI data.
To provide flexibility for the end users to use their own preprocessing pipeline of choice, minimally preprocessed data are also made available for download in compressed NIfTI format (Fig. 1c, data output A). With the minimally preprocessed data, only the gradient nonlinearity correction has been performed without any other preprocessing corrections. The rationale in performing this correction is that the gradient nonlinearity coefficients needed in this correction are considered proprietary information by Siemens, and are not made publicly available. Because of this, the correction for spatial variation of b-matrices due to gradient nonlinearity77,78,79,80,81 was not performed in this work.
T1 and T2-weighted images are available for download in compressed NIfTI format (Fig. 1c, data output A). After online reconstruction, the DICOM data were converted to NIfTI format using dcm2nii82 tool of MRIcron (http://www.mricro.com), and MRI gradient nonlinearity correction was performed. Subsequently, face and ear regions were masked to de-identify the high-resolution structural images using the defacing tool provided by FSL62,63,64. The averaged images of the three 0.7-mm T1 and T2-weighted images are also provided after registration, averaging and de-identification. The low-resolution field maps are provided after gradient nonlinearity correction.
Data Records
All data records listed in this section are publicly available in dryad83,84. The raw k-space data (~7 TB) can also be made available upon request.
dMRI data
-
i)
The preprocessed dMRI data are saved in zip files with a prefix of ‘processedDWI_’83, encoded by labels identifying scan session 1–9 and data subset number 01–18. The images of different data subsets are saved in separate files in the compressed NIfTI format, each including 78 volumes with PE in AP and another 78 volumes in PA. The corresponding b-value and b-vector (reoriented after motion correction) files are also provided. Data subset 01–06 were acquired at b = 1000 s/mm2, and data subset 07–18 were acquired at b = 2500 s/mm2.
-
ii)
The minimally preprocessed dMRI data are saved in zip files with a prefix of ‘reconDWI_’84. They are encoded and organized in the same format as the preprocessed dMRI data mentioned above.
Structural data
-
i)
T1-weighted images are saved under ‘T1.zip’83 (~ 350 MB) in compressed NIfTI format, containing one 1-mm MEMPRAGE image (echo combined), three 0.7-mm MEMPRAGE images (echo combined) and their average image.
-
ii)
T2-weighted images are saved under ‘T2.zip’83 (~300 MB) in compressed NIfTI format, containing three 0.7-mm T2-SPACE images and their average image.
-
iii)
Field maps for all 9 sessions are saved under ‘FieldMap.zip’83 (~14 MB) in compressed NIfTI format, labeled by numbers identifying scan session.
Supplementary to the above data, a readme text file83 is included in the repository with more information of the datasets.
Technical Validation
Assessment of DW image quality and head motion
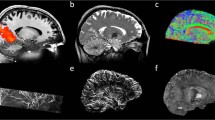
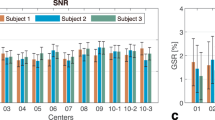
The single and mean DWIs at b = 1000 s/mm2 and b = 2500 s/mm2 in three orthogonal views are shown in Fig. 3a, which highlight the high image quality of this isotropic resolution dataset in all three spatial orientations with minimal reconstruction and ghosting artifacts. The high isotropic spatial resolution of the data reveals detailed structures of the brain (e.g., thalamus, globus pallidus, cerebellum) and sharp delineation of the gray-white matter boundaries (apparent in mean DWIs).
DW images and participant movement measurements. (a) Single DWI and mean DWI images at different b-values shown in three orthogonal views. (b) participant movement measurements of all 2808 dMRI volumes acquired in 9 two-hour sessions. From top to bottom are translations along left-right (LR), anterior-posterior (AP), head-foot (HF) and rotations around LR (Pitch), AP (Roll), HF (Yaw) axis. The motion parameters were estimated relative to the first b = 0 s/mm2 volume in the first session. dMRI volumes acquired in different sessions on different dates are separated by black dashed lines. As shown, both within-session movements and across-session head position changes were kept small with the use of the personalized motion stabilizer.
Participant motion between the 2808 dMRI volumes across the 9 two-hour sessions was evaluated in Fig. 3b. The motion parameter estimates from different sessions acquired on different dates were concatenated together in a chronological order and separated by dashed lines. Within each two-hour session, the translations along the LR and the HF directions as well as the rotations around the AP (roll) and the HF axes (yaw) were all confined within a ± 1 mm/degree range. The rotations around the LR axis was slight larger but was still within a ± 2-degree range. The estimates of translational motion along the AP direction appeared to be the highest and contained fluctuations and large jumps between the 25-minute acquisition blocks with opposing phase-encoding directions. This likely resulted from the inclusion of some eddy-current induced translation along the phase-encoding direction, which is known to be difficult to differentiate from actual participant motion since they affect the data in the same way. However, whether this translation is included in the eddy current-induced estimate or the participant movement-induced estimate will not affect the image motion correction. Across different scan sessions, the abrupt changes in motion parameters represented differences in head position placements across sessions. With the use of the personalized head motion stabilizer, the position changes across different sessions were minimized to within ± 2 mm/degrees.
Assessment of diffusion modelling outcomes of the multi-session dataset
The quality of the multi-session dataset was also assessed by evaluating the diffusion modelling outcomes. Specifically, the diffusion tensor (DT) model and a multi-shell extension of the ball & stick model85,86,87 implemented in the bedpostx tool in FSL were used. The necessity and benefits of the multi-session data acquisition were illustrated by comparing the above diffusion outcomes obtained using data from 1, 3 and 9 sessions (total dMRI data acquisition time of 1.6, 4.8 and 14.5 hours, with 140, 420 and 1260 q-space sampling points respectively). As shown in Fig. 4, increasing the amount of data significantly increased the SNR and quality of the fractional anisotropy (FA) maps. The FA maps obtained using data from all 9 sessions revealed exquisite detail within fine-scale structures, including the radiating fiber bundles within the corpus callosum, highlighted as red stripes on the FA map (Fig. 4, zoomed-in area, arrow-A); the ~1-voxel-width line of reduced-FA separating the corticospinal tract from an adjacent tract (Fig. 4, zoomed-in area, arrow-B), highlighting the low level of blurring and high spatial accuracy of this data acquired across 9 two-hour sessions; and the gray matter bridges that span the internal capsule, giving rise to the characteristic stripes seen in the striatum (Fig. 4, zoomed-in area, arrow-C). In addition, the increased SNR and the denser q-space sampling from the multi-session data dramatically increased the ability to resolve three-way crossing fibers (see Fig. 4 for increased volume fraction of the third fiber shown in red-yellow on top left) and revealed complex intermingling fiber bundles within the centrum semiovale (see Fig. 4 fiber orientation maps modelling 3 fiber compartments per voxel), where the callosal fibers, the corticospinal tract and the superior longitudinal fasciculus intersect.
Comparison of FA maps and crossing fiber analysis results using data from 1, 3 and 9 acquisition sessions. On the left, colored FA maps in three orthogonal views are shown for data acquired in 1 session (top), 3 sessions (middle) and 9 sessions (bottom). The 9-session data provides high SNR FA and reveals exquisite fine-scale structures as shown in the zoomed-in areas (yellow boxes), including (A) the radiating fiber bundles within the corpus callosum; (B) the 1-voxel-width line of reduced-FA separating the corticospinal tract from an adjacent tract, highlighting the low level of blurring and high spatial accuracy of this data; and (C) the gray matter bridges that span the internal capsule. On the right, zoomed-in fiber orientations (three fiber compartments per voxel) within centrum semiovale (yellow box) overlaid on gray-scale maps of the total anisotropic volume fraction are shown for those data as well. In addition, the volume fraction of the third fiber are shown in red-yellow on the top left of each fiber orientation map to visualize the sensitivity of solving 3-way crossing fibers. Both fiber orientation and third-fiber volume fraction are shown when the respective volume fraction is higher than 5%. The denser q-space sampling and higher SNR of the 9-session data increases its ability to solve three-way crossing fibers and to reveal complex intermingling fiber bundles within the centrum semiovale.
Assessment of spatial resolution
A comparison of diffusion results at different spatial resolutions was performed to validate the benefit of using high spatial resolution acquisition. The dMRI data at 0.76 mm isotropic resolution was compared with downsampled data at 1-mm, 1.25-mm and 1.5-mm isotropic resolution (by cropping in the k-space of the dMRI data along all three dimensions). Figure 5 shows the colored-FA maps of four zoomed-in areas (yellow squares) at those spatial resolutions. Improved delineation of fiber bundles and more detailed structures can be captured under higher spatial resolution. Figure 6 shows the comparison of DTI tensors at 1.5 mm and 0.76 mm isotropic resolution, which consistently highlights the improved capability of the submillimeter dMRI data to visualize subcortical white matter as it turns into the cortex. A clear dark band with reduced FA at the gray-white junction can be seen at 0.76-mm resolution (red arrows, top and middle rows). In addition, 0.76-mm data detected more sharp-turning fibers such as those connecting cortical regions between adjacent gyri (red arrows, bottom row). These features of high spatial resolution could potentially help improve the accuracy of tractography results (e.g., reducing gyral bias).
Comparison of DTI principal directions at 1.5-mm and 0.76-mm resolution. Zoomed-in FA and DTI principal vector maps are shown in the area highlighted by yellow boxes. High resolution dMRI data shows improved ability to visualize subcortical white matters as it turns into the cortex. A clear dark-band with reduced FA at the grey-white junction can be seen (red arrows, top and middle rows) at 0.76 mm resolution, and more sharping-turning fibers such as those connecting cortical regions between adjacent gyri (red arrows, bottom row) can be detected.
Code availability
The MATLAB code for the preprocessing steps described above are included in the released dataset83. The code make use of the functions in the FSL62,63,64 toolbox and Freesurfer76 that are freely available for download at https://fsl.fmrib.ox.ac.uk/fsl/fslwiki/FslInstallation and https://surfer.nmr.mgh.harvard.edu/fswiki/DownloadAndInstall.
References
Le Bihan, D. & Breton, E. Imagerie de diffusion in vivo par résonance magnétique nucléaire. Mécanique, Physique, Chimie, Sciences de l’univers, Sciences de la Terre. 301, 1109–1112 (1985).
Moseley, M. et al. Diffusion-weighted MR imaging of acute stroke: correlation with T2-weighted and magnetic susceptibility-enhanced MR imaging in cats. Am. J. Neuroradiol. 11, 423–429 (1990).
Warach, S., Chien, D., Li, W., Ronthal, M. & Edelman, R. R. Fast magnetic resonance diffusion‐weighted imaging of acute human stroke. Neurology 42, 1717–1717 (1992).
Mangeat, G. et al. Changes in structural network are associated with cortical demyelination in early multiple sclerosis. Hum. Brain Mapp. 39, 2133–2146 (2018).
Malayeri, A. A. et al. Principles and applications of diffusion-weighted imaging in cancer detection, staging, and treatment follow-up. Radiographics 31, 1773–1791 (2011).
Jones, D. K. et al. Age effects on diffusion tensor magnetic resonance imaging tractography measures of frontal cortex connections in schizophrenia. Hum. Brain Mapp. 27, 230–238 (2006).
Sussmann, J. E. et al. White matter abnormalities in bipolar disorder and schizophrenia detected using diffusion tensor magnetic resonance imaging. Bipolar Disord. 11, 11–18 (2009).
Pasternak, O., Kelly, S., Sydnor, V. J. & Shenton, M. E. Advances in microstructural diffusion neuroimaging for psychiatric disorders. Neuroimage 182, 259–282 (2018).
Goveas, J. et al. Diffusion-MRI in neurodegenerative disorders. Magn. Reson. Imaging 33, 853–876 (2015).
Bozzali, M. et al. White matter damage in Alzheimer’s disease assessed in vivo using diffusion tensor magnetic resonance imaging. J. Neurol. Neurosurg. Psychiatry 72, 742–746 (2002).
Eickhoff, S., Nichols, T. E., Van Horn, J. D. & Turner, J. A. Sharing the wealth: neuroimaging data repositories. Neuroimage 124, 1065 (2016).
Walker, L. et al. The diffusion tensor imaging (DTI) component of the NIH MRI study of normal brain development (PedsDTI). Neuroimage 124, 1125–1130 (2016).
Hodge, M. R. et al. ConnectomeDB—sharing human brain connectivity data. Neuroimage 124, 1102–1107 (2016).
Hsu, Y. C., Lo, Y. C., Chen, Y. J. & Wedeen, V. J. & Isaac Tseng, W. Y. NTU‐DSI‐122: A diffusion spectrum imaging template with high anatomical matching to the ICBM‐152 space. Hum. Brain Mapp. 36, 3528–3541 (2015).
Varentsova, A., Zhang, S. & Arfanakis, K. Development of a high angular resolution diffusion imaging human brain template. NeuroImage 91, 177–186 (2014).
Mori, S., Wakana, S., Van Zijl, P. C. & Nagae-Poetscher, L. MRI atlas of human white matter. Elsevier (2005).
Poldrack, R. A. et al. Long-term neural and physiological phenotyping of a single human. Nat. Commun 6, 1–15 (2015).
Froeling, M., Tax, C. M., Vos, S. B., Luijten, P. R. & Leemans, A. “MASSIVE” brain dataset: Multiple acquisitions for standardization of structural imaging validation and evaluation. Magn. Reson. Med. 77, 1797–1809 (2017).
Uğurbil, K. et al. Pushing spatial and temporal resolution for functional and diffusion MRI in the Human Connectome Project. NeuroImage 80, 80–104 (2013).
Setsompop, K. et al. Pushing the limits of in vivo diffusion MRI for the Human Connectome Project. NeuroImage 80, 220–233 (2013).
McNab, J. A. et al. The Human Connectome Project and beyond: initial applications of 300 mT/m gradients. NeuroImage 80, 234–245 (2013).
Van Essen, D. C. et al. The WU-Minn human connectome project: an overview. Neuroimage 80, 62–79 (2013).
Miller, K. L. et al. Multimodal population brain imaging in the UK Biobank prospective epidemiological study. Nat. Neurosci. 19, 1523–1536 (2016).
Miller, K. L. et al. Diffusion imaging of whole, post-mortem human brains on a clinical MRI scanner. Neuroimage 57, 167–181 (2011).
Dyrby, T. B. et al. An ex vivo imaging pipeline for producing high‐quality and high‐resolution diffusion‐weighted imaging datasets. Hum. Brain Mapp. 32, 544–563 (2011).
Leuze, C. W. et al. Layer-specific intracortical connectivity revealed with diffusion MRI. Cereb. Cortex 24, 328–339 (2014).
Fan, Q. et al. HIgh b-value and high Resolution Integrated Diffusion (HIBRID) imaging. NeuroImage 150, 162–176 (2017).
McNab, J. A. et al. Surface based analysis of diffusion orientation for identifying architectonic domains in the in vivo human cortex. NeuroImage 69, 87–100 (2013).
McNab, J. A. et al. The Human Connectome Project and beyond: initial applications of 300mT/m gradients. Neuroimage 80, 234–245 (2013).
Keil, B. et al. A 64‐channel 3T array coil for accelerated brain MRI. Magn. Reson. Med. 70, 248–258 (2013).
Chen, N.-k, Guidon, A., Chang, H.-C. & Song, A. W. A robust multi-shot scan strategy for high-resolution diffusion weighted MRI enabled by multiplexed sensitivity-encoding (MUSE). Neuroimage 72, 41–47 (2013).
Wu, W. et al. High-resolution diffusion MRI at 7T using a three-dimensional multi-slab acquisition. NeuroImage 143, 1–14 (2016).
Dong, Z. et al. Interleaved EPI diffusion imaging using SPIR i T‐based reconstruction with virtual coil compression. Magn. Reson. Med. 79, 1525–1531 (2018).
Dong, Z. et al. Tilted‐CAIPI for highly accelerated distortion‐free EPI with point spread function (PSF) encoding. Magn. Reson. Med. 81, 377–392 (2019).
Frost, R., Jezzard, P., Porter, D. A., Tijssen, R. & Miller, K. In Proceedings of the 21st Annual Meeting of ISMRM, Salt Lake City, USA, p 3176.
Engström, M. & Skare, S. Diffusion‐weighted 3D multislab echo planar imaging for high signal‐to‐noise ratio efficiency and isotropic image resolution. Magn. Reson. Med. 70, 1507–1514 (2013).
Song, A. W., Chang, H.-C., Petty, C., Guidon, A. & Chen, N.-K. Improved delineation of short cortical association fibers and gray/white matter boundary using whole-brain three-dimensional diffusion tensor imaging at submillimeter spatial resolution. Brain Connect. 4, 636–640 (2014).
Chang, H.-C. et al. Human brain diffusion tensor imaging at submillimeter isotropic resolution on a 3Tesla clinical MRI scanner. NeuroImage 118, 667–675 (2015).
Setsompop, K. et al. High‐resolution in vivo diffusion imaging of the human brain with generalized slice dithered enhanced resolution: Simultaneous multislice (gSlider-SMS). Magn. Reson. Med. 79, 141–151 (2018).
Wang, F. et al. Motion‐robust sub‐millimeter isotropic diffusion imaging through motion corrected generalized slice dithered enhanced resolution (MC-gSlider) acquisition. Magn. Reson. Med. 80, 1891–1906 (2018).
Haldar, J. P., Liu, Y., Liao, C., Fan, Q. & Setsompop, K. Fast submillimeter diffusion MRI using gSlider‐SMS and SNR‐enhancing joint reconstruction. Magn. Reson. Med. 84, 762–776 (2020).
Fan, Q. et al. MGH–USC Human Connectome Project datasets with ultra-high b-value diffusion MRI. Neuroimage 124, 1108–1114 (2016).
Hoge, W. S. & Polimeni, J. R. Dual‐polarity GRAPPA for simultaneous reconstruction and ghost correction of echo planar imaging data. Magn. Reson. Med. 76, 32–44 (2016).
Hoge, W. S., Setsompop, K. & Polimeni, J. R. Dual‐polarity slice‐GRAPPA for concurrent ghost correction and slice separation in simultaneous multi‐slice EPI. Magn. Reson. Med. 80, 1364–1375 (2018).
Calabrese, E. et al. Investigating the tradeoffs between spatial resolution and diffusion sampling for brain mapping with diffusion tractography: Time well spent? Hum. Brain Mapp. 35, 5667–5685 (2014).
Zhan, L. et al. Angular versus spatial resolution trade-offs for diffusion imaging under time constraints. Hum. Brain Mapp. 34, 2688–2706 (2013).
Vos, S. B. et al. Trade-off between angular and spatial resolutions in in vivo fiber tractography. NeuroImage 129, 117–132 (2016).
Ades-Aron, B. et al. Evaluation of the accuracy and precision of the diffusion parameter EStImation with Gibbs and NoisE removal pipeline. NeuroImage 183, 532–543 (2018).
Veraart, J. et al. Denoising of diffusion MRI using random matrix theory. Neuroimage 142, 394–406 (2016).
St-Jean, S., Coupé, P. & Descoteaux, M. Non Local Spatial and Angular Matching: Enabling higher spatial resolution diffusion MRI datasets through adaptive denoising. Med. Image Anal. 32, 115–130 (2016).
Setsompop, K. et al. Blipped‐controlled aliasing in parallel imaging for simultaneous multislice echo planar imaging with reduced g‐factor penalty. Magn. Reson. Med. 67, 1210–1224 (2012).
Caruyer, E., Lenglet, C., Sapiro, G. & Deriche, R. Design of multishell sampling schemes with uniform coverage in diffusion MRI. Magn. Reson. Med. 69, 1534–1540 (2013).
Van der Kouwe, A. J., Benner, T., Salat, D. H. & Fischl, B. Brain morphometry with multiecho MPRAGE. Neuroimage 40, 559–569 (2008).
Lichy, M. P. et al. Magnetic resonance imaging of the body trunk using a single-slab, 3-dimensional, T2-weighted turbo-spin-echo sequence with high sampling efficiency (SPACE) for high spatial resolution imaging: initial clinical experiences. Investig. Radiol. 40, 754–760 (2005).
Sotiropoulos, S. N. et al. Effects of image reconstruction on fiber orientation mapping from multichannel diffusion MRI: reducing the noise floor using SENSE. Magn. Reson. Med. 70, 1682–1689 (2013).
Uecker, M. et al. ESPIRiT—an eigenvalue approach to autocalibrating parallel MRI: where SENSE meets GRAPPA. Magn. Reson. Med. 71, 990–1001 (2014).
Haacke, E., Lindskogj, E. & Lin, W. A fast, iterative, partial-Fourier technique capable of local phase recovery. J. Magn. Reson. 92, 126–145 (1991).
Eichner, C. et al. Real diffusion-weighted MRI enabling true signal averaging and increased diffusion contrast. NeuroImage 122, 373–384 (2015).
Liao, C. et al. High‐fidelity, high‐isotropic‐resolution diffusion imaging through gSlider acquisition with and T1 corrections and integrated ΔB0/Rx shim array. Magn. Reson. Med. 83, 56–67 (2019).
Glasser, M. F. et al. The minimal preprocessing pipelines for the Human Connectome Project. Neuroimage 80, 105–124 (2013).
Eichner, C. et al. A Joint Recommendation for Optimized Preprocessing of Connectom Diffusion MRI Data. In Proceedings of the 27th Annual Meeting of ISMRM, Montreal, Canada. p 1047.
Smith, S. M. et al. Advances in functional and structural MR image analysis and implementation as FSL. Neuroimage 23, S208–S219 (2004).
Woolrich, M. W. et al. Bayesian analysis of neuroimaging data in FSL. Neuroimage 45, S173–S186 (2009).
Jenkinson, M., Beckmann, C. F., Behrens, T. E., Woolrich, M. W. & Smith, S. M. Fsl. Neuroimage 62, 782–790 (2012).
Vos, S. B. et al. The importance of correcting for signal drift in diffusion MRI. Magn. Reson. Med. 77, 285–299 (2017).
Jeurissen, B., Leemans, A., Tournier, J. D., Jones, D. K. & Sijbers, J. Investigating the prevalence of complex fiber configurations in white matter tissue with diffusion magnetic resonance imaging. Hum. Brain Mapp. 34, 2747–2766 (2013).
Andersson, J. L., Skare, S. & Ashburner, J. How to correct susceptibility distortions in spin-echo echo-planar images: application to diffusion tensor imaging. Neuroimage 20, 870–888 (2003).
Andersson, J. L. & Sotiropoulos, S. N. An integrated approach to correction for off-resonance effects and subject movement in diffusion MR imaging. Neuroimage 125, 1063–1078 (2016).
Andersson, J. L., Graham, M. S., Zsoldos, E. & Sotiropoulos, S. N. Incorporating outlier detection and replacement into a non-parametric framework for movement and distortion correction of diffusion MR images. NeuroImage 141, 556–572 (2016).
Andersson, J. L., Graham, M. S., Drobnjak, I., Zhang, H. & Campbell, J. Susceptibility-induced distortion that varies due to motion: correction in diffusion MR without acquiring additional data. Neuroimage 171, 277–295 (2018).
Jovicich, J. et al. Reliability in multi-site structural MRI studies: effects of gradient non-linearity correction on phantom and human data. Neuroimage 30, 436–443 (2006).
Jenkinson, M., Bannister, P., Brady, M. & Smith, S. Improved optimization for the robust and accurate linear registration and motion correction of brain images. Neuroimage 17, 825–841 (2002).
Jenkinson, M. & Smith, S. A global optimisation method for robust affine registration of brain images. Med. Image Anal. 5, 143–156 (2001).
Greve, D. N. & Fischl, B. Accurate and robust brain image alignment using boundary-based registration. Neuroimage 48, 63–72 (2009).
Leemans, A. & Jones, D. K. The B‐matrix must be rotated when correcting for subject motion in DTI data. Magn. Reson. Med. 61, 1336–1349 (2009).
Fischl, B. FreeSurfer. Neuroimage 62, 774–781 (2012).
Bammer, R. et al. Analysis and generalized correction of the effect of spatial gradient field distortions in diffusion-weighted imaging. Magn. Reson. Med. 50, 560–569 (2003).
Rudrapatna, U., Parker, G. D., Roberts, J. & Jones, D. K. A comparative study of gradient nonlinearity correction strategies for processing diffusion data obtained with ultra-strong gradient MRI scanners. Magn. Reson. Med. 85, 1104–1113 (2021).
Mohammadi, S. et al. The effect of local perturbation fields on human DTI: characterisation, measurement and correction. Neuroimage 60, 562–570 (2012).
Malyarenko, D. I., Ross, B. D. & Chenevert, T. L. Analysis and correction of gradient nonlinearity bias in apparent diffusion coefficient measurements. Magn. Reson. Med. 71, 1312–1323 (2014).
Borkowski, K., Kłodowski, K., Figiel, H. & Krzyżak, A. T. A theoretical validation of the B-matrix spatial distribution approach to diffusion tensor imaging. Magn. Reson. Imaging 36, 1–6 (2017).
Li, X., Morgan, P. S., Ashburner, J., Smith, J. & Rorden, C. The first step for neuroimaging data analysis: DICOM to NIfTI conversion. J. Neurosci. Methods 264, 47–56 (2016).
Wang, F. et al. Data from: In vivo human whole-brain Connectom diffusion MRI dataset at 760 µm isotropic resolution (PART I). Dryad https://doi.org/10.5061/dryad.nzs7h44q2 (2021).
Wang, F. et al. Data from: In vivo human whole-brain Connectom diffusion MRI dataset at 760 µm isotropic resolution (PART II). Dryad https://doi.org/10.5061/dryad.rjdfn2z8g (2021).
Behrens, T. E., Berg, H. J., Jbabdi, S., Rushworth, M. F. & Woolrich, M. W. Probabilistic diffusion tractography with multiple fibre orientations: What can we gain? Neuroimage 34, 144–155 (2007).
Jbabdi, S., Sotiropoulos, S. N., Savio, A. M., Graña, M. & Behrens, T. E. Model‐based analysis of multishell diffusion MR data for tractography: how to get over fitting problems. Magn. Reson. Med. 68, 1846–1855 (2012).
Hernández, M. et al. Accelerating fibre orientation estimation from diffusion weighted magnetic resonance imaging using GPUs. PloS one 8, e61892 (2013).
Acknowledgements
This work was supported by the NIH (R01-EB020613, R01-EB019437, R01-MH116173, U01-EB025162, and U01-EB026996, U01-MH093765, P41-EB015896, P41-EB030006, K23-NS096056) and NIH shared instrumentation grants (S10-RR023401, S10-RR023043, and S10-RR019307). We thank Dr. Cornelius Eichner for sharing his python-based code for optimized preprocessing of dMRI data with one-step resampling.
Author information
Authors and Affiliations
Contributions
F.W. conceptualized and conducted the study, acquired and processed the data, and wrote the manuscript. Z.D. helped with the data acquisition and the data processing, and contributed to the manuscript. Q.T., C.L. and Q.F. helped with the data processing. W.S.H. helped with the D.P.G. reconstruction. B.K. provided the 64-channel phased-array coil. J.R.P., L.L.W. and S.Y.H. provided the conceptual discussion and contributed to the manuscript. K.S. conceptualized and supervised the study, and contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files associated with this article.
About this article
Cite this article
Wang, F., Dong, Z., Tian, Q. et al. In vivo human whole-brain Connectom diffusion MRI dataset at 760 µm isotropic resolution. Sci Data 8, 122 (2021). https://doi.org/10.1038/s41597-021-00904-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-021-00904-z
This article is cited by
-
A multimodal submillimeter MRI atlas of the human cerebellum
Scientific Reports (2024)
-
Optimal deep brain stimulation sites and networks for stimulation of the fornix in Alzheimer’s disease
Nature Communications (2022)
-
Anatomy and physiology of word-selective visual cortex: from visual features to lexical processing
Brain Structure and Function (2021)