Abstract
Transcription factors control gene expression; among these, transcriptional repressors must liberate the promoter for derepression to occur. Toxin–antitoxin (TA) modules are bacterial elements that autoregulate their transcription by binding the promoter in a T:A ratio-dependent manner, known as conditional cooperativity. The molecular basis of how excess toxin triggers derepression has remained elusive, largely because monitoring the rearrangement of promoter–repressor complexes, which underpin derepression, is challenging. Here, we dissect the autoregulation of the Salmonella enterica tacAT3 module. Using a combination of assays targeting DNA binding and promoter activity, as well as structural characterization, we determine the essential TA and DNA elements required to control transcription, and we reconstitute a repression-to-derepression path. We demonstrate that excess toxin triggers molecular stripping of the repressor complex off the DNA through multiple allosteric changes causing DNA distortion and ultimately leading to derepression. Thus, our work provides important insight into the mechanisms underlying conditional cooperativity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The atomic coordinates for TacA1 and TacA3–TacT3–DNA were deposited in the PDB under accession codes 7ZG6 and 7ZG5, respectively. Source data are provided with this paper.
References
Rojo, F. Repression of transcription initiation in bacteria. J. Bacteriol. 181, 2987–2991 (1999).
Zhang, R. G. et al. The crystal structure of trp aporepressor at 1.8 A shows how binding tryptophan enhances DNA affinity. Nature 327, 591–597 (1987).
Lewis, M. et al. Crystal structure of the lactose operon repressor and its complexes with DNA and inducer. Science 271, 1247–1254 (1996).
Matthews, K. S. & Nichols, J. C. Lactose repressor protein: functional properties and structure. Prog. Nucleic Acid Res. Mol. Biol. 58, 127–150 (1998).
Reedstrom, R. J., Brown, M. P., Grillo, A., Roen, D. & Royer, C. A. Affinity and specificity of trp repressor–DNA interactions studied with fluorescent oligonucleotides. J. Mol. Biol. 273, 572–585 (1997).
Jurėnas, D., Fraikin, N., Goormaghtigh, F. & Van Melderen, L. Biology and evolution of bacterial toxin–antitoxin systems. Nat. Rev. Microbiol. 20, 335–350 (2022).
Page, R. & Peti, W. Toxin–antitoxin systems in bacterial growth arrest and persistence. Nat. Chem. Biol. 12, 208–214 (2016).
Afif, H., Allali, N., Couturier, M. & Van Melderen, L. The ratio between CcdA and CcdB modulates the transcriptional repression of the ccd poison–antidote system. Mol. Microbiol. 41, 73–82 (2001).
Magnuson, R. & Yarmolinsky, M. B. Corepression of the P1 addiction operon by Phd and Doc. J. Bacteriol. 180, 6342–6351 (1998).
Overgaard, M., Borch, J., Jørgensen, M. G. & Gerdes, K. Messenger RNA interferase RelE controls relBE transcription by conditional cooperativity. Mol. Microbiol. 69, 841–857 (2008).
Boggild, A. et al. The crystal structure of the intact E. coli RelBE toxin–antitoxin complex provides the structural basis for conditional cooperativity. Structure 20, 1641–1648 (2012).
Garcia-Pino, A. et al. Allostery and intrinsic disorder mediate transcription regulation by conditional cooperativity. Cell 142, 101–111 (2010).
Rycroft, J. A. et al. Activity of acetyltransferase toxins involved in Salmonella persister formation during macrophage infection. Nat. Commun. 9, 1993 (2018).
Cheverton, A. M. et al. A Salmonella toxin promotes persister formation through acetylation of tRNA. Mol. Cell 63, 86–96 (2016).
Bikmetov, D. et al. GNAT toxins evolve toward narrow tRNA target specificities. Nucleic Acids Res. 50, 5807–5817 (2022).
Grabe, G. J. et al. Auxiliary interfaces support the evolution of specific toxin–antitoxin pairing. Nat. Chem. Biol. 17, 1296–1304 (2021).
Jurėnas, D., Van Melderen, L. & Garcia-Pino, A. Mechanism of regulation and neutralization of the AtaR–AtaT toxin–antitoxin system. Nat. Chem. Biol. 15, 285–294 (2019).
Qian, H. et al. Toxin–antitoxin operon kacAT of Klebsiella pneumoniae is regulated by conditional cooperativity via a W-shaped KacA–KacT complex. Nucleic Acids Res. 47, 7690–7702 (2019).
LeRoux, M., Culviner, P. H., Liu, Y. J., Littlehale, M. L. & Laub, M. T. Stress can induce transcription of toxin–antitoxin systems without activating toxin. Mol. Cell 79, 280–292 (2020).
Gollan, B., Grabe, G., Michaux, C. & Helaine, S. Bacterial persisters and infection: past, present, and progressing. Annu. Rev. Microbiol. 73, 359–385 (2019).
Leroux, M. & Laub, M. T. Toxin–antitoxin systems as phage defense elements. Annu. Rev. Microbiol. 76, 21–43 (2022).
Schreiter, E. R. & Drennan, C. L. Ribbon–helix–helix transcription factors: variations on a theme. Nat. Rev. Microbiol. 5, 710–720 (2007).
Shin, Y. et al. Structural basis of ribosomal RNA transcription regulation. Nat. Commun. 12, 528 (2021).
Xavier Gomis-Rüth, F. et al. The structure of plasmid-encoded transcriptional repressor CopG unliganded and bound to its operator. EMBO J. 17, 7404–7415 (1998).
Chen, Y., Yang, E., Liu, Z., Tian, Z. & Wang, Y. The role of CopG mediated DNA bending on the regulation of the σ54-dependent promoters in E. coli. Chin. Sci. Bull. 51, 934–940 (2006).
Lavery, R. et al. A systematic molecular dynamics study of nearest-neighbor effects on base pair and base pair step conformations and fluctuations in B-DNA. Nucleic Acids Res. 38, 299–313 (2010).
Lankaš, F., Lavery, R. & Maddocks, J. H. Kinking occurs during molecular dynamics simulations of small DNA minicircles. Structure 14, 1527–1534 (2006).
Chen, S. et al. Indirect readout of DNA sequence at the primary-kink site in the CAP–DNA complex: DNA binding specificity based on energetics of DNA kinking. J. Mol. Biol. 314, 63–74 (2001).
McNamara, P. T., Bolshoy, A., Trifonov, E. A. & Harrington, R. E. Sequence-dependent kinks induced in curved DNA. J. Biomol. Struct. Dyn. 8, 529–538 (1990).
Nagaich, A. K., Bhattacharyya, D., Brahmachari, S. K. & Bansal, M. CA/TG sequence at the 5′ end of oligo(A)-tracts strongly modulates DNA curvature. J. Biol. Chem. 269, 7824–7833 (1994).
De Bruyn, P., Girardin, Y. & Loris, R. Prokaryote toxin–antitoxin modules: complex regulation of an unclear function. Protein Sci. 30, 1103–1113 (2021).
Yashiro, Y., Yamashita, S. & Tomita, K. Crystal structure of the enterohemorrhagic Escherichia coli AtaT–AtaR toxin–antitoxin complex. Structure 27, 476–484 (2019).
Yashiro, Y., Zhang, C., Sakaguchi, Y., Suzuki, T. & Tomita, K. Molecular basis of glycyl-tRNAGly acetylation by TacT from Salmonella Typhimurium. Cell Rep. 37, 110130 (2021).
Potoyan, D. A., Zheng, W., Komives, E. A. & Wolynes, P. G. Molecular stripping in the NF-κB/IκB/DNA genetic regulatory network. Proc. Natl Acad. Sci. USA 113, 110–115 (2016).
Bendtsen, K. L. et al. Toxin inhibition in C. crescentus VapBC1 is mediated by a flexible pseudo-palindromic protein motif and modulated by DNA binding. Nucleic Acids Res. 45, 2875 (2017).
Scheich, C., Kümmel, D., Soumailakakis, D., Heinemann, U. & Büssow, K. Vectors for co-expression of an unrestricted number of proteins. Nucleic Acids Res. 35, e43 (2007).
Clabbers, M. T. B., Gruene, T., Parkhurst, J. M., Abrahams, J. P. & Waterman, D. G. Electron diffraction data processing with DIALS. Acta Crystallogr. D Struct. Biol. 74, 506–518 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Solovyev V. & Salamov A. in Metagenomics and Its Applications in Agriculture, Biomedicine and Environmental Studies 1st edn, Vol. 1 (ed. Li, R. W.) Ch. 4 (Nova Science Publishers, 2011).
Kröger, C. et al. An infection-relevant transcriptomic compendium for Salmonella enterica serovar Typhimurium. Cell Host Microbe 14, 683–695 (2013).
McGibbon, R. T. et al. MDTraj: a modern open library for the analysis of molecular dynamics trajectories. Biophys. J. 109, 1528–1532 (2015).
Wieczór, M., Hospital, A., Bayarri, G., Czub, J. & Orozco, M. Molywood: streamlining the design and rendering of molecular movies. Bioinformatics 36, 4660–4661 (2020).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Lu, X. J. & Olson, W. K. 3DNA: a software package for the analysis, rebuilding and visualization of three‐dimensional nucleic acid structures. Nucleic Acids Res. 31, 5108–5121 (2003).
Tribello, G. A., Bonomi, M., Branduardi, D., Camilloni, C. & Bussi, G. PLUMED 2: new feathers for an old bird. Comput. Phys. Commun. 185, 604–613 (2014).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78, 1950–1958 (2010).
Ivani, I. et al. Parmbsc1: a refined force field for DNA simulations. Nat. Methods 13, 55–58 (2015).
Barducci, A., Bussi, G. & Parrinello, M. Well-tempered metadynamics: a smoothly converging and tunable free-energy method. Phys. Rev. Lett. 100, 020603 (2008).
Zwier, M. C. et al. WESTPA: an interoperable, highly scalable software package for weighted ensemble simulation and analysis. J. Chem. Theory Comput. 11, 800–809 (2015).
Jurasz, J., Bagiński, M., Czub, J. & Wieczór, M. Molecular mechanism of proton-coupled ligand translocation by the bacterial efflux pump EmrE. PLoS Comput. Biol. 17, e1009454 (2021).
Sfriso, P., Hospital, A., Emperador, A. & Orozco, M. Exploration of conformational transition pathways from coarse-grained simulations. Bioinformatics 29, 1980–1986 (2013).
Branduardi, D., Gervasio, F. L. & Parrinello, M. From A to B in free energy space. J. Chem. Phys. 126, 054103 (2007).
Acknowledgements
We thank all the members of the Helaine laboratory, A. Hochschild and D. Mavridou for critical reading of the manuscript. We also thank Harvard Medical School’s Center for Macromolecular Interactions for access to SEC–MALS instrumentation. The work was supported by the National Institutes of Health (R01AI155552) to S.H. In memoriam of Stephen A. Hare who died prematurely and unexpectedly during the preparation of this manuscript. We are indebted to him for his substantial contribution to the work and for his friendship.
Author information
Authors and Affiliations
Contributions
G.J.G. performed protein purification and crystallization and flow cytometry experiments. R.T.G., B.G. and M.S. conducted EMSA experiments. G.J.G. and S.A.H. determined the crystal structures of protein complexes. M.W. and M.O. performed molecular simulations and energy calculations. S.H. and G.J.G. wrote the manuscript. All authors contributed to data interpretation and the writing of the manuscript. S.H. and G.J.G. initiated and orchestrated the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Abel Garcia-Pino and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Dimitris Typas, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Transcriptional control and DNA binding by TacAT3.
a, Promoter activity measured by quantification of GFP intensity in E. coli BL21(DE3) cells carrying pBAD33 (Empty) or pBAD33-tacAT3 (+tacAT3); pFPV25 containing gfp-lva fused to wild-type (black) or mutated (OP1*, red; OP2*, green) tacAT3 promoter. The GFP intensity was normalized to that obtained for the wild-type promoter in the E. coli carrying pBAD33 empty. Statistical analysis (unpaired t test) with p values and average activity are shown for N= 3 biologically independent experiments. b, A representative EMSA (top left) and quantification of DNA binding (top right) by the purified TacA3-TacT3Y143F complex are shown for wild-type S. Typhimurium and related operator sequences in K. pneumoniae (kacAT) and E. coli (ataRT) (two technical repeats per combination). Average binding and ±SEM are shown for N= 3 independent experiments. (bottom) Identity in the OP1 inverted repeat sequences (black dashed boxes) of E. coli ataRT, K. pneumoniae kacAT, and wild-type S. Typhimurium. Bold asterisks point to nucleotide identity. Mutated OP1* sequence with mutation sites (red bold letter) is also shown. c, Operators in closely related TA systems localize to different promoter regions. Start codon of the antitoxin is represented in blue. Bold black letters and half arrows depict operator inverted repeat sequences. Red triangles depict TSS.
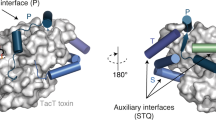
Extended Data Fig. 2 Toxin plasticity displayed near the operator minor groove.
a, A representative EMSA of DNA operator binding by different combinations of wild-type and mutant purified TacA3 and TacT3 (for quantification see Fig. 2b). The asterisk indicates the use of a TacT3Y143F catalytic mutant. b, Cartoon representation of the structure of the KacA-KacT-DNA interface in the minor groove of the operator (light violet), where KacA antitoxin (orange) stabilizes the charged region of KacT toxin (blue) and orientates the positively charged residues of its α1 helix (blue sticks) towards the operator (PDB: 5ZGN). Black dashed lines indicate hydrogen bonds. c, Structures of KacT (left) and TacT3 (right) toxin-DNA interfaces. Polar and charged amino acids are shown as sticks. Toxins are each superimposed with their respective toxin-only states (grey). Superimpositions reveal rearrangement of the α1 helix in the toxin (from grey to blue) when in the toxin-antitoxin-operator complex. Representations are based on structures from PDB: 5XUN, 5ZGN (left) and PDB: 6G96 and TacAT3 hexamer-DNA model (right).
Extended Data Fig. 3 Comparison of P and P’ interfaces.
a, Cartoon representation of the structure of TacT3 molecules with their respective TacA3 antitoxins bound in a P (left) or P’ (right) interface. b, Cartoon representation of the structure of TacA3 α3 helix (top left), α4 helix (top right), and β2 strand (bottom) regions. Superimposed toxins forming the P and P’ interface are overlayed in top and split in bottom respective panels. The main amino acids contributing to the interface are shown as sticks. Black sticks indicate coenzyme A. Black dashed lines depict hydrogen bonds.
Extended Data Fig. 4 Two types of TacT3 dimers.
a, Two side views (top and bottom) of TacT3 condensed (left; PDB 6G96) and elongated (right; PDB 7ZG5) dimers superimposed on blue monomer 1. Black dashed line represents the distance between the centers of mass between each TacT3 monomer within respective dimer. Red dashed circle (top left) depicts putative tRNA binding site of the condensed dimer. b, Surface contribution [Å2] of each secondary structure element of TacT3 to the dimerization interface for the condensed (orange) and elongated (pink) dimers. c, Side view representation of the spatial transition of monomer 2 from a condensed to elongated dimer state. d, Cα distance [Å] of each amino acid in monomer 1 (blue line) and monomer 2 (green line) measured between the two TacT3 dimer states superimposed on blue monomer 1 as shown in A. Secondary structure elements relative to amino acid position are depicted. e, SEC-MALS chromatograms of TacT3Y143F (top) and TacT3Y143F,Loop (bottom) purified proteins. f, SEC-MALS chromatograms of TacA3-TacT3Y143F,Loop (top; Theoretical mass of hexameric TacA3-TacT3 is 79 kDa, measured mass was 80 kDa) and TacA3-TacT3Y143F,Loop-OP complex (bottom; Theoretical mass of hexameric TacA3-TacT3-DNA is 110 kDa, measured mass was 104 kDa). (e and f) Molecular mass, relative UV280, and relative Rayleigh Ratio measurements are shown in green, violet, and red, respectively. g, Free energy profile along the conformation transition between the extended (ξ = −1) and native (ξ = 1) geometries. The value of 0 refers to the dissociated state, in which the CoMs of the TacT3 monomers are separated by more than 5.5 nm. The free energy difference between the two states is ca. 3 kcal/mol.
Extended Data Fig. 5 Conformational changes in octamer vs. hexamer TacAT3 complex.
a, Architecture of secondary structure elements (top) in hexameric and octameric TacA3 antitoxins forming either P (dark yellow) or S/P’ (orange) interfaces. Cartoon representation of the TacA3 antitoxin molecules superimposed on their RHH regions (bottom) from the hexameric (shaded) and octameric (non-shaded) TacAT3 complex forming either S/P’ (bottom left) or P (bottom right) interfaces. Gray dashed lines indicate disordered regions. b, Cα distances [Å] of amino acids involved in the P interfaces measured between the octameric and hexameric complexes relative to antitoxin dimer region and the DNA. The TacA3 dimer was superimposed at the RHH region (amino acids 10–52). The dashed grey line indicates the disordered TacT3 loop 4 region in the hexameric complex. The red number specifies the change in distance between TacT3 R32 from the hexamer to octamer complexes (See Fig. 6a and Supplementary Movie 2). c, Hexamer-to-octamer TacA3 dimer repositioning. Cα distances [Å] of each amino acid of the TacA3 RHH domain superimposed on antitoxin dimer 1 (amino acids 10–52; dashed orange and yellow lines). Distances of TacA3 molecules constituting dimer 2 are shown as solid yellow and orange lines. (See Fig. 6b and Supplementary Movie 3). Secondary structure elements are shown (top) for orientation. d, Relative B factor distribution of antitoxins forming the P and P’ interface in related GNAT TA modules. Two (P and P’) TacA3 molecules from TacAT3-DNA, KacA (P) from KacAT-DNA (PDB 5ZGN), and AtaR (P) from AtaRT-DNA (PDB 6GTS) are shown.
Extended Data Fig. 6 Operator kink and flexibility of hexameric and octameric TacAT3 complex.
a, C5-C5 distances between consecutive bases in equilibrium simulations of octamer with the 25- and 26-bp DNA molecules in which Watson-Crick base-pairing was enforced. For each setup 4 replicates are shown, 400 ns each. In all the systems kink formation consistently occurs at the 5′-TpG-3′ step. b, A representative EMSAs (top) and quantification (bottom) of two tacAT3 operator binding by purified TacA3-TacT3Y143F and TacA3-TacT3Y143F,Loop protein complexes. Statistical analysis (unpaired t test) with p values for change in free DNA fraction are shown for N = 3 independent experiments. c, Conditional probability density of finding the second DNA-binding interface at a given relative position with respect to X-ray DNA structure in pure protein systems (hexamer and octamer on the left and right, respectively), after aligning the first DNA-binding interface with the experimental binding site. The distance is measured from the center of the DNA major groove of each base pair to the center of the β-sheet interface. The position along the DNA is calculated as a continuous argmin function of the distance vector and is offset so that the experimental value (red dot) be equal to 0. The distance shown corresponds to the minimum value of the vector, or minimal distance between the β-sheet and the major groove. Probability densities are calculated using kernel density estimation. d, Free energy profile for the bending of a naked 25-bp double-stranded DNA, with the native operator sequence capable of forming the kink (blue) and a mutated sequence devoid of a central 5′-YR-3′ motif. The dashed line represents the angle found in the crystal structure; an angle of π would correspond to a perfectly straight DNA helix.
Supplementary information
Supplementary Information
Supplementary Tables 1–3, experimental repeats with FACS histograms for Figs. 1b, 2a,e, 3e, 4a and 5d and Extended Data Fig. 1a, and gating strategy.
Supplementary Video 1
Toxin dimer shifts from elongated to condensed state. Movie represents the transition between two TacT3 dimer (green and blue) states (elongated and condensed; Extended Data Fig. 4a–d), generated with adaptive weighted ensemble simulations. In the inset, the red dot marks the position of the system on the 1D free-energy plot analogous to Extended Data Fig. 4g.
Supplementary Video 2
P interface rotation leads to loss of association between the toxin and operator minor groove. Morph movie of a transition of P interface (blue toxin wrapped by a yellow antitoxin) rearranging starting from hexamer and ending on octameric state. Operator from hexameric state (light violet) is fixed throughout.
Supplementary Video 3
Rearrangement of antitoxin dimers in the octamer. Morph movie of RHH regions (amino acids 10–52) of both TacA3 dimers (yellow and orange) starting from hexameric and ending on octameric state. Operator from hexameric state (light violet) is fixed throughout.
Supplementary Video 4
Repression-to-derepression shift. Movie representing all stages of the derepression cycle discussed in this work, from the conformational change of a free hexamer through DNA binding establishing the repression, recruitment of additional TacT3 monomers, formation of the octameric complex, and DNA bending and kinking to the eventual dissociation of the low-affinity complex from the DNA strand.
Supplementary Video 5
Dimer-mediated derepression. Movie representing external TacT3 toxin dimer (gray and green) interacting with one side of the hexameric TacA3 (yellow and orange)–TacT3 (blue) complex bound to the operator (sticks). The interaction is mediated by flexible toxin loop 4 regions (blue and green) forming D interface and loose C-terminal end of the TacA3 antitoxin (orange) that switches from S to P′ interface, ultimately leading to increase in toxin–antitoxin stoichiometry and dissociation of residual toxin monomer (gray).
Source data
Source Data Fig. 1
Unprocessed EMSA gels.
Source Data Fig. 2
Unprocessed EMSA gels.
Source Data Fig. 3
Unprocessed EMSA gels.
Source Data Fig. 4
Unprocessed EMSA gels.
Source Data Fig. 5
Unprocessed EMSA gels.
Source Data Fig. 7
Unprocessed EMSA gels.
Source Data Extended Data Fig. 1
Unprocessed EMSA gels.
Source Data Extended Data Fig. 2
Unprocessed EMSA gels.
Source Data Extended Data Fig. 6
Unprocessed EMSA gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Grabe, G.J., Giorgio, R.T., Wieczór, M. et al. Molecular stripping underpins derepression of a toxin–antitoxin system. Nat Struct Mol Biol (2024). https://doi.org/10.1038/s41594-024-01253-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41594-024-01253-2