Abstract
Germline stem cells are germ cells at an early developmental stage, so their development is key to ensuring human reproduction. There is increasing evidence that long noncoding RNA (lncRNA) and circular RNA (circRNA) play important roles in the development of germ cells. This data descriptor provides unique lncRNA and circRNA transcriptomic information for mouse germline stem cells. Using the Illumina HiSeqx 2000 system, a total of 511,836,732 raw reads were generated. High-quality transcripts, lncRNAs, and circRNAs were identificated and quantified using the reads, and more precise annotations of lncRNAs (especially 9357 novel lncRNAs) and circRNAs were performed in the germline stem cells. We then analyzed the transcript structures, genetic variants, and the interaction between circRNA and microRNA to provide the basis for subsequent functional experiments. This comprehensive dataset will help advance data sharing and deepen our understanding of mouse germline stem cells, providing a theoretical foundation for research on germ cell development and human reproduction, among others.
Design Type(s) | transcription profiling design • cell type comparison design |
Measurement Type(s) | transcription profiling assay |
Technology Type(s) | RNA sequencing |
Factor Type(s) | technical replicate |
Sample Characteristic(s) | Mus musculus • testis • ovary |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
There is growing recognition that cells, especially in mammals, produce many thousands of noncoding transcripts. long noncoding RNA (lncRNA), a group of noncoding RNAs that are longer than 200 bp, are emerging as potent regulators of gene expression, and have dramatically altered our understanding of cell biology under pathological conditions1,2. Owing to the strong time- and tissue-specificity of lncRNA expression, there is great potential for novel lncRNA prediction. Circular RNAs (circRNAs) are a recently discovered group of noncoding RNAs identified by the presence of a special circular structure formed by covalent bonds3,4. In eukaryotic organisms, circRNAs are mostly present in the cytoplasm, but a few intron-cyclized circRNAs are localized in the nucleus5. They are widespread in mammalian cells, sometimes being more common than linear RNA. CircRNAs are also highly conserved and are not easily degraded by RNase. As millions of transcripts are generated by next-generation sequencing, many lncRNAs and circRNAs have been identified, but few have been functionally characterized. In this context, it is important to be able to determine the connections between lncRNAs/circRNAs and their target mRNAs, and to clarify their potential functions. RNA-seq is an effective technology that utilizes the features of next-generation sequencing to study lncRNA and circRNA6. It can be used in combination with reference annotation-based assembly, which facilitates the detection of novel isoforms being applied in fundamental scientific research, clinical diagnosis, pharmacogenomics research, and drug R&D, among others7.
It has been reported that approximately 48.5 million couples experience problems with infertility each year globally8. In recent years, with rapid economic growth and the acceleration of economic globalization, there have been increases in the proportion of women pursuing careers and the proportion of marriages that end in divorce. As such, childbearing has been put off until a later age, but reproductive capacity also declines with age. Taking all of these factors together, it is understandable that infertility rates continue to grow. Germline stem cells are germ cells at an early developmental stage, so their development is key to ensuring successful human reproduction. Germ stem cells are a group of cells that can transmit genetic information from parent to offspring and have the characteristics of both germ cells and stem cells9,10. Main examples of them include spermatogonial stem cells (SSCs) and female germline stem cells (FGSCs)11. Comprehensive transcriptome analyses can provide a solid foundation to understand the functions of lncRNAs and circRNAs in sex determination and the differentiation of male and female germline stem cells. Specifically, transcriptomic profiles of germline stem cells as reported in this paper should make a major contribution to the discovery of molecular markers, help to uncover the regulatory mechanisms of germline stem cells, and be useful as a reference for studies on the self-renewal and differentiation of such cells.
Although the expression profiles of lncRNAs and circRNAs in the mouse germline stem cells were provided in our previous study12, the current data descriptor provided more detailed descriptions of these unique lncRNA and circRNA transcriptomic datasets of the mouse germline stem cells, including both the methods used to collect the data and technical analyses supporting the quality of the measurements. Moreover, lncRNAs (especially, 9357 novel lncRNAs) were analyzed and annotated in a more precise manner (including interaction analysis of complementary lncRNA-mRNA13, up/down stream lncRNA of a gene14,15,16, pre-miRNA prediction17, and lncRNA family prediction18). More importantly, in view of the most important regulation mode of circRNA (circRNA as an endogenous competitive RNA (ceRNA) affecting the post-transcriptional regulation function of microRNAs), we made a thorough analysis of the interaction between circRNAs and microRNAs to provide the basis for subsequent functional experiments. In addition, tens of thousands of examples of single-nucleotide polymorphisms (SNPs), insertion–deletions (indels), alternative splicing (AS), and differential exon usage (DEU) were identified in these cells. This data descriptor will help advance data sharing and reuse to support reproducible research. The obtained data should also serve as an important reference for studying germ cell development and human reproduction, among others.
Methods
Animals
We purchased Mvh-Cre mice (FVB-Tg(Ddx4-Cre)1Dcas/J) and mT/mG mice (B6.129(Cg)-Gt(Rosa)26Sortm4(ACTB-tdTomato,−EGFP)Luo/J) from Model Animal Research Center of Nanjing University. Mvh-Cre; mT/mG mice were produced using Mvh-Cre and mT/mG mice as mentioned previously19. Briefly, male Mvh-Cre mice were bred with female wild type females, while homozygous mT/mG mice were bred together. The mT/mG mice have two kinds of membrane-targeted fluorescent proteins (namely tdTomato and EGFP) at the Rosa26 locus. Each side of the membrane-targeted tdTomato (mT) cassette has a loxP site, which shows red fluorescence in all the tissues. When mT/mG mice were hybridized with CRE-expressing mice, the tdTomato cassette was removed by CRE-mediated recombination and immediately began to express downstream membrane-targeted EGFP (mG) cassette in the CRE-expressing cells of the offspring. To obtain the Mvh -Cre; mT/mG mice, male Mvh -Cre mice (younger than 63 days old) were hybridized with female mT/mG mice. The Mvh -driven CRE was expressed in the germline lineage and led to a change in expression from tdTomato to EGFP in the germ cells of this strain. All procedures involving animals were approved by the Institutional Animal Care and Use Committee of Shanghai, and were conducted on the basis of the National Research Council Guide for Care and Use of Laboratory Animals.
Mouse germline stem cell collection
To isolate and purify SSCs from mice, the testes of 50 male mice (Mvh-Cre; mT/mG mice, 6 days old) were collected and cut into small pieces. Then, SSCs were isolated by two-step enzymatic digestion, as mentioned previously20. After passing through a 13-µm nylon cell filter, the GFP-positive cells were sorted by flow cytometry (Fig. 1a). Lastly, GFP-positive cells were suspended in PBS and plated in 35 mm cell culture plate precoated with mouse laminin (4.4 µg/cm2). After incubated for 45 min at 37 °C, unbound cells were removed from bound cells by pipetting, and stored at −80 °C with proper amount of TRIzol reagent added. Fifty male mice were divided into 3 groups (16 or 17 mice for each group). All SSCs samples were collected in one day. To isolate and purify FGSCs from mice, the ovaries of 500 female mice (Mvh-Cre; mT/mG mice, 6 days old) were collected and cut into small pieces. Then, FGSCs were isolated by two-step enzymatic digestion, as mentioned previously10,21. After passing through a 13-µm nylon cell filter, the GFP-positive cells were sorted by flow cytometry (Fig. 1b). Lastly, GFP-positive cells were suspended in PBS and plated in 35 mm cell culture plate precoated with mouse laminin (4.4 µg/cm2). After incubated for 45 min at 37 °C, unbound cells were removed from bound cells by pipetting, and stored at −80 °C with proper amount of TRIzol reagent added. Five hundred female mice were divided into 3 groups (160–170 mice for each group). Each group of FGSCs samples were collected in 1 day.
Isolation and purification of mouse SSCs and FGSCs. (a) Representative examples of SSCs purification with FACS and morphology of SSCs under fluorescence microscopy after purification. (b) Representative examples of FGSCs purification with FACS and morphology of FGSCs under fluorescence microscopy after purification. Scale bars: 20 μm.
Characterization of mouse SSCs and FGSCs
To characterize these cells, we assessed marker gene expression of SSCs and FGSCs: Gfrα1 (GDNF family receptor alpha 1), Oct4 (also known as Pou5f1, POU domain class 5, transcription factor 1), Etv5 (ets variant 5), Dazl (deleted in azoospermia-like), Fragilis (also termed Ifitm3, interferon induced transmembrane protein 3), and Mvh (also known as Ddx4, DEAD (Asp-Glu-Ala-Asp) box polypeptide 4). First, we determined the expression of these marker genes using reverse transcription PCR (RT-PCR). Total RNA was extracted from SSCs and FGSCs using Trizol reagent, according to the manufacturer’s protocol. Approximately 1000 ng RNA was used to synthesize cDNA (Complementary Deoxyribonucleic acid) using M-MLV reverse transcriptase in a 20 µl volume. PCR analysis was carried out with Taq DNA polymerase with primer sets specific for each gene22. The PCR conditions were 95 °C for 2 min, followed by 30 cycles comprising 95 °C for 30 s, 58 °C for 30 s, and 72 °C for 1 min, finally followed by 72 °C for 10 min, and storage at 4 °C. The glyceraldehyde −3- phosphate dehydrogenase (GAPDH) gene was amplified in each sample as a loading control. Samples were resolved through 1.5% agarose gels and run under the same conditions. DNA bands were detected using nucleic acid gel stain. RT-PCR results showed that the cells expressed Gfrα, Oct4, Etv5, Dazl, Fragilis, and Mvh (Fig. 2a). Then, we confirmed the expression of marker genes using immunofluorescence analysis. For this analysis, the cells cultured in 48-well plates were washed with 1× phosphate-buffered saline (PBS), fixed in 4% formaldehyde for 30 min at room temperature, and then washed three times with PBS for 5 min each. Then, the cells were incubated at 37 °C for 10 min in blocking buffer (PBS containing 10% goat serum). Next, the cells were incubated overnight at 4 °C with primary rabbit anti-GFRA1 antibody (1:100, ABclonal), or anti-OCT4 antibody (1:150, Santa Cruz). After washing three times with PBS, the cells were incubated at 37 °C for 30 min with a 1:150 diluted tetramethylrhodamine isothiocyanate (TRITC)-conjugated secondary antibody (goat anti-rabbit IgG; ProteinTech). The cells were incubated at 37 °C for 10 min with 500 ng/mL 4′,6-diamidino-2-phenylindole (DAPI; Sigma). Images were acquired using a Leica digital camera under a fluorescence microscope (DM2500, DMI3000B; Leica). Positive results were shown for GFRA1, and OCT4 proteins (Fig. 2b). All the characteristics clearly demonstrated the cells were actually pure population of SSCs and FGSCs.
Characterization of mouse SSCs and FGSCs. (a) RT-RCR analysis of germline stem cell markers in SSCs and FGSCs. M, 250 bp DNA maker; lane 1, SSCs; lane 2, FGSCs; lane 3, no template control. (b) The SSCs was detected by immunofluorescence analysis with the antibodies against GFRA1 and OCT4. Scale bars: 10 μm. (c) The FGSCs was detected by immunofluorescence analysis with the antibodies against GFRA1 and OCT4. Scale bars: 10 μm.
RNA extraction, cDNA library establishment, and Illumina sequencing
Total RNA was extracted from SSCs and FGSCs using Trizol reagent (Invitrogen, CA, USA), following the manufacturer’s protocol, and the RNA integrity number (RIN) was determined to evaluate RNA integrity using an Agilent Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA, USA). Ribosomal RNA was removed from total RNA to maximize the retention of all ncRNA. The obtained RNA was randomly cut into short fragments, and then this fragmented RNA was used as a template to synthesize the first strand of cDNA using random hexamers. The second strand was synthesized by adding buffer, dNTPs, RNase H, and DNA polymerase I. After purification using the QiaQuick PCR kit, end repair with EB buffer, single nucleotide A (adenine) addition, and connection with adapters, the second strand was finally degraded using uracil-N-glycosylase (UNG)23. Then, agarose gel electrophoresis was used to select fragment size. The suitable fragments were selected as templates for PCR amplification. Finally, a sequencing library was constructed and sequenced using the Illumina HiSeq 2000 platform (Table 1).
Filtering out “dirty” raw reads and alignment of reads to ribosomal RNA
To ensure the quality of data, raw data should be quality-controlled before information analysis, and data noise could be reduced by filtering. We here refer to reads containing an adapter, excessive “N,” or a large number of low-quality bases as “dirty” raw reads, which need to be removed before information analysis. The steps of filtering are as follows: 1) remove reads with adapters; 2) remove reads containing more than 10% “N”; and 3) remove low-quality reads (bases with Q < 10 constitute more than 50% of all reads). After filtering, 233,978,622 and 246,157,442 clean reads remained in the SSC and FGSC libraries, respectively, and were used for downstream bioinformatic analysis.
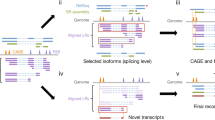
lncRNA identification
We used the transcriptome data comparison software TopHat224 to compare the filtered reads to the reference genome (UCSC mm10). Reads were assembled with Cufflinks25 after mapping to the genome. Considering the incomplete assembly of transcripts due to read coverage gaps, we performed reference annotation-based transcript (BRAT)26 assembly. The final assembled transcripts were compared with the reference gene, and the fragments that were roughly identical to the known transcripts were removed. After the assembly, we obtained the whole parsimonious set of transcripts; to detect the novel transcripts from the initial assembly, we compared the assembly transcripts to the reference annotation by utilizing Cuffcompare25. We utilized Cuffmege to merge several assemblies together; it automatically filtered a number of transfrags that were probably artifacts and produced a single annotation file for use in downstream differential analysis27. We evaluated and compared several software programs for lncRNA prediction, and chose Coding-Non-Coding Index (CNCI)28, Coding Potential Calculator (CPC)29, and iSeeRNA30, which performed well compared with the other software in both accuracy and efficiency. The findings that matched across all of the software were used to define novel lncRNA transcripts. For known lncRNA identification, we used the Gencode (GRCm38, M19) (https://www.gencodegenes.org/mouse/). The information on the annotation of lncRNAs in the Gencode database is currently more comprehensive than that in other databases, and the classification information of lncRNAs is more detailed. At the same time, we integrated 13 other databases to further verify the reliability of the lncRNAs: noncode (V4.0), RefSeq, UCSC (mm10), Ensemble, Hox_ncRNAs, Antisense_ncRNA_pipeline, fRNAdb, Affymetrix, TUCP, lncRNAdisease, eRNAs, ncRNA_imprint, and Hox_cluster_ncRNAs. The final potential lncRNAs were obtained by filtering the above basic properties and coding potential. Overall, 25704 expressed lncRNAs (16347 known and 9357 novel lncRNAs), 14270 expressed protein coding transcripts were identified and subjected to further analysis (Fig. 3a). The lncRNA identification data were deposited in figshare22. To display the distribution of lncRNA candidates along the chromosomes more intuitively, chromosome density distributions over the regions that were annotated as lncRNAs were analyzed statistically22. Moreover, we presented examples of RNA-seq read density over randomly selected mRNA, known lncRNA and novel lncRNA (Fig. 3b). Based on the fragments per kilobase of transcript per million mapped reads (FPKM) of each transcript, calculated by “Cufflinks” “abundance estimation mode” across SSCs and FGSCs, we compared the differences in expression among known lncRNAs, novel lncRNAs, and protein-coding genes. The average expression levels of lncRNAs were lower than those for protein coding genes while those of novel lncRNAs were similar to those of known lncRNAs (Fig. 3c). According to its location relative to nearby protein-coding genes, a lncRNA could be classified as sense overlap lncRNA, bidirectional lncRNA, antisense lncRNA, or intergenic lncRNA. When comparing the expression levels of the different classes of the lncRNAs, we found that the expression levels of lncRNAs of each type were similar (Fig. 3d).
Identification and expression analysis of lncRNA candidates in germline stem cells. (a) A pie diagram showing the number of novel lncRNAs, known lncRNAs and mRNAs. (b) Examples of RNA-seq read density over random select mRNA, known lncRNA and novel lncRNA. (c) Represents the density of expression of known lncRNAs, novel lncRNAs and mRNAs in germline stem cells. N = 39974, Bandwidth = 0.2745. (d) Expression level of four types of lncRNAs.
In order to analyze the structural charateristics of lncRNAs in the germline stem cells, we analyzed and compared the transcript length distribution, ORF length, and exon number between lncRNAs and mRNAs. Our analyses showed that transcript length distribution, ORF length, and exon number of lncRNAs were different from those of protein- coding transcripts. (Fig. 4a–c). The average transcript lengths of known and novel lncRNAs were much shorter than those of protein- coding transcripts. (Fig. 4a). Next, we compared the ORF length between the lncRNAs and the mRNAs. The principle of ORF analysis is based mainly on the six-frame translation of nucleic acids. Our results showed that the average ORF lengths of the lncRNAs and the mRNAs were 86.24 bp and 394.84 bp, respectively (Fig. 3b). This indicated that the ORF length of mRNAs were significantly longer than those of the lncRNAs. The main reason behind this is that lncRNAs do not encode proteins. Finally, the number of exons of lncRNAs and mRNAs was compared and analyzed, and the results showed that known and novel lncRNAs had fewer exons than mRNAs (Fig. 3c).
Sequence features and expression analysis of identified lncRNAs. (a) Length distribution of known and novel lncRNAs, and protein coding transcripts. (b) ORF distribution of lncRNAs and protein coding transcripts. (c) Exon number distribution of known and novel lncRNAs, and protein coding transcripts.
lncRNA annotation and functional prediction
The annotation and functional prediction of lncRNAs were performed based on the mechanism of lncRNA function17: (1) Interaction analysis of complementary lncRNA–mRNA. To reveal the interaction between antisense lncRNA and RNA, we used RNAplex31, software for finding short interactions between two long strands of RNA, to predict complementary binding between antisense lncRNA and mRNA. The program includes the Vienna RNA package, and calculates the minimum free energy according to the thermodynamic structure to predict the best base pairing relationship. The results showed the best lncRNA–mRNA base pairing sites and the minimum free energy of antisense lncRNA and its corresponding mRNA. (2) For lncRNAs up/downstream of a gene, we annotated those classified as being located in an “unknown region” in the former analysis, if they were upstream or downstream of a gene. These lncRNAs could potentially overlap with cis-regulatory elements that are probably involved in transcriptional regulation. (3) Pre-miRNA prediction. We aligned lncRNAs to miRBase32 to detect potential pre-miRNAs, with those showing hit coverage higher than 90% being selected. Support vector machine (SVM)-based software miRPara33 was also used to predict probable miRNAs. It classified sequences from miRBase into animal, plant, and overall categories and used an SVM to train three models based on the physical properties of pre-miRNA and its miRNAs. (4) lncRNA family prediction. To better annotate lncRNA at the level of evolution, we used INFERNAL to classify all predicted lncRNAs according to their conserved sequence and secondary structure through multiple sequence alignment, secondary structure, and a covariance model33. The lncRNA annotation data were deposited in figshare22.
CircRNA identification
In accordance with the structural characteristics and splicing sequence characteristics of circRNAs, we used the following methods to identify them34,35,36: (1) Given the circular character of circRNAs, a database of junction reads was first established by using clean data that could not be compared with the mouse genome. With these junction reads as anchors, alternatively spliced transcripts were assembled by extending Cufflinks to both ends. (2) The assembled transcripts were divided into two sections centered on the junction reads and BLAST localization of the genome was performed; if the positions of the two sections in the mouse genome were reversed, the transcripts were considered as candidate circRNAs. (3) The candidate circRNAs were further filtered for protein-coding potential. The filtering parameters were as follows: Pysocsf (score < 100), CPC (score < 10), CNCI (score < 0), and Pofam (score < 0). Overall, 18822 expressed circRNAs derived from 5334 host genes were identified and subjected to further analysis (Fig. 5a). Among these 18822 circRNAs, 98% were exonic circRNAs (Fig. 5b). Moreover, the vast majority of exonic circRNAs identified were processed from exons in the middle of their hosting genes, with only a few including the first or the last exons (0.1% for the first exon, and none for the last exon) (Fig. 5c). The circRNA identification data were deposited in figshare22.
Sequence features of circRNAs. (a) Histogram showing the number of circRNAs and their hosting genes. (b) Ninety-eight percent of circRNAs were exonic circRNAs. (c) Distribution of the back-spliced exons in circRNAs. Nearly all (99.9%) back-spliced exons that contribute to circRNAs were located in the middle of their hosting genes, whereas 19 were in the first exon and none was in the last exon, as annotated. (d) Distribution of circRNA candidates in chromosomes. Each chromosome was mapped setting 25 MB as the basic unit. The expression of circRNAs in each segment was analyzed when the full array of circRNAs in different samples was visualized. SSC-1, SSC-2, SSC-3, FGSC-1, FGSC-2, and FGSC-3 were presented from the center of the circle outward, in this order. (e) Represents the density of expression of circRNAs in germline stem cells.
To display the distribution of circRNA candidates along the chromosomes more intuitively, we used Circos software (http://circos.ca/) to map the genomic locations of circRNAs screened as described above. We performed genome mapping according to different samples of circRNAs (Fig. 5d). We also analyzed the expression density of circRNAs in the germline stem cells. The results showed that the expression density of each sample conformed to the normal distribution and the average expression levels of circRNAs were lower than those of the protein- coding genes (Fig. 5e).
CircRNA annotation and functional prediction
On the basis of the relationship between the locations of circRNAs in the genome and protein-coding genes, the screened circRNA candidates were annotated, which mainly involved categorical and functional annotations. The categorical annotations of circRNAs were mainly based on their positional relationships in the genome, which can be divided into five types: intronic circRNA, exonic circRNA, antisense circRNA, sense overlapping circRNA, and intergenic circRNA. The functional annotation of circRNAs was based on the mechanism by which the circRNAs form. This mainly involves the annotation of circRNA function according to the corresponding circRNA-hosting gene function, including gene function annotation, gene ontology (GO) annotation, and Kyoto Encyclopedia of Genes and Genomes (KEGG) annotation35. The circRNA annotation data were deposited in figshare22.
Analysis of circRNA–miRNA interaction network
The analysis of interactions between circRNAs and miRNAs was mainly based on Targetscan 7.0 software (http://www.targetscan.org/) and miRanda software (http://www.microrna.org/microrna/home.do). The former software predicts the target of microRNAs based on the seed region, while the latter is mainly based on the size of the binding free energy between circRNA and miRNA. The smaller the binding free energy, the stronger the binding ability, and the screening threshold is max. energy ≤ −20. An entire circRNA–miRNA interaction network of germline stem cells was constructed22, part of which was delineated by Cytoscape (Fig. 6).
Identification of SNPs, indels, AS, and DEU
Based on data at the transcriptome level, the SNP loci in coding regions were analyzed. According to the results of the TopHat comparison between each sample and the reference genome, Samtools software37 was used to mpileup the possible SNP and indel information of each sample, and Annovar software38 was used to annotate it22. We selected Alternative Splicing Detector (ASD; available at http://www.novelbio.com/asd/ASD.html) as a tool to detect the cases with differential alternative splicing based on a bam file after mapping and using the mouse genome sequence as a reference, based on a P-value threshold of <0.05 (Fig. 7a). DEU analysis is currently used for alternative exon usage in alternative splicing22.In this study, DEXSeq software39 was used for DEU analysis (Fig. 7b). DEXSeq uses a generalized linear model to detect differentially expressed genes at the exon level22.
Analysis of alternative splicing and differential exon usage. (a) Classes and statistics of AS profile in germline stem cells. SKIP: exon skipping; MSKIP: cassette exons; IR: retention of single introns; MIR: retention of multiple introns; AE: alternative exon ends; TSS: alternative transcription start site; TTS: alternative transcription termination site. X in front of an abbreviation represents a blurred boundary. (b) An example of alternative exon usage analysis.
Data Records
The identification, annotation data (circRNAs and lncRNAs) as well as SNPs, indels, AS, DEU and circRNA–miRNA interaction network were uploaded to figshare22. The original and normalized data associated with the samples analyzed in this study are deposited at Gene Expression Omnibus (GEO) datasets40.
Technical Validation
Quality control–RNA integrity
The RNA integrity number (RIN) of total RNA in germ stem cells was determined to evaluate RNA integrity using an Agilent Bioanalyzer 2100 (Agilent Technologies). The results showed that the RIN value was more than 7.0 for each sample. The values met the requirements for a noncoding RNA sequencing library.
Quality validation and analyses
We applied FastQC v0.11.5 software to determine data quality and analyzed several variables reflecting this41. A representative summary plot is depicted (FGSC-3). Here, the per base sequence quality was high, with a median quality score above 30, suggesting high-quality sequences across all bases (Fig. 8a). We also created a figure of base composition and base quality for clean data. The results showed satisfactory base composition because the T curve is in accordance with the A curve; meanwhile, the C curve is in accordance with the G curve (Fig. 8b). The per sequence GC content was examined. The pattern of GC composition was similar to the theoretical distribution, indicating that the samples were free from contamination (Fig. 8c). In addition, the sequence length distribution also corresponded to the theoretical curve (Fig. 8d). All other FastQC files were shown to have similar quality metrics compared to sample FGSC-3.
The data provided in these experimental datasets are the first reported genome-wide lncRNA and circRNA transcriptome resources for male and female mouse germline stem cells, which include SNPs, indels, AS, and DEU analyzed by Illumina high-throughput sequencing technology. These findings are useful for the identification of lncRNAs and circRNAs related to the self-renewal and sex-specific properties required for differentiation into gametes, as well as the development of polymorphic genetic markers in germline stem cells and other related research.
References
Kapranov, P. et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316, 1484–1488 (2007).
Sun, J., Lin, Y. & Wu, J. Long non-coding RNA expression profiling of mouse testis during postnatal development. PLoS One 8, e75750 (2013).
Enuka, Y. et al. Circular RNAs are long-lived and display only minimal early alterations in response to a growth factor. Nucleic Acids Res 44, 1370–1383 (2016).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338 (2013).
Chen, L. L. The biogenesis and emerging roles of circular RNAs. Nat Rev Mol Cell Biol 17, 205–211 (2016).
Cabili, M. N. et al. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev 25, 1915–1927 (2011).
Bao, Z. S. et al. RNA-seq of 272 gliomas revealed a novel, recurrent PTPRZ1-MET fusion transcript in secondary glioblastomas. Genome Res 24, 1765–1773 (2014).
Mascarenhas, M. N., Flaxman, S. R., Boerma, T., Vanderpoel, S. & Stevens, G. A. National, regional, and global trends in infertility prevalence since 1990: a systematic analysis of 277 health surveys. PLoS Med 9, e1001356 (2012).
Kubota, H., Avarbock, M. R. & Brinster, R. L. Spermatogonial stem cells share some, but not all, phenotypic and functional characteristics with other stem cells. Proc Natl Acad Sci USA 100, 6487–6492 (2003).
Zou, K. et al. Production of offspring from a germline stem cell line derived from neonatal ovaries. Nat Cell Biol 11, 631–636 (2009).
Wu, J., Luo, H. & Wang, H. Germline stem cells. Curr Top Dev Biol 102, 97–126 (2013).
Li, X., Ao, J. & Wu, J. Systematic identification and comparison of expressed profiles of lncRNAs and circRNAs with associated co-expression and ceRNA networks in mouse germline stem cells. Oncotarget 8, 26573–26590 (2017).
Carrieri, C. et al. Long non-coding antisense RNA controls Uchl1 translation through an embedded SINEB2 repeat. Nature 491, 454–457 (2012).
Mercer, T. R., Dinger, M. E. & Mattick, J. S. Long non-coding RNAs: insights into functions. Nat Rev Genet 10, 155–159 (2009).
Yazgan, O. & Krebs, J. E. Noncoding but nonexpendable: transcriptional regulation by large noncoding RNA in eukaryotes. Biochem Cell Biol 85, 484–496 (2007).
Carninci, P. et al. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005).
Wilusz, J. E., Sunwoo, H. & Spector, D. L. Long noncoding RNAs: functional surprises from the RNA world. Genes Dev 23, 1494–1504 (2009).
Burge, S. W. et al. Rfam 11.0: 10 years of RNA families. Nucleic Acids Res 41, D226–D232 (2013).
Zhang, C. & Wu, J. Production of offspring from a germline stem cell line derived from prepubertal ovaries of germline reporter mice. Mol Hum Reprod 22, 457–464 (2016).
Kanatsu-Shinohara, M. et al. Long-term proliferation in culture and germline transmission of mouse male germline stem cells. Biol Reprod 69, 612–616 (2003).
Li, X. et al. Dosage compensation in the process of inactivation/reactivation during both germ cell development and early embryogenesis in mouse. Sci Rep 7, 3729 (2017).
Li, X., Tian, G. G., Wu, J. Genome-wide identification and characterization of long noncoding and circular RNAs in germline stem cells. figshare https://doi.org/10.6084/m9.figshare.7434089.v3 (2019).
Parkhomchuk, D. et al. Transcriptome analysis by strand-specific sequencing of complementary DNA. Nucleic Acids Res 37, e123 (2009).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14, R36 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28, 511–515 (2010).
Roberts, A., Pimentel, H., Trapnell, C. & Pachter, L. Identification of novel transcripts in annotated genomes using RNA-Seq. Bioinformatics 27, 2325–2329 (2011).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7, 562–578 (2012).
Sun, L. et al. Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res 41, e166 (2013).
Kong, L. et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res 35, W345–W349 (2007).
Sun, K. et al. iSeeRNA: identification of long intergenic non-coding RNA transcripts from transcriptome sequencing data. BMC Genomics 14(Suppl 2), S7 (2013).
Tafer, H. & Hofacker, I. L. RNAplex: a fast tool for RNA-RNA interaction search. Bioinformatics 24, 2657–2663 (2008).
Griffiths-Jones, S., Grocock, R. J., van Dongen, S., Bateman, A. & Enright, A. J. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 34, D140–D144 (2006).
Wu, Y., Wei, B., Liu, H., Li, T. & Rayner, S. MiRPara: a SVM-based software tool for prediction of most probable microRNA coding regions in genome scale sequences. BMC Bioinformatics 12, 107 (2011).
Gao, Y. et al. Comprehensive identification of internal structure and alternative splicing events in circular RNAs. Nat Commun 7, 12060 (2016).
Ahmed, I. et al. Altered expression pattern of circular RNAs in primary and metastatic sites of epithelial ovarian carcinoma. Oncotarget 7, 36366–36381 (2016).
Dang, Y. et al. Tracing the expression of circular RNAs in human pre-implantation embryos. Genome Biol 17, 130 (2016).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Yang, H. & Wang, K. Genomic variant annotation and prioritization with ANNOVAR and wANNOVAR. Nat Protoc 10, 1556–1566 (2015).
Li, Y., Rao, X., Mattox, W. W., Amos, C. I. & Liu, B. RNA-Seq Analysis of Differential Splice Junction Usage and Intron Retentions by DEXSeq. PLoS One 10, e0136653 (2015).
Wu, J. & Li, X. Systematic identification and comparison of expressed profiles of lncRNAs and circRNAs in male and female germline stem cells. Gene Expression Omnibus http://identifiers.org/geo:GSE87824 (2017).
Wang, L., Wang, S. & Li, W. RSeQC: quality control of RNA-seq experiments. Bioinformatics 28, 2184–2185 (2012).
Acknowledgements
This work was supported by National Basic Research Program of China (2018YFC1003501, 2017YFA0504201), Projects of International Cooperation and Exchanged, National Nature Science Foundation of China (81720108017), the National Major Scientific Instruments and Equipment Development Project, National Nature Science Foundation of China (61827814), and Shanghai Jiao Tong University Medicine-Engineering Fund (YG2017ZD11).
Author information
Authors and Affiliations
Contributions
J.W. conceived and supervised the study. X.L. performed the experiments. G.T. performed analysis of circRNA-miRNA interaction network. Y.Z. performed RT-PCR analysis of marker genes for SSCs and FGSCs. X.L. and J.W. analyzed data and wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
ISA-Tab metadata file
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files associated with this article.
About this article
Cite this article
Li, X., Tian, G.G., Zhao, Y. et al. Genome-wide identification and characterization of long noncoding and circular RNAs in germline stem cells. Sci Data 6, 8 (2019). https://doi.org/10.1038/s41597-019-0014-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-019-0014-9
This article is cited by
-
Circular RNAs: Novel Biomarkers in Spermatogenesis Defects and Male Infertility
Reproductive Sciences (2023)
-
Serum- and Feeder-Free Culture of Juvenile Monkey Female Germline Stem Cells and Testosterone Regulation of their Self-Renewal
Stem Cell Reviews and Reports (2022)
-
Ubiquitin-Specific-Processing Protease 7 Regulates Female Germline Stem Cell Self-Renewal Through DNA Methylation
Stem Cell Reviews and Reports (2021)